

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.3
(149 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
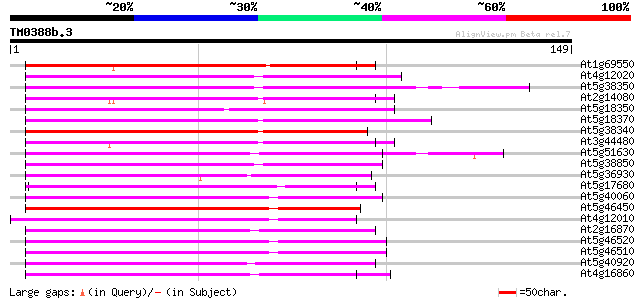
Sequences producing significant alignments: (bits) Value
At1g69550 putative disease resistance protein 70 4e-13
At4g12020 putative disease resistance protein 69 7e-13
At5g38350 disease resistance protein-like 66 6e-12
At2g14080 putative disease resistance protein 65 1e-11
At5g18350 disease resistance protein -like 64 3e-11
At5g18370 disease resistance protein -like 63 7e-11
At5g38340 disease resistance - like protein 62 9e-11
At3g44480 disease resistance protein -like 62 1e-10
At5g51630 putative protein 61 3e-10
At5g38850 disease resistance protein - like 61 3e-10
At5g36930 disease resistance like protein 61 3e-10
At5g17680 disease resistance protein RPP1-WsB - like protein 61 3e-10
At5g40060 disease resistance -like protein 60 4e-10
At5g46450 disease resistance protein-like 60 6e-10
At4g12010 like disease resistance protein (TMV N-like) 60 6e-10
At2g16870 putative disease resistance protein 60 6e-10
At5g46520 disease resistance protein-like 59 7e-10
At5g46510 disease resistance protein-like 59 7e-10
At5g40920 disease resistance protein-like 59 7e-10
At4g16860 disease resistance RPP5 like protein 59 1e-09
>At1g69550 putative disease resistance protein
Length = 1398
Score = 70.1 bits (170), Expect = 4e-13
Identities = 41/93 (44%), Positives = 60/93 (64%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS L+E P+ G+ L++LD +GC++L+++ SIG L L L L GCSSLV
Sbjct: 1076 LKTLNLSGCSSLVELPSSIGNLNLKKLDLSGCSSLVELPSSIGNLINLKKLDLSGCSSLV 1135
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 1136 ELPL-SIGNLINLQELYLSECSSLVELPSSIGN 1167
Score = 62.4 bits (150), Expect = 9e-11
Identities = 40/94 (42%), Positives = 59/94 (62%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+LDLS L+E P+ G+ L++LD +GC++L+++ SIG L L L L CSSL
Sbjct: 1099 LKKLDLSGCSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSL 1158
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 1159 VELP-SSIGNLINLQELYLSECSSLVELPSSIGN 1191
Score = 62.0 bits (149), Expect = 1e-10
Identities = 41/94 (43%), Positives = 59/94 (62%), Gaps = 3/94 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L+ L LS L+E P+ G+ L++LD +GC++L+++ SIG L L L+L GCSSL
Sbjct: 1028 LQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSGCSSL 1087
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ LN L+ L LSGC+ L P+ IGN
Sbjct: 1088 VEL-PSSIGNLN-LKKLDLSGCSSLVELPSSIGN 1119
Score = 59.7 bits (143), Expect = 6e-10
Identities = 39/94 (41%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L RLDL L+E P+ G+ LE F GC++LL++ SIG L L L L+ SSL
Sbjct: 788 LPRLDLMGCSSLVELPSSIGNLINLEAFYFHGCSSLLELPSSIGNLISLKILYLKRISSL 847
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V + ++ L +L++L+LSGC+ L P+ IGN
Sbjct: 848 VEI-PSSIGNLINLKLLNLSGCSSLVELPSSIGN 880
Score = 58.2 bits (139), Expect = 2e-09
Identities = 34/93 (36%), Positives = 53/93 (56%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DL S +L E PN + L + + C++L+++ SIG T + L ++GCSSL+
Sbjct: 693 LKVMDLRYSSHLKELPNLSTAINLLEMVLSDCSSLIELPSSIGNATNIKSLDIQGCSSLL 752
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L ++ L +L L L GC+ L P+ IGN
Sbjct: 753 KL-PSSIGNLITLPRLDLMGCSSLVELPSSIGN 784
