

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.5
(359 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
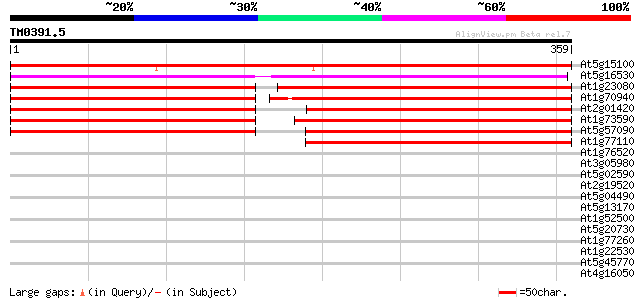
Sequences producing significant alignments: (bits) Value
At5g15100 auxin transport protein - like 404 e-113
At5g16530 auxin efflux carrier -like protein 278 3e-75
At1g23080 auxin transport protein (PIN7) 220 1e-57
At1g70940 auxin transport like protein 219 2e-57
At2g01420 auxin transporter splice variant b (PIN4) 212 2e-55
At1g73590 auxin transporter splice variant b, putative 206 2e-53
At5g57090 root gravitropism control protein (PIN2) 199 2e-51
At1g77110 auxin transport protein (PIN6) 193 1e-49
At1g76520 unknown protein 37 0.017
At3g05980 unknown protein 32 0.41
At5g02590 unknown protein 32 0.54
At2g19520 WD-40 repeat protein MSI4 (MSI4) 31 0.92
At5g04490 unknown protein 30 2.0
At5g13170 senescence-associated protein (SAG29) 29 4.5
At1g52500 putative formamidopyrimidine-DNA glycosylase 1 (fpg1) 29 4.5
At5g20730 auxin response factor 7 (ARF7) 28 5.9
At1g77260 unknown protein 28 5.9
At1g22530 unknown protein 28 5.9
At5g45770 predicted GPI-anchored protein 28 7.8
At4g16050 hypothetical protein 28 7.8
>At5g15100 auxin transport protein - like
Length = 367
Score = 404 bits (1037), Expect = e-113
Identities = 207/367 (56%), Positives = 275/367 (74%), Gaps = 8/367 (2%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MIS D+YHVV+ATVPLYV+M L F+S + KLF+P+QC+GINKFVA FSIPLLSFQ+IS
Sbjct: 1 MISWLDIYHVVSATVPLYVSMTLGFLSARHLKLFSPEQCAGINKFVAKFSIPLLSFQIIS 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKI----SARGG-LKWIITGLSLSTLPNTLI 115
NN +KMS KLI +D +QK L ++ L ++++ RGG L W+ITGLS+S LPNTLI
Sbjct: 61 ENNPFKMSPKLILSDILQKFLVVVVLAMVLRFWHPTGGRGGKLGWVITGLSISVLPNTLI 120
Query: 116 LGIPLMKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAAIPARTMPATPPSQGTGES 175
LG+P++ A+Y DEA ++L QI+ LQS+IWY +LLFL EL+AA + A+ G +
Sbjct: 121 LGMPILSAIYGDEAASILEQIVVLQSLIWYTILLFLFELNAARALPSSGASLEHTGNDQE 180
Query: 176 ETSQELQSKEEEEAEHRP---GRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFRW 232
E + E + KEEE+ E R + + IL K +KLI NPNTYAT IG+IW+++HFR
Sbjct: 181 EANIEDEPKEEEDEEEVAIVRTRSVGTMKILLKAWRKLIINPNTYATLIGIIWATLHFRL 240
Query: 233 GVHMPDVVNQSITILSSGGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPALM 292
G ++P+++++SI +LS GGLGMA FSLGLFMAS S+IIACG +M ++ + LKF++GPALM
Sbjct: 241 GWNLPEMIDKSIHLLSDGGLGMAMFSLGLFMASQSSIIACGTKMAIITMLLKFVLGPALM 300
Query: 293 AVASIVIGLRDTMLKVAIVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVALT 352
