

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
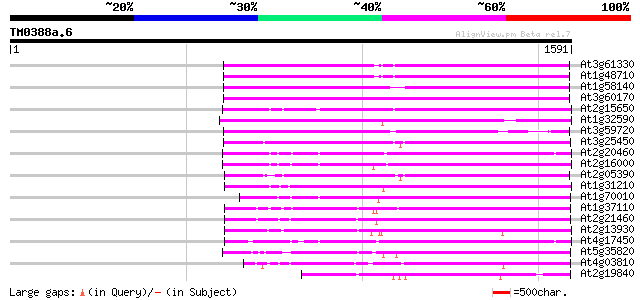
Query= TM0388a.6
(1591 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g61330 copia-type polyprotein 738 0.0
At1g48710 hypothetical protein 733 0.0
At1g58140 hypothetical protein 732 0.0
At3g60170 putative protein 722 0.0
At2g15650 putative retroelement pol polyprotein 694 0.0
At1g32590 hypothetical protein, 5' partial 641 0.0
At3g59720 copia-type reverse transcriptase-like protein 632 0.0
At3g25450 hypothetical protein 622 e-178
At2g20460 putative retroelement pol polyprotein 603 e-172
At2g16000 putative retroelement pol polyprotein 595 e-170
At2g05390 putative retroelement pol polyprotein 566 e-161
At1g31210 putative reverse transcriptase 566 e-161
At1g70010 hypothetical protein 562 e-160
At1g37110 539 e-153
At2g21460 putative retroelement pol polyprotein 530 e-150
At2g13930 putative retroelement pol polyprotein 528 e-149
At4g17450 retrotransposon like protein 513 e-145
At5g35820 copia-like retrotransposable element 503 e-142
At4g03810 putative retrotransposon protein 478 e-134
At2g19840 copia-like retroelement pol polyprotein 454 e-127
>At3g61330 copia-type polyprotein
Length = 1352
Score = 738 bits (1906), Expect = 0.0
Identities = 398/992 (40%), Positives = 565/992 (56%), Gaps = 26/992 (2%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICV----DSSPC 661
++ WYLDSG S HM G + MF EL G V G K ++ G G I +
Sbjct: 331 ENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF 390
Query: 662 IYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLS 721
I NV + + N+LS+ QL +KGYD+ + Q + KN ++ + +
Sbjct: 391 ISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIR 450
Query: 722 ELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQK 781
AQ +K + EE W+WH R GH + + LS+ +VRGLP + + +CE C
Sbjct: 451 NDIAQCLK--MCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQ-VCEGCLL 507
Query: 782 GKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTR 841
GK K+ F ++ +PLEL+H D+ GP+K +S+G Y ++ +DD+SR TWV FL
Sbjct: 508 GKQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKE 567
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K E +F F A V+ E I +RSD GGEF + +F + GI + PR+PQQ
Sbjct: 568 KSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQ 627
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNI 961
NGVVERKNRT+ EMAR+ML+ + K W EAV A Y+ NR + + KTP E W
Sbjct: 628 NGVVERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGR 687
Query: 962 KPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIH 1021
KP +S+ FG + + ++ K D KS K + +GY SKG++ YN D K S +
Sbjct: 688 KPGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRN 747
Query: 1022 VRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRIT 1081
+ FD++ + D + N D P E +E EP E PS+ T + T
Sbjct: 748 IVFDEEGEWDWNS----------NEEDYNFFPH-FEEDEPEPTREEPPSEEPTTPPTSPT 796
Query: 1082 AAHPMELIMGNKDEPVRTRSAFRPSEETL----LSLKGLVSLIEPKSIDEALQDKDWILA 1137
++ E + + R RS E T L+L L + EP +A++ K W A
Sbjct: 797 SSQIEE---SSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQKAIEKKTWRNA 853
Query: 1138 MEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGI 1197
M+EE+ KND W L P IG KWV++ K N KG+V R KARLVA+GYSQ+ GI
Sbjct: 854 MDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRVGI 913
Query: 1198 DYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKK 1257
DY E FAP+ARLE +RL+IS + + +HQMDVKSAFLNG + EEVY+ QP G+ + +
Sbjct: 914 DYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGE 973
Query: 1258 PDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVD 1317
D V +LKK LYGLKQAPRAW R+ + E +F++ + L+ K K+DIL+ +YVD
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVD 1033
Query: 1318 DIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKK 1377
D+IF N S+ +EF K M EFEM+ +G ++Y+LGI+V Q G +I Q Y KE+LKK
