

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.1
(949 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
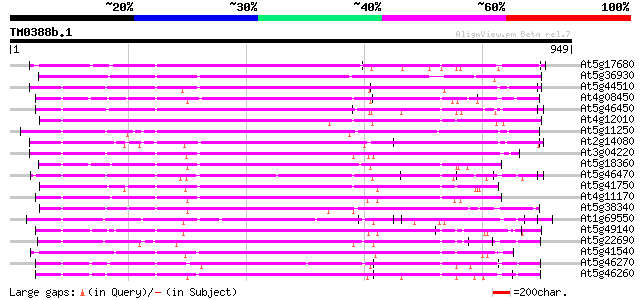
Sequences producing significant alignments: (bits) Value
At5g17680 disease resistance protein RPP1-WsB - like protein 550 e-156
At5g36930 disease resistance like protein 503 e-142
At5g44510 disease resistance protein-like 441 e-124
At4g08450 putative protein 440 e-123
At5g46450 disease resistance protein-like 439 e-123
At4g12010 like disease resistance protein (TMV N-like) 438 e-123
At5g11250 RPP1 disease resistance protein - like 429 e-120
At2g14080 putative disease resistance protein 427 e-119
At3g04220 putative disease resistance protein 425 e-119
At5g18360 disease resistance protein -like 423 e-118
At5g46470 disease resistance protein-like 418 e-117
At5g41750 disease resistance protein-like 417 e-116
At4g11170 RPP1-WsA-like disease resistance protein 410 e-114
At5g38340 disease resistance - like protein 407 e-113
At1g69550 putative disease resistance protein 406 e-113
At5g49140 disease resistance protein-like 402 e-112
At5g22690 disease resistance protein-like 402 e-112
At5g41540 disease resistance protein-like 399 e-111
At5g46270 disease resistance protein-like 399 e-111
At5g46260 disease resistance protein-like 397 e-110
>At5g17680 disease resistance protein RPP1-WsB - like protein
Length = 1294
Score = 550 bits (1416), Expect = e-156
Identities = 333/905 (36%), Positives = 505/905 (55%), Gaps = 63/905 (6%)
Query: 34 STASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGE 93
S++SSS+TV +K DVF+SFRG D R TFV HL+ R GI F+DD LQ+G+
Sbjct: 7 SSSSSSSTV-------WKTDVFVSFRGEDVRKTFVSHLFCEFDRMGIKAFRDDLDLQRGK 59
Query: 94 SISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPV 153
SIS +L+ AI+ SR +IVV S+NYA S WCLDE+ I EC +D T+ P+FY+VDPS V
Sbjct: 60 SISPELIDAIKGSRFAIVVVSRNYAASSWCLDELLKIMECNKD---TIVPIFYEVDPSDV 116
Query: 154 RNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENIVEAVI 213
R Q G + H D ++V +WK A++ LA +G D RN + + I+ IV+ +
Sbjct: 117 RRQRGSFGEDVESHS-----DKEKVGKWKEALKKLAAISGEDSRNWDDSKLIKKIVKDIS 171
Query: 214 EALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDR 273
+ L + LIG+ ++ L++++ + + D +++GIWGMGG+GKTT+A LY++
Sbjct: 172 DKLVSTSWDDSKGLIGMSSHMDFLQSMISIVDK--DVRMLGIWGMGGVGKTTIAKYLYNQ 229
Query: 274 ISHLFEARCFVENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLK 333
+S F+ CF+ENV +V GV +Q + L + E + E +S I+++R R
Sbjct: 230 LSGQFQVHCFMENVKEVCNRYGVRRLQVEFLCRMFQERDKEAWSSVSCCNIIKERFRHKM 289
Query: 334 VLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDAR 393
V +VLD+VD+ QL E G F GSR+I+TTRD H+L +G ++VY+V + +A
Sbjct: 290 VFIVLDDVDRSEQLNELVKETGWFGPGSRIIVTTRDRHLLLSHGINLVYKVKCLPKKEAL 349
Query: 394 ELFYRKGFKSDN-LSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKN 452
+LF F+ + L EL + + YA GLPLA+RV GSFL R+ ++W L RLK
Sbjct: 350 QLFCNYAFREEIILPHGFEELSVQAVNYASGLPLALRVLGSFLYRRSQIEWESTLARLKT 409
Query: 453 NPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIE 512
P + +M+VL++S++GL ++K IFL+I+CF+ ++ +YV+++LD CG IGI + E
Sbjct: 410 YPHSDIMEVLRVSYDGLDEQEKAIFLYISCFYNMKQVDYVRKLLDLCGYAAEIGITILTE 469
Query: 513 RSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKA 572
+SLI N + +H++++ +G+++VRQQ P LW + H+L GT V+
Sbjct: 470 KSLIVESNGCVKIHDLLEQMGRELVRQQAVNNPAQRLLLWDPEDICHLLSENSGTQLVEG 529
Query: 573 IVLDQNEDISEYPQLRA-EGLSIMRGLIILILHHQNFSG--------SLHFLSNNLQYLL 623
I L+ +E + RA EGLS L +L + +F G L +L L+YL
Sbjct: 530 ISLNLSEISEVFASDRAFEGLS---NLKLLNFYDLSFDGETRVHLPNGLSYLPRKLRYLR 586
Query: 624 WHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEG 683
W GYP ++PS F P LVEL M S++++LW+G + L LK+MDLS KYL E P+
Sbjct: 587 WDGYPLKTMPSRFFPEFLVELCMSNSNLEKLWDGIQPLRNLKKMDLSRCKYLVEVPDLSK 646
Query: 684 SRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSG 743
+ LE L+L+ C +L++V PSI L L+ +C L + +G +L SL + +SG
Sbjct: 647 ATNLEELNLSYCQSLVEVTPSIKNLKGLSCFYLTNCIQLKDIPIG--IILKSLETVGMSG 704
Query: 744 CTKLESTPNF-----------TGVENLE----------YLDIDQCVSLSTVDQSIGVLTR 782
C+ L+ P T +E L LD+ C L T+ +G L
Sbjct: 705 CSSLKHFPEISWNTRRLYLSSTKIEELPSSISRLSCLVKLDMSDCQRLRTLPSYLGHLVS 764
Query: 783 LEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLG 842
L+ L+L C L N+P ++ N+ SL TL+ GCL + P S+ L +
Sbjct: 765 LKSLNLDGCRRLENLPDTLQNLTSLETLEVSGCLNVNEFPR--------VSTSIEVLRIS 816
Query: 843 FCSLSEVPHALGEIECLERLNL-EGNNFVSLPSSLRWLSSLAYLNLAHCSKLEFLSELQL 901
S+ E+P + + L L++ E SLP S+ L SL L L+ CS LE L++
Sbjct: 817 ETSIEEIPARICNLSQLRSLDISENKRLASLPVSISELRSLEKLKLSGCSVLESF-PLEI 875
Query: 902 CDIAS 906
C S
Sbjct: 876 CQTMS 880
Score = 79.0 bits (193), Expect = 1e-14
Identities = 102/342 (29%), Positives = 156/342 (44%), Gaps = 53/342 (15%)
Query: 597 GLIILILHHQNFSGSL---HF--LSNNLQYLLWHGYPFASLPSNFEPFR-LVELNMPYSS 650
G+I+ L SG HF +S N + L LPS+ LV+L+M S
Sbjct: 691 GIILKSLETVGMSGCSSLKHFPEISWNTRRLYLSSTKIEELPSSISRLSCLVKLDM--SD 748
Query: 651 IQRLWEGRKDLPF-------LKRMDLSNSKYLTETPN-FEGSRRLERLDLTGCTNLLQ-- 700
QRL + LP LK ++L + L P+ + LE L+++GC N+ +
Sbjct: 749 CQRL----RTLPSYLGHLVSLKSLNLDGCRRLENLPDTLQNLTSLETLEVSGCLNVNEFP 804
Query: 701 -VHPSIGLL----TKLAFLSFESC--SSLVSLDLG----------SLCVLYSLAVLHLSG 743
V SI +L T + + C S L SLD+ S+ L SL L LSG
Sbjct: 805 RVSTSIEVLRISETSIEEIPARICNLSQLRSLDISENKRLASLPVSISELRSLEKLKLSG 864
Query: 744 CTKLESTPN--FTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSV 801
C+ LES P + L + D+D+ S+ + ++IG L LE L + + P S+
Sbjct: 865 CSVLESFPLEICQTMSCLRWFDLDR-TSIKELPENIGNLVALEVLQASRTV-IRRAPWSI 922
Query: 802 NNMESLLTLDFCGCLKLKHLPLGL-----PSLSPFTLQSLIFLDLGFCSLSEVPHALGEI 856
+ L L P GL P LS F L L L +++E+P+++G +
Sbjct: 923 ARLTRLQVLAIGNSF---FTPEGLLHSLCPPLSRF--DDLRALSLSNMNMTEIPNSIGNL 977
Query: 857 ECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCSKLEFLSE 898
L L+L GNNF +P+S++ L+ L LNL +C +L+ L +
Sbjct: 978 WNLLELDLSGNNFEFIPASIKRLTRLNRLNLNNCQRLQALPD 1019
>At5g36930 disease resistance like protein
Length = 1164
Score = 503 bits (1294), Expect = e-142
Identities = 307/871 (35%), Positives = 475/871 (54%), Gaps = 54/871 (6%)
Query: 49 RYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRV 108
R+ YDVF+SFRG+D R F+ HLY L R GI F DD +LQ+GE IS +LL AI S++
Sbjct: 11 RWTYDVFVSFRGADVRKNFLSHLYDSLRRCGISTFMDDVELQRGEYISPELLNAIETSKI 70
Query: 109 SIVVFSKNYAESRWCLDEMAAIAECCEDF-KQTVFPVFYDVDPSPVRNQNGVYENAFVFH 167
IVV +K+YA S WCLDE+ I + ++ VFP+F VDPS +R Q G Y +F H
Sbjct: 71 LIVVLTKDYASSAWCLDELVHIMKSHKNNPSHMVFPIFLYVDPSDIRWQQGSYAKSFSKH 130
Query: 168 MLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENIVEAVIEALGRKFSGFADDL 227
+ H +++ W+ A+ +A +GWD++N+ E I +I +++ L ++
Sbjct: 131 --KNSHPLNKLKDWREALTKVANISGWDIKNRNEAECIADITREILKRLPCQYLHVPSYA 188
Query: 228 IGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENV 287
+G++ R++ + +LL + S+ +VI I+GMGGIGKTTLA V ++ SHLFE F+EN
Sbjct: 189 VGLRSRLQHISSLLSIGSD--GVRVIVIYGMGGIGKTTLAKVAFNEFSHLFEGSSFLENF 246
Query: 288 SKVYRDG-GVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLVQ 346
+ + G T +Q Q+L + ++E + V++R RS +VL+V+D+VD + Q
Sbjct: 247 REYSKKPEGRTHLQHQLLSDILRRNDIEFKG---LDHAVKERFRSKRVLLVVDDVDDVHQ 303
Query: 347 LQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDNL 406
L A++ F GSR+IITTR+ H+LK A Y ++ +++ ELF F++
Sbjct: 304 LNSAAIDRDCFGHGSRIIITTRNMHLLKQLRAEGSYSPKELDGDESLELFSWHAFRTSEP 363
Query: 407 SSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISF 466
+ EV+ Y GLPLA+ V G+FL R+ +W L LK P++ + LQISF
Sbjct: 364 PKEFLQHSEEVVTYCAGLPLAVEVLGAFLIERSIREWESTLKLLKRIPNDNIQAKLQISF 423
Query: 467 EGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLITIRNQEIHMH 526
L E K++FL IACFF G YV ILD C L+P I + ++ER LITI I MH
Sbjct: 424 NALTIEQKDVFLDIACFFIGVDSYYVACILDGCNLYPDIVLSLLMERCLITISGNNIMMH 483
Query: 527 EMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLDQNEDISEYPQ 586
++++D+G++IVR+ P++ G SRLW + VL + GTN ++ + L D+ ++
Sbjct: 484 DLLRDMGRQIVREISPKKCGERSRLWSHNDVVGVLKKKSGTNAIEGLSL--KADVMDFQY 541
Query: 587 LRAEGLSIMRGLIILILHHQNFSGSLHFLSNNLQYLLWHGYPFASLPSNFEPFRLVELNM 646
E + M+ L +L L + + +GS +L++L WHG+ P N L L++
Sbjct: 542 FEVEAFAKMQELRLLELRYVDLNGSYEHFPKDLRWLCWHGFSLECFPINLSLESLAALDL 601
Query: 647 PYSSIQRLWEGR---KDLPFLKRMDLSNSKYLTETPNFEGSRRLERLDLTGCTNLLQVHP 703
YS+++R W+ + + +K +DLS+S YL ETP+F +E+L L C +L+ VH
Sbjct: 602 QYSNLKRFWKAQSPPQPANMVKYLDLSHSVYLRETPDFSYFPNVEKLILINCKSLVLVHK 661
Query: 704 SIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTP-NFTGVENLEYL 762
SIG+L K L +L+LS C +L+ P +++LE L
Sbjct: 662 SIGILDK------------------------KLVLLNLSSCIELDVLPEEIYKLKSLESL 697
Query: 763 DIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKL---- 818
+ C L +D ++G L L L L D L IP ++N ++ L L GC L
Sbjct: 698 FLSNCSKLERLDDALGELESLTTL-LADFTALREIPSTINQLKKLKRLSLNGCKGLLSDD 756
Query: 819 -----KHLPLGLPSLSPFTLQSLIF---LDLGFCSLSE--VPHALGEIECLERLNLEGNN 868
+ L P +L L + L LG+C+LS+ +P +G + L L+L GN+
Sbjct: 757 IDNLYSEKSHSVSLLRPVSLSGLTYMRILSLGYCNLSDELIPEDIGSLSFLRDLDLRGNS 816
Query: 869 FVSLPSSLRWLSSLAYLNLAHCSKLEFLSEL 899
F +LP+ L +L L L+ CSKL+ + L
Sbjct: 817 FCNLPTDFATLPNLGELLLSDCSKLQSILSL 847
Score = 33.9 bits (76), Expect = 0.45
Identities = 33/118 (27%), Positives = 49/118 (40%), Gaps = 7/118 (5%)
Query: 687 LERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTK 746
L LDL G + + L L L CS L S+ L + SL L + C
Sbjct: 807 LRDLDLRG-NSFCNLPTDFATLPNLGELLLSDCSKLQSI----LSLPRSLLFLDVGKCIM 861
Query: 747 LESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNM 804
L+ TP+ + L L ++ C+SL + I L F+ L C L + ++N M
Sbjct: 862 LKRTPDISKCSALFKLQLNDCISLFEI-PGIHNHEYLSFIVLDGC-KLASTDTTINTM 917
>At5g44510 disease resistance protein-like
Length = 1187
Score = 441 bits (1134), Expect = e-124
Identities = 304/908 (33%), Positives = 470/908 (51%), Gaps = 57/908 (6%)
Query: 34 STASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGE 93
S++ SS++ S+ ++ + + VF+SFRG D R + H+ R GI F D++ +++G
Sbjct: 22 SSSLSSSSPPSSLSQNWLHPVFLSFRGEDVRKGLLSHIQKEFQRNGITPFIDNE-MKRGG 80
Query: 94 SISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPV 153
SI +LLQAIR S+++I++ S+NY S+WCLDE+ I +C E+ QTV VFYDVDPS V
Sbjct: 81 SIGPELLQAIRGSKIAIILLSRNYGSSKWCLDELVEIMKCREELGQTVMTVFYDVDPSDV 140
Query: 154 RNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAV 212
R Q G + VF + V RWK+A+ S A G D RN + E I I + V
Sbjct: 141 RKQKGDFGK--VFKKTCVGRPEEMVQRWKQALTSAANILGEDSRNWENEADMIIKISKDV 198
Query: 213 IEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYD 272
+ L S D+ +GI+ + +LL+L+ E + ++IGIWG GIGKTT++ VLY+
Sbjct: 199 SDVLSFTPSKDFDEFVGIEAHTTEITSLLQLDLE--EVRMIGIWGPAGIGKTTISRVLYN 256