++ I L+ T+ KVAI+QAALPQG+VPFVFA+EYN+HP I+STGV+ GMLIALP L
Sbjct: 301 IASAYCIRLKSTLFKVAILQAALPQGVVPFVFAKEYNLHPEIISTGVIFGMLIALPTTLA 360
Query: 353 YYVLLAL 359
YY LL L
Sbjct: 361 YYFLLDL 367
>At5g16530 auxin efflux carrier -like protein
Length = 351
Score = 278 bits (712), Expect = 3e-75
Identities = 144/357 (40%), Positives = 213/357 (59%), Gaps = 10/357 (2%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MI+ GDVY V+ A VPLYV +IL + SVKWW +FT DQC IN+ V F++PL + + +
Sbjct: 1 MINCGDVYKVIEAMVPLYVALILGYGSVKWWHIFTRDQCDAINRLVCYFTLPLFTIEFTA 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKISARGGLKWIITGLSLSTLPNTLILGIPL 120
+ + M+ + I AD + K++ + L L K S +G W IT SL TL N+L++G+PL
Sbjct: 61 HVDPFNMNYRFIAADVLSKVIIVTVLALWAKYSNKGSYCWSITSFSLCTLTNSLVVGVPL 120
Query: 121 MKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAAIPARTMPATPPSQGTGESETSQE 180
KAMY +A L+ Q Q+++W LLLF+ E A S+ +
Sbjct: 121 AKAMYGQQAVDLVVQSSVFQAIVWLTLLLFVLEFRKA----------GFSSNNISDVQVD 170
Query: 181 LQSKEEEEAEHRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFRWGVHMPDVV 240
+ E + E + L +++ V KL NPN Y+ +G+ W+ I RW + +P ++
Sbjct: 171 NINIESGKRETVVVGEKSFLEVMSLVWLKLATNPNCYSCILGIAWAFISNRWHLELPGIL 230
Query: 241 NQSITILSSGGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPALMAVASIVIG 300
SI I+S G G A F++G+FMA +I CG +T++ + LKF+ GPA MA+ SIV+G
Sbjct: 231 EGSILIMSKAGTGTAMFNMGIFMALQEKLIVCGTSLTVMGMVLKFIAGPAAMAIGSIVLG 290
Query: 301 LRDTMLKVAIVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVALTYYVLL 357
L +L+VAI+QAALPQ I F+FA+EY +H +LST V+ GML++LPV + YY L
Sbjct: 291 LHGDVLRVAIIQAALPQSITSFIFAKEYGLHADVLSTAVIFGMLVSLPVLVAYYAAL 347
>At1g23080 auxin transport protein (PIN7)
Length = 619
Score = 220 bits (560), Expect = 1e-57
Identities = 109/188 (57%), Positives = 137/188 (71%)
Query: 172 TGESETSQELQSKEEEEAEHRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFR 231
T E + +++ E +H P + LIL V +KLI+NPNTY++ IGLIW+ + FR
Sbjct: 432 TAELNPKEAIETGETVPVKHMPPASVMTRLILIMVWRKLIRNPNTYSSLIGLIWALVAFR 491
Query: 232 WGVHMPDVVNQSITILSSGGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPAL 291
W V MP ++ QSI+ILS GLGMA FSLGLFMA +IACG A+ ++F GPA+
Sbjct: 492 WDVAMPKIIQQSISILSDAGLGMAMFSLGLFMALQPKLIACGNSTATFAMAVRFFTGPAV 551
Query: 292 MAVASIVIGLRDTMLKVAIVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVAL 351
MAVA++ IGLR +L+VAIVQAALPQGIVPFVFA+EYNVHPAILSTGV+ GMLIALP+ L
Sbjct: 552 MAVAAMAIGLRGDLLRVAIVQAALPQGIVPFVFAKEYNVHPAILSTGVIFGMLIALPITL 611
Query: 352 TYYVLLAL 359
YY+LL L
Sbjct: 612 VYYILLGL 619
Score = 182 bits (462), Expect = 2e-46