Sbjct: 1034 DLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKK 1093
Query: 1378 FNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARF 1437
F + +S TPM L K++ V ++ ++GSL YLT +RPDIL++V + +R+
Sbjct: 1094 FKIDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRY 1153
Query: 1438 QSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQ 1497
P TH A KRILRY+K T N GL Y TS+YKL GY D+D+ GD +RKSTSG
Sbjct: 1154 MEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVF 1213
Query: 1498 FLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQI-LESNIPIYCDN 1556
++G +W SK+Q + LST EAEY++A C +W+++ L++ + E I+ DN
Sbjct: 1214 YIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDN 1273
Query: 1557 TAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
+AI+L+KNP+ H R+KHI+ +YH+IR+ V K
Sbjct: 1274 KSAIALAKNPVFHDRSKHIDTRYHYIRECVSK 1305
>At1g48710 hypothetical protein
Length = 1352
Score = 733 bits (1892), Expect = 0.0
Identities = 396/992 (39%), Positives = 562/992 (55%), Gaps = 26/992 (2%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICV----DSSPC 661
++ WYLDSG S HM G + MF EL G V G K ++ G G I +
Sbjct: 331 ENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF 390
Query: 662 IYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLS 721
I NV + + N+LS+ QL +KGYD+ + Q + KN ++ + +
Sbjct: 391 ISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIR 450
Query: 722 ELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQK 781
AQ +K + EE W+WH R GH + + LS+ +VRGLP + + +CE C
Sbjct: 451 NDIAQCLK--MCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQ-VCEGCLL 507
Query: 782 GKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTR 841
GK K+ F ++ + LEL+H D+ GP+K +S+G Y ++ +DD+SR TWV FL
Sbjct: 508 GKQFKMSFPKESSSRAQKSLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKE 567
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K E +F F A V+ E I +RSD GGEF + +F + GI + PR+PQQ
Sbjct: 568 KSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQ 627
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNI 961
NGV ERKNRT+ EMAR+ML+ + K W EAV A Y+ NR + + KTP E W
Sbjct: 628 NGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGR 687
Query: 962 KPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIH 1021
K +S+ FG + + ++ K D KS K + +GY SKG++ YN D K S +
Sbjct: 688 KSGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRN 747
Query: 1022 VRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRIT 1081
+ FD++ + D + N D P E +E EP E PS+ T + T
Sbjct: 748 IVFDEEGEWDWNS----------NEEDYNFFPH-FEEDEPEPTREEPPSEEPTTPPTSPT 796
Query: 1082 AAHPMELIMGNKDEPVRTRSAFRPSEETL----LSLKGLVSLIEPKSIDEALQDKDWILA 1137
++ E + + R RS E T L+L L + EP EA++ K W A
Sbjct: 797 SSQIEE---SSSERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQEAIEKKTWRNA 853
Query: 1138 MEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGI 1197
M+EE+ KND W L P IG KWV++ K N KG+V R KARLVA+GY Q+ GI
Sbjct: 854 MDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKGEVERYKARLVAKGYIQRAGI 913
Query: 1198 DYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKK 1257
DY E FAP+ARLE +RL+IS + + +HQMDVKSAFLNG + EEVY+ QP G+ + +
Sbjct: 914 DYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGE 973
Query: 1258 PDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVD 1317
D V +LKK+LYGLKQAPRAW R+ + E +F++ + L+ K K+DIL+ +YVD
Sbjct: 974 EDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVD 1033
Query: 1318 DIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKK 1377
D+IF N S+ +EF K M EFEM+ +G ++Y+LGI+V Q G +I Q Y KE+LKK
Sbjct: 1034 DLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKK 1093
Query: 1378 FNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARF 1437
F M +S TPM L K++ V ++ ++GSL YLT +RPDIL++V + +R+
Sbjct: 1094 FKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRY 1153
