

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.14
(76 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
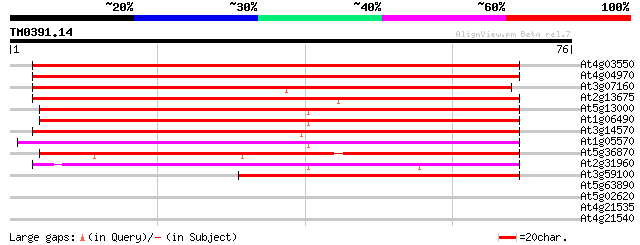
Sequences producing significant alignments: (bits) Value
At4g03550 putative glucan synthase component 95 5e-21
At4g04970 94 1e-20
At3g07160 putative glucan synthase 73 2e-14
At2g13675 callose synthase (1,3-beta-glucan synthase) like protein 73 2e-14
At5g13000 callose synthase catalytic subunit -like protein 67 2e-12
At1g06490 glucan synthase, putative 65 5e-12
At3g14570 hypothetical protein 59 4e-10
At1g05570 putative glucan synthase 55 8e-09
At5g36870 putative glucan synthase 49 3e-07
At2g31960 glucan synthase like protein 49 6e-07
At3g59100 putative protein 45 7e-06
At5g63890 histidinol dehydrogenase 27 2.4
At5g02620 ankyrin - like protein 26 4.1
At4g21535 predicted protein 25 5.3
At4g21540 unknown protein 25 7.0
>At4g03550 putative glucan synthase component
Length = 1780
Score = 95.1 bits (235), Expect = 5e-21
Identities = 41/66 (62%), Positives = 53/66 (80%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
F+FSYFLQ++P+I S+++ LKDV+Y+WH F + N V LLW+P VLIYLMDIQIWY
Sbjct: 502 FTFSYFLQVKPMIKPSKLLWNLKDVDYEWHQFYGDSNRFSVALLWLPVVLIYLMDIQIWY 561
Query: 64 AIYSSL 69
AIYSS+
Sbjct: 562 AIYSSI 567
>At4g04970
Length = 1768
Score = 94.0 bits (232), Expect = 1e-20
Identities = 39/66 (59%), Positives = 52/66 (78%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWY 63
F FSYFLQI+P+IA +R +L LKD Y WH F + + + VG+LW+P +L+YLMD+QIWY
Sbjct: 493 FIFSYFLQIRPLIAPTRALLNLKDATYNWHEFFGSTHRIAVGMLWLPVILVYLMDLQIWY 552
Query: 64 AIYSSL 69
+IYSSL
Sbjct: 553 SIYSSL 558
>At3g07160 putative glucan synthase
Length = 1931
Score = 73.2 bits (178), Expect = 2e-14
Identities = 31/67 (46%), Positives = 48/67 (71%), Gaps = 2/67 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFI--KNGNTVYVGLLWIPAVLIYLMDIQI 61
FSF+YFLQI+P++ +R++++ ++ Y WH F+ KN N + V LW P V IYL+DI I
Sbjct: 696 FSFAYFLQIKPLVGPTRMIVKQNNIPYSWHDFVSRKNYNALTVASLWAPVVAIYLLDIHI 755
Query: 62 WYAIYSS 68
+Y I+S+
Sbjct: 756 FYTIFSA 762
>At2g13675 callose synthase (1,3-beta-glucan synthase) like protein
Length = 1939
Score = 73.2 bits (178), Expect = 2e-14
Identities = 29/68 (42%), Positives = 47/68 (68%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVY--VGLLWIPAVLIYLMDIQI 61
F+FSYFLQ++ ++ + ++ ++ V+Y+WH F N Y V LW+P +L+Y MD QI
Sbjct: 676 FAFSYFLQVKLLVKPTNAIMSIRHVKYKWHEFFPNAEHNYGAVVSLWLPVILVYFMDTQI 735
Query: 62 WYAIYSSL 69
WYAI+S++
Sbjct: 736 WYAIFSTI 743
>At5g13000 callose synthase catalytic subunit -like protein
Length = 1963
Score = 67.0 bits (162), Expect = 2e-12
Identities = 27/67 (40%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+++I+P++A ++ +++ + +QWH F N V LW P +L+Y MD QIW
Sbjct: 684 AFSYYIEIRPLVAPTQAIMKARVTNFQWHEFFPRAKNNIGVVIALWAPIILVYFMDSQIW 743
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 744 YAIFSTL 750
>At1g06490 glucan synthase, putative
Length = 1933
Score = 65.5 bits (158), Expect = 5e-12
Identities = 27/67 (40%), Positives = 45/67 (66%), Gaps = 2/67 (2%)
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDIQIW 62
+FSY+++I P++ ++++ ++ V Y+WH F N N + +W P VL+Y MD QIW
Sbjct: 693 AFSYYVEILPLVNPTKLIWDMHVVNYEWHEFFPNATHNIGVIIAIWGPIVLVYFMDTQIW 752
Query: 63 YAIYSSL 69
YAI+S+L
Sbjct: 753 YAIFSTL 759
>At3g14570 hypothetical protein
Length = 1973
Score = 58.9 bits (141), Expect = 4e-10
Identities = 22/68 (32%), Positives = 42/68 (61%), Gaps = 2/68 (2%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKN--GNTVYVGLLWIPAVLIYLMDIQI 61