Score = 58.2 bits (139), Expect = 2e-09
Identities = 38/94 (40%), Positives = 58/94 (61%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L+ L LS L+E P+ G+ L++LD +GC++L+++ SIG L L L+L CSSL
Sbjct: 956 LQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSL 1015
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 1016 VEL-PSSIGNLINLQELYLSECSSLVELPSSIGN 1048
Score = 58.2 bits (139), Expect = 2e-09
Identities = 36/94 (38%), Positives = 53/94 (56%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
+K LD+ L++ P+ G+ L RLD GC++L+++ SIG L L L L GCSSL
Sbjct: 740 IKSLDIQGCSSLLKLPSSIGNLITLPRLDLMGCSSLVELPSSIGNLINLPRLDLMGCSSL 799
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L + GC+ L P+ IGN
Sbjct: 800 VEL-PSSIGNLINLEAFYFHGCSSLLELPSSIGN 832
Score = 57.8 bits (138), Expect = 2e-09
Identities = 39/94 (41%), Positives = 57/94 (60%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK L L L+E P+ G+ L+ L+ +GC++L+++ SIG L L L L GCSSL
Sbjct: 836 LKILYLKRISSLVEIPSSIGNLINLKLLNLSGCSSLVELPSSIGNLINLKKLDLSGCSSL 895
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 896 VELPL-SIGNLINLQELYLSECSSLVELPSSIGN 928
Score = 56.2 bits (134), Expect = 6e-09
Identities = 38/94 (40%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+LDLS L+E P G+ L+ L + C++L+++ SIG L L L+L CSSL
Sbjct: 884 LKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLKTLNLSECSSL 943
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 944 VEL-PSSIGNLINLQELYLSECSSLVELPSSIGN 976
Score = 51.2 bits (121), Expect = 2e-07
Identities = 36/89 (40%), Positives = 52/89 (57%), Gaps = 2/89 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+LDLS L+E P G+ L+ L + C++L+++ SIG L L L L CSSL
Sbjct: 1123 LKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLQELYLSECSSL 1182
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTP 92
V L ++ L +L+ L L+ CT+L S P
Sbjct: 1183 VEL-PSSIGNLINLKKLDLNKCTKLVSLP 1210
>At4g12020 putative disease resistance protein
Length = 1895
Score = 69.3 bits (168), Expect = 7e-13
Identities = 37/100 (37%), Positives = 57/100 (57%), Gaps = 2/100 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++ LS S L + P + LE +D GC +LL + SI L +L FL+L+GCS L
Sbjct: 1260 LKKMRLSYSDQLTKIPRLSSATNLEHIDLEGCNSLLSLSQSISYLKKLVFLNLKGCSKLE 1319
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCG 104
+ + ++ +L SL +L+LSGC++L + P N G
Sbjct: 1320 N--IPSMVDLESLEVLNLSGCSKLGNFPEISPNVKELYMG 1357
Score = 30.0 bits (66), Expect = 0.48
Identities = 19/54 (35%), Positives = 30/54 (55%), Gaps = 1/54 (1%)
Query: 5 LKRLDLSNSKYLMETP-NFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSL 57
L++LDL NS++L P + + LE L+ +GC +L + S + L FL L
Sbjct: 1374 LEKLDLENSRHLKNLPTSIYKLKHLETLNLSGCISLERFPDSSRRMKCLRFLDL 1427
Score = 27.3 bits (59), Expect = 3.1
Identities = 23/68 (33%), Positives = 33/68 (47%), Gaps = 2/68 (2%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTR 87
LE+LD +L + SI L L L+L GC SL + + + LR L LS T
Sbjct: 1374 LEKLDLENSRHLKNLPTSIYKLKHLETLNLSGCISL-ERFPDSSRRMKCLRFLDLSR-TD 1431
Query: 88 LESTPNFI 95
++ P+ I
Sbjct: 1432 IKELPSSI 1439
>At5g38350 disease resistance protein-like
Length = 833
Score = 66.2 bits (160), Expect = 6e-12
Identities = 45/134 (33%), Positives = 69/134 (50%), Gaps = 9/134 (6%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DLS SK+L E P+ + LE L +GC +L+++ SIG L +L LSL GCS L