Query: 273 RISHLFEARCFVENVSKVY------RDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVR 326
++ H F+ ++N+ Y +QK++L Q +++ ++ G+ +
Sbjct: 257 KLFHQFQLGAIIDNIKVRYPRPCHDEYSAKLQLQKELLSQMINQKDMVVPH----LGVAQ 312
Query: 327 DRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPL 386
+RL+ KVL+VLD+VD LVQL A + F GSR+I+ T+D +LK +G +Y+V
Sbjct: 313 ERLKDKKVLLVLDDVDGLVQLDAMAKDVQWFGLGSRIIVVTQDLKLLKAHGIKYIYKVDF 372
Query: 387 MNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDA 446
+++A E+F F + ++ V A LPL +RV GS+L + +W +
Sbjct: 373 PTSDEALEIFCMYAFGEKSPKVGFEQIARTVTTLAGKLPLGLRVMGSYLRRMSKQEWAKS 432
Query: 447 LDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIG 506
+ RL+ + D+ + VL+ S+ L ++K++FLHI CFF+ E+ ++ L + G
Sbjct: 433 IPRLRTSLDDDIESVLKFSYNSLAEQEKDLFLHITCFFRRERIETLEVFLAKKSVDMRQG 492
Query: 507 IQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMG 566
+Q + ++SL+++ I MH ++ LG IVR+Q +PG L + VL + G
Sbjct: 493 LQILADKSLLSLNLGNIEMHNLLVQLGLDIVRKQSIHKPGKRQFLVDTEDICEVLTDDTG 552
Query: 567 TNKVKAIVLDQNEDISEYPQLRAEGLSIMRGLIILILHHQ---------NFSGSLHFLSN 617
T + I L+ + I + M L L HH L +S
Sbjct: 553 TRTLIGIDLELSGVIEGVINISERAFERMCNLQFLRFHHPYGDRCHDILYLPQGLSHISR 612
Query: 618 NLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTE 677
L+ L W YP LP F P LV++NM S +++LW+G + + LK MDLS L E
Sbjct: 613 KLRLLHWERYPLTCLPPKFNPEFLVKINMRDSMLEKLWDGNEPIRNLKWMDLSFCVNLKE 672
Query: 678 TPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSL--DLGSLCVL-- 733
P+F + L+ L L C +L+++ SIG T L L CSSLV L +G+L L
Sbjct: 673 LPDFSTATNLQELRLINCLSLVELPSSIGNATNLLELDLIDCSSLVKLPSSIGNLTNLKK 732
Query: 734 -------------------YSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTV 773
SL L+LSGC+ L P+ G + NL+ + D C SL +
Sbjct: 733 LFLNRCSSLVKLPSSFGNVTSLKELNLSGCSSLLEIPSSIGNIVNLKKVYADGCSSLVQL 792
Query: 774 DQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLP-LGLPSLSPFT 832
SIG T L+ L L +C +L P S+ N+ L L+ GCL L LP +G +
Sbjct: 793 PSSIGNNTNLKELHLLNCSSLMECPSSMLNLTRLEDLNLSGCLSLVKLPSIG----NVIN 848
Query: 833 LQSLIFLDLGFCSLSEVPHALGEIECLERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCS 891
LQSL D SL E+P + L+ L L+G +N + LPSS+ +++L L L CS
Sbjct: 849 LQSLYLSDCS--SLMELPFTIENATNLDTLYLDGCSNLLELPSSIWNITNLQSLYLNGCS 906
Query: 892 KLEFLSEL 899
L+ L L
Sbjct: 907 SLKELPSL 914
Score = 79.7 bits (195), Expect = 7e-15
Identities = 81/289 (28%), Positives = 132/289 (45%), Gaps = 13/289 (4%)
Query: 611 SLHFLSNNLQYLLWHGYPFASLPSNF-EPFRLVELNMP-YSSIQRLWEGRKDLPFLKRMD 668
S+ L+N + L LPS+F L ELN+ SS+ + ++ LK++
Sbjct: 723 SIGNLTNLKKLFLNRCSSLVKLPSSFGNVTSLKELNLSGCSSLLEIPSSIGNIVNLKKVY 782
Query: 669 LSNSKYLTETPNFEGSR-RLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDL 727
L + P+ G+ L+ L L C++L++ S+ LT+L L+ C SLV L
Sbjct: 783 ADGCSSLVQLPSSIGNNTNLKELHLLNCSSLMECPSSMLNLTRLEDLNLSGCLSLVKLP- 841
Query: 728 GSLCVLYSLAVLHLSGCTKLESTP-NFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFL 786
S+ + +L L+LS C+ L P NL+ L +D C +L + SI +T L+ L
Sbjct: 842 -SIGNVINLQSLYLSDCSSLMELPFTIENATNLDTLYLDGCSNLLELPSSIWNITNLQSL 900
Query: 787 SLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSL 846
L C +L +P V N +L +L C L LP + + + +L +LD+ CS
Sbjct: 901 YLNGCSSLKELPSLVENAINLQSLSLMKCSSLVELPSSI-----WRISNLSYLDVSNCSS 955
Query: 847 SEVPHALGEIECLERLNLEGNNFVSLPSSLR--WLSSLAYLNLAHCSKL 893
+ + + L L+ + SL L + + LN A+C KL
Sbjct: 956 LLELNLVSHPVVPDSLILDAGDCESLVQRLDCFFQNPKIVLNFANCFKL 1004
>At4g08450 putative protein
Length = 1234
Score = 440 bits (1131), Expect = e-123
Identities = 309/923 (33%), Positives = 487/923 (52%), Gaps = 97/923 (10%)
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ + + YDVF SF G D R TF+ H L RK I FKD++ +++G SI +L+QAI
Sbjct: 3 SSSSHNWVYDVFTSFSGEDIRVTFLTHFLKELDRKMIIAFKDNE-IERGNSIGTELIQAI 61
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENA 163
++SR+++VVFSK Y+ S WCL+E+ I C K+ V PVFYD+DPS VR Q G + +
Sbjct: 62 KDSRIAVVVFSKKYSSSSWCLNELVEIVNC----KEIVIPVFYDLDPSDVRKQEGEFGES 117
Query: 164 FVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKP--EFREIENIVEAVIEALGRKF- 220
F + + D + + RW +A+ ++A AG+ R KP E + IE I V++ L +
Sbjct: 118 FK-ETCKNRTDYE-IQRWGQALTNVANIAGYHTR-KPNNEAKLIEEITNDVLDKLMKLTP 174
Query: 221 SGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEA 280
S D+ GI+ ++ L LL L SE + +++GIWG GIGKTT+A L++RI F+
Sbjct: 175 SKDFDEFFGIEDHIKELSLLLCLESE--EVRMVGIWGPTGIGKTTIARALFNRIYRHFQG 232
Query: 281 RCFVEN--VSK---VYRDGGVTA------VQKQVLRQTVDEMNLETYSPSEISGIVRDRL 329
R F++ +SK +Y +Q+++L + +D+ NLE V++RL
Sbjct: 233 RVFIDRAFISKSMAIYSRANSDDYNLKLHLQEKLLSKLLDKKNLEINHLDA----VKERL 288
Query: 330 RSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNN 389
R +KVL+ +D++D V L+ A F GSR+I+ T+D+H+L+ YG +YEV L +
Sbjct: 289 RQMKVLIFIDDLDDQVVLEALACQTQWFGHGSRIIVITKDKHLLRAYGIDHIYEVLLPSK 348
Query: 390 NDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDR 449
+ A ++F R F+ D+ + EL +V+K A LPL + + GS+L R+ W D +
Sbjct: 349 DLAIKMFCRSAFRKDSPPNGFIELAYDVVKRAGSLPLGLNILGSYLRGRSKEDWIDMMPG 408
Query: 450 LKNNPDNKVMDVLQISFEGLHSEDKE-IFLHIACFFKGEKENYVKRILDACGLHPHIGIQ 508
L+N D K+ L++S++GL SED + IF HIAC F E + +K++L+ GL+ G+
Sbjct: 409 LRNKLDGKIQKTLRVSYDGLASEDDQAIFRHIACIFNFEACSDIKKLLEDSGLNVTNGLI 468
Query: 509 NMIERSLITI--RNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMG 566
N++++SLI I + + + MH ++Q+ ++I+R Q ++PG L + VL + G
Sbjct: 469 NLVDKSLIRIEPKQKTVEMHCLLQETAREIIRAQSFDDPGKREFLVDGKDIADVLDNCSG 528
Query: 567 TNKVKAIVLDQNEDISEYPQLRAEGLSIMRGLIILILH-HQNFS---------GSLHFLS 616
T KV I LD +E E L+ + M L L L+ + N S ++L
Sbjct: 529 TRKVLGISLDMDE--IEELHLQVDAFKKMLNLRFLKLYTNTNISEKEDKLLLPKEFNYLP 586
Query: 617 NNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLT 676
N L+ L W +P +PS+F P LV+L MP S +++LW+G L LK M+L S+ L
Sbjct: 587 NTLRLLSWQRFPMRCMPSDFFPKYLVKLLMPGSKLEKLWDGVMPLQCLKNMNLFGSENLK 646
Query: 677 ETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSL 736
E PN + LE L L C +L++V +IG L KL +L+ C +L L SL
Sbjct: 647 EFPNLSLATNLETLSLGFCLSLVEVPSTIGNLNKLTYLNMSGCHNLEKFPAD--VNLKSL 704
Query: 737 AVLHLSGCTKL--------------------ESTPNFTGVENLEYLDIDQCVSLSTVDQS 776
+ L L+GC++L E P+ +ENL YL I S+ D
Sbjct: 705 SDLVLNGCSRLKIFPAISSNISELCLNSLAVEEFPSNLHLENLVYLLIWGMTSVKLWD-G 763
Query: 777 IGVLTRLEFLSLRD-----------------------CLNLTNIPLSVNNMESLLTLDFC 813
+ VLT L+ + LRD C+++ +P S+ N+ +L+ LD
Sbjct: 764 VKVLTSLKTMHLRDSKNLKEIPDLSMASNLLILNLEQCISIVELPSSIRNLHNLIELDMS 823
Query: 814 GCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFVSLP 873
GC L+ P G+ LQSL ++L CS ++ + + L+L +P
Sbjct: 824 GCTNLETFPTGI------NLQSLKRINLARCSRLKIFPDIS--TNISELDLSQTAIEEVP 875
Query: 874 SSLRWLSSLAYLNLAHCSKLEFL 896
+ S L YL + C+ LE++
Sbjct: 876 LWIENFSKLKYLIMGKCNMLEYV 898
Score = 63.5 bits (153), Expect = 5e-10
Identities = 55/198 (27%), Positives = 90/198 (44%), Gaps = 23/198 (11%)
Query: 615 LSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKY 674
+S+N+ L + PSN LV L + + +LW+G K L LK M L +SK
Sbjct: 721 ISSNISELCLNSLAVEEFPSNLHLENLVYLLIWGMTSVKLWDGVKVLTSLKTMHLRDSKN 780
Query: 675 LTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLY 734
L E P+ + L L+L C +++++ SI L L L C++L + G L
Sbjct: 781 LKEIPDLSMASNLLILNLEQCISIVELPSSIRNLHNLIELDMSGCTNLETFPTG--INLQ 838
Query: 735 SLAVLHLSGCTKLESTPNFTG------------------VEN---LEYLDIDQCVSLSTV 773
SL ++L+ C++L+ P+ + +EN L+YL + +C L V
Sbjct: 839 SLKRINLARCSRLKIFPDISTNISELDLSQTAIEEVPLWIENFSKLKYLIMGKCNMLEYV 898
Query: 774 DQSIGVLTRLEFLSLRDC 791
+I L L+ + DC
Sbjct: 899 FLNISKLKHLKSVDFSDC 916
>At5g46450 disease resistance protein-like
Length = 1123
Score = 439 bits (1130), Expect = e-123
Identities = 297/916 (32%), Positives = 470/916 (50%), Gaps = 86/916 (9%)
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ +R + YDVF SF G D R TF+ H L RK I FKD++ +++ +SI+ +L++AI
Sbjct: 5 SSSSRNWSYDVFPSFSGEDVRKTFLSHFLRELERKSIITFKDNE-MERSQSIAPELVEAI 63
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENA 163
++SR++++VFSKNYA S WCL+E+ I C + Q V PVFY +DPS +R Q+G + A
Sbjct: 64 KDSRIAVIVFSKNYASSSWCLNELLEIMRCNKYLGQQVIPVFYYLDPSHLRKQSGEFGEA 123
Query: 164 FVFHMLRFKHDADRV-DRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALGRKFS 221
F ++ + V ++WK+A+ ++ G+ +N E IE I ++ L S
Sbjct: 124 F---KKTCQNQTEEVKNQWKQALTDVSNILGYHSKNCNSEATMIEEISSHILGKLSLTPS 180
Query: 222 GFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEAR 281
++ +GI+ +E + LL L S+ + +++GIWG GIGKTT+A L+ +S F++
Sbjct: 181 NDFEEFVGIKDHIEKVRLLLHLESD--EVRMVGIWGTSGIGKTTIARALFSNLSSQFQSS 238
Query: 282 CFVEN--VSKVYRDGGVTAVQKQVLRQTVDEMNL-ETYSPSEIS-GIVRDRLRSLKVLVV 337
+++ +SK G ++ + E L E + G + +RL+ KVL++
Sbjct: 239 VYIDRAFISKSMEGYGRANPDDYNMKLRLRENFLFEILGKKNMKIGAMEERLKHQKVLII 298
Query: 338 LDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFY 397
+D++D L F GSR+I+ T+++H L+ +G VYE L + A E+F
Sbjct: 299 IDDLDDQDVLDALVGRTQWFGSGSRIIVVTKNKHFLRAHGIDHVYEACLPSEELALEMFC 358
Query: 398 RKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNK 457
R F+ ++ EL EV A LPL ++V GS+L R+ W D + RL+N+ D K
Sbjct: 359 RYAFRKNSPPDGFMELSSEVALRAGNLPLGLKVLGSYLRGRDIEDWMDMMPRLQNDLDGK 418
Query: 458 VMDVLQISFEGLHS-EDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLI 516
+ L++S++GL++ +D+ IF HIAC F GEK N +K +L L +IG++N++++SLI
Sbjct: 419 IEKTLRVSYDGLNNKKDEAIFRHIACLFNGEKVNDIKLLLAESDLDVNIGLKNLVDKSLI 478
Query: 517 TIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLD 576
+R I MH ++QD+GK+IVR Q EPG L +H + VL GT KV I LD
Sbjct: 479 FVREDTIEMHRLLQDMGKEIVRAQ-SNEPGEREFLVDSKHIYDVLEDNTGTKKVLGIALD 537
Query: 577 QNEDISEYPQLRAEGLSIMRGLIILILHHQ-------NFSGSLHFLSNNLQYLLWHGYPF 629
NE Y + MR L+ L + + + S L L+ L W YP
Sbjct: 538 INETDGLY--IHESAFKGMRNLLFLNFYTKQKKDVTWHLSEGFDHLPPKLRLLSWEKYPL 595
Query: 630 ASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSRRLER 689
+PSNF P LV+L M S +++LW+G L L+ MDL S+ L E P+ + L++
Sbjct: 596 RCMPSNFRPENLVKLQMCESKLEKLWDGVHSLTGLRNMDLRGSENLKEIPDLSLATNLKK 655
Query: 690 LDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLES 749
LD++ CT+L+++ +I L +L L E C +L +L +G L SL L+L+GC+KL S
Sbjct: 656 LDVSNCTSLVELSSTIQNLNQLEELQMERCENLENLPIG--INLESLYCLNLNGCSKLRS 713
Query: 750 TPNFT--------------------GVENLEYLD-------------------------- 763
P+ + +ENL YL
Sbjct: 714 FPDISTTISELYLSETAIEEFPTELHLENLYYLGLYDMKSEKLWKRVQPLTPLMTMLSPS 773
Query: 764 -----IDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKL 818
+ SL + S L LE L++ C NL +P V N+E L LDF GC +L
Sbjct: 774 LTKLFLSDIPSLVELPSSFQNLHNLEHLNIARCTNLETLPTGV-NLELLEQLDFSGCSRL 832
Query: 819 KHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEG-NNFVSLPSSLR 877
+ P +S ++ L L + EVP + + L L++ G NN + ++