Query: 1438 QSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQ 1497
P TH A KRILRY+K T N GL Y TS+YKL GY D+D+ GD +RKSTSG
Sbjct: 1154 MEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVF 1213
Query: 1498 FLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQI-LESNIPIYCDN 1556
++G +W SK+Q + LST EAEY++A C +W+++ L++ + E I+ DN
Sbjct: 1214 YIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDN 1273
Query: 1557 TAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
+AI+L+KNP+ H R+KHI+ +YH+IR+ V K
Sbjct: 1274 KSAIALAKNPVFHDRSKHIDTRYHYIRECVSK 1305
>At1g58140 hypothetical protein
Length = 1320
Score = 732 bits (1889), Expect = 0.0
Identities = 388/988 (39%), Positives = 550/988 (55%), Gaps = 50/988 (5%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICV----DSSPC 661
++ WYLDSG S HM G + MF EL G V G K ++ G G I +
Sbjct: 331 ENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF 390
Query: 662 IYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLS 721
I NV + + N+LS+ QL +KGYD+ + Q + KN ++ + +
Sbjct: 391 ISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIR 450
Query: 722 ELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQK 781
AQ +K + EE W+WH R GH + + LS+ +VRGLP + + +CE C
Sbjct: 451 NDIAQCLK--MCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQ-VCEGCLL 507
Query: 782 GKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTR 841
GK K+ F ++ +PLEL+H D+ GP+K +S+G Y ++ +DD+SR TWV FL
Sbjct: 508 GKQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKE 567
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K E +F F A V+ E I +RSD GGEF + +F + GI + PR+PQQ
Sbjct: 568 KSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQ 627
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNI 961
NGV ERKNRT+ EMAR+ML+ + K W EAV A Y+ NR + + KTP E W
Sbjct: 628 NGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGR 687
Query: 962 KPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIH 1021
KP +S+ FG + + ++ K D KS K + +GY SKG++ YN D K S +
Sbjct: 688 KPGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRN 747
Query: 1022 VRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRIT 1081
+ FD++ + D + E + DK + E P E+ P+ SQ +K
Sbjct: 748 IVFDEEGEWDWNSNEEDYNFFPHFEEDKPEPTREEPPSEEPTTPPTSPTSSQIEEKC--- 804
Query: 1082 AAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEE 1141
EP EA++ K W AM+EE
Sbjct: 805 ---------------------------------------EPMDFQEAIEKKTWRNAMDEE 825
Query: 1142 LNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTE 1201
+ KND W L P IG KWV++ K N KG+V R KARLVA+GYSQ+ GIDY E
Sbjct: 826 IKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGIDYDE 885
Query: 1202 TFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHV 1261
FAP+ARLE +RL+IS + + +HQMDVKSAFLNG + EEVY+ QP G+ + + D V
Sbjct: 886 VFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKV 945
Query: 1262 FKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIF 1321
+LKK+LYGLKQAPRAW R+ + E +F++ + L+ K K+DIL+ +YVDD+IF
Sbjct: 946 LRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIF 1005
Query: 1322 GSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNML 1381
N S+ +EF K M EFEM+ +G ++Y+LGI+V Q G +I Q Y KE+LKKF M
Sbjct: 1006 TGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMD 1065
Query: 1382 ESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDP 1441
+S TPM L K++ V ++ ++GSL YLT +RPDIL++V + +R+ P
Sbjct: 1066 DSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHP 1125
Query: 1442 RETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGS 1501
TH A KRILRY+K T N GL Y TS+YKL GY D+D+ GD +RKSTSG ++G
Sbjct: 1126 TTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGD 1185
Query: 1502 NLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQI-LESNIPIYCDNTAAI 1560
+W SK+Q + LST EAEY++A C +W+++ L++ + E I+ DN +AI