Sbjct: 480 LKRMDLSESKHLKELPDLSTATNLEYLIMSGCISLVELPSSIGKLRKLLMLSLRGCSKLE 539
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKG 124
+ L T L SL L L+ C ++ P N + + ++P S K
Sbjct: 540 A--LPTNINLESLDYLDLTDCLLIKKFPEISTNIKDLKLTKTAI---KEVP----STIKS 590
Query: 125 GSRIREVDHVYEQD 138
S +R+++ Y ++
Sbjct: 591 WSHLRKLEMSYSEN 604
>At2g14080 putative disease resistance protein
Length = 1215
Score = 65.1 bits (157), Expect = 1e-11
Identities = 37/98 (37%), Positives = 58/98 (58%), Gaps = 1/98 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DL +SK L E P+ + LE L+ GC++L+++ SIG T+L L L GCSSL+
Sbjct: 676 LKRMDLFSSKNLKELPDLSSATNLEVLNLNGCSSLVELPFSIGNATKLLKLELSGCSSLL 735
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L ++ +L+ + S C L P+ IGN ++ +
Sbjct: 736 EL-PSSIGNAINLQTIDFSHCENLVELPSSIGNATNLK 772
Score = 57.8 bits (138), Expect = 2e-09
Identities = 36/99 (36%), Positives = 57/99 (57%), Gaps = 2/99 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L +L+LS L+E P+ G+ L+ +DF+ C NL+++ SIG T L L L CSSL
Sbjct: 723 LLKLELSGCSSLLELPSSIGNAINLQTIDFSHCENLVELPSSIGNATNLKELDLSCCSSL 782
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L ++ +L+ LHL C+ L+ P+ IGN ++ +
Sbjct: 783 KEL-PSSIGNCTNLKKLHLICCSSLKELPSSIGNCTNLK 820
Score = 51.2 bits (121), Expect = 2e-07
Identities = 37/99 (37%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK LDLS L E P+ G+ L++L C++L ++ SIG T L L L CSSL
Sbjct: 771 LKELDLSCCSSLKELPSSIGNCTNLKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSL 830
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+ L +N L L L+GC L P+FIG ++ +
Sbjct: 831 IKLPSSIGNAIN-LEKLILAGCESLVELPSFIGKATNLK 868
Score = 48.5 bits (114), Expect = 1e-06
Identities = 36/96 (37%), Positives = 53/96 (54%), Gaps = 6/96 (6%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+L L L E P+ G+ L+ L T C++L+++ SIG L L L GC SL
Sbjct: 795 LKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSLIKLPSSIGNAINLEKLILAGCESL 854
Query: 64 VSL--YLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++G + +L+IL+L + L P+FIGN
Sbjct: 855 VELPSFIG---KATNLKILNLGYLSCLVELPSFIGN 887
Score = 30.4 bits (67), Expect = 0.37
Identities = 31/99 (31%), Positives = 44/99 (44%), Gaps = 11/99 (11%)
Query: 3 PFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCT--NLLQVHPSIGLLTELAFLSLEGC 60
P L+ L + S+ L E S LER+ + N+ ++ P + +T L L L GC
Sbjct: 956 PRLEDLQMLYSENLSEF-----SHVLERITVLELSDINIREMTPWLNRITRLRRLKLSGC 1010
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
LVSL + +SL IL C LE NP+
Sbjct: 1011 GKLVSLPQLS----DSLIILDAENCGSLERLGCSFNNPN 1045
>At5g18350 disease resistance protein -like
Length = 1193
Score = 63.9 bits (154), Expect = 3e-11
Identities = 36/98 (36%), Positives = 54/98 (54%), Gaps = 1/98 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DLS+SK L E P+ + LE LD + C+ LL++ SIG T L L L C SL+
Sbjct: 647 LKRMDLSHSKDLKEIPDLSNATNLEELDLSSCSGLLELTDSIGKATNLKRLKL-ACCSLL 705
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
++ + +L++L L C E P IG ++ +
Sbjct: 706 KKLPSSIGDATNLQVLDLFHCESFEELPKSIGKLTNLK 743
>At5g18370 disease resistance protein -like
Length = 1210
Score = 62.8 bits (151), Expect = 7e-11
Identities = 38/108 (35%), Positives = 59/108 (54%), Gaps = 1/108 (0%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR++L +++ L E P+ + LE L + CT+LL++ SI T L L L GC+SLV