Sbjct: 833 R----SFPDIS----TNIFSLVLDGTGIEEVPWWIEDFYRLSFLSMIGCNNLQGVSLNIS 884
Query: 878 WLSSLAYLNLAHCSKL 893
L L ++ + C L
Sbjct: 885 KLEKLETVDFSDCEAL 900
Score = 58.9 bits (141), Expect = 1e-08
Identities = 81/344 (23%), Positives = 140/344 (40%), Gaps = 52/344 (15%)
Query: 581 ISEYPQLRAEGL-SIMRGLIILILHHQNFSG-----SLHFLSNNLQYLLWHGYPFASLPS 634
+ E R E L ++ G+ + L+ N +G S +S + L P+
Sbjct: 677 LEELQMERCENLENLPIGINLESLYCLNLNGCSKLRSFPDISTTISELYLSETAIEEFPT 736
Query: 635 NFEPFRLVELNMPYSSIQRLWEGRKDL--------PFLKRMDLSNSKYLTETPN-FEGSR 685
L L + ++LW+ + L P L ++ LS+ L E P+ F+
Sbjct: 737 ELHLENLYYLGLYDMKSEKLWKRVQPLTPLMTMLSPSLTKLFLSDIPSLVELPSSFQNLH 796
Query: 686 RLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCT 745
LE L++ CTNL E+ + V+L+L L L SGC+
Sbjct: 797 NLEHLNIARCTNL------------------ETLPTGVNLEL--------LEQLDFSGCS 830
Query: 746 KLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNME 805
+L S P+ + N+ L +D + V I RL FLS+ C NL + L+++ +E
Sbjct: 831 RLRSFPDIS--TNIFSLVLDG-TGIEEVPWWIEDFYRLSFLSMIGCNNLQGVSLNISKLE 887
Query: 806 SLLTLDFCGCLKLKH-----LPLGLPSLSPFTLQSLIFLDLGFCSLSEVPH--ALGEIEC 858
L T+DF C L H +P + +++ + S + + + F + + H L +
Sbjct: 888 KLETVDFSDCEALSHANWDTIPSAV-AMATENIHSKLPVCIKFSNCFNLDHKAVLLQQSI 946
Query: 859 LERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCSKLEFLSELQLC 902
++L L G S + +SL + L H S + + C
Sbjct: 947 FKQLILSGGEMFSYFTHRTTGTSLTNIPLLHISPCQPFFRFRAC 990
Score = 36.2 bits (82), Expect = 0.090
Identities = 40/116 (34%), Positives = 55/116 (46%), Gaps = 16/116 (13%)
Query: 801 VNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFC-SLSEVPHALGEIECL 859
V+++ L +D G LK +P LS T +L LD+ C SL E+ + + L
Sbjct: 624 VHSLTGLRNMDLRGSENLKEIP----DLSLAT--NLKKLDVSNCTSLVELSSTIQNLNQL 677
Query: 860 ERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSKLE-------FLSELQLCDIASE 907
E L +E N +LP + L SL LNL CSKL +SEL L + A E
Sbjct: 678 EELQMERCENLENLPIGIN-LESLYCLNLNGCSKLRSFPDISTTISELYLSETAIE 732
>At4g12010 like disease resistance protein (TMV N-like)
Length = 1219
Score = 438 bits (1127), Expect = e-123
Identities = 296/922 (32%), Positives = 474/922 (51%), Gaps = 84/922 (9%)
Query: 51 KYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSI 110
++DVF+SFRG DTRN F HL L +GI F DD+ L++G++++A L I S+++I
Sbjct: 10 EFDVFLSFRGFDTRNNFTGHLQKALRLRGIDSFIDDR-LRRGDNLTA-LFDRIEKSKIAI 67
Query: 111 VVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLR 170
+VFS NYA S WCL E+ I EC +Q V P+FY VD S V Q + F L
Sbjct: 68 IVFSTNYANSAWCLRELVKILECRNSNQQLVVPIFYKVDKSDVEKQRNSFAVPFKLPELT 127
Query: 171 FKH-DADRVDRWKRAMRSLAGSAGWDVR--NKPEFREIENIVEAVIEALGRKFSGFADDL 227
F + + WK A+ S + G+ V+ + E + ++ I + L + L
Sbjct: 128 FPGVTPEEISSWKAALASASNILGYVVKEISTSEAKLVDEIAVDTFKKLNDLAPSGNEGL 187
Query: 228 IGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENV 287
+GI+ R++ LE LL + +IGI GM GIGKTTLA LY R+ F+ CF+ N+
Sbjct: 188 VGIESRLKNLEKLLSWE-DLDTVHIIGIVGMVGIGKTTLADCLYGRMRGQFDGSCFLTNI 246
Query: 288 SKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLVQL 347
+ G+ ++ +++ +++ +LE +P RL+S ++L+VLD+V+ Q+
Sbjct: 247 RENSGRSGLESLLQKLFSTVLNDRDLEIGAPGNAHERFERRLKSKRLLIVLDDVNDEKQI 306
Query: 348 QEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDNLS 407
+ + +Q GSR+IITTRD +++ Y +P +N+ +A +LF F +
Sbjct: 307 RYLMGHCKWYQGGSRIIITTRDSKLIETIKGR-KYVLPKLNDREALKLFSLNAFSNSFPL 365
Query: 408 SRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISFE 467
L VL YA+G PLA++V GS LC R+ + W LDRLK+ + +VL+ S+E
Sbjct: 366 KEFEGLTNMVLDYAKGHPLALKVLGSDLCERDDLYWEAKLDRLKSRSHGDIYEVLETSYE 425
Query: 468 GLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLITIRNQEIHMHE 527
L +E K +FL IACFF+ E +YV +L++ G+ +++++++ LIT+ + I MH+
Sbjct: 426 ELTTEQKNVFLDIACFFRSENVDYVTSLLNSHGVDVSGVVKDLVDKCLITLSDNRIEMHD 485
Query: 528 MVQDLGKKIVRQ---------QFPEEPGS---WS-RLWLYQHFHHVLMSEMGTNKVKAIV 574
M+Q + K+I + ++ G+ W RLW + +L +GT+K++ I
Sbjct: 486 MLQTMAKEISLKVETIGIRDCRWLSRHGNQCQWHIRLWDSEDICDLLTEGLGTDKIRGIF 545
Query: 575 LDQNEDISEYPQLRAEGLSIMRGLIILILHHQNFSGS------------LHFLSNNLQYL 622
LD ++ +L A+ M L L ++ + S L FL N L YL
Sbjct: 546 LDTSK--LRAMRLSAKAFQGMYNLKYLKIYDSHCSRGCEAEFKLHLRRGLSFLPNELTYL 603
Query: 623 LWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFE 682
WHGYP S+P +F+P LV+L +P+S ++ +W+ KD+ LK +DLS+S L +
Sbjct: 604 HWHGYPLQSIPLDFDPKNLVDLKLPHSQLEEIWDDEKDVGMLKWVDLSHSINLRQCLGLA 663
Query: 683 GSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLS 742
+ LERL+L GCT+L ++ +I L KL +L+ C+SL SL G SL L LS
Sbjct: 664 NAHNLERLNLEGCTSLKKLPSTINCLEKLIYLNLRDCTSLRSLPKG--IKTQSLQTLILS 721
Query: 743 GCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVN 802
GC+ L+ P + EN+E L +D V + ++ +SI RL L+L++C L ++ +
Sbjct: 722 GCSSLKKFPLIS--ENVEVLLLDGTV-IKSLPESIQTFRRLALLNLKNCKKLKHLSSDLY 778
Query: 803 NMESLLTLDFCGCLKLKHLP-------------------------LGLPSLSPFTL---- 833
++ L L GC +L+ P + L ++ F+L
Sbjct: 779 KLKCLQELILSGCSQLEVFPEIKEDMESLEILLMDDTSITEMPKMMHLSNIKTFSLCGTS 838
Query: 834 ----------------QSLIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFVSLPSSLR 877
L L L CSL ++P +G + L+ L L GNN +LP S
Sbjct: 839 SHVSVSMFFMPPTLGCSRLTDLYLSRCSLYKLPDNIGGLSSLQSLCLSGNNIENLPESFN 898
Query: 878 WLSSLAYLNLAHCSKLEFLSEL 899
L++L + +L C L+ L L
Sbjct: 899 QLNNLKWFDLKFCKMLKSLPVL 920
Score = 35.0 bits (79), Expect = 0.20
Identities = 53/219 (24%), Positives = 85/219 (38%), Gaps = 56/219 (25%)
Query: 615 LSNNLQYLLWHGYPFASLPSNFEPFR-LVELNMPY------------------------- 648
+S N++ LL G SLP + + FR L LN+
Sbjct: 732 ISENVEVLLLDGTVIKSLPESIQTFRRLALLNLKNCKKLKHLSSDLYKLKCLQELILSGC 791
Query: 649 SSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSRRLERLDLTGCTNLLQVH-----P 703
S ++ E ++D+ L+ + L + +TE P ++ L G ++ + V P
Sbjct: 792 SQLEVFPEIKEDMESLEIL-LMDDTSITEMPKMMHLSNIKTFSLCGTSSHVSVSMFFMPP 850
Query: 704 SIGLLTKLAFLSFESCSSLVSLD-------LGSLCV--------------LYSLAVLHLS 742
++G ++L L CS D L SLC+ L +L L
Sbjct: 851 TLGC-SRLTDLYLSRCSLYKLPDNIGGLSSLQSLCLSGNNIENLPESFNQLNNLKWFDLK 909
Query: 743 GCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLT 781
C L+S P +NL+YLD +C SL T+ + LT
Sbjct: 910 FCKMLKSLPVLP--QNLQYLDAHECESLETLANPLTPLT 946
>At5g11250 RPP1 disease resistance protein - like
Length = 1189
Score = 429 bits (1103), Expect = e-120
Identities = 306/899 (34%), Positives = 466/899 (51%), Gaps = 41/899 (4%)
Query: 18 SRRHSSHHLGPLAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVR 77
++ S L P + S + SS S+ + + VF SF G D R F+ H+ R
Sbjct: 29 NKEKKSTSLSPSSSPSSLSPSSVPPPSSSCI-WTHQVFPSFSGEDVRRDFLSHIQMEFQR 87
Query: 78 KGIFVFKDDKKLQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDF 137
GI F D++ +++GESI +LL+AIR S+++I++ S+NYA S+WCLDE+ I +C E++
Sbjct: 88 MGITPFVDNE-IKRGESIGPELLRAIRGSKIAIILLSRNYASSKWCLDELVEIMKCREEY 146
Query: 138 KQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVR 197
QTV +FY VDPS V+N G + VF + RW++A +A AG+
Sbjct: 147 GQTVMAIFYKVDPSDVKNLTGDFGK--VFRKTCAGKPKKDIGRWRQAWEKVATVAGYHSI 204
Query: 198 N--------KPEFREIENIVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYD 249
N K +I NI+ + R F G L+G++ +E ++ LL L+++ +
Sbjct: 205 NWDNEAAMIKKIATDISNIL--INSTPSRDFDG----LVGMRAHLEKMKPLLCLDTD--E 256
Query: 250 CQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENVSKVY-RDGGVTAVQKQVLRQTV 308
++IGIWG GIGKTT+A V+Y+++SH F+ F+EN+ Y R G ++ Q +
Sbjct: 257 VRIIGIWGPPGIGKTTIARVVYNQLSHSFQLSVFMENIKANYTRPTGSDDYSAKLQLQQM 316
Query: 309 DEMNLETYSPSEIS--GIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIIT 366
+ EI G+ +DRL+ KVLVVLD V+Q VQL A F GSR+IIT
Sbjct: 317 FMSQITKQKDIEIPHLGVAQDRLKDKKVLVVLDGVNQSVQLDAMAKEAWWFGPGSRIIIT 376
Query: 367 TRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPL 426
T+D+ + + +G + +Y+V +A ++F F ++ L +V+ A LPL
Sbjct: 377 TQDQKLFRAHGINHIYKVDFPPTEEALQIFCMYAFGQNSPKDGFQNLAWKVINLAGNLPL 436
Query: 427 AIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKG 486
+R+ GS+ + +W+ +L RL+++ D + +L+ S++ L EDK +FLHIACFF G
Sbjct: 437 GLRIMGSYFRGMSREEWKKSLPRLESSLDADIQSILKFSYDALDDEDKNLFLHIACFFNG 496
Query: 487 EKENYVKRILDACGLHPHIGIQNMIERSLITIRN-QEIHMHEMVQDLGKKIVRQQFPEEP 545
++ ++ L + + + E+SLI+ N I MH+++ LG +IVR Q EP
Sbjct: 497 KEIKILEEHLAKKFVEVRQRLNVLAEKSLISFSNWGTIEMHKLLAKLGGEIVRNQSIHEP 556
Query: 546 GSWSRLWLYQHFHHVLMSE-MGTNKVKAI----VLDQNEDISEYPQLRAEGLSIMRGLII 600
G L+ + VL + G+ V I ++++ D++E EG+S ++ L
Sbjct: 557 GQRQFLFDGEEICDVLNGDAAGSKSVIGIDFHYIIEEEFDMNERV---FEGMSNLQFLRF 613
Query: 601 LILHHQ-NFSGSLHFLSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRK 659
H S L +LS LQ L W +P LPS L+ELN+ +S + LWEG K
Sbjct: 614 DCDHDTLQLSRGLSYLSRKLQLLDWIYFPMTCLPSTVNVEFLIELNLTHSKLDMLWEGVK 673
Query: 660 DLPFLKRMDLSNSKYLTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESC 719
L L++MDLS S L E P+ + L +L L+ C++L+++ IG L L C
Sbjct: 674 PLHNLRQMDLSYSVNLKELPDLSTAINLRKLILSNCSSLIKLPSCIGNAINLEDLDLNGC 733
Query: 720 SSLVSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTVDQSIG 778
SSLV +L S +L L L C+ L P+ G NL LD+ C SL + SIG
Sbjct: 734 SSLV--ELPSFGDAINLQKLLLRYCSNLVELPSSIGNAINLRELDLYYCSSLIRLPSSIG 791
Query: 779 VLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIF 838
L L L C NL +P S+ N +L LD C KL LP + + LQ+L+
Sbjct: 792 NAINLLILDLNGCSNLLELPSSIGNAINLQKLDLRRCAKLLELPSSIG--NAINLQNLLL 849
Query: 839 LDLGFCSLSEVPHALGEIECLERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSKLEFL 896
D SL E+P ++G L +NL +N V LP S+ L L L L CSKLE L
Sbjct: 850 DDCS--SLLELPSSIGNATNLVYMNLSNCSNLVELPLSIGNLQKLQELILKGCSKLEDL 906
>At2g14080 putative disease resistance protein
Length = 1215
Score = 427 bits (1099), Expect = e-119
Identities = 312/911 (34%), Positives = 468/911 (51%), Gaps = 68/911 (7%)
Query: 34 STASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGE 93
+++S T+ S+ + K+DVF SF G+D R +F+ H+ RKGI F D+ +++ +
Sbjct: 38 TSSSPPTSPQSSLSCNQKHDVFPSFHGADVRKSFLSHILKEFKRKGIDTFIDNN-IERSK 96
Query: 94 SISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPV 153
SI +L++AI+ S++++V+ SK+YA S WCL+E+ I +C + QTV +FY+VDP+ V
Sbjct: 97 SIGPELIEAIKGSKIAVVLLSKDYASSSWCLNELVEIMKCRKMLDQTVMTIFYEVDPTDV 156
Query: 154 RNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAG-----WDVRNKPEFREIENI 208
+ Q G + F + + R +W A+ +A AG WD E IE I
Sbjct: 157 KKQTGDFGKVFKKTCMGKTNAVSR--KWIEALSEVATIAGEHSINWDT----EAAMIEKI 210
Query: 209 VEAVIEALG-----RKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGK 263
+ L R F G L+G+ +E LE LL L+S + ++IGIWG GIGK
Sbjct: 211 STDISNKLNNSTPLRDFDG----LVGMGAHMEKLELLLCLDS--CEVRMIGIWGPPGIGK 264
Query: 264 TTLATVLYDRISHLFEARCFVENVSKVYRD-------GGVTAVQKQVLRQTVDEMNLETY 316
TT+ LY+++S FE F+EN+ ++ +Q+Q L + +D ++E
Sbjct: 265 TTIVRFLYNQLSSSFELSIFMENIKTMHTILASSDDYSAKLILQRQFLSKILDHKDIEIP 324