Sbjct: 1186 TAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAI 1245
Query: 1561 SLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
+L+KNP+ H R+KHI+ +YH+IR+ V K
Sbjct: 1246 ALAKNPVFHDRSKHIDTRYHYIRECVSK 1273
>At3g60170 putative protein
Length = 1339
Score = 722 bits (1863), Expect = 0.0
Identities = 400/1001 (39%), Positives = 581/1001 (57%), Gaps = 22/1001 (2%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSS---PCI 662
+ + W+LDSGCS HMTG + F EL+ V G + + +VG G++ V + I
Sbjct: 296 RDEVWFLDSGCSNHMTGSKEWFSELEEGFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQVI 355
Query: 663 YNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSE 722
V V L +NLLS+ QL ++G ++ +C+ G+++ + N ++ + S+
Sbjct: 356 PEVYYVPELRNNLLSLGQLQERGLAILIRDGTCKVYHPSKGAIMETNMSGNRMFFLLASK 415
Query: 723 LEAQNVKCLLS---VNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEAC 779
+ +N CL + +++E +WH R GH + + L+ +V GLP LK A+ +C C
Sbjct: 416 PQ-KNSLCLQTEEVMDKENHLWHCRFGHLNQEGLKLLAHKKMVIGLPILK-ATKEICAIC 473
Query: 780 QKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFL 839
GK + K +S L+L+H D+ GP+ S GKRY + +DD++R TWV FL
Sbjct: 474 LTGKQHRESMSKKTSWKSSTQLQLVHSDICGPITPISHSGKRYILSFIDDFTRKTWVYFL 533
Query: 840 TRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTP 899
K E+ A F F A V+ E + +R+D GGEF +++F S+GI+ + TP
Sbjct: 534 HEKSEAFATFKIFKASVEKEIGAFLTCLRTDRGGEFTSNEFGEFCRSHGISRQLTAAFTP 593
Query: 900 QQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWK 959
QQNGV ERKNRT+ R+ML E + K FW+EA + +IQNR + TP E W
Sbjct: 594 QQNGVAERKNRTIMNAVRSMLSERQVPKMFWSEATKWSVHIQNRSPTAAVEGMTPEEAWS 653
Query: 960 NIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEES 1019
KP + YF FGC+ YV + K D KS KC+ LG SE SK +R Y+ K I S
Sbjct: 654 GRKPVVEYFRVFGCIGYVHIPDQKRSKLDDKSKKCVFLGVSEESKAWRLYDPVMKKIVIS 713
Query: 1020 IHVRFDD--KLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSD---SQT 1074
V FD+ D DQ+ + K L D K E EP G + S
Sbjct: 714 KDVVFDEDKSWDWDQADVEAKEVTLECGDEDDEKNSEVVEPIAVASPNHVGSDNNVSSSP 773
Query: 1075 LKKSRITAAHPMELIMGNKDEPVRTRSAFRPSE----ETLLSLKGLVSLIE--PKSIDEA 1128
+ A P+ + + P + + E E LS+ L+ + E P D+A
Sbjct: 774 ILAPSSPAPSPVAAKVTRERRPPGWMADYETGEGEEIEENLSVMLLMMMTEADPIQFDDA 833
Query: 1129 LQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVA 1188
++DK W AME E+ KN+ W L P+ IG KWV++ KLNE G+V + KARLVA
Sbjct: 834 VKDKIWREAMEHEIESIVKNNTWELTTLPKGFTPIGVKWVYKTKLNEDGEVDKYKARLVA 893
Query: 1189 QGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQ 1248
+GY+Q GIDYTE FAP+ARL+ +R +++ S N + Q+DVKSAFL+G + EEVYV Q
Sbjct: 894 KGYAQCYGIDYTEVFAPVARLDTVRTILAISSQFNWEIFQLDVKSAFLHGELKEEVYVRQ 953
Query: 1249 PPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDD 1308
P GF E + + V+KL+K+LYGLKQAPRAWY R+ ++ L+ EF R + TLF KT +
Sbjct: 954 PEGFIREGEEEKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFTKTRVGN 1013
Query: 1309 ILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQS 1368
IL+V +YVDD+IF +++++C EF K M EFEMS +G++ +FLGI+V Q+ G +I Q
Sbjct: 1014 ILIVSLYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFICQR 1073
Query: 1369 KYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDIL 1428
+Y +E+L +F M ES K P+ P L K++ KV + +++ ++GSL+YLT +RPD++
Sbjct: 1074 RYAREVLARFGMDESNAVKNPIVPGTKLTKDENGEKVDETMFKQLVGSLMYLTVTRPDLM 1133
Query: 1429 FSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMY--KKTSEYKLSGYCDADYAGDR 1486
+ V L +RF S+PR +H A KRILRYLK T LG+ Y +K KL + D+DYAGD
Sbjct: 1134 YGVCLISRFMSNPRMSHWLAAKRILRYLKGTVELGIFYRRRKNRSLKLMAFTDSDYAGDL 1193
Query: 1487 TERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQIL 1546
+R+STSG + S + W SK+Q +ALST EAEYI+AA C+ Q +W++ LE
Sbjct: 1194 NDRRSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLGAE 1253
Query: 1547 E-SNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV 1586
E S I CDN++ I LSK+P+LH ++KHIEV++H++RD V