Sbjct: 688 LKRMELGDARNLKEIPDLSNATNLESLLLSFCTSLLEIPSSIRGTTNLKELDLGGCASLV 747
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGN 112
L +C SL L+LS C+ L P + S+ R +++ G+
Sbjct: 748 KL-SSCICNATSLEELNLSACSNLVELPCALPGDSNMRSLSKLLLNGS 794
>At5g38340 disease resistance - like protein
Length = 1059
Score = 62.4 bits (150), Expect = 9e-11
Identities = 37/91 (40%), Positives = 57/91 (61%), Gaps = 1/91 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK ++LSNS+ L E P+ + +L+ L+ T C++L+++ SIG T L L+L C+SLV
Sbjct: 681 LKWMNLSNSRNLKELPDLSTATKLQDLNLTRCSSLVEIPFSIGNTTNLEKLNLVMCTSLV 740
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
L ++ L+ LR L L GC++LE P I
Sbjct: 741 EL-PSSIGSLHKLRELRLRGCSKLEVLPTNI 770
Score = 28.5 bits (62), Expect = 1.4
Identities = 30/117 (25%), Positives = 45/117 (37%), Gaps = 9/117 (7%)
Query: 11 SNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLG 69
SN + E P + RLE L GC NL+ + L L+ + + C SL L
Sbjct: 845 SNDTKMQELPRWVKKISRLETLMLEGCKNLVTLPE---LPDSLSNIGVINCESLERLDCS 901
Query: 70 TVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKGGS 126
N + C +L + S C ++PG ++P F + GGS
Sbjct: 902 FYKHPN--MFIGFVNCLKLNKEARELIQTSSSTCS---ILPGRRVPSNFTYRKTGGS 953
>At3g44480 disease resistance protein -like
Length = 1220
Score = 61.6 bits (148), Expect = 1e-10
Identities = 38/98 (38%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DLS S YL E PN + LE L C++L+++ SI LT L L LE CSSL
Sbjct: 716 LKWMDLSYSSYLKELPNLSTATNLEELKLRNCSSLVELPSSIEKLTSLQILDLENCSSLE 775
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L + LR L L C+ L P IG ++ +
Sbjct: 776 K--LPAIENATKLRELKLQNCSSLIELPLSIGTATNLK 811
Score = 61.2 bits (147), Expect = 2e-10
Identities = 33/93 (35%), Positives = 53/93 (56%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ LDL N L + P + + +L L C++L+++ SIG T L L++ GCSSLV
Sbjct: 763 LQILDLENCSSLEKLPAIENATKLRELKLQNCSSLIELPLSIGTATNLKQLNISGCSSLV 822
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L ++ ++ L + LS C+ L + P+ IGN
Sbjct: 823 KL-PSSIGDITDLEVFDLSNCSSLVTLPSSIGN 854
Score = 50.4 bits (119), Expect = 3e-07
Identities = 34/99 (34%), Positives = 54/99 (54%), Gaps = 3/99 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+L++S L++ P+ G LE D + C++L+ + SIG L L L + GCS L
Sbjct: 810 LKQLNISGCSSLVKLPSSIGDITDLEVFDLSNCSSLVTLPSSIGNLQNLCKLIMRGCSKL 869
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+L + L SL L+L+ C++L+S P + S R
Sbjct: 870 EALPIN--INLKSLDTLNLTDCSQLKSFPEISTHISELR 906
Score = 26.9 bits (58), Expect = 4.1
Identities = 27/95 (28%), Positives = 43/95 (44%), Gaps = 10/95 (10%)
Query: 5 LKRLDLSNSKYLMETPN-FDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L +S + LME P+ FD +L ++ +V P + ++ L LSL C++L
Sbjct: 925 LADFQISYFESLMEFPHAFDIITKLHL-----SKDIQEVPPWVKRMSRLRDLSLNNCNNL 979
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNP 98
VSL + +SL ++ C LE NP
Sbjct: 980 VSLPQLS----DSLDYIYADNCKSLERLDCCFNNP 1010
>At5g51630 putative protein
Length = 1239
Score = 60.8 bits (146), Expect = 3e-10
Identities = 44/134 (32%), Positives = 63/134 (46%), Gaps = 12/134 (8%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L +DLS S+ L E PN L+ LD GC +L+ V SI L++L L++ C+ L
Sbjct: 786 LVNIDLSLSEKLKEFPNLSKVTNLDTLDLYGCKSLVTVPSSIQSLSKLTELNMRRCTGLE 845
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQF-- 122