Identities = 89/157 (56%), Positives = 117/157 (73%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MI+ D+Y V+TA +PLYV MILA+ SV+WWK+F+PDQCSGIN+FVA F++PLLSF IS
Sbjct: 1 MITWHDLYTVLTAVIPLYVAMILAYGSVRWWKIFSPDQCSGINRFVAIFAVPLLSFHFIS 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKISARGGLKWIITGLSLSTLPNTLILGIPL 120
SNN Y M+L+ I AD +QKL+ + L + + G L+W IT SLSTLPNTL++GIPL
Sbjct: 61 SNNPYAMNLRFIAADTLQKLIMLTLLIIWANFTRSGSLEWSITIFSLSTLPNTLVMGIPL 120
Query: 121 MKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAA 157
+ AMY + + +L+ QI+ LQ +IWY LLLFL E A
Sbjct: 121 LIAMYGEYSGSLMVQIVVLQCIIWYTLLLFLFEYRGA 157
>At1g70940 auxin transport like protein
Length = 640
Score = 219 bits (558), Expect = 2e-57
Identities = 113/193 (58%), Positives = 141/193 (72%), Gaps = 2/193 (1%)
Query: 167 PPSQGTGESETSQELQSKEEEEAEHRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWS 226
P S +S+T L E + ++ P + LIL V +KLI+NPNTY++ IGLIW+
Sbjct: 450 PNSTAALQSKTG--LGGAEASQRKNMPPASVMTRLILIMVWRKLIRNPNTYSSLIGLIWA 507
Query: 227 SIHFRWGVHMPDVVNQSITILSSGGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFL 286
+ FRW V MP ++ QSI+ILS GLGMA FSLGLFMA +IACG + A+ ++FL
Sbjct: 508 LVAFRWHVAMPKIIQQSISILSDAGLGMAMFSLGLFMALQPKLIACGNSVATFAMAVRFL 567
Query: 287 VGPALMAVASIVIGLRDTMLKVAIVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIA 346
GPA+MAVA+I IGLR +L+VAIVQAALPQGIVPFVFA+EYNVHPAILSTGV+ GMLIA
Sbjct: 568 TGPAVMAVAAIAIGLRGDLLRVAIVQAALPQGIVPFVFAKEYNVHPAILSTGVIFGMLIA 627
Query: 347 LPVALTYYVLLAL 359
LP+ L YY+LL L
Sbjct: 628 LPITLVYYILLGL 640
Score = 182 bits (463), Expect = 2e-46
Identities = 89/157 (56%), Positives = 117/157 (73%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MIS D+Y V+TA +PLYV MILA+ SV+WWK+F+PDQCSGIN+FVA F++PLLSF IS
Sbjct: 1 MISWHDLYTVLTAVIPLYVAMILAYGSVRWWKIFSPDQCSGINRFVAIFAVPLLSFHFIS 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKISARGGLKWIITGLSLSTLPNTLILGIPL 120
+NN Y M+L+ I AD +QK++ + L L + G L+W IT SLSTLPNTL++GIPL
Sbjct: 61 TNNPYAMNLRFIAADTLQKIIMLSLLVLWANFTRSGSLEWSITIFSLSTLPNTLVMGIPL 120
Query: 121 MKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAA 157
+ AMY + + +L+ QI+ LQ +IWY LLLFL E A
Sbjct: 121 LIAMYGEYSGSLMVQIVVLQCIIWYTLLLFLFEFRGA 157
>At2g01420 auxin transporter splice variant b (PIN4)
Length = 612
Score = 212 bits (540), Expect = 2e-55
Identities = 104/169 (61%), Positives = 128/169 (75%)
Query: 191 HRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFRWGVHMPDVVNQSITILSSG 250
H P + LIL V +KLI+NPNTY++ IGLIW+ + +RW V MP ++ QSI+ILS
Sbjct: 444 HMPPTSVMTRLILIMVWRKLIRNPNTYSSLIGLIWALVAYRWHVAMPKILQQSISILSDA 503