Query: 317 SPSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVY 376
++++RL + KVLVVLD+VDQ VQL A F SR++ITT+D +LK +
Sbjct: 325 HLR----VLQERLYNKKVLVVLDDVDQSVQLDALAKETRWFGPRSRILITTQDRKLLKAH 380
Query: 377 GAHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLC 436
+ +Y+V L N++DA ++F F +L +V PL +RV GS+
Sbjct: 381 RINNIYKVDLPNSDDALQIFCMYAFGQKTPYDGFYKLARKVTWLVGNFPLGLRVVGSYFR 440
Query: 437 TRNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRIL 496
+ +WR + RL+ D K+ VL+ S++ L EDK++FLHIACFF E ++ L
Sbjct: 441 EMSKQEWRKEIPRLRARLDGKIESVLKFSYDALCDEDKDLFLHIACFFNHESIEKLEDFL 500
Query: 497 DACGLHPHIGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQH 556
L + E+SLI+I + + MH+ + LGK+IVR+Q EPG L +
Sbjct: 501 GKTFLDIAQRFHVLAEKSLISINSNFVEMHDSLAQLGKEIVRKQSVREPGQRQFLVDARD 560
Query: 557 FHHVLMSE-MGTNKVKAIVLD--QNEDISEYPQLRAEGLSIMRGL----------IILIL 603
VL + G V I LD +N+D+ + EG+S ++ L I+ L
Sbjct: 561 ISEVLADDTAGGRSVIGIYLDLHRNDDVFNISEKAFEGMSNLQFLRVKNFGNLFPAIVCL 620
Query: 604 HHQNFSGSLHFLSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPF 663
H L ++S L+ L W +P PS F P LVELNM S +++LWE + L
Sbjct: 621 PH-----CLTYISRKLRLLDWMYFPMTCFPSKFNPEFLVELNMWGSKLEKLWEEIQPLRN 675
Query: 664 LKRMDLSNSKYLTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLV 723
LKRMDL +SK L E P+ + LE L+L GC++L+++ SIG TKL L CSSL+
Sbjct: 676 LKRMDLFSSKNLKELPDLSSATNLEVLNLNGCSSLVELPFSIGNATKLLKLELSGCSSLL 735
Query: 724 SLDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTVDQSIGVLTR 782
L S+ +L + S C L P+ G NL+ LD+ C SL + SIG T
Sbjct: 736 ELP-SSIGNAINLQTIDFSHCENLVELPSSIGNATNLKELDLSCCSSLKELPSSIGNCTN 794
Query: 783 LEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLG 842
L+ L L C +L +P S+ N +L L C L LP + + L+ LI G
Sbjct: 795 LKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSLIKLPSSIG--NAINLEKLIL--AG 850
Query: 843 FCSLSEVPHALGEIECLERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSK--------- 892
SL E+P +G+ L+ LNL + V LPS + L L+ L L C K
Sbjct: 851 CESLVELPSFIGKATNLKILNLGYLSCLVELPSFIGNLHKLSELRLRGCKKLQVLPTNIN 910
Query: 893 LEFLSELQLCD 903
LEFL+EL L D
Sbjct: 911 LEFLNELDLTD 921
Score = 70.9 bits (172), Expect = 3e-12
Identities = 75/252 (29%), Positives = 114/252 (44%), Gaps = 22/252 (8%)
Query: 649 SSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSR-RLERLDLTGCTNLLQVHPSIGL 707
SS++ L + LK + L+ L + P+ G+ LE+L L GC +L+++ IG
Sbjct: 804 SSLKELPSSIGNCTNLKELHLTCCSSLIKLPSSIGNAINLEKLILAGCESLVELPSFIGK 863
Query: 708 LTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTGVENLEYLDIDQC 767
T L L+ S LV L + L+ L+ L L GC KL+ P +E L LD+ C
Sbjct: 864 ATNLKILNLGYLSCLVELP-SFIGNLHKLSELRLRGCKKLQVLPTNINLEFLNELDLTDC 922
Query: 768 VSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPS 827
+ L T + T ++ L LR + +P S+ + L L L +
Sbjct: 923 ILLKTFPV---ISTNIKRLHLRG-TQIEEVPSSLRSWPRLEDLQM----------LYSEN 968
Query: 828 LSPFT--LQSLIFLDLGFCSLSEVPHALGEIECLERLNLEG-NNFVSLPSSLRWLSSLAY 884
LS F+ L+ + L+L ++ E+ L I L RL L G VSLP + SL
Sbjct: 969 LSEFSHVLERITVLELSDINIREMTPWLNRITRLRRLKLSGCGKLVSLP---QLSDSLII 1025
Query: 885 LNLAHCSKLEFL 896
L+ +C LE L
Sbjct: 1026 LDAENCGSLERL 1037
>At3g04220 putative disease resistance protein
Length = 896
Score = 425 bits (1093), Expect = e-119
Identities = 291/863 (33%), Positives = 445/863 (50%), Gaps = 50/863 (5%)
Query: 34 STASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGE 93
ST S + S+ + KYDVF SFRG D R F+ H+ R+GI F D+ +++GE
Sbjct: 45 STQPSPSPPLSSLSLTRKYDVFPSFRGEDVRKDFLSHIQKEFQRQGITPFVDNN-IKRGE 103
Query: 94 SISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPV 153
SI +L++AIR S+++I++ SKNYA S WCLDE+ I +C E+ QTV +FY VDPS V
Sbjct: 104 SIGPELIRAIRGSKIAIILLSKNYASSSWCLDELVEIIKCKEEMGQTVIVIFYKVDPSLV 163
Query: 154 RNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAV 212
+ G + VF + + ++RW+ A + +A AG+D R E IE IV +
Sbjct: 164 KKLTGDFGK--VFRNTCKGKERENIERWREAFKKVATIAGYDSRKWDNESGMIEKIVSDI 221
Query: 213 IEALGRKF-SGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLY 271
E L S DDLIG+ +E ++ LL ++S+ + + IGIWG G+GKTT+A LY
Sbjct: 222 SEMLNHSTPSRDFDDLIGMGDHMEKMKPLLDIDSD--EMKTIGIWGPPGVGKTTIARSLY 279
Query: 272 DRISHLFEARCFVENVSKVYRDGGVT-------AVQKQVLRQTVDEMNLETYSPSEISGI 324
++ S F+ F+E++ Y + +Q++ L Q ++ N++ G+
Sbjct: 280 NQHSDKFQLSVFMESIKTAYTIPACSDDYYEKLQLQQRFLSQITNQENVQIPH----LGV 335
Query: 325 VRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEV 384
++RL KVLVV+D+V+Q VQ+ A GSR+IITT+D IL+ +G +YEV
Sbjct: 336 AQERLNDKKVLVVIDDVNQSVQVDALAKENDWLGPGSRIIITTQDRGILRAHGIEHIYEV 395
Query: 385 PLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWR 444
N +A ++F F + EL +V + LPL ++V GS+ +W
Sbjct: 396 DYPNYEEALQIFCMHAFGQKSPYDGFEELAQQVTTLSGRLPLGLKVMGSYFRGMTKQEWT 455
Query: 445 DALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPH 504
AL R++ + D K+ +L++S++ L DK +FLH+AC F + V++ L
Sbjct: 456 MALPRVRTHLDGKIESILKLSYDALCDVDKSLFLHLACSFHNDDTELVEQQLGKKFSDLR 515
Query: 505 IGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSE 564
G+ + E+SLI + + I MH ++ LG++IVR+Q EPG L VL +
Sbjct: 516 QGLHVLAEKSLIHMDLRLIRMHVLLAQLGREIVRKQSIHEPGQRQFLVDATDIREVLTDD 575
Query: 565 MGTNKVKAIVLDQNE-----DISEYPQLRAEGLSIMRGLIILILHH-------------Q 606
G+ V I D N DISE L +R L H
Sbjct: 576 TGSRSVIGIDFDFNTMEKELDISEKAFRGMSNLQFIRIYGDLFSRHGVYYFGGRGHRVSL 635
Query: 607 NFSGSLHF------LSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKD 660
++ LHF L L+ L W +P SLPS F LV+L MPYS +++LWEG +
Sbjct: 636 DYDSKLHFPRGLDYLPGKLRLLHWQQFPMTSLPSEFHAEFLVKLCMPYSKLEKLWEGIQP 695
Query: 661 LPFLKRMDLSNSKYLTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCS 720
L L+ +DL+ S+ L E P+ + L+RL + C++L+++ SIG T L ++ C
Sbjct: 696 LRNLEWLDLTCSRNLKELPDLSTATNLQRLSIERCSSLVKLPSSIGEATNLKKINLRECL 755
Query: 721 SLVSLDLGSLCVLYSLAVLHLSGCTKLESTP-NFTGVENLEYLDIDQCVSLSTVDQSIGV 779
SLV L S L +L L L C+ L P +F + N+E L+ +C SL + + G
Sbjct: 756 SLVELP-SSFGNLTNLQELDLRECSSLVELPTSFGNLANVESLEFYECSSLVKLPSTFGN 814
Query: 780 LTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFL 839
LT L L LR+C ++ +P S N+ +L L+ C L LP +L+ +L L
Sbjct: 815 LTNLRVLGLRECSSMVELPSSFGNLTNLQVLNLRKCSTLVELPSSFVNLT-----NLENL 869
Query: 840 DLGFCSLSEVPHALGEIECLERL 862
DL CS S +P + G + L+RL
Sbjct: 870 DLRDCS-SLLPSSFGNVTYLKRL 891
>At5g18360 disease resistance protein -like
Length = 900
Score = 423 bits (1088), Expect = e-118
Identities = 283/841 (33%), Positives = 442/841 (51%), Gaps = 75/841 (8%)
Query: 50 YKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVS 109
+++ VF SF G D R TF+ HL RKGI F D+ +++ + IS++L++AIR SR++
Sbjct: 14 WRHHVFPSFSGKDVRRTFLSHLLKEFRRKGIRTFIDND-IKRSQMISSELVRAIRESRIA 72
Query: 110 IVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHML 169
+VV S+ YA S WCL+E+ I + Q + PVFY+VDPS VR + G + AF
Sbjct: 73 VVVLSRTYASSSWCLNELVEIKKV----SQMIMPVFYEVDPSDVRKRTGEFGKAFEEACE 128
Query: 170 RFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALGRKFSGFADDLI 228
R + D + +W+ A+ +A AG +N E I+ I ++ L S + +L+
Sbjct: 129 R-QPDEEVKQKWREALVYIANIAGESSQNWDNEADLIDKIAMSISYELNSTLSRDSYNLV 187
Query: 229 GIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENVS 288
GI + L++LL L S + +++GIWG GIGKTT+A L++R+S F+ F+ENV
Sbjct: 188 GIDNHMRELDSLLCLEST--EVKMVGIWGPAGIGKTTIARALFNRLSENFQHTIFMENVK 245
Query: 289 KVYRDGGVTA------VQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVD 342
R + A +Q+Q L + +D +++ + G+V++RL+ LKVLVVLD+VD
Sbjct: 246 GSSRTSELDAYGFQLRLQEQFLSEVIDHKHMKIHD----LGLVKERLQDLKVLVVLDDVD 301
Query: 343 QLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFK 402
+L QL F GSR+I+TT ++ +L+ +G +YE+ + +D+ ++F + F
Sbjct: 302 KLEQLDALVKQSQWFGSGSRIIVTTENKQLLRAHGITCIYELGFPSRSDSLQIFCQYAFG 361
Query: 403 SDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVL 462
+ C EL E+ K A LPLA++V GS L + + + AL RL+ + + + +VL
Sbjct: 362 ESSAPDGCIELATEITKLAGYLPLALKVLGSSLRGMSKDEQKSALPRLRTSLNEDIRNVL 421
Query: 463 QISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLITIR--N 520
++ ++G+H +DK IFLHIAC F GE +YVK+IL + GL G+Q + RSLI I N
Sbjct: 422 RVGYDGIHDKDKVIFLHIACLFNGENVDYVKQILASSGLDVTFGLQVLTSRSLIHISRCN 481
Query: 521 QEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLDQNED 580
+ I MH +++ LG++IV +Q EPG L + VL GT V I LD ++
Sbjct: 482 RTITMHNLLEQLGREIVCEQSIAEPGKRQFLMDASEIYDVLADNTGTGAVLGISLDISKI 541
Query: 581 ISEYPQLRAEGLSIMRGLIILILHHQNFS---------GSLHFLSNNLQYLLWHGYPFAS 631
+ RA G M L+ L + + S L +L L+ L W +P S
Sbjct: 542 NELFLNERAFG--GMHNLLFLRFYKSSSSKDQPELHLPRGLDYLPRKLRLLHWDAFPMTS 599
Query: 632 LPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSRRLERLD 691
+P +F P LV +N+ S +++LWEG + L LK+MDLS S+ L E P+ + +E L
Sbjct: 600 MPLSFCPQFLVVINIRESQLEKLWEGTQPLRSLKQMDLSKSENLKEIPDLSKAVNIEELC 659
Query: 692 LTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTP 751
L+ C +L+ + SI L KL L + CS L + L SL++L+L GC++LES P
Sbjct: 660 LSYCGSLVMLPSSIKNLNKLVVLDMKYCSKLEIIPCN--MDLESLSILNLDGCSRLESFP 717
Query: 752 NF-----------TGVEN----------LEYLDIDQCVSLST------------------ 772
T +E L LD+ C +L T
Sbjct: 718 EISSKIGFLSLSETAIEEIPTTVASWPCLAALDMSGCKNLKTFPCLPKTIEWLDLSRTEI 777
Query: 773 --VDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSP 830
V I L++L L + C+ L +I ++ +E + TLDF GC + P+ + S
Sbjct: 778 EEVPLWIDKLSKLNKLLMNSCMKLRSISSGISTLEHIKTLDFLGCKNIVSFPVEIFESSR 837
Query: 831 F 831
F
Sbjct: 838 F 838
Score = 38.1 bits (87), Expect = 0.024
Identities = 52/204 (25%), Positives = 85/204 (41%), Gaps = 25/204 (12%)
Query: 690 LDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSL-----VSLDLGSLCVLYSLAVLHLSG- 743
LD++ L + G + L FL F SS + L G + L +LH
Sbjct: 536 LDISKINELFLNERAFGGMHNLLFLRFYKSSSSKDQPELHLPRGLDYLPRKLRLLHWDAF 595
Query: 744 ---CTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLS 800
L P F V N+ +++ L Q + L +++ + + ++ +
Sbjct: 596 PMTSMPLSFCPQFLVVINIRESQLEK---LWEGTQPLRSLKQMDLSKSENLKEIPDLSKA 652
Query: 801 VNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLE 860
VN E L L +CG L + LPS S L L+ LD+ +CS E+ ++E L
Sbjct: 653 VNIEE--LCLSYCGSLVM------LPS-SIKNLNKLVVLDMKYCSKLEIIPCNMDLESLS 703
Query: 861 RLNLEG----NNFVSLPSSLRWLS 880
LNL+G +F + S + +LS
Sbjct: 704 ILNLDGCSRLESFPEISSKIGFLS 727
Score = 33.5 bits (75), Expect = 0.58
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 1/65 (1%)
Query: 833 LQSLIFLDLGFC-SLSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCS 891
L+SL +DL +L E+P + E + V LPSS++ L+ L L++ +CS
Sbjct: 629 LRSLKQMDLSKSENLKEIPDLSKAVNIEELCLSYCGSLVMLPSSIKNLNKLVVLDMKYCS 688
Query: 892 KLEFL 896
KLE +
Sbjct: 689 KLEII 693
>At5g46470 disease resistance protein-like
Length = 1160