+ L T L SL L LSGC++L + P N + ++P W D F
Sbjct: 846 A--LPTDVNLESLHTLDLSGCSKLTTFPKISRNIERLLLDDTAI---EEVPSWIDDFFEL 900
Query: 123 -----KGGSRIREV 131
KG R+R +
Sbjct: 901 TTLSMKGCKRLRNI 914
Score = 54.7 bits (130), Expect = 2e-08
Identities = 33/95 (34%), Positives = 51/95 (52%), Gaps = 2/95 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++DLS S+ L E P+ + LE +D C +L+ + S+ L +L L + CS++
Sbjct: 626 LKKMDLSKSENLKEIPDLSYAVNLEEMDLCSCKSLVTLPSSVRNLDKLRVLRMSSCSNVE 685
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
L T L SL +L+L C++L S P N S
Sbjct: 686 --VLPTDLNLESLDLLNLEDCSQLRSFPQISRNIS 718
Score = 35.8 bits (81), Expect = 0.009
Identities = 28/84 (33%), Positives = 41/84 (48%), Gaps = 4/84 (4%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L LDLS L P S+ +ERL T + +V I EL LS++GC L
Sbjct: 856 LHTLDLSGCSKLTTFPKI--SRNIERL-LLDDTAIEEVPSWIDDFFELTTLSMKGCKRLR 912
Query: 65 SLYLGTVCELNSLRILHLSGCTRL 88
++ ++CEL + + + S C RL
Sbjct: 913 NIST-SICELKCIEVANFSDCERL 935
Score = 33.1 bits (74), Expect = 0.057
Identities = 37/144 (25%), Positives = 54/144 (36%), Gaps = 15/144 (10%)
Query: 17 METPNFDGSQRLERLDFTGCTN--LLQVHPSIGLLTELAFLS--LEGCSSLVSLYLGTVC 72
+E NF +RL D L + I L E +FL C LVS+
Sbjct: 924 IEVANFSDCERLTEFDDASMVRRILRTIDDLIALYEEASFLHAIFVLCRKLVSICAMVFK 983
Query: 73 ELNSLRI--------LHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKG 124
+L L + C+ L+ + S+ C V+PG K+P F +Q G
Sbjct: 984 YPQALSYFFNSPEADLIFANCSSLDRDAETLILESNHGCA---VLPGGKVPNCFMNQACG 1040
Query: 125 GSRIREVDHVYEQDHWLGFAFCAV 148
S + Y + +LGF C V
Sbjct: 1041 SSVSIPLHESYYSEEFLGFKACIV 1064
>At5g38850 disease resistance protein - like
Length = 986
Score = 60.8 bits (146), Expect = 3e-10
Identities = 33/95 (34%), Positives = 54/95 (56%), Gaps = 2/95 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++ LS+S YL + P+ + LE LD C NL+++ S L +L +L++ GC L
Sbjct: 616 LKKMSLSSSWYLKKLPDLSNATNLEELDLRACQNLVELPSSFSYLHKLKYLNMMGCRRLK 675
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
+ L SL ++++ GC+RL+S P+ N S
Sbjct: 676 E--VPPHINLKSLELVNMYGCSRLKSFPDISTNIS 708
>At5g36930 disease resistance like protein
Length = 1164
Score = 60.8 bits (146), Expect = 3e-10
Identities = 37/93 (39%), Positives = 54/93 (57%), Gaps = 2/93 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLL-TELAFLSLEGCSSL 63
+K LDLS+S YL ETP+F +E+L C +L+ VH SIG+L +L L+L C L
Sbjct: 622 VKYLDLSHSVYLRETPDFSYFPNVEKLILINCKSLVLVHKSIGILDKKLVLLNLSSCIEL 681
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
+ + +L SL L LS C++LE + +G
Sbjct: 682 -DVLPEEIYKLKSLESLFLSNCSKLERLDDALG 713
Score = 29.3 bits (64), Expect = 0.82
Identities = 31/108 (28%), Positives = 47/108 (42%), Gaps = 9/108 (8%)
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS-SLVSL 66
LD+ L TP+ L +L C +L ++ P I L+F+ L+GC +
Sbjct: 854 LDVGKCIMLKRTPDISKCSALFKLQLNDCISLFEI-PGIHNHEYLSFIVLDGCKLASTDT 912
Query: 67 YLGTVCELNSLRILHLSGCTRLE-STPNFIGNPSHF---RCGFDIVVP 110
+ T+ E N L+ H C + PN I N +F + F I VP
Sbjct: 913 TINTMLE-NWLKRNH--ECIYIPVDRPNVIPNWVYFEEEKRSFSITVP 957
Score = 28.1 bits (61), Expect = 1.8
Identities = 35/125 (28%), Positives = 50/125 (40%), Gaps = 32/125 (25%)
Query: 17 METPNFDGSQRLERLD------------FTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+E+ +LERLD T L ++ +I L +L LSL GC L+
Sbjct: 694 LESLFLSNCSKLERLDDALGELESLTTLLADFTALREIPSTINQLKKLKRLSLNGCKGLL 753