Query: 251 GLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPALMAVASIVIGLRDTMLKVAI 310
GLGMA FSLGLFMA IIACG + A+ ++F+ GPA+MAVA I IGL +L++AI
Sbjct: 504 GLGMAMFSLGLFMALQPKIIACGNSVATFAMAVRFITGPAIMAVAGIAIGLHGDLLRIAI 563
Query: 311 VQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVALTYYVLLAL 359
VQAALPQGIVPFVFA+EYNVHP ILSTGV+ GMLIALP+ L YY+LL L
Sbjct: 564 VQAALPQGIVPFVFAKEYNVHPTILSTGVIFGMLIALPITLVYYILLGL 612
Score = 180 bits (457), Expect = 9e-46
Identities = 86/157 (54%), Positives = 118/157 (74%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MI+ D+Y V+TA VPLYV MILA+ SV+WWK+F+PDQCSGIN+FVA F++PLLSF IS
Sbjct: 1 MITWHDLYTVLTAVVPLYVAMILAYGSVQWWKIFSPDQCSGINRFVAIFAVPLLSFHFIS 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKISARGGLKWIITGLSLSTLPNTLILGIPL 120
+N+ Y M+ + + AD +QK++ ++ L L ++ G L+W+IT SLSTLPNTL++GIPL
Sbjct: 61 TNDPYAMNFRFVAADTLQKIIMLVLLALWANLTKNGSLEWMITIFSLSTLPNTLVMGIPL 120
Query: 121 MKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAA 157
+ AMY A +L+ Q++ LQ +IWY LLLFL E A
Sbjct: 121 LIAMYGTYAGSLMVQVVVLQCIIWYTLLLFLFEYRGA 157
>At1g73590 auxin transporter splice variant b, putative
Length = 622
Score = 206 bits (523), Expect = 2e-53
Identities = 102/177 (57%), Positives = 129/177 (72%)
Query: 183 SKEEEEAEHRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFRWGVHMPDVVNQ 242
S + +A+ P + LIL V +KLI+NPN+Y++ G+ WS I F+W + MP ++ +
Sbjct: 446 SNKTTQAKVMPPTSVMTRLILIMVWRKLIRNPNSYSSLFGITWSLISFKWNIEMPALIAK 505
Query: 243 SITILSSGGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPALMAVASIVIGLR 302
SI+ILS GLGMA FSLGLFMA N IIACG R A ++F+VGPA+M VAS +GLR
Sbjct: 506 SISILSDAGLGMAMFSLGLFMALNPRIIACGNRRAAFAAAMRFVVGPAVMLVASYAVGLR 565
Query: 303 DTMLKVAIVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVALTYYVLLAL 359
+L VAI+QAALPQGIVPFVFA+EYNVHP ILST V+ GMLIALP+ L YY+LL L
Sbjct: 566 GVLLHVAIIQAALPQGIVPFVFAKEYNVHPDILSTAVIFGMLIALPITLLYYILLGL 622
Score = 188 bits (477), Expect = 4e-48
Identities = 91/157 (57%), Positives = 117/157 (73%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MI+ D YHV+TA VPLYV MILA+ SVKWWK+FTPDQCSGIN+FVA F++PLLSF I+
Sbjct: 1 MITAADFYHVMTAMVPLYVAMILAYGSVKWWKIFTPDQCSGINRFVALFAVPLLSFHFIA 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKISARGGLKWIITGLSLSTLPNTLILGIPL 120
+NN Y M+L+ + AD +QK++ + L L K+S G L W IT SLSTLPNTL++GIPL
Sbjct: 61 ANNPYAMNLRFLAADSLQKVIVLSLLFLWCKLSRNGSLDWTITLFSLSTLPNTLVMGIPL 120
Query: 121 MKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAA 157
+K MY + + L+ QI+ LQ +IWY L+LFL E A
Sbjct: 121 LKGMYGNFSGDLMVQIVVLQCIIWYTLMLFLFEYRGA 157