Sbjct: 1254 EKSATVINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLRDLV 1294
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 694 bits (1790), Expect = 0.0
Identities = 384/1007 (38%), Positives = 568/1007 (56%), Gaps = 38/1007 (3%)
Query: 605 LKHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICV---DSSPC 661
L+ W +DSGC+ HMT E R F + + + G G I V
Sbjct: 321 LREDVWLVDSGCTNHMTKEERYFSNINKSIKVPIRVRNGDIVMTAGKGDITVMTRHGKRI 380
Query: 662 IYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLS 721
I NV LV GL NLLS+ Q+ GY V F K C + +G + N + + +KI+LS
Sbjct: 381 IKNVFLVPGLEKNLLSVPQIISSGYWVRFQDKRC-IIQDANGKEIMNIEMTDKSFKIKLS 439
Query: 722 ELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQK 781
+E + + + E WH+RLGH S +++ Q+ LV GLP K + C+AC
Sbjct: 440 SVEEEAMTANVQTEE---TWHKRLGHVSNKRLQQMQDKELVNGLPRFKVTKET-CKACNL 495
Query: 782 GKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTR 841
GK ++ F ++ T LE++H D+ GP++ +SI G RY ++ +DDY+ WV FL +
Sbjct: 496 GKQSRKSFPKESQTKTREKLEIVHTDVCGPMQHQSIDGSRYYVLFLDDYTHMCWVYFLKQ 555
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K E+ A F F A V+ + C I +R E + GI + P +PQQ
Sbjct: 556 KSETFATFKKFKALVEKQSNCSIKTLRP----------MEVFCEDEGINRQVTLPYSPQQ 605
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNK-TPYELWKN 960
NG ERKNR+L EMAR+ML E + W EAV T+ Y+QNR+ + I + TP E W
Sbjct: 606 NGAAERKNRSLVEMARSMLVEQDLPLKLWAEAVYTSAYLQNRLPSKAIEDDVTPMEKWCG 665
Query: 961 IKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESI 1020
KPN+S+ FG +CYV + K DAK+ +L+GYS ++KG+R + + + +E S
Sbjct: 666 HKPNVSHLRIFGSICYVHIPDQKRRKLDAKAKCGILIGYSNQTKGYRVFLLEDEKVEVSR 725
Query: 1021 HVRF--DDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEE-----DEPEEEAGPSDSQ 1073
V F D K D D+ + V+K +SIN + + +E + D G + S
Sbjct: 726 DVVFQEDKKWDWDKQEEVKKTFVMSINDIQESRDQQETSSHDLSQIDDHANNGEGETSSH 785
Query: 1074 TL-----KKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKG-LVSLIEPKSIDE 1127
L ++ R T+ P + + + ++ ++E ++ LV+ EP++ DE
Sbjct: 786 VLSQVNDQEERETSESPKKY---KSMKEILEKAPRMENDEAAQGIEACLVANEEPQTYDE 842
Query: 1128 ALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLV 1187
A DK+W AM EE+ KN W LV KPE +VI KW+++ K + G+ V++KARLV
Sbjct: 843 ARGDKEWEEAMNEEIKVIEKNRTWKLVDKPEKKNVISVKWIYKIKTDASGNHVKHKARLV 902
Query: 1188 AQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVH 1247
A+G+SQ+ GIDY ETFAP++R + IR L++++ L+QMDVKSAFLNG + EEVYV
Sbjct: 903 ARGFSQEYGIDYLETFAPVSRYDTIRALLAYAAQMKWRLYQMDVKSAFLNGELEEEVYVT 962
Query: 1248 QPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKD 1307
QPPGF E K + V +L K+LYGLKQAPRAWYER+ S+ ++N F R D L+ K +
Sbjct: 963 QPPGFVIEGKEEKVLRLYKALYGLKQAPRAWYERIDSYFIQNGFARSMNDAALYSKKKGE 1022
Query: 1308 DILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQ 1367
D+L+V +YVDD+I N L F K M+ EFEM+ +G L YFLG++V+Q G ++ Q
Sbjct: 1023 DVLIVSLYVDDLIITGNNTHLINTFKKNMKDEFEMTDLGLLNYFLGMEVNQDDSGIFLSQ 1082
Query: 1368 SKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGK--VCQKLYRGMIGSLLYLTASRP 1425
KY +L+ KF M ES TP+ P + + K YR ++G LLYL ASRP
Sbjct: 1083 EKYANKLIDKFGMKESKSVSTPLTPQGKRKGVEGDDKEFADPTKYRRIVGGLLYLCASRP 1142
Query: 1426 DILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGD 1485
D++++ +R+ S P H KR+LRY+K T+N G+++ +L GY D+D+ G
Sbjct: 1143 DVMYASSYLSRYMSSPSIQHYQEAKRVLRYVKGTSNFGVLFTSKETPRLVGYSDSDWGGS 1202
Query: 1486 RTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQI 1545
++KST+G LG + W+S +Q T+A STAEAEYI+ + Q +W++ ED+ +
Sbjct: 1203 LEDKKSTTGYVFTLGLAMFCWQSCKQQTVAQSTAEAEYIAVCAATNQAIWLQRLFEDFGL 1262
Query: 1546 -LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
+ IPI CDN +AI++ +NP+ H R KHIE+KYHF+R+ KG++
Sbjct: 1263 KFKEGIPILCDNKSAIAIGRNPVQHRRTKHIEIKYHFVREAEHKGLI 1309
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 641 bits (1653), Expect = 0.0