Query: 65 S-----LYLGTVCELNSLRILHLSGCTRL------------ESTPNFIGNPSHFRCGFDI 107
S LY ++ LR + LSG T + E P IG+ S R D+
Sbjct: 754 SDDIDNLYSEKSHSVSLLRPVSLSGLTYMRILSLGYCNLSDELIPEDIGSLSFLR---DL 810
Query: 108 VVPGN 112
+ GN
Sbjct: 811 DLRGN 815
>At5g17680 disease resistance protein RPP1-WsB - like protein
Length = 1294
Score = 60.8 bits (146), Expect = 3e-10
Identities = 34/93 (36%), Positives = 52/93 (55%), Gaps = 2/93 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++DLS KYL+E P+ + LE L+ + C +L++V PSI L L+ L C L
Sbjct: 627 LKKMDLSRCKYLVEVPDLSKATNLEELNLSYCQSLVEVTPSIKNLKGLSCFYLTNCIQLK 686
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
+ +G + L SL + +SGC+ L+ P N
Sbjct: 687 DIPIGII--LKSLETVGMSGCSSLKHFPEISWN 717
Score = 40.4 bits (93), Expect = 4e-04
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 1/87 (1%)
Query: 6 KRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVS 65
+RL LS++K + L +LD + C L + +G L L L+L+GC L +
Sbjct: 719 RRLYLSSTKIEELPSSISRLSCLVKLDMSDCQRLRTLPSYLGHLVSLKSLNLDGCRRLEN 778
Query: 66 LYLGTVCELNSLRILHLSGCTRLESTP 92
L T+ L SL L +SGC + P
Sbjct: 779 L-PDTLQNLTSLETLEVSGCLNVNEFP 804
Score = 36.6 bits (83), Expect = 0.005
Identities = 41/153 (26%), Positives = 58/153 (37%), Gaps = 23/153 (15%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L LDLS + + + RL RL+ C L Q P L L ++ + C+SLV
Sbjct: 980 LLELDLSGNNFEFIPASIKRLTRLNRLNLNNCQRL-QALPD-ELPRGLLYIYIHSCTSLV 1037
Query: 65 SLY--LGTVCELNSLRILHLSGCTRLESTPNFI---------GNPSHFRCGFDIVVPGNK 113
S+ C LR L S C +L+ + P H PG+
Sbjct: 1038 SISGCFNQYC----LRKLVASNCYKLDQAAQILIHRNLKLESAKPEHS------YFPGSD 1087
Query: 114 IPYWFDSQFKGGSRIREVDHVYEQDHWLGFAFC 146
IP F+ Q G S ++ LGF+ C
Sbjct: 1088 IPTCFNHQVMGPSLNIQLPQSESSSDILGFSAC 1120
Score = 33.9 bits (76), Expect = 0.033
Identities = 32/109 (29%), Positives = 48/109 (43%), Gaps = 22/109 (20%)
Query: 5 LKRLDLSNSKYLMETPN-FDGSQRLERLDFTGC--------------------TNLLQVH 43
LK L+L + L P+ LE L+ +GC T++ ++
Sbjct: 765 LKSLNLDGCRRLENLPDTLQNLTSLETLEVSGCLNVNEFPRVSTSIEVLRISETSIEEIP 824
Query: 44 PSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTP 92
I L++L L + L SL + ++ EL SL L LSGC+ LES P
Sbjct: 825 ARICNLSQLRSLDISENKRLASLPV-SISELRSLEKLKLSGCSVLESFP 872
>At5g40060 disease resistance -like protein
Length = 1088
Score = 60.1 bits (144), Expect = 4e-10
Identities = 36/95 (37%), Positives = 54/95 (55%), Gaps = 2/95 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DL SK L E P+ + L+ L+ C++L+++ SI L +L L++EGC++L
Sbjct: 558 LKDMDLEKSKNLKEIPDLSMATNLKTLNLKYCSSLVKISSSIQNLNKLTKLNMEGCTNLE 617
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
+L G L SL L L GC+RL P+ N S
Sbjct: 618 TLPAG--INLKSLHRLDLRGCSRLRMFPDISNNIS 650
Score = 30.0 bits (66), Expect = 0.48
Identities = 20/56 (35%), Positives = 30/56 (52%), Gaps = 2/56 (3%)
Query: 38 NLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTPN 93
+L+++ I L +L LS+ C +L SL G L L LSGC++L S P+
Sbjct: 716 SLVELPCGIQNLKKLMELSIRRCKNLESLPTGA--NFKYLDYLDLSGCSKLRSFPD 769
>At5g46450 disease resistance protein-like
Length = 1123
Score = 59.7 bits (143), Expect = 6e-10
Identities = 32/89 (35%), Positives = 55/89 (60%), Gaps = 2/89 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ +DL S+ L E P+ + L++LD + CT+L+++ +I L +L L +E C +L
Sbjct: 630 LRNMDLRGSENLKEIPDLSLATNLKKLDVSNCTSLVELSSTIQNLNQLEELQMERCENLE 689