Score = 418 bits (1075), Expect = e-117
Identities = 301/902 (33%), Positives = 456/902 (50%), Gaps = 76/902 (8%)
Query: 36 ASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESI 95
ASSS S+ +R + Y VF SF G D RNTF+ H L RK I FKD++ +++ +S+
Sbjct: 2 ASSS----SSSSRNWSYHVFPSFSGEDVRNTFLSHFLKELDRKLIISFKDNE-IERSQSL 56
Query: 96 SAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRN 155
+L IRNSR+++VVFSK YA S WCL+E+ I +C ++F Q V P+FY++DPS VR
Sbjct: 57 DPELKHGIRNSRIAVVVFSKTYASSSWCLNELLEIVKCKKEFGQLVIPIFYNLDPSHVRK 116
Query: 156 QNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIE 214
Q G + +F D RWK A+ +A G+ + E IE I ++
Sbjct: 117 QTGDFGK--IFEKTCRNKTVDEKIRWKEALTDVANILGYHIVTWDNEASMIEEIANDILG 174
Query: 215 ALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRI 274
+ S +DL+GI+ + + +LL L SE + +++GIWG GIGKTT+A L+ R+
Sbjct: 175 KMNISPSNDFEDLVGIEDHITKMSSLLHLESE--EVRMVGIWGPSGIGKTTIARALFSRL 232
Query: 275 SHLFEARCFVENV-----SKVYRDGGVT------AVQKQVLRQTVDEMNLETYSPSEISG 323
S F++ F++ V +VY + +Q+ L + D+ +++ + G
Sbjct: 233 SCQFQSSVFIDKVFISKSMEVYSGANLVDYNMKLHLQRAFLAEIFDKKDIKIH-----VG 287
Query: 324 IVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYE 383
+ ++ K L+V+D++D L A F GSR+I+ T ++H L+ +Y+
Sbjct: 288 AMEKMVKHRKALIVIDDLDDQDVLDALADQTQWFGSGSRIIVVTENKHFLRANRIDHIYK 347
Query: 384 VPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQW 443
V L +N A E+F R FK ++ EL EV A LPL + V GS L N W
Sbjct: 348 VCLPSNALALEMFCRSAFKKNSPPDDFLELSSEVALRAGNLPLGLNVLGSNLRGINKGYW 407
Query: 444 RDALDRLKNNPDNKVMDVLQISFEGLHS-EDKEIFLHIACFFKGEKENYVKRILDACGLH 502
D L RL+ D K+ L++S++GL++ +D+ IF HIAC F GEK + +K +L L
Sbjct: 408 IDMLPRLQ-GLDGKIGKTLRVSYDGLNNRKDEAIFRHIACIFNGEKVSDIKLLLANSNLD 466
Query: 503 PHIGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLM 562
+IG++N+++RSLI R + MH ++Q+LGK+IVR Q +PG L + VL
Sbjct: 467 VNIGLKNLVDRSLICERFNTLEMHSLLQELGKEIVRTQ-SNQPGEREFLVDLKDICDVLE 525
Query: 563 SEMGTNKVKAIVLDQNEDISEYPQLRAEGLSIMRGLIILILHHQNFSGS----------L 612
GT KV I LD +E ++ + M L+ L ++ +
Sbjct: 526 HNTGTKKVLGITLDIDE--TDELHIHESSFKGMHNLLFLKIYTKKLDQKKKVRWHLPERF 583
Query: 613 HFLSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNS 672
+L + L+ L + YP LPSNF P LV+L M S +++LW+G L L+ MDL S
Sbjct: 584 DYLPSRLRLLRFDRYPSKCLPSNFHPENLVKLQMQQSKLEKLWDGVHSLAGLRNMDLRGS 643
Query: 673 KYLTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCV 732
+ L E P+ + LE L L+ C++L+++ SI L KL L C L ++ G
Sbjct: 644 RNLKEIPDLSMATNLETLKLSSCSSLVELPSSIQYLNKLNDLDMSYCDHLETIPSG--VN 701
Query: 733 LYSLAVLHLSGCTKLES------------------TPNFTGVENLEYLDIDQCVSLST-- 772
L SL L+LSGC++L+S P+ ++NL+ L + + V L T
Sbjct: 702 LKSLDRLNLSGCSRLKSFLDIPTNISWLDIGQTADIPSNLRLQNLDELILCERVQLRTPL 761
Query: 773 VDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFT 832
+ LTRL F + + +P S+ N+ L L+ C L LP G+
Sbjct: 762 MTMLSPTLTRLTF---SNNPSFVEVPSSIQNLYQLEHLEIMNCRNLVTLPTGI------N 812
Query: 833 LQSLIFLDLGFCS-LSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCS 891
L SLI LDL CS L P I LNL +P S+ LS L YL++ CS
Sbjct: 813 LDSLISLDLSHCSQLKTFPDISTNI---SDLNLSYTAIEEVPLSIEKLSLLCYLDMNGCS 869
Query: 892 KL 893
L
Sbjct: 870 NL 871
Score = 53.1 bits (126), Expect = 7e-07
Identities = 50/172 (29%), Positives = 78/172 (45%), Gaps = 24/172 (13%)
Query: 662 PFLKRMDLSNSKYLTETPN-FEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCS 720
P L R+ SN+ E P+ + +LE L++ C NL+ + I L L L CS
Sbjct: 767 PTLTRLTFSNNPSFVEVPSSIQNLYQLEHLEIMNCRNLVTLPTGINL-DSLISLDLSHCS 825
Query: 721 SLVSL-DLGSLCVLYSLAVLHLSGCTKLESTPNFTGVENLE---YLDIDQCVSLSTVDQS 776
L + D+ + +++ L+LS T +E P +E L YLD++ C +L V +
Sbjct: 826 QLKTFPDIST-----NISDLNLS-YTAIEEVP--LSIEKLSLLCYLDMNGCSNLLCVSPN 877
Query: 777 IGVLTRLEFLSLRDCLNLTNIPLSVNNME----------SLLTLDFCGCLKL 818
I L LE DC+ LT + ++ E S + L+F C KL
Sbjct: 878 ISKLKHLERADFSDCVELTEASWNGSSSEMVKLLPADNFSTVKLNFINCFKL 929
Score = 50.1 bits (118), Expect = 6e-06
Identities = 43/151 (28%), Positives = 69/151 (45%), Gaps = 11/151 (7%)
Query: 757 ENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCL 816
ENL L + Q L + + L L + LR NL IP ++ +L TL C
Sbjct: 610 ENLVKLQMQQS-KLEKLWDGVHSLAGLRNMDLRGSRNLKEIP-DLSMATNLETLKLSSCS 667
Query: 817 KLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEG----NNFVSL 872
L LP + L+ L LD+ +C E + ++ L+RLNL G +F+ +
Sbjct: 668 SLVELPSSIQYLN-----KLNDLDMSYCDHLETIPSGVNLKSLDRLNLSGCSRLKSFLDI 722
Query: 873 PSSLRWLSSLAYLNLAHCSKLEFLSELQLCD 903
P+++ WL ++ +L+ L EL LC+
Sbjct: 723 PTNISWLDIGQTADIPSNLRLQNLDELILCE 753
Score = 32.0 bits (71), Expect = 1.7
Identities = 23/70 (32%), Positives = 30/70 (42%), Gaps = 9/70 (12%)
Query: 690 LDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDL-GSLCVL--------YSLAVLH 740
LD+ GC+NLL V P+I L L F C L GS + +S L+
Sbjct: 863 LDMNGCSNLLCVSPNISKLKHLERADFSDCVELTEASWNGSSSEMVKLLPADNFSTVKLN 922
Query: 741 LSGCTKLEST 750
C KL+ T
Sbjct: 923 FINCFKLDLT 932
>At5g41750 disease resistance protein-like
Length = 1068
Score = 417 bits (1071), Expect = e-116
Identities = 279/814 (34%), Positives = 442/814 (54%), Gaps = 59/814 (7%)
Query: 51 KYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSI 110
+Y VF SF G D R F+ HL++ KGI F +D+K+ +G++I +L+Q IR +RVSI
Sbjct: 12 RYQVFSSFHGPDVRKGFLSHLHSVFASKGITTF-NDQKIDRGQTIGPELIQGIREARVSI 70
Query: 111 VVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLR 170
VV SK YA S WCLDE+ I +C E Q V VFY+VDPS V+ Q+GV+ AF +
Sbjct: 71 VVLSKKYASSSWCLDELVEILKCKEALGQIVMTVFYEVDPSDVKKQSGVFGEAFE-KTCQ 129
Query: 171 FKHDADRVDRWKRAMRSLAGSAG-----WDVRNKPEFREIENIVEAVIEALGRKFSGFAD 225
K++ ++ RW+ A+ +A AG WD E + I+ IV V + L S +
Sbjct: 130 GKNEEVKI-RWRNALAHVATIAGEHSLNWD----NEAKMIQKIVTDVSDKLNLTPSRDFE 184
Query: 226 DLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVE 285
++G++ ++ L +LL L S+ + ++IGIWG GIGKTT+A L+++IS +F +CF+E
Sbjct: 185 GMVGMEAHLKRLNSLLCLESD--EVKMIGIWGPAGIGKTTIARTLFNKISSIFPFKCFME 242
Query: 286 NVSKVYRDGGV----TAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNV 341
N+ + G ++QKQ+L + + + N++ + G ++ L KVL++LD+V
Sbjct: 243 NLKGSIKGGAEHYSKLSLQKQLLSEILKQENMKIHH----LGTIKQWLHDQKVLIILDDV 298
Query: 342 DQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGF 401
D L QL+ A +P F GSR+I+TT D++ILK + +Y V + +A E+ F
Sbjct: 299 DDLEQLEVLAEDPSWFGSGSRIIVTTEDKNILKAHRIQDIYHVDFPSEEEALEILCLSAF 358
Query: 402 KSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDV 461
K ++ EL +V + LPL + V G+ L ++ +W L R++++ D + ++
Sbjct: 359 KQSSIPDGFEELANKVAELCGNLPLGLCVVGASLRRKSKNEWERLLSRIESSLDKNIDNI 418
Query: 462 LQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLITIR-- 519
L+I ++ L +ED+ +FLHIACFF EK +Y+ +L L G + +RSL+ I
Sbjct: 419 LRIGYDRLSTEDQSLFLHIACFFNNEKVDYLTALLADRKLDVVNGFNILADRSLVRISTD 478
Query: 520 NQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLDQNE 579
+ H ++Q LG++IV +Q+P EPG L + VL GT VK I D +
Sbjct: 479 GHVVMHHYLLQKLGRRIVHEQWPNEPGKRQFLIEAEEIRDVLTKGTGTESVKGISFDTSN 538
Query: 580 DISEYPQLRAEGLSIMRGLIILILHHQNFS--GSLHFLSNNLQY------LLWHGYPFAS 631
E + MR L L ++ +F+ G+L + +++Y L W YP S
Sbjct: 539 --IEEVSVGKGAFEGMRNLQFLRIYRDSFNSEGTLQ-IPEDMEYIPPVRLLHWQNYPRKS 595
Query: 632 LPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSRRLERLD 691
LP F P LV++ MP S +++LW G + LP LK +D+S S L E PN + LE L
Sbjct: 596 LPQRFNPEHLVKIRMPSSKLKKLWGGIQPLPNLKSIDMSFSYSLKEIPNLSKATNLEILS 655
Query: 692 LTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTP 751
L C +L+++ SI L KL L+ E+CS L + L SL L ++GC++L + P
Sbjct: 656 LEFCKSLVELPFSILNLHKLEILNVENCSMLKVIPTN--INLASLERLDMTGCSELRTFP 713
Query: 752 NFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFL-----SLR-----DCL--------N 793
+ + N++ L++ + + V S+G +RL+ L SL+ C+ N
Sbjct: 714 DIS--SNIKKLNLGDTM-IEDVPPSVGCWSRLDHLYIGSRSLKRLHVPPCITSLVLWKSN 770
Query: 794 LTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPS 827
+ +IP S+ + L L+ C KLK + LGLPS
Sbjct: 771 IESIPESIIGLTRLDWLNVNSCRKLKSI-LGLPS 803
>At4g11170 RPP1-WsA-like disease resistance protein
Length = 1174
Score = 410 bits (1053), Expect = e-114
Identities = 276/826 (33%), Positives = 426/826 (51%), Gaps = 49/826 (5%)
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ + ++YDVF SFRG D RN F+ HL KGI F+DD +++ +I +L AI
Sbjct: 3 SSSSNSWRYDVFPSFRGEDVRNNFLSHLLKEFESKGIVTFRDDH-IKRSHTIGHELRAAI 61
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENA 163
R S++S+V+FS+NYA S WCLDE+ I +C E+ V PVFY VDPS +R Q G + +
Sbjct: 62 RESKISVVLFSENYASSSWCLDELIEIMKCKEEQGLKVMPVFYKVDPSDIRKQTGKFGMS 121
Query: 164 FVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALGRKFSG 222
F+ +R W+RA+ A G +N E +I I + V+E L S
Sbjct: 122 FLETCCG--KTEERQHNWRRALTDAANILGDHPQNWDNEAYKITTISKDVLEKLNATPSR 179
Query: 223 FADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARC 282
+DL+G++ + +E+LL L S+ +++GIWG G+GKTT+A LY++ F
Sbjct: 180 DFNDLVGMEAHIAKMESLLCLESQ--GVRIVGIWGPAGVGKTTIARALYNQYHENFNLSI 237
Query: 283 FVENVSKVYRDGGVTA------VQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLV 336
F+ENV + Y + G+ +Q++ L + +D+ +L G + +RL+S KVL+
Sbjct: 238 FMENVRESYGEAGLDDYGLKLHLQQRFLSKLLDQKDLRVRH----LGAIEERLKSQKVLI 293
Query: 337 VLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELF 396
+LD+VD + QL+ A F SR+++TT+++ +L + + +Y+V + +A +F
Sbjct: 294 ILDDVDNIEQLKALAKENQWFGNKSRIVVTTQNKQLLVSHDINHMYQVAYPSKQEALTIF 353
Query: 397 YRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDN 456
+ FK + S L E A LPLA+RV GSF+ + +W +L LK+ D
Sbjct: 354 CQHAFKQSSPSDDLKHLAIEFTTLAGHLPLALRVLGSFMRGKGKEEWEFSLPTLKSRLDG 413
Query: 457 KVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACG-LHPHIGIQNMIERSL 515
+V VL++ ++GLH +K++FLHIAC F G+ ENY+K+++ A + G+Q + ++SL
Sbjct: 414 EVEKVLKVGYDGLHDHEKDLFLHIACIFSGQHENYLKQMIIANNDTYVSFGLQVLADKSL 473
Query: 516 I-TIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIV 574
I N I MH +++ LGK++VR+Q EPG L + VL + GT V I
Sbjct: 474 IQKFENGRIEMHSLLRQLGKEVVRKQSIYEPGKRQFLMNAKETCGVLSNNTGTGTVLGIS 533
Query: 575 LDQNEDISEYPQLRAEGLSIMRGLIILILH-----HQNFSGSLHFLSNNLQY------LL 623
LD E I E + + MR L+ L + L L Y L
Sbjct: 534 LDMCE-IKEELYISEKTFEEMRNLVYLKFYMSSPIDDKMKVKLQLPEEGLSYLPQLRLLH 592
Query: 624 WHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEG 683
W YP PS+F P LVELNM +S +++LW G + L L+ M+L++S+ L PN
Sbjct: 593 WDAYPLEFFPSSFRPECLVELNMSHSKLKKLWSGVQPLRNLRTMNLNSSRNLEILPNLME 652
Query: 684 SRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSG 743
+ +L RLDL C +L+++ SI L L L C L + L SL VLH
Sbjct: 653 ATKLNRLDLGWCESLVELPSSIKNLQHLILLEMSCCKKLEIIPTN--INLPSLEVLHFRY 710
Query: 744 CTKLESTP---------NFTGVENLE-------YLDIDQ-CVSLSTVDQSIGVLTRLEFL 786
CT+L++ P N G E + ID+ C+ + V + + V LE L