Identities = 360/1018 (35%), Positives = 546/1018 (53%), Gaps = 54/1018 (5%)
Query: 594 LSMLQISFIAPLKHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGT 653
L M + I + Q W+LDSGCS HM G R F EL V G + + + G G
Sbjct: 242 LLMAHVEQIGDEEKQIWFLDSGCSNHMCGTREWFLELDSGFKQNVRLGDDRRMAVEGKGK 301
Query: 654 ICVDSS---PCIYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSK 710
+ ++ I +V V GL +NL S+ QL KG I C + + ++ +S
Sbjct: 302 LRLEVDGRIQVISDVYFVPGLKNNLFSVGQLQQKGLRFIIEGDVCEVWHKTEKRMVMHST 361
Query: 711 RKNN----IYKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLP 766
N ++ E + +CL + + +WH+R GH + + + L++ +V+GLP
Sbjct: 362 MTKNRMFVVFAAVKKSKETEETRCLQVIGKANNMWHKRFGHLNHQGLRSLAEKEMVKGLP 421
Query: 767 NLKFASD-ALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMI 825
+ A+C+ C KGK + ++ +++ L+L+H D+ GP+ S GKRY +
Sbjct: 422 KFDLGEEEAVCDICLKGKQIRESIPKESAWKSTQVLQLVHTDICGPINPASTSGKRYILN 481
Query: 826 IVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFD 885
+DD+SR W L+ K E+ F F A+V+ E ++V +RSD GGE+ + +F+
Sbjct: 482 FIDDFSRKCWTYLLSEKSETFQFFKEFKAEVERESGKKLVCLRSDRGGEYNSREFDEYCK 541
Query: 886 SYGIAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRIS 945
+GI + TPQQNGV ERKNR++ M R ML E + + FW EAV A YI NR
Sbjct: 542 EFGIKRQLTAAYTPQQNGVAERKNRSVMNMTRCMLMEMSVPRKFWPEAVQYAVYILNRSP 601
Query: 946 VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKG 1005
+ + + TP E W + KP++ + FG + Y L + K D KS KC++ G S+ SK
Sbjct: 602 SKALNDITPEEKWSSWKPSVEHLRIFGSLAYALVPYQKRIKLDEKSIKCVMFGVSKESKA 661
Query: 1006 FRFYNTDAKTIEESIHVRFDDKLDSD-QSKLVEKFADLSINVSDKGKAPEEA-------- 1056
+R Y+ I S V+FD++ + + K +E+ +L + SD A EE
Sbjct: 662 YRLYDPATGKILISRDVQFDEERGWEWEDKSLEE--ELVWDNSDHEPAGEEGPEINHNGQ 719
Query: 1057 --EPEEDEPEEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLK 1114
+ E +E EE + Q L + + KD V +E L
Sbjct: 720 QDQEETEEEEETVAETVHQNLPAVGTGGVRQRQQPVWMKDYVVGNARVLITQDEEDEVLA 779
Query: 1115 GLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLN 1174
+ +P +EA Q + W AME E+ +N+ W LV+ PE VIG KW+F+ K N
Sbjct: 780 LFIGPDDPVCFEEAAQLEVWRKAMEAEITSIEENNTWELVELPEEAKVIGLKWIFKTKFN 839
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
EKG+V + KARLVA+GY Q+ G+D+ E FAP+A+ + IRL++ + + Q+DVKSA
Sbjct: 840 EKGEVDKFKARLVAKGYHQRYGVDFYEVFAPVAKWDTIRLILGLAAEKGWSVFQLDVKSA 899
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRG 1294
FL+G + E+V+V QP GFE E++ V+KLKK+LYGLKQAPRAWY R+ F + F +
Sbjct: 900 FLHGDLKEDVFVEQPKGFEVEEESSKVYKLKKALYGLKQAPRAWYSRIEEFFGKEGFEKC 959
Query: 1295 KVDTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGI 1354
+ TLF K + D LVV +YVDD+I+ ++ + + F M EF M+ +G++ YFLG+
Sbjct: 960 YCEHTLFVKKERSDFLVVSVYVDDLIYTGSSMEMIEGFKNSMMEEFAMTDLGKMKYFLGV 1019
Query: 1355 QVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMI 1414
+V Q G +I+Q KY E++KK+ M K P+ P L K A
Sbjct: 1020 EVIQDERGIFINQRKYAAEIIKKYGMEGCNSVKNPIVPGQKLTKAGA------------- 1066
Query: 1415 GSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKL 1474
+R+ P E HL AVKRILRY++ T +LG+ Y++ +L
Sbjct: 1067 -------------------VSRYMESPNEQHLLAVKRILRYVQGTLDLGIQYERGGATEL 1107
Query: 1475 SGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKL 1534
G+ D+DYAGD +RKSTSG LG ++W SK+Q + LST EAE++SA+ + Q +
Sbjct: 1108 VGFVDSDYAGDVDDRKSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAV 1167
Query: 1535 WMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
W+++ LE+ E ++CDN++ I LSKNP+LH R+KHI V+YHF+R+ V +G +
Sbjct: 1168 WLRNVLEEIGCRQEGGTLVFCDNSSTIKLSKNPVLHGRSKHIHVRYHFLRELVKEGTI 1225
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 632 bits (1629), Expect = 0.0
Identities = 362/992 (36%), Positives = 515/992 (51%), Gaps = 106/992 (10%)
Query: 606 KHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICV----DSSPC 661
++ WYLDSG S HM G + MF EL G V G K ++ G G I +
Sbjct: 331 ENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF 390