F+FSY +I+P+I +R+++++ Y+WH N + +W P +++Y MD QI
Sbjct: 706 FAFSYAFEIKPLIEPTRLIMKVGVRNYEWHEIFPEVKSNAAAIVAVWAPIMVVYFMDTQI 765
Query: 62 WYAIYSSL 69
WY++Y ++
Sbjct: 766 WYSVYCTI 773
>At1g05570 putative glucan synthase
Length = 1858
Score = 54.7 bits (130), Expect = 8e-09
Identities = 25/70 (35%), Positives = 39/70 (55%), Gaps = 2/70 (2%)
Query: 2 IFFSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLIYLMDI 59
I ++ QI+P++ ++ ++ + Y WH F + N V LW P +L+Y MD
Sbjct: 634 IMMLMMWWSQIKPLVGPTKDIMRIHISVYSWHEFFPHAKNNLGVVIALWSPVILVYFMDT 693
Query: 60 QIWYAIYSSL 69
QIWYAI S+L
Sbjct: 694 QIWYAIVSTL 703
>At5g36870 putative glucan synthase
Length = 1662
Score = 49.3 bits (116), Expect = 3e-07
Identities = 25/69 (36%), Positives = 43/69 (62%), Gaps = 5/69 (7%)
Query: 5 SFSYFL-QIQPIIASSRVVLELKDVEY---QWHAFIKNGNTVYVGLLWIPAVLIYLMDIQ 60
+FSY++ QI+P++ ++ ++ + Y ++ +KN V + LW P +L+Y MD Q
Sbjct: 537 AFSYYVEQIKPLMGPTKEIMSVPMPGYWLPEFFPHVKNNRGVVI-TLWSPVILVYFMDTQ 595
Query: 61 IWYAIYSSL 69
IWYAI S+L
Sbjct: 596 IWYAIVSTL 604
>At2g31960 glucan synthase like protein
Length = 1956
Score = 48.5 bits (114), Expect = 6e-07
Identities = 28/76 (36%), Positives = 41/76 (53%), Gaps = 11/76 (14%)
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNG--NTVYVGLLWIPAVLI------- 54
FSF Y QI+P++ ++ ++ + Y+WH F + N V LW P +L+
Sbjct: 673 FSF-YAEQIKPLVKPTKDIMRVHISVYRWHEFFPHAKSNMGVVIALWSPVILVSRHIFLA 731
Query: 55 -YLMDIQIWYAIYSSL 69
Y MD QIWYAI S+L
Sbjct: 732 VYFMDTQIWYAIVSTL 747
>At3g59100 putative protein
Length = 1808
Score = 45.1 bits (105), Expect = 7e-06
Identities = 20/38 (52%), Positives = 24/38 (62%)
Query: 32 WHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWYAIYSSL 69
W A N V +W P VL+YLMD QIWYAI+S+L
Sbjct: 619 WWAQATTNNIGVVIAIWAPIVLVYLMDTQIWYAIFSTL 656
>At5g63890 histidinol dehydrogenase
Length = 466
Score = 26.6 bits (57), Expect = 2.4
Identities = 11/36 (30%), Positives = 21/36 (57%), Gaps = 2/36 (5%)
Query: 15 IIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIP 50
+ A +++ +KD E +W I+N +V++G W P
Sbjct: 352 LYAPEHLIINVKDAE-KWEGLIENAGSVFIG-PWTP 385
>At5g02620 ankyrin - like protein
Length = 524
Score = 25.8 bits (55), Expect = 4.1
Identities = 12/65 (18%), Positives = 32/65 (48%), Gaps = 7/65 (10%)
Query: 1 IIFFSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQ 60
++F SF+ F+ + ++ + VV+ + + Q A I L+W+ ++I + +
Sbjct: 403 VVFDSFALFISLAVVVVQTSVVVIERRAKKQMMAIINK-------LMWMACIMISVAFVS 455
Query: 61 IWYAI 65
+ + +
Sbjct: 456 LSFVV 460
>At4g21535 predicted protein
Length = 455
Score = 25.4 bits (54), Expect = 5.3
Identities = 10/30 (33%), Positives = 21/30 (69%), Gaps = 2/30 (6%)
Query: 11 QIQPIIASSRVVLELKDVEYQWHA--FIKN 38
+++P+ + V LE+++ +YQ HA F+K+
Sbjct: 136 EVKPLFEDADVQLEIQETKYQLHAKEFVKS 165
>At4g21540 unknown protein
Length = 485
Score = 25.0 bits (53), Expect = 7.0
Identities = 7/24 (29%), Positives = 17/24 (70%)
Query: 11 QIQPIIASSRVVLELKDVEYQWHA 34
+++P+ + + LE+++ +YQ HA
Sbjct: 141 EVKPLFEDANIQLEIQETKYQLHA 164
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.333 0.146 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,660,787
Number of Sequences: 26719
Number of extensions: 54132
Number of successful extensions: 213
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 192
Number of HSP's gapped (non-prelim): 16
length of query: 76
length of database: 11,318,596
effective HSP length: 52
effective length of query: 24
effective length of database: 9,929,208
effective search space: 238300992
effective search space used: 238300992
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0391.14