>At5g57090 root gravitropism control protein (PIN2)
Length = 647
Score = 199 bits (506), Expect = 2e-51
Identities = 98/170 (57%), Positives = 124/170 (72%)
Query: 190 EHRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFRWGVHMPDVVNQSITILSS 249
+ P + LIL V +KLI+NPNTY++ GL WS + F+W + MP +++ SI+ILS
Sbjct: 478 QQMPPASVMTRLILIMVWRKLIRNPNTYSSLFGLAWSLVSFKWNIKMPTIMSGSISILSD 537
Query: 250 GGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPALMAVASIVIGLRDTMLKVA 309
GLGMA FSLGLFMA IIACG + A+ ++FL GPA++A SI IG+R +L +A
Sbjct: 538 AGLGMAMFSLGLFMALQPKIIACGKSVAGFAMAVRFLTGPAVIAATSIAIGIRGDLLHIA 597
Query: 310 IVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVALTYYVLLAL 359
IVQAALPQGIVPFVFA+EYNVHP ILST V+ GML+ALPV + YYVLL L
Sbjct: 598 IVQAALPQGIVPFVFAKEYNVHPDILSTAVIFGMLVALPVTVLYYVLLGL 647
Score = 181 bits (458), Expect = 7e-46
Identities = 90/157 (57%), Positives = 117/157 (74%)
Query: 1 MISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVIS 60
MI+ D+Y V+ A VPLYV MILA+ SV+WW +FTPDQCSGIN+FVA F++PLLSF IS
Sbjct: 1 MITGKDMYDVLAAMVPLYVAMILAYGSVRWWGIFTPDQCSGINRFVAVFAVPLLSFHFIS 60
Query: 61 SNNVYKMSLKLIYADCIQKLLAILALTLIIKISARGGLKWIITGLSLSTLPNTLILGIPL 120
SN+ Y M+ + AD +QK++ + AL L S RG L+W+IT SLSTLPNTL++GIPL
Sbjct: 61 SNDPYAMNYHFLAADSLQKVVILAALFLWQAFSRRGSLEWMITLFSLSTLPNTLVMGIPL 120
Query: 121 MKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAA 157
++AMY D + L+ QI+ LQS+IWY L+LFL E A
Sbjct: 121 LRAMYGDFSGNLMVQIVVLQSIIWYTLMLFLFEFRGA 157
>At1g77110 auxin transport protein (PIN6)
Length = 429
Score = 193 bits (491), Expect = 1e-49
Identities = 93/170 (54%), Positives = 126/170 (73%)
Query: 190 EHRPGRKLRILLILTKVGKKLIKNPNTYATFIGLIWSSIHFRWGVHMPDVVNQSITILSS 249
+ P + + LILT VG+KL +NPNTY++ +GL+WS I F+W + MP++V+ SI I+S
Sbjct: 260 QEMPSAIVMMRLILTVVGRKLSRNPNTYSSLLGLVWSLISFKWNIPMPNIVDFSIKIISD 319
Query: 250 GGLGMATFSLGLFMASNSTIIACGPRMTMVAIGLKFLVGPALMAVASIVIGLRDTMLKVA 309
GLGMA FSLGLFMA +I CG + + + ++F+ GP MA AS+++GLR + L A
Sbjct: 320 AGLGMAMFSLGLFMALQPKMIPCGAKKATMGMLIRFISGPLFMAGASLLVGLRGSRLHAA 379
Query: 310 IVQAALPQGIVPFVFAREYNVHPAILSTGVLLGMLIALPVALTYYVLLAL 359
IVQAALPQGIVPFVFAREYN+HP +LST V+ GM+++LPV + YYVLL L
Sbjct: 380 IVQAALPQGIVPFVFAREYNLHPDLLSTLVIFGMIVSLPVTILYYVLLGL 429
>At1g76520 unknown protein
Length = 390
Score = 37.0 bits (84), Expect = 0.017
Identities = 49/178 (27%), Positives = 80/178 (44%), Gaps = 18/178 (10%)
Query: 166 TPPSQGTGESETSQELQSKEEEEAEHRPGR----KLRILLILTKVGKKLIKNPNTYATF- 220
TPPS + L S +EEE + GR K R++ + KV K I P+T A