Query: 662 IYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLS 721
I NV + + N+LS+ QL +KGYD+ + + + KN ++ + +
Sbjct: 391 ISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDKESNLITKVPMSKNRMFVLNIR 450
Query: 722 ELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQK 781
AQ +K + EE W+WH R GH + + LS+ +VRGLP + + +CE C
Sbjct: 451 NDIAQCLK--MCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQ-VCEGCLL 507
Query: 782 GKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTR 841
G K+ F ++ +PLEL+H D+ GP+K +S+G Y ++ +DD+SR TWV FL
Sbjct: 508 GNQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKE 567
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K E +F F A V+ E I +RSD GGEF + +F + GI + PR+PQQ
Sbjct: 568 KSEVFEIFKKFKAHVEKESGLVIKTMRSDSGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQ 627
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNI 961
NGV ERKNRT+ EMAR+ML+ + K W EAV A Y+ NR + + KTP E W
Sbjct: 628 NGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGR 687
Query: 962 KPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIH 1021
KP +S+ FG + + ++ +K D KS K + +GY SKG++ YN D K S +
Sbjct: 688 KPGVSHLRVFGSIAHAHVPDEKRNKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRN 747
Query: 1022 VRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSRIT 1081
+ FD++ + D + E + DK + E P E+ P+ SQ + S
Sbjct: 748 IVFDEEGEWDWNSNEEDYNFFPHFEEDKPEPTREEPPSEEPTTPPTSPTSSQIEESS--- 804
Query: 1082 AAHPMELIMGNKDEPVRTRSAFRPSEETL----LSLKGLVSLIEPKSIDEALQDKDWILA 1137
+ R RS E T L+L L + EP EA++ K W A
Sbjct: 805 -----------SERTPRFRSIQELYEVTENQENLTLFCLFAECEPMDFQEAIEKKTWRNA 853
Query: 1138 MEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGI 1197
M+EE+ KND W L P IG KWV++ K N KG+V R KARLVA+GYSQ+ GI
Sbjct: 854 MDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGI 913
Query: 1198 DYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKK 1257
DY E FAP+ARLE +RL+IS + + +HQMDVKSAFLNG + EEVY+ QP G+ + +
Sbjct: 914 DYDEIFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGE 973
Query: 1258 PDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVD 1317
D V +LKK LYGLKQAPRAW R+ + E +F++ + L+ K K+DIL+ +YVD
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVD 1033
Query: 1318 DIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKK 1377
D+IF N S+ +EF K M EFEM+ +G ++Y+LGI+V Q G +I Q Y KE+LKK
Sbjct: 1034 DLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKK 1093
Query: 1378 FNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARF 1437
F M +S + ++GSL YLT +RPDIL++V + +R+
Sbjct: 1094 FKMDDSNPS--------------------------LVGSLRYLTCTRPDILYAVGVVSRY 1127
Query: 1438 QSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQ 1497
P TH A KRILRY+K T N GL Y TS DY
Sbjct: 1128 MEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTS----------DY--------------- 1162
Query: 1498 FLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQI-LESNIPIYCDN 1556
+C +W+++ L++ + E I+ DN
Sbjct: 1163 ---------------------------KLVVCHA--IWLRNLLKELSLPQEEPTKIFVDN 1193
Query: 1557 TAAISLSKNPILHSRAKHIEVKYHFIRDYV*K 1588
+AI+L+KNP+ H R+KHI+ +YH+IR+ V K
Sbjct: 1194 KSAIALAKNPVFHDRSKHIDTRYHYIRECVSK 1225
>At3g25450 hypothetical protein
Length = 1343
Score = 622 bits (1604), Expect = e-178
Identities = 360/1012 (35%), Positives = 553/1012 (54%), Gaps = 41/1012 (4%)
Query: 607 HQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSS----PCI 662
+ +WYLD+G S HMTG R F +L G+V FG + I G G+I S +
Sbjct: 289 NNAWYLDNGASNHMTGNRAWFCKLDEMITGKVRFGDDSCINIKGKGSIPFISKGGERKIL 348
Query: 663 YNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKR-KNNIYKIRLS 721
++V + L N+LS+ Q + G D+ + + +G++L ++R +N +YK+
Sbjct: 349 FDVYYIPDLKSNILSLGQATESGCDIRMREDYL-TLHDREGNLLIKAQRSRNRLYKV--- 404
Query: 722 ELEAQNVKCL-LSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQ 780