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPN 93
+L +G L SL L+L+GC++L S P+
Sbjct: 690 NLPIG--INLESLYCLNLNGCSKLRSFPD 716
Score = 31.2 bits (69), Expect = 0.22
Identities = 27/105 (25%), Positives = 44/105 (41%), Gaps = 23/105 (21%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGL---------------- 48
L+ L+++ L P + LE+LDF+GC+ L+ P I
Sbjct: 798 LEHLNIARCTNLETLPTGVNLELLEQLDFSGCSR-LRSFPDISTNIFSLVLDGTGIEEVP 856
Query: 49 -----LTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRL 88
L+FLS+ GC++L + L + +L L + S C L
Sbjct: 857 WWIEDFYRLSFLSMIGCNNLQGVSL-NISKLEKLETVDFSDCEAL 900
>At4g12010 like disease resistance protein (TMV N-like)
Length = 1219
Score = 59.7 bits (143), Expect = 6e-10
Identities = 37/92 (40%), Positives = 54/92 (58%), Gaps = 2/92 (2%)
Query: 1 DVPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGC 60
DV LK +DLS+S L + + LERL+ GCT+L ++ +I L +L +L+L C
Sbjct: 641 DVGMLKWVDLSHSINLRQCLGLANAHNLERLNLEGCTSLKKLPSTINCLEKLIYLNLRDC 700
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTP 92
+SL SL G + SL+ L LSGC+ L+ P
Sbjct: 701 TSLRSLPKG--IKTQSLQTLILSGCSSLKKFP 730
>At2g16870 putative disease resistance protein
Length = 1109
Score = 59.7 bits (143), Expect = 6e-10
Identities = 35/93 (37%), Positives = 54/93 (57%), Gaps = 2/93 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++DLS S +L E P+ + LERL+ C L+++ SIG L +L L + C SL
Sbjct: 625 LKKMDLSRSVHLKELPDLSNATNLERLELCDCRALVELPKSIGNLHKLENLVMANCISLE 684
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
+ T L SL + ++GC+RL++ P+F N
Sbjct: 685 --VIPTHINLASLEHITMTGCSRLKTFPDFSTN 715
>At5g46520 disease resistance protein-like
Length = 1298
Score = 59.3 bits (142), Expect = 7e-10
Identities = 36/96 (37%), Positives = 54/96 (55%), Gaps = 2/96 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK LD+ SKYL E P+ + +E+LDF C +L+++ SI L +L L++E C L
Sbjct: 670 LKELDMWASKYLKEIPDLSKATNIEKLDFGHCWSLVELPSSIRNLNKLLELNMEYCGELE 729
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSH 100
+L G L SL L+ + C +L + P F N S+
Sbjct: 730 TLPTG--FNLKSLDYLNFNECWKLRTFPEFATNISN 763
Score = 33.5 bits (75), Expect = 0.043
Identities = 25/62 (40%), Positives = 31/62 (49%), Gaps = 2/62 (3%)
Query: 3 PFLKRLDLSNSKYLME-TPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
P L L+L N L+E + +F LERLD C NL + I L L L+L GCS
Sbjct: 810 PTLTLLELWNIPNLVELSSSFQNLNNLERLDICYCRNLESLPTGIN-LESLVSLNLFGCS 868
Query: 62 SL 63
L
Sbjct: 869 RL 870
>At5g46510 disease resistance protein-like
Length = 1353
Score = 59.3 bits (142), Expect = 7e-10
Identities = 36/96 (37%), Positives = 54/96 (55%), Gaps = 2/96 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK LD+ SKYL E P+ + +E+LDF C +L+++ SI L +L L++E C L
Sbjct: 631 LKELDMWASKYLKEIPDLSKATNIEKLDFGHCWSLVELPSSIRNLNKLLELNMEYCGELE 690
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSH 100
+L G L SL L+ + C +L + P F N S+
Sbjct: 691 TLPTG--FNLKSLDYLNFNECWKLRTFPEFATNISN 724
Score = 33.5 bits (75), Expect = 0.043
Identities = 25/62 (40%), Positives = 31/62 (49%), Gaps = 2/62 (3%)
Query: 3 PFLKRLDLSNSKYLME-TPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
P L L+L N L+E + +F LERLD C NL + I L L L+L GCS
Sbjct: 771 PTLTLLELWNIPNLVELSSSFQNLNNLERLDICYCRNLESLPTGIN-LESLVSLNLFGCS 829
Query: 62 SL 63
L
Sbjct: 830 RL 831
>At5g40920 disease resistance protein-like
Length = 876