Sbjct: 711 CTRLQTFPEISTNIRLLNLIGTAITEVPPSVKYWSKIDEICMERAKVKRLVHVPYVLEKL 770
Query: 787 SLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFT 832
LR+ L IP + + L +D C+ + LP S+S T
Sbjct: 771 CLRENKELETIPRYLKYLPRLQMIDISYCINIISLPKLPGSVSALT 816
Score = 32.0 bits (71), Expect = 1.7
Identities = 46/157 (29%), Positives = 66/157 (41%), Gaps = 17/157 (10%)
Query: 749 STPNFTGVENLEYL--------DIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLS 800
S F + NL YL D V L ++ + L +L L D L P S
Sbjct: 546 SEKTFEEMRNLVYLKFYMSSPIDDKMKVKLQLPEEGLSYLPQLRLLHW-DAYPLEFFPSS 604
Query: 801 VNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLE 860
E L+ L+ KLK L G+ L ++L ++L E+ L E L
Sbjct: 605 FRP-ECLVELNMSHS-KLKKLWSGVQPL-----RNLRTMNLNSSRNLEILPNLMEATKLN 657
Query: 861 RLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSKLEFL 896
RL+L + V LPSS++ L L L ++ C KLE +
Sbjct: 658 RLDLGWCESLVELPSSIKNLQHLILLEMSCCKKLEII 694
>At5g38340 disease resistance - like protein
Length = 1059
Score = 407 bits (1046), Expect = e-113
Identities = 297/870 (34%), Positives = 449/870 (51%), Gaps = 51/870 (5%)
Query: 51 KYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSI 110
KYDVF SF G+D R TF+ H+ RKGI F D+ + + +SI +L +AIR S+++I
Sbjct: 56 KYDVFPSFHGADVRKTFLSHMLKEFKRKGIVPFIDND-IDRSKSIGPELDEAIRGSKIAI 114
Query: 111 VVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLR 170
V+ SKNYA S WCL+E+ I +C +D QTV +FY VDP+ V+ Q G + VF
Sbjct: 115 VMLSKNYASSSWCLNELVEITKCRKDLNQTVMTIFYGVDPTDVKKQTGEFGK--VFERTC 172
Query: 171 FKHDADRVDRWKRAMRSLAGSAG--WDVRNKPEFREIENIVEAVIEALGRKF-SGFADDL 227
++V W+ + A AG W + + E IE I V L R S DDL
Sbjct: 173 ESKTEEQVKTWREVLDGAATIAGEHWHIWDN-EASMIEKISIDVSNILNRSSPSRDFDDL 231
Query: 228 IGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENV 287
IG++ +E +++LL L+S + ++IGIWG GIGKTT+A VLY+R S F F++N+
Sbjct: 232 IGMEAHMEKMKSLLSLHSN--EVKMIGIWGPSGIGKTTIARVLYNRFSGDFGLSVFMDNI 289
Query: 288 SKVYRDGGVTA----VQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQ 343
++ V + + + Q + E+ + G+V DRL+ KVL+VLD++DQ
Sbjct: 290 KELMHTRPVGSDDYSAKLHLQNQLMSEITNHKETKITHLGVVPDRLKDNKVLIVLDSIDQ 349
Query: 344 LVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKS 403
+QL A F GSR+IITT+D+ +L+ + + +Y+V + +A ++F F
Sbjct: 350 SIQLDAIAKETQWFGPGSRIIITTQDQKLLEAHDINNIYKVEFPSKYEAFQIFCTYAFGQ 409
Query: 404 DNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQ 463
+ +L EV LPL +RV GS + W AL RLK D + +L+
Sbjct: 410 NFPKDGFEKLAWEVTDLLGELPLGLRVMGSHFRRMSKDDWVIALPRLKTRLDANIQSILK 469
Query: 464 ISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLITIRN--- 520
S++ L EDK++FLHIAC F E+ V+ L L G+ + E+SLI +
Sbjct: 470 FSYDALSPEDKDLFLHIACLFNNEEIVKVEDYLALDFLDARHGLHLLAEKSLIDLEGVNY 529
Query: 521 QEIHMHEMVQDLGKKIVR----QQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLD 576
+ + MH +++ LGK+IVR EP L + VL G+ +K I D
Sbjct: 530 KVLKMHNLLEQLGKEIVRYHPAHHSIREPEKRQFLVDTKDICEVLADGTGSKSIKGICFD 589
Query: 577 QNE-----DISEYPQLRA-EGLSIMRGLIILILHHQN--FSGSLHFLSNNLQYLLWHGYP 628
+ +ISE RA EG++ ++ L +L + L++L L+ + W +P
Sbjct: 590 LDNLSGRLNISE----RAFEGMTNLKFLRVLRDRSEKLYLPQGLNYLPKKLRLIEWDYFP 645
Query: 629 FASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSRRLE 688
SLPSNF LV L+M S +++LWEG++ L LK M+LSNS+ L E P+ + +L+
Sbjct: 646 MKSLPSNFCTTYLVNLHMRKSKLEKLWEGKQPLGNLKWMNLSNSRNLKELPDLSTATKLQ 705
Query: 689 RLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLE 748
L+LT C++L+++ SIG T L L+ C+SLV L S+ L+ L L L GC+KLE
Sbjct: 706 DLNLTRCSSLVEIPFSIGNTTNLEKLNLVMCTSLVELP-SSIGSLHKLRELRLRGCSKLE 764
Query: 749 STPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSL-RDCLNLTNIPLSVNNMESL 807
P +E+L+ LDI C L + + T ++ LSL R +N +P + + L
Sbjct: 765 VLPTNISLESLDNLDITDCSLLKSFPD---ISTNIKHLSLARTAIN--EVPSRIKSWSRL 819
Query: 808 LTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEG- 866
LK SP L ++ L + E+P + +I LE L LEG
Sbjct: 820 RYFVVSYNENLKE--------SPHALDTITMLSSNDTKMQELPRWVKKISRLETLMLEGC 871
Query: 867 NNFVSLPSSLRWLSSLAYLNLAHCSKLEFL 896
N V+LP LS++ +N C LE L
Sbjct: 872 KNLVTLPELPDSLSNIGVIN---CESLERL 898
>At1g69550 putative disease resistance protein
Length = 1398
Score = 406 bits (1044), Expect = e-113
Identities = 313/912 (34%), Positives = 477/912 (51%), Gaps = 70/912 (7%)
Query: 29 LAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKK 88
L+ S SSST+ S+ + + VF SFRG D R F+ H+ RKGI F D++
Sbjct: 55 LSSSTSHPSSSTSPPSSLSCTGTHHVFPSFRGDDVRRNFLSHIQKEFRRKGITPFIDNE- 113
Query: 89 LQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDV 148
+++GESI +L++AIR S+++IV+ S+NYA S+WCL+E+ I +C ++F TVF +FY+V
Sbjct: 114 IRRGESIGPELIKAIRESKIAIVLLSRNYASSKWCLEELVEIMKCKKEFGLTVFAIFYEV 173
Query: 149 DPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIEN 207
DPS V+ G E VF + + RW++A +A AG+D RN + E IE
Sbjct: 174 DPSHVKKLTG--EFGAVFQKTCKGRTKENIMRWRQAFEEVATIAGYDSRNWENEAAMIEE 231
Query: 208 IVEAVIEAL--GRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTT 265
I + + L FSGF + LIG++ +E ++ LL L+S + + +GI G GIGK+T
Sbjct: 232 IAIEISKRLINSSPFSGF-EGLIGMKAHIEKMKQLLCLDSTD-ERRTVGISGPSGIGKST 289
Query: 266 LATVLYDRISHLFEARCFVENVSKVYR-----DGGVTA-VQKQVLRQTVDEMNLETYSPS 319
+A VL+++IS F+ F++ R D V +++Q L Q +++ +++ +
Sbjct: 290 IARVLHNQISDGFQMSVFMKFKPSYTRPICSDDHDVKLQLEQQFLAQLINQEDIKIHQ-- 347
Query: 320 EISGIVRDRLRSLKVLVVLDNVDQLVQLQEF--AVNPGLFQKGSRMIITTRDEHILKVYG 377
G ++ + KVL+VLD VDQLVQL AV G GSR+IITT+D+ +LK +
Sbjct: 348 --LGTAQNFVMGKKVLIVLDGVDQLVQLLAMPKAVCLG---PGSRIIITTQDQQLLKAFQ 402
Query: 378 AHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCT 437
+Y V +++A ++F F D+ +L +V + A LPL +RV GS
Sbjct: 403 IKHIYNVDFPPDHEALQIFCIHAFGHDSPDDGFEKLATKVTRLAGNLPLGLRVMGSHFRG 462
Query: 438 RNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEK-ENYVKRIL 496
+ W+ L RL+ D ++ +L+ S++ L EDK++FLHIACFF E ++ + L
Sbjct: 463 MSKEDWKGELPRLRIRLDGEIGSILKFSYDVLDDEDKDLFLHIACFFNDEGIDHTFEDTL 522
Query: 497 DACGLHPHIGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQH 556
+ G+Q +++RSLI+ + MH ++ LG++IVR Q EPG L +
Sbjct: 523 RHKFSNVQRGLQVLVQRSLIS-EDLTQPMHNLLVQLGREIVRNQSVYEPGKRQFLVDGKE 581
Query: 557 FHHVLMSEMGTNKVKAIVLDQNEDISE--YPQLRAEGLSIMRGLIILILHHQNFSGSLH- 613
VL S G+ V I + + E EG+S ++ +N G LH
Sbjct: 582 ICEVLTSHTGSESVIGINFEVYWSMDELNISDRVFEGMSNLQ----FFRFDENSYGRLHL 637
Query: 614 -----FLSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMD 668
+L L+ L W YP SLPS F LV++ + +S +++LWEG + L LK MD
Sbjct: 638 PQGLNYLPPKLRILHWDYYPMTSLPSKFNLKFLVKIILKHSELEKLWEGIQPLVNLKVMD 697
Query: 669 LSNSKYLTETPNFE------------------------GSRRLERLDLTGCTNLLQVHPS 704
L S +L E PN + ++ LD+ GC++LL++ S
Sbjct: 698 LRYSSHLKELPNLSTAINLLEMVLSDCSSLIELPSSIGNATNIKSLDIQGCSSLLKLPSS 757
Query: 705 IGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLD 763
IG L L L CSSLV L S+ L +L L L GC+ L P+ G + NLE
Sbjct: 758 IGNLITLPRLDLMGCSSLVELP-SSIGNLINLPRLDLMGCSSLVELPSSIGNLINLEAFY 816
Query: 764 IDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPL 823
C SL + SIG L L+ L L+ +L IP S+ N+ +L L+ GC L LP
Sbjct: 817 FHGCSSLLELPSSIGNLISLKILYLKRISSLVEIPSSIGNLINLKLLNLSGCSSLVELPS 876
Query: 824 GLPSLSPFTLQSLIFLDLGFC-SLSEVPHALGEIECLERLNL-EGNNFVSLPSSLRWLSS 881
+ +L +L LDL C SL E+P ++G + L+ L L E ++ V LPSS+ L +
Sbjct: 877 SIGNLI-----NLKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLIN 931
Query: 882 LAYLNLAHCSKL 893
L LNL+ CS L
Sbjct: 932 LKTLNLSECSSL 943
Score = 122 bits (307), Expect = 7e-28
Identities = 110/335 (32%), Positives = 168/335 (49%), Gaps = 35/335 (10%)
Query: 590 EGLSIMRGLIILILHHQNFSGSLHFLS---NNLQYLLWHGYPFASLPSNF-EPFRLVELN 645
EG+ + L ++ L + + L LS N L+ +L LPS+ + L+
Sbjct: 685 EGIQPLVNLKVMDLRYSSHLKELPNLSTAINLLEMVLSDCSSLIELPSSIGNATNIKSLD 744
Query: 646 MP-YSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSR-RLERLDLTGCTNLLQVHP 703
+ SS+ +L +L L R+DL L E P+ G+ L RLDL GC++L+++
Sbjct: 745 IQGCSSLLKLPSSIGNLITLPRLDLMGCSSLVELPSSIGNLINLPRLDLMGCSSLVELPS 804
Query: 704 SIGLLTKLAFLSFESCSSLVSL-----DLGSLCVLY------------------SLAVLH 740
SIG L L F CSSL+ L +L SL +LY +L +L+
Sbjct: 805 SIGNLINLEAFYFHGCSSLLELPSSIGNLISLKILYLKRISSLVEIPSSIGNLINLKLLN 864
Query: 741 LSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPL 799
LSGC+ L P+ G + NL+ LD+ C SL + SIG L L+ L L +C +L +P
Sbjct: 865 LSGCSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPS 924
Query: 800 SVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECL 859
S+ N+ +L TL+ C L LP + +L LQ L + SL E+P ++G + L
Sbjct: 925 SIGNLINLKTLNLSECSSLVELPSSIGNL--INLQELYLSECS--SLVELPSSIGNLINL 980
Query: 860 ERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSKL 893
++L+L G ++ V LP S+ L +L LNL+ CS L
Sbjct: 981 KKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSL 1015
Score = 117 bits (292), Expect = 4e-26
Identities = 88/234 (37%), Positives = 133/234 (56%), Gaps = 11/234 (4%)
Query: 664 LKRMDLSNSKYLTETPNFEGSR-RLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSL 722
L+ + LS L E P+ G+ L+ L+L+ C++L+++ SIG L L L CSSL
Sbjct: 908 LQELYLSECSSLVELPSSIGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSECSSL 967
Query: 723 VSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTVDQSIGVLT 781
V L S+ L +L L LSGC+ L P G + NL+ L++ +C SL + SIG L
Sbjct: 968 VELP-SSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSLVELPSSIGNLI 1026
Query: 782 RLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDL 841
L+ L L +C +L +P S+ N+ +L LD GC L LPL + +L +L L+L
Sbjct: 1027 NLQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLI-----NLKTLNL 1081
Query: 842 GFC-SLSEVPHALGEIECLERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSKL 893
C SL E+P ++G + L++L+L G ++ V LPSS+ L +L L+L+ CS L
Sbjct: 1082 SGCSSLVELPSSIGNLN-LKKLDLSGCSSLVELPSSIGNLINLKKLDLSGCSSL 1134
Score = 117 bits (292), Expect = 4e-26
Identities = 100/275 (36%), Positives = 145/275 (52%), Gaps = 11/275 (4%)
Query: 649 SSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSR-RLERLDLTGCTNLLQVHPSIGL 707
SS+ L +L LK++DLS L E P G+ L+ L+L+ C++L+++ SIG
Sbjct: 965 SSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSLVELPSSIGN 1024
Query: 708 LTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQ 766
L L L CSSLV L S+ L +L L LSGC+ L P G + NL+ L++
Sbjct: 1025 LINLQELYLSECSSLVELP-SSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSG 1083
Query: 767 CVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLP 826
C SL + SIG L L+ L L C +L +P S+ N+ +L LD GC L LPL +
Sbjct: 1084 CSSLVELPSSIGNLN-LKKLDLSGCSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIG 1142
Query: 827 SLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNL-EGNNFVSLPSSLRWLSSLAYL 885
+L LQ L + SL E+P ++G + L+ L L E ++ V LPSS+ L +L L