Sbjct: 179 TPPSVESNYDSYKVPLISSKEEENNQKAGRWEKVKRRLVSLSQKVNLKTIFAPSTIAAMI 238
Query: 221 ---IGLIWSSIHFRWGVHMP-DVVNQSITILSSGGLGMATFSLGLFMASNSTIIACGPRM 276
IGLI G P V+ S+T++ G + T +G + + + G +M
Sbjct: 239 ALVIGLITPLRKLIIGTEAPLRVLQDSVTLVGDGAVPAMTMIIGGNLLKG--LRSSGMKM 296
Query: 277 TMVAIGLKFLVGPALMAVASIVIGLRDTMLKVAIVQAALPQGIVPFVFAREYNVHPAI 334
+ + IG+ LV ++ S V+ +R K+ +V + + + FV +Y V PA+
Sbjct: 297 SSI-IGV--LVARYVLLPMSGVLIVRGA-YKLDLVTS---EPLYQFVLLLQYAVPPAM 347
>At3g05980 unknown protein
Length = 245
Score = 32.3 bits (72), Expect = 0.41
Identities = 16/59 (27%), Positives = 32/59 (54%), Gaps = 3/59 (5%)
Query: 141 SMIWYNLLLFLHELDAA---IPARTMPATPPSQGTGESETSQELQSKEEEEAEHRPGRK 196
+++W LL + + + + ART+ + PS T S + ++ +EE+E E + G+K
Sbjct: 140 TVLWKELLRLKKQRNPSSSPVTARTVSSLSPSSSTSSSSSLEDAAKREEKEKEGKRGKK 198
>At5g02590 unknown protein
Length = 326
Score = 32.0 bits (71), Expect = 0.54
Identities = 24/96 (25%), Positives = 38/96 (39%), Gaps = 12/96 (12%)
Query: 151 LHELDAAIPARTMPATPPSQGTGESETSQELQSKEEEEAEHRPGRKLRILLILTKVGKKL 210
LH + A T+ + PP T TS ++ S+ +EE E + K L
Sbjct: 75 LHLSTKPVTAATLVSPPPPPSTAADLTSDQISSRPKEEEEE------------AALEKHL 122
Query: 211 IKNPNTYATFIGLIWSSIHFRWGVHMPDVVNQSITI 246
KNPN L+ + H +++N+ I I
Sbjct: 123 TKNPNDVEALQSLMKIKFQTKNIDHALEILNRLIEI 158
>At2g19520 WD-40 repeat protein MSI4 (MSI4)
Length = 507
Score = 31.2 bits (69), Expect = 0.92
Identities = 22/85 (25%), Positives = 38/85 (43%), Gaps = 16/85 (18%)
Query: 165 ATPPSQGTGESETSQELQSKEEEEAEHRPGRKLRILLILTKVGKKLIKNPNT---YATFI 221
AT PS GTG S + + + +E P + + + + + GKK ++P+ Y+ +
Sbjct: 13 ATTPSGGTGASGPKKRGRKPKTKEDSQTPSSQQQSDVKMKESGKKTQQSPSVDEKYSQWK 72
Query: 222 G-------------LIWSSIHFRWG 233
G L+W S+ RWG
Sbjct: 73 GLVPILYDWLANHNLVWPSLSCRWG 97
>At5g04490 unknown protein
Length = 304
Score = 30.0 bits (66), Expect = 2.0
Identities = 30/112 (26%), Positives = 54/112 (47%), Gaps = 18/112 (16%)
Query: 53 LLSFQVISSNNVYKMSLKLIYADCIQKLLAILALTLII-KISAR---------GGLKWII 102
+LSF+ ++ NV + SL + LL +LA + AR GL+ +I
Sbjct: 85 VLSFESLTKRNVIQQSLSRKLVHILSGLLFVLAWPIFSGSTEARYFAAFVPLVNGLRLVI 144
Query: 103 TGLSLSTLPNTLILGIPLMKAMYKD-EADALLPQIIFLQSMIWYNLLLFLHE 153
GLS+S PN++ L+K++ ++ A+ LL +F + ++ + F E
Sbjct: 145 NGLSIS--PNSM-----LIKSVTREGRAEELLKGPLFYVLALLFSAVFFWRE 189
>At5g13170 senescence-associated protein (SAG29)
Length = 292
Score = 28.9 bits (63), Expect = 4.5
Identities = 15/62 (24%), Positives = 31/62 (49%)
Query: 98 LKWIITGLSLSTLPNTLILGIPLMKAMYKDEADALLPQIIFLQSMIWYNLLLFLHELDAA 157
L WI +S+S L++ ++K + L + + +++W+ LFL+++ A
Sbjct: 136 LGWICVAISVSVFAAPLMIVARVIKTKSVEYMPFTLSFFLTISAVMWFAYGLFLNDICIA 195