Score = 59.3 bits (142), Expect = 7e-10
Identities = 34/93 (36%), Positives = 51/93 (54%), Gaps = 2/93 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK+++L S L E PN + LE L TGC +L+++ SI L +L L GCS L
Sbjct: 407 LKKINLEYSSNLKEIPNLSKATNLETLRLTGCESLMEIPSSISNLHKLEVLDASGCSKL- 465
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
+ T L+SL+++ + C+RL S P+ N
Sbjct: 466 -HVIPTKINLSSLKMVGMDDCSRLRSFPDISTN 497
Score = 27.7 bits (60), Expect = 2.4
Identities = 32/122 (26%), Positives = 48/122 (39%), Gaps = 14/122 (11%)
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVH---PSIGLLTELAFLSLEGCSSLV 64
LDLS+S M G L+ L C L+ + PS+ + +SLE S+
Sbjct: 542 LDLSHSDIKMIPDYVIGLPHLQHLTIGNCRKLVSIEGHSPSLESIVAYRCISLE---SMC 598
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKG 124
+ + +L L L ES I + H I + GN++P F Q +G
Sbjct: 599 CSFHRPILKLEFYNCLKLDN----ESKRRIILHSGHRI----IFLTGNEVPAQFTHQTRG 650
Query: 125 GS 126
S
Sbjct: 651 NS 652
>At4g16860 disease resistance RPP5 like protein
Length = 1256
Score = 58.9 bits (141), Expect = 1e-09
Identities = 36/88 (40%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DLS S+ L E P+ + L+RL GC +L+ + +IG L L L ++ C+ L
Sbjct: 917 LKRMDLSESENLTEIPDLSKATNLKRLYLNGCKSLVTLPSTIGNLHRLVRLEMKECTGLE 976
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTP 92
L T L+SL IL LSGC+ L + P
Sbjct: 977 --LLPTDVNLSSLIILDLSGCSSLRTFP 1002
Score = 53.1 bits (126), Expect = 5e-08
Identities = 36/97 (37%), Positives = 48/97 (49%), Gaps = 2/97 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK + L SKYL E P+ + LERL GC +L+ + SI T+L L + C L
Sbjct: 757 LKEMYLHGSKYLKEIPDLSLAINLERLYLFGCESLVTLPSSIQNATKLINLDMRDCKKLE 816
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHF 101
S T L SL L+L+GC L + P S+F
Sbjct: 817 S--FPTDLNLESLEYLNLTGCPNLRNFPAIKMGCSYF 851
Score = 33.5 bits (75), Expect = 0.043
Identities = 31/128 (24%), Positives = 54/128 (41%), Gaps = 4/128 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++DL S L E P+ + LE L+ + C +L+ + SI +L L G +
Sbjct: 620 LKKMDLGCSNNLKEIPDLSLAINLEELNLSKCESLVTLPSSIQNAIKLRTLYCSGVLLID 679
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKG 124
L +C L L + + +E T I P + + P ++P F +++
Sbjct: 680 LKSLEGMCNLEYLSV----DWSSMEGTQGLIYLPRKLKRLWWDYCPVKRLPSNFKAEYLV 735
Query: 125 GSRIREVD 132
R+ D
Sbjct: 736 ELRMENSD 743
Score = 29.3 bits (64), Expect = 0.82
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 7/69 (10%)
Query: 31 LDFTGCTN--LLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRL 88
LD +GC + L + S+G L + E + + L T +L+ L+L+GC L
Sbjct: 897 LDVSGCKHEKLWEGIQSLGSLKRMDLSESENLTEIPDLSKAT-----NLKRLYLNGCKSL 951
Query: 89 ESTPNFIGN 97
+ P+ IGN
Sbjct: 952 VTLPSTIGN 960
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.143 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,606,037
Number of Sequences: 26719
Number of extensions: 146045
Number of successful extensions: 1130
Number of sequences better than 10.0: 171
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 491
Number of HSP's gapped (non-prelim): 448
length of query: 149
length of database: 11,318,596
effective HSP length: 90
effective length of query: 59
effective length of database: 8,913,886
effective search space: 525919274
effective search space used: 525919274
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0388b.3