Sbjct: 1143 NL--INLQELYLSECS--SLVELPSSIGNLINLQELYLSECSSLVELPSSIGNLINLKKL 1198
Query: 886 NLAHCSKLEFLSEL--QLCDIASEGGRYFRTLSGS 918
+L C+KL L +L L + +E TL+ S
Sbjct: 1199 DLNKCTKLVSLPQLPDSLSVLVAESCESLETLACS 1233
Score = 99.4 bits (246), Expect = 9e-21
Identities = 81/224 (36%), Positives = 116/224 (51%), Gaps = 5/224 (2%)
Query: 649 SSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSR-RLERLDLTGCTNLLQVHPSIGL 707
SS+ L +L LK++DLS L E P G+ L+ L+L+GC++L+++ SIG
Sbjct: 1037 SSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSGCSSLVELPSSIGN 1096
Query: 708 LTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQ 766
L L L CSSLV L S+ L +L L LSGC+ L P G + NL+ L + +
Sbjct: 1097 LN-LKKLDLSGCSSLVELP-SSIGNLINLKKLDLSGCSSLVELPLSIGNLINLQELYLSE 1154
Query: 767 CVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLP 826
C SL + SIG L L+ L L +C +L +P S+ N+ +L LD C KL LP
Sbjct: 1155 CSSLVELPSSIGNLINLQELYLSECSSLVELPSSIGNLINLKKLDLNKCTKLVSLPQLPD 1214
Query: 827 SLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFV 870
SLS +S L+ CS L I+C +LN +G + +
Sbjct: 1215 SLSVLVAESCESLETLACSFPNPQVWLKFIDCW-KLNEKGRDII 1257
>At5g49140 disease resistance protein-like
Length = 980
Score = 402 bits (1034), Expect = e-112
Identities = 293/862 (33%), Positives = 453/862 (51%), Gaps = 60/862 (6%)
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S + +KYDVF SFRG D R F+ HL KGI FKDD +++ ++I +L +A+
Sbjct: 7 SGSSSCWKYDVFPSFRGEDVRGNFLSHLMKEFESKGIVTFKDDL-IERSQTIGLELKEAV 65
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENA 163
R S++ +V+FSKNYA S WCLDE+ I +C E+ + + P+FY V+PS VRNQ G +
Sbjct: 66 RQSKIFVVIFSKNYASSSWCLDELVEILKCKEE--RRLIPIFYKVNPSDVRNQTGKFGRG 123
Query: 164 FVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALGRKFSG 222
F K+D + ++WK A+ A AG D ++ K E + I + ++ L S
Sbjct: 124 FR-ETCEGKNDETQ-NKWKAALTEAANIAGEDSQSWKNEADFLTKIAKDILAKLNGTPSN 181
Query: 223 FADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARC 282
+++IGI+ +E + LL LN + D +++GIWG GIGKTT+A VL+ R S F
Sbjct: 182 DFENIIGIESHMEKMVQLLCLNDD--DVRMVGIWGPAGIGKTTIARVLHSRFSGDFRFTV 239
Query: 283 FVENVSKVYR----DGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVL 338
F+ENV Y+ GG +Q ++ ++ + + + + +RL+ KVL+VL
Sbjct: 240 FMENVRGNYQRIVDSGGEYNLQARLQKEFLPIIFNQKDRKINHLWKIEERLKKQKVLIVL 299
Query: 339 DNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYR 398
+VD++ QL+ A F GSR+I+TT+D+ IL + + +YEV L A E+
Sbjct: 300 GDVDKVEQLEALANETRWFGPGSRIIVTTKDKQILVGHEINHIYEVKLPCRKTALEILCL 359
Query: 399 KGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKV 458
FK + ++V EV + + LPL +RV GS + ++ +W+ L RL + D KV
Sbjct: 360 YAFKQNVAPDDFMDVVVEVAELSGHLPLGLRVLGSHMRGKSKDRWKLELGRLTTSLDEKV 419
Query: 459 MDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIERSLITI 518
+L+IS++ LH DK +FLHIAC F GE + VK++L L +G+Q ++++SLI I
Sbjct: 420 EKILKISYDDLHIRDKALFLHIACMFNGENIDLVKQMLVNSDLDVSLGLQLLLDKSLIQI 479
Query: 519 R-NQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLDQ 577
++EI MH ++ +GK++V Q EPG L+ + ++L + G+ V I LD
Sbjct: 480 NDDREIVMHSLLLKMGKEVVCQH-SSEPGKRQFLFNTKETCNILSNNTGSEAVLGISLDT 538
Query: 578 NEDISEYPQLRAEGLSIMRGLIILILHH----QNFSGSLHFLSNNLQY------LLWHGY 627
+E I + MR L L ++ +N S LH L L Y L W Y
Sbjct: 539 SE-IQNDVFMSERVFEDMRNLKFLRFYNKKIDENPSLKLH-LPRGLNYLPAVRLLHWDSY 596
Query: 628 PFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFEGSRRL 687
P +PS F P LVEL M +S + +LWEG + L +LK +DLS S L E P+ + L
Sbjct: 597 PMKYIPSQFRPECLVELRMMHSKVVKLWEGTQTLAYLKTIDLSFSNNLVEVPDLSKAISL 656
Query: 688 ERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLSGCTKL 747
E L L GC +L ++ S+ L +L +L C L + L L SL VL + GC KL
Sbjct: 657 ETLCLEGCQSLAELPSSVLNLHRLKWLRLTMCEKLEVIPLH--INLASLEVLDMEGCLKL 714
Query: 748 ESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNL---TNIPLSV--- 801
+S P+ + +N+E + + + + SI +RLE L + CLNL +++P SV
Sbjct: 715 KSFPDIS--KNIERIFMKN-TGIEEIPPSISQWSRLESLDISGCLNLKIFSHVPKSVVYI 771
Query: 802 ----NNMESL------------LTLDFCGCL-KLKHLPLGLPSLSPFTLQSLIFLDLGFC 844
+ +E L L +D C L L LP + LS +SL + F
Sbjct: 772 YLTDSGIERLPDCIKDLTWLHYLYVDNCRKLVSLPELPSSIKILSAINCESLERISSSF- 830
Query: 845 SLSEVPHALGEIECLERLNLEG 866
+ P+A ++E + +N +G
Sbjct: 831 ---DCPNA--KVEFSKSMNFDG 847
Score = 63.2 bits (152), Expect = 7e-10
Identities = 59/201 (29%), Positives = 93/201 (45%), Gaps = 31/201 (15%)
Query: 720 SSLVSLDLGSLCVLYSLAVLHLSGCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGV 779
S +V L G+ + Y L + LS L P+ + +LE L ++ C SL+ + S+
Sbjct: 618 SKVVKLWEGTQTLAY-LKTIDLSFSNNLVEVPDLSKAISLETLCLEGCQSLAELPSSVLN 676
Query: 780 LTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFL 839
L RL++L L C L IPL +N + SL LD GCLKLK P +S ++ +
Sbjct: 677 LHRLKWLRLTMCEKLEVIPLHIN-LASLEVLDMEGCLKLK----SFPDISK-NIERIFMK 730
Query: 840 DLGFCSLSEVPHALGEIECLERLNLEG---------------------NNFVSLPSSLRW 878
+ G + E+P ++ + LE L++ G + LP ++
Sbjct: 731 NTG---IEEIPPSISQWSRLESLDISGCLNLKIFSHVPKSVVYIYLTDSGIERLPDCIKD 787
Query: 879 LSSLAYLNLAHCSKLEFLSEL 899
L+ L YL + +C KL L EL
Sbjct: 788 LTWLHYLYVDNCRKLVSLPEL 808
>At5g22690 disease resistance protein-like
Length = 1008
Score = 402 bits (1033), Expect = e-112
Identities = 277/794 (34%), Positives = 411/794 (50%), Gaps = 41/794 (5%)
Query: 48 RRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSR 107
R + YDVF SF G D R TF+ H L RK I VFKD+ +Q+ +S+ +L AIR+SR
Sbjct: 6 RNWLYDVFPSFSGEDVRVTFLSHFLKELDRKLISVFKDND-IQRSQSLDPELKLAIRDSR 64
Query: 108 VSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFH 167
++IVVFSKNYA S WCLDE+ I +C E+F Q V PVFY +DP VR Q+G E VF
Sbjct: 65 IAIVVFSKNYAASSWCLDELLEIVKCKEEFGQIVIPVFYGLDPCHVRKQSG--EFGIVFE 122
Query: 168 MLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALG-----RKFS 221
D + +W+RA+ +A G+ N E +E+I V+ L F
Sbjct: 123 NTCQTKTDDEIQKWRRALTDVANILGFHSSNWDNEATMVEDIANDVLAKLNLTTTSNDFE 182
Query: 222 GFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEA- 280
GF +GI+ + + +L L E ++ GIWG GIGKTT+A L+ RIS F+
Sbjct: 183 GF----VGIEGHIAKISLMLCL--ECKQVRMFGIWGPSGIGKTTIARALFSRISRHFQGS 236
Query: 281 ----RCFVENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEIS--GIVRDRLRSLKV 334
R FV ++Y G V ++ Q + +IS G+V +RL+ +KV
Sbjct: 237 VFLDRAFVSKSMEIYSGGNVDNYNAKLHLQGKFLSEILRAKDIKISNLGVVGERLKHMKV 296
Query: 335 LVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARE 394
L+ +D++D V L A P F GSR+I+ T+D+ + +G + YEV L ++ A E
Sbjct: 297 LIFIDDLDDQVVLDALASKPHWFGCGSRIIVITKDKQFFRAHGIGLFYEVGLPSDKLALE 356
Query: 395 LFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNP 454
+F + F+ ++ EL EV K + LPLA+ V GS L R+ W D L RL+
Sbjct: 357 MFSQSAFRQNSPPPGFTELASEVSKRSGNLPLALNVLGSHLRGRDKEDWIDMLPRLRKGL 416
Query: 455 DNKVMDVLQISFEGL-HSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNMIER 513
D K+ +L++ ++ L + +DK IF IAC F G + +Y+K +L L IG++N++++
Sbjct: 417 DGKIEKILRVGYDELSNKDDKAIFRLIACLFNGAEISYIKLLLADSNLGVTIGLKNLVDK 476
Query: 514 SLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAI 573
SLI I + MH M+Q++G++IVR+Q EPG L VL GT KV I
Sbjct: 477 SLIRIGCDTVEMHSMLQEMGREIVREQSIYEPGEREFLVDSTDILDVLNDNTGTKKVLGI 536
Query: 574 VLDQNE----DISEYPQLRAEGLSIMRGLIILILHHQNFSGSLH-------FLSNNLQYL 622
D +E I + R L +R L Q+ LH F L+ L
Sbjct: 537 SFDMSEIEELHIHKRAFKRMPNLRFLR--FYKKLGKQSKEARLHLQEGFDKFFPPKLKLL 594
Query: 623 LWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFE 682
W YP +PSNF LV L M +S +++LW+G + L L+ M L SK L E P+
Sbjct: 595 SWDDYPMRRMPSNFHAGYLVVLRMQHSKLEKLWQGVQPLTCLREMQLWGSKKLKEIPDLS 654
Query: 683 GSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLS 742
+ LE L L C++L+++ SI L KL L + C L L L SL L L
Sbjct: 655 LATNLETLYLNDCSSLVELPSSIKNLNKLWDLGMKGCEKLELLPTD--INLKSLYRLDLG 712
Query: 743 GCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVN 802
C++L+S P+ + N+ L +++ ++ V I +RL+ L +R+C L I +++
Sbjct: 713 RCSRLKSFPDIS--SNISELYLNR-TAIEEVPWWIQKFSRLKRLRMRECKKLKCISPNIS 769
Query: 803 NMESLLTLDFCGCL 816
++ L LDF C+
Sbjct: 770 KLKHLEMLDFSNCI 783
Score = 55.1 bits (131), Expect = 2e-07
Identities = 41/122 (33%), Positives = 56/122 (45%), Gaps = 10/122 (8%)
Query: 777 IGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSL 836
+ + T LE L L DC +L +P S+ N+ L L GC KL+ LP + L+SL
Sbjct: 653 LSLATNLETLYLNDCSSLVELPSSIKNLNKLWDLGMKGCEKLELLP------TDINLKSL 706
Query: 837 IFLDLGFCS-LSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCSKLEF 895
LDLG CS L P I L L +P ++ S L L + C KL+
Sbjct: 707 YRLDLGRCSRLKSFPDISSNI---SELYLNRTAIEEVPWWIQKFSRLKRLRMRECKKLKC 763
Query: 896 LS 897
+S
Sbjct: 764 IS 765
>At5g41540 disease resistance protein-like
Length = 1038
Score = 399 bits (1025), Expect = e-111
Identities = 295/853 (34%), Positives = 443/853 (51%), Gaps = 67/853 (7%)
Query: 36 ASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESI 95
ASSST V KY VF SF GSD R F+ HL H KGI FKD +++++G+ I
Sbjct: 2 ASSSTHV-------RKYHVFPSFHGSDVRRKFLSHLRFHFAIKGIVAFKD-QEIERGQRI 53
Query: 96 SAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRN 155
+L+QAIR SRVS+VV SKNY S WCLDE+ I +C ED +Q V P+FY++DPS VR
Sbjct: 54 GPELVQAIRESRVSLVVLSKNYPSSSWCLDELVEILKCKEDQEQIVMPIFYEIDPSDVRK 113
Query: 156 QNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIE 214
Q+G + AF + + + RW A+ A G N E IE IV V
Sbjct: 114 QSGDFGKAFGKTCVGKTKEVKQ--RWTNALTEAANIGGEHSLNWTDEAEMIEKIVADVSN 171
Query: 215 ALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRI 274
L S ++++G+ + L++LL LNS+ + ++IGIWG GIGKTT+A LY+++
Sbjct: 172 KLNVIPSRDFEEMVGLDAHLRKLDSLLCLNSD--EVKMIGIWGPAGIGKTTIARALYNQL 229
Query: 275 SHLFEARCFVENVSKVYRDGGV------TAVQKQVLRQTVDEMNLETYSPSEISGIVRDR 328
S F+ +CF+ N+ Y+ GV +Q Q+L + +++ +++T + G ++D
Sbjct: 230 STNFQFKCFMGNLKGSYKSIGVDNYDWKLNLQNQLLSKILNQNDVKT----DHLGGIKDW 285
Query: 329 LRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILK--VYGAHIVYEVPL 386
L KVL+V+D+VD L QL A P F GSR+I+TT+D+ I+K + + Y V
Sbjct: 286 LEDKKVLIVIDDVDDLEQLLALAKEPSWFGSGSRIIVTTKDKTIMKTLLVNDNNFYHVGY 345
Query: 387 MNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDA 446
N A E+ F+ EL +V LPL + V GS L ++ +W+
Sbjct: 346 PTNKVALEILCLSAFQKSFPRDGFEELARKVAYLCGNLPLCLSVVGSSLRGQSKHRWKLQ 405
Query: 447 LDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIG 506
DRL+ + D K+ DVL+ ++E L +++ +FLHIACFF + VK +L L G
Sbjct: 406 SDRLETSLDRKIEDVLKSAYEKLSKKEQVLFLHIACFFNNTYISVVKTLLADSNLDVRNG 465
Query: 507 IQNMIERSLITI-RNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEM 565
++ + ++ L+ I R I MH ++Q LG+ IV +Q +EP L + VL +E
Sbjct: 466 LKTLADKCLVHISRVDRIFMHPLLQQLGRYIVLEQ-SDEPEKRQFLVEAEEIRDVLANET 524
Query: 566 GTNKVKAIVLDQNEDISEYPQLRAEGLSIMRGLIILILHHQNFSG--SLHFLSN-----N 618