Query: 158 IP 159
IP
Sbjct: 196 IP 197
>At1g52500 putative formamidopyrimidine-DNA glycosylase 1 (fpg1)
Length = 390
Score = 28.9 bits (63), Expect = 4.5
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 3/51 (5%)
Query: 151 LHELDAAIPARTMPA---TPPSQGTGESETSQELQSKEEEEAEHRPGRKLR 198
L+ DA A+ PA P + G+ E ++ KE+E A+ + G+K R
Sbjct: 278 LYGKDAEKAAKVRPAKRGVKPKEDDGDGEEDEQETEKEDESAKSKKGQKPR 328
>At5g20730 auxin response factor 7 (ARF7)
Length = 1164
Score = 28.5 bits (62), Expect = 5.9
Identities = 15/46 (32%), Positives = 24/46 (51%), Gaps = 3/46 (6%)
Query: 152 HELDAAIPARTMPA---TPPSQGTGESETSQELQSKEEEEAEHRPG 194
H L + + +M + TPPS +S Q+ QSK+ ++A H G
Sbjct: 632 HHLQPQLVSGSMASSVITPPSSSLNQSFQQQQQQSKQLQQAHHHLG 677
>At1g77260 unknown protein
Length = 655
Score = 28.5 bits (62), Expect = 5.9
Identities = 22/63 (34%), Positives = 29/63 (45%), Gaps = 8/63 (12%)
Query: 2 ISLGDVYHVVTATVPLYVTMILAFISVKWWKLFTPDQCSGINKFVANFSIPLLSFQVISS 61
+ L DV V T L + AF+S+ LF N F +FS P L F + SS
Sbjct: 1 MKLSDVGLDVVKTPRLVKLIAFAFLSISTIFLF--------NHFSDSFSYPSLPFPISSS 52
Query: 62 NNV 64
+NV
Sbjct: 53 SNV 55
>At1g22530 unknown protein
Length = 683
Score = 28.5 bits (62), Expect = 5.9
Identities = 12/40 (30%), Positives = 21/40 (52%)
Query: 155 DAAIPARTMPATPPSQGTGESETSQELQSKEEEEAEHRPG 194
+ +P T PA P + T E E + + ++ +EE + PG
Sbjct: 228 EKVVPVETTPAAPVTTETKEEEKAAPVTTETKEEEKAAPG 267
>At5g45770 predicted GPI-anchored protein
Length = 425
Score = 28.1 bits (61), Expect = 7.8
Identities = 11/24 (45%), Positives = 16/24 (65%)
Query: 173 GESETSQELQSKEEEEAEHRPGRK 196
G E ++ L++KEEEE EH+ K
Sbjct: 373 GNEEKTENLKTKEEEEEEHKGSNK 396
>At4g16050 hypothetical protein
Length = 900
Score = 28.1 bits (61), Expect = 7.8
Identities = 15/49 (30%), Positives = 27/49 (54%)
Query: 170 QGTGESETSQELQSKEEEEAEHRPGRKLRILLILTKVGKKLIKNPNTYA 218
Q E+++S + +++EE+ E RKL I + K +++K NT A
Sbjct: 833 QKNNENKSSNGVAAEKEEDDERLKQRKLAIKELALKTEARMLKVENTLA 881
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.140 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,176,001
Number of Sequences: 26719
Number of extensions: 281269
Number of successful extensions: 1280
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1253
Number of HSP's gapped (non-prelim): 29
length of query: 359
length of database: 11,318,596
effective HSP length: 100
effective length of query: 259
effective length of database: 8,646,696
effective search space: 2239494264
effective search space used: 2239494264
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0391.5