GT V I D ++ +SE+ + MR L L ++ ++ S +L + +
Sbjct: 525 GTGSVLGISFDMSK-VSEF-SISGRAFEAMRNLRFLRIYRRSSSKKVTLRIVEDMKYLPR 582
Query: 619 LQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTET 678
L+ L W YP SLP F+P RLV L+MP+S++++LW G + L LK +DLS S+ L E
Sbjct: 583 LRLLHWEHYPRKSLPRRFQPERLVVLHMPHSNLEKLWGGIQSLTNLKNIDLSFSRKLKEI 642
Query: 679 PNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAV 738
PN + LE L L C++L+++ SI L KL L C L + L SL
Sbjct: 643 PNLSNATNLETLTLIKCSSLVELPSSISNLQKLKALMMFGCKMLKVVPTN--INLVSLEK 700
Query: 739 LHLSGCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSL--RDCLNLTN 796
+ ++ C++L S P+ + N++ LD+ + + +RL+ LSL R LT
Sbjct: 701 VSMTLCSQLSSFPDIS--RNIKSLDVGKTKIEEVPPSVVKYWSRLDQLSLECRSLKRLTY 758
Query: 797 IP-------LSVNNMES----------LLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFL 839
+P LS +++E+ L TL C KL LP GLP SL FL
Sbjct: 759 VPPSITMLSLSFSDIETIPDCVIRLTRLRTLTIKCCRKLVSLP-GLP-------PSLEFL 810
Query: 840 DLGFCSLSEVPHA 852
C E H+
Sbjct: 811 CANHCRSLERVHS 823
>At5g46270 disease resistance protein-like
Length = 1145
Score = 399 bits (1024), Expect = e-111
Identities = 292/906 (32%), Positives = 451/906 (49%), Gaps = 84/906 (9%)
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ +R + YDVF+SFRG D R TF H L RK I F+D++ +++ S+ L QAI
Sbjct: 4 SSSSRNWLYDVFLSFRGGDVRVTFRSHFLKELDRKLITAFRDNE-IERSHSLWPDLEQAI 62
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENA 163
++SR+++V+FSKNYA S WCL+E+ I C + + V PVFY VDPS VR+Q G +
Sbjct: 63 KDSRIAVVIFSKNYASSSWCLNELLEIVNCND---KIVIPVFYGVDPSQVRHQIGDFGKI 119
Query: 164 FVFHMLRFKHDADRV-DRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALGRKFS 221
F K ++V ++WK+A+ +A G+D E + IE I V+ L
Sbjct: 120 FE---KTCKRQTEQVKNQWKKALTDVANMLGFDSATWDDEAKMIEEIANDVLAKLLLTTP 176
Query: 222 GFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEAR 281
++ +GI+ + + LLKL +E + +++GIWG GIGKTT+A L++++S F
Sbjct: 177 KDFENFVGIEDHIANMSVLLKLEAE--EVRMVGIWGSSGIGKTTIARALFNQLSRHFPVS 234
Query: 282 CFVENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEIS--------------GIVRD 327
F++ VY+ + + R D+ N++ + ++ G++ +
Sbjct: 235 KFIDRAF-VYKSREIFS------RANPDDHNMKLHLQEKLLSEILRMPDIKIDHLGVLGE 287
Query: 328 RLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLM 387
RL+ KVL+++D++D V L F GSR+I T ++H L+ + +YEV L
Sbjct: 288 RLQHQKVLIIVDDLDDQVILDSLVGQTQWFGSGSRIIAVTNNKHFLRAHEIDHIYEVSLP 347
Query: 388 NNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDAL 447
A + + F+ + LV +V ++ LPL + V GS+L R+ W + L
Sbjct: 348 TQQHALAMLCQSAFRKKSPPEGFEMLVVQVARHVDSLPLGLNVLGSYLRGRDKEYWMEML 407
Query: 448 DRLKNNPDNKVMDVLQISFEGLHS-EDKEIFLHIACFFKGEKENYVKRILDACGLHPHIG 506
RL+N +K+ +L+IS++GL S EDK IF HIAC F + + +L G+ +IG
Sbjct: 408 PRLENGLHDKIEKILRISYDGLGSEEDKAIFRHIACLFNHMEVTTITSLLTDLGI--NIG 465
Query: 507 IQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMG 566
++N++++S+I +R + MH M+Q++G+KIVR Q ++PG L VL +G
Sbjct: 466 LKNLVDKSIIHVRRGCVEMHRMLQEMGRKIVRTQSIDKPGKREFLVDPNDISDVLSEGIG 525
Query: 567 TNKVKAIVLDQNEDISEYPQLRA-EGLSIMRGLIILILHHQNFS--------GSLHFLSN 617
T KV I L+ E Y A +G+S +R L + +NF SL +L
Sbjct: 526 TQKVLGISLNTGEIDELYVHESAFKGMSNLR---FLEIDSKNFGKAGRLYLPESLDYLPP 582
Query: 618 NLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTE 677
L+ L W +P +PSNF P LV L MP S + +LWEG L LK MD+ S L E
Sbjct: 583 RLKLLCWPNFPMRCMPSNFRPENLVTLKMPNSKLHKLWEGVASLTCLKEMDMVGSSNLKE 642
Query: 678 TPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLA 737
P+ LE L L C +L+++ SI L KL L E C SL L G L SL
Sbjct: 643 IPDLSMPTNLEILKLGFCKSLVELPSSIRNLNKLLKLDMEFCHSLEILPTG--FNLKSLD 700
Query: 738 VLHLSGCTKLESTPNFT---------GVENLEYLDIDQCVSLS-TVDQSIG--------- 778
L+ C++L + P F+ G E+ +++ V LS + ++S G
Sbjct: 701 HLNFRYCSELRTFPEFSTNISVLMLFGTNIEEFPNLENLVELSLSKEESDGKQWDGVKPL 760
Query: 779 ------VLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFT 832
+ L+ L L + +L +P S N+ L L C L+ LP G+
Sbjct: 761 TPFLEMLSPTLKSLKLENIPSLVELPSSFQNLNQLKELSITYCRNLETLPTGI------N 814
Query: 833 LQSLIFLDLGFCS-LSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCS 891
L+SL +L CS L P I LNLE +P + +L L + CS
Sbjct: 815 LKSLNYLCFKGCSQLRSFPEISTNISV---LNLEETGIEEVPWQIENFFNLTKLTMRSCS 871
Query: 892 KLEFLS 897
KL+ LS
Sbjct: 872 KLKCLS 877
Score = 55.5 bits (132), Expect = 1e-07
Identities = 62/215 (28%), Positives = 96/215 (43%), Gaps = 27/215 (12%)
Query: 616 SNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYS-SIQRLWEGRKDL-PFLKRMDLSNSK 673
S N+ L+ G P N E LVEL++ S + W+G K L PFL+ +
Sbjct: 717 STNISVLMLFGTNIEEFP-NLE--NLVELSLSKEESDGKQWDGVKPLTPFLEML------ 767
Query: 674 YLTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVL 733
S L+ L L +L+++ S L +L LS C +L +L G L
Sbjct: 768 ----------SPTLKSLKLENIPSLVELPSSFQNLNQLKELSITYCRNLETLPTG--INL 815
Query: 734 YSLAVLHLSGCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLN 793
SL L GC++L S P + N+ L++++ + V I L L++R C
Sbjct: 816 KSLNYLCFKGCSQLRSFPEIS--TNISVLNLEE-TGIEEVPWQIENFFNLTKLTMRSCSK 872
Query: 794 LTNIPLSVNNMESLLTLDFCGCLKLKHLPL-GLPS 827
L + L++ M++L +DF C L + L G PS
Sbjct: 873 LKCLSLNIPKMKTLWDVDFSDCAALTVVNLSGYPS 907
Score = 40.4 bits (93), Expect = 0.005
Identities = 33/101 (32%), Positives = 53/101 (51%), Gaps = 11/101 (10%)
Query: 801 VNNMESLLTLDFCGCLKLKHLP-LGLPSLSPFTLQSLIFLDLGFC-SLSEVPHALGEIEC 858
V ++ L +D G LK +P L +P+ +L L LGFC SL E+P ++ +
Sbjct: 623 VASLTCLKEMDMVGSSNLKEIPDLSMPT-------NLEILKLGFCKSLVELPSSIRNLNK 675
Query: 859 LERLNLEG-NNFVSLPSSLRWLSSLAYLNLAHCSKLEFLSE 898
L +L++E ++ LP+ L SL +LN +CS+L E
Sbjct: 676 LLKLDMEFCHSLEILPTGFN-LKSLDHLNFRYCSELRTFPE 715
>At5g46260 disease resistance protein-like
Length = 1205
Score = 397 bits (1019), Expect = e-110
Identities = 289/905 (31%), Positives = 451/905 (48%), Gaps = 77/905 (8%)
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ +R + YDVF+SFRG D R TF H RK I F+D++ +++ S+ L QAI
Sbjct: 4 SSSSRNWLYDVFLSFRGGDVRVTFRSHFLKEFDRKLITAFRDNE-IERSHSLWPDLEQAI 62
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENA 163
+ SR+++VVFSKNYA S WCL+E+ I C + + + PVFY VDPS VR Q G +
Sbjct: 63 KESRIAVVVFSKNYASSSWCLNELLEIVNCND---KIIIPVFYGVDPSQVRYQIGEFGK- 118
Query: 164 FVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRN-KPEFREIENIVEAVIEALGRKFSG 222
+F + + ++WK+A+ +A G+D E + IE I V+ L S
Sbjct: 119 -IFEKTCKRQTEEVKNQWKKALTHVANMLGFDSSKWDDEAKMIEEIANDVLRKLLLTTSK 177
Query: 223 FADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARC 282
+D +G++ + + LL L S+ + +++GIWG GIGKTT+A L++ + F+ R
Sbjct: 178 DFEDFVGLEDHIANMSALLDLESK--EVKMVGIWGSSGIGKTTIARALFNNLFRHFQVRK 235
Query: 283 FVENVSKVYRDGGVTA------------VQKQVLRQTVDEMNLETYSPSEISGIVRDRLR 330
F++ S Y+ + + +Q+ L + + N++ + G++ +RL+
Sbjct: 236 FIDR-SFAYKSREIHSSANPDDHNMKLHLQESFLSEILRMPNIKI----DHLGVLGERLQ 290
Query: 331 SLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNN 390
KVL+++D+VD V L F GSR+I+ T ++H L +G +YEV L
Sbjct: 291 HQKVLIIIDDVDDQVILDSLVGKTQWFGNGSRIIVVTNNKHFLTAHGIDRMYEVSLPTEE 350
Query: 391 DARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRL 450
A + + FK + LV +V +YA LPL ++V GS+L ++ W D L RL
Sbjct: 351 HALAMLCQSAFKKKSPPEGFEMLVVQVARYAGSLPLVLKVLGSYLSGKDKEYWIDMLPRL 410
Query: 451 KNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIGIQNM 510
+N ++K+ +L+IS++GL SED+ IF HIAC F + +K +L ++G+QN+
Sbjct: 411 QNGLNDKIERILRISYDGLESEDQAIFRHIACIFNHMEVTTIKSLLANSIYGANVGLQNL 470
Query: 511 IERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKV 570
+++S+I +R + MH ++Q++G+KIVR Q +P L VL + T KV
Sbjct: 471 VDKSIIHVRWGHVEMHPLLQEMGRKIVRTQSIGKPRKREFLVDPNDICDVLSEGIDTQKV 530
Query: 571 KAIVLDQNEDISEYPQLRAEGLSIMRGLIILIL--------HHQNFSGSLHFLSNNLQYL 622
I L+ ++ I E + MR L L + + + S +L L+ L
Sbjct: 531 LGISLETSK-IDEL-CVHESAFKRMRNLRFLKIGTDIFGEENRLHLPESFDYLPPTLKLL 588
Query: 623 LWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLKRMDLSNSKYLTETPNFE 682
W +P +PSNF P LV L M S + +LWEG L LK MDL S L E P+
Sbjct: 589 CWSEFPMRCMPSNFCPKNLVTLKMTNSKLHKLWEGAVPLTCLKEMDLDGSVNLKEIPDLS 648
Query: 683 GSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGSLCVLYSLAVLHLS 742
+ LE L+ C +L+++ I L KL L+ C+SL +L G L SL + +
Sbjct: 649 MATNLETLNFENCKSLVELPSFIQNLNKLLKLNMAFCNSLETLPTG--FNLKSLNRIDFT 706
Query: 743 GCTKLESTPNF-----------TGVENLE---YLD--IDQCVSLSTVD--QSIGVLTRLE 784
C+KL + P+F T +E L +L+ ID +S +D Q GV+ L+
Sbjct: 707 KCSKLRTFPDFSTNISDLYLTGTNIEELPSNLHLENLIDLRISKKEIDGKQWEGVMKPLK 766
Query: 785 -----------FLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTL 833
L L++ NL +P S N+ L LD C L+ LP G+ L
Sbjct: 767 PLLAMLSPTLTSLQLQNIPNLVELPCSFQNLIQLEVLDITNCRNLETLPTGI------NL 820
Query: 834 QSLIFLDLGFCS-LSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCSK 892
QSL L CS L P I LNLE +P + S+L L++ CS+
Sbjct: 821 QSLDSLSFKGCSRLRSFPEISTNI---SSLNLEETGIEEVPWWIDKFSNLGLLSMDRCSR 877
Query: 893 LEFLS 897
L+ +S
Sbjct: 878 LKCVS 882
Score = 65.1 bits (157), Expect = 2e-10
Identities = 80/261 (30%), Positives = 113/261 (42%), Gaps = 42/261 (16%)
Query: 616 SNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQ-RLWEG-RKDL--------PFLK 665
S N+ L G LPSN L++L + I + WEG K L P L
Sbjct: 718 STNISDLYLTGTNIEELPSNLHLENLIDLRISKKEIDGKQWEGVMKPLKPLLAMLSPTLT 777
Query: 666 RMDLSNSKYLTETP-NFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVS 724
+ L N L E P +F+ +LE LD+T C NL + I L + L LSF+ CS L S
Sbjct: 778 SLQLQNIPNLVELPCSFQNLIQLEVLDITNCRNLETLPTGINLQS-LDSLSFKGCSRLRS 836
Query: 725 LDLGSLCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTVDQSIGVLTRL 783
S +++ L+L T +E P + NL L +D+C L V I L RL
Sbjct: 837 FPEIST----NISSLNLEE-TGIEEVPWWIDKFSNLGLLSMDRCSRLKCVSLHISKLKRL 891
Query: 784 EFLSLRDCLNLTNIPL---------SVNNME--SLLTLDFCGCLKLKHLPLGLPSLSPFT 832
+ +DC LT + L NN++ S + LDF C +L P T
Sbjct: 892 GKVDFKDCGALTIVDLCGCPIGMEMEANNIDTVSKVKLDFRDCF----------NLDPET 941
Query: 833 L---QSLIFLDLGFCSLSEVP 850
+ +S+IF + F E+P
Sbjct: 942 VLHQESIIFKYMLFPGKEEMP 962
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,208,490
Number of Sequences: 26719
Number of extensions: 924633
Number of successful extensions: 5860
Number of sequences better than 10.0: 441
Number of HSP's better than 10.0 without gapping: 249
Number of HSP's successfully gapped in prelim test: 195
Number of HSP's that attempted gapping in prelim test: 2468
Number of HSP's gapped (non-prelim): 1403
length of query: 949
length of database: 11,318,596
effective HSP length: 109
effective length of query: 840
effective length of database: 8,406,225
effective search space: 7061229000
effective search space used: 7061229000
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0388b.1