LE +N KCL L+ E +WH RLGH S I + K LV G+ + C +C
Sbjct: 405 SLEVENSKCLQLTTTNESTIWHARLGHISFETIKAMIKKELVIGISSSVPQEKETCGSCL 464
Query: 781 KGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLT 840
GK + F ++ LEL+H DL GP+ + KRY +++DD+SR+ W L
Sbjct: 465 FGKQARHSFPKATSYRAAQVLELIHGDLCGPISPSTAAKKRYVFVLIDDHSRYMWSILLK 524
Query: 841 RKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQ 900
K E+ F F A V+ E I R+D GGEF + +F+ GI + P TPQ
Sbjct: 525 EKSEAFGKFKEFKALVEQECGAIIKTFRTDRGGEFLSHEFQEFCAKEGINRHLTAPYTPQ 584
Query: 901 QNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKN 960
QNGVVER+NRTL M R++L+ M + W EAV + Y+ NR+ R + N+TPYE++K+
Sbjct: 585 QNGVVERRNRTLLGMTRSILKHMNMPNYLWGEAVRHSTYLINRVGTRSLSNQTPYEVFKH 644
Query: 961 IKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESI 1020
KPN+ + FGCV Y L K D +S + LG SK +R + + I S
Sbjct: 645 KKPNVEHLRVFGCVSYAKVEVPNLKKLDDRSRMLVYLGTEPGSKAYRLLDPTKRRIFVSR 704
Query: 1021 HVRFDDKLD----SDQSKLVEKFADLSINVSD---KGKAPEEAEPEEDEPEEEAGPSDSQ 1073
V FD+ S+ ++ +I +S+ G + E +E EE + +
Sbjct: 705 DVVFDENRSWMWQESSSETDKESGTFTITLSEFGNNGVTENDISTEPEETEEAEINGEDE 764
Query: 1074 TLKKSRITAAHPMELIMGNKDEPVRT--RSAFRPS-------------EETLLSLKGLVS 1118
+ + T H + +PVR R RP+ E LL++
Sbjct: 765 NIIEEAETEEHDQSQ---EEPQPVRRSQRQVIRPNYLKDYVLCAEIEAEHLLLAVND--- 818
Query: 1119 LIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGD 1178
EP EA + K+W A +EE+ KN WSLV P IG KWVF+ K N G
Sbjct: 819 --EPWDFKEANKSKEWRDACKEEIQSIEKNRTWSLVDLPVGSKAIGVKWVFKLKHNSDGS 876
Query: 1179 VVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNG 1238
+ + KARLVA+GY Q+ G+D+ E FAP+AR+E +RL+I+ + ++ +H +DVK+AFL+G
Sbjct: 877 INKYKARLVAKGYVQRHGVDFEEVFAPVARIETVRLIIALAASNGWEIHHLDVKTAFLHG 936
Query: 1239 YISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDT 1298
+ E+VYV QP GF +++ + V+KL K+LYGL+QAPRAW +L+ L E +F + +
Sbjct: 937 ELREDVYVSQPEGFTNKESKEKVYKLHKALYGLRQAPRAWNTKLNEILKELKFEKCHKEP 996
Query: 1299 TLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQ 1358
+L+ K ++ILVV +YVDD++ +N + F K M +FEMS +G+LTY+LGI+V Q
Sbjct: 997 SLYRKQEGENILVVAVYVDDLLVTGSNLDIILNFKKGMVGKFEMSDLGKLTYYLGIEVLQ 1056
Query: 1359 TPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLL 1418
+ +G + Q +Y K++L++ M + TPM + L K ++ + YR IG L
Sbjct: 1057 SKDGITLKQERYAKKILEEAGMSKCNTVNTPMIASLELSKAQDEKRIDETDYRRNIGCLR 1116
Query: 1419 YLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYC 1478
YL +RPD+ ++V + +R+ +PRE+H A+K+ILRYL+ TT+ GL +KK L GY
Sbjct: 1117 YLLHTRPDLSYNVGILSRYLQEPRESHGAALKQILRYLQGTTSHGLYFKKGENAGLIGYS 1176
Query: 1479 DADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKH 1538
D+ + D + KST G+ +L ++W S++Q + LS+ EAE+++A + Q +W++
Sbjct: 1177 DSSHNVDLDDGKSTGGHIFYLNDCPITWCSQKQQVVTLSSCEAEFMAATEAAKQAIWLQE 1236
Query: 1539 QLEDYQILE-SNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KG 1589
L + E + I DN +AI+L+KNP+ H R+KHI +YHFIR+ V G
Sbjct: 1237 LLAEVIGTECEKVTIRVDNKSAIALTKNPVFHGRSKHIHRRYHFIRECVENG 1288
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 603 bits (1556), Expect = e-172
Identities = 349/1004 (34%), Positives = 544/1004 (53%), Gaps = 37/1004 (3%)
Query: 605 LKHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYN 664
L +W +DSG + H++ +R++FQ L V +I G GT+ ++ + N
Sbjct: 437 LSSDTWVIDSGATHHVSHDRKLFQTLDTSIVSFVNLPTGPNVRISGVGTVLINKDIILQN 496
Query: 665 VLLVDGLTHNLLSISQLA-DKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSEL 723
VL + NL+SIS L D G VIF+ C+ G L KR N+Y + ++
Sbjct: 497 VLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGLTLGEGKRIGNLYVLD-TQS 555
Query: 724 EAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
A +V ++ V+ VWH+RLGH S ++ LS+ V G K A C C K
Sbjct: 556 PAISVNAVVDVS----VWHKRLGHPSFSRLDSLSE---VLGTTRHKNKKSAYCHVCHLAK 608
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
K+ F + N + S ELLHID++GP E++ G +Y + IVDD+SR TW+ L K
Sbjct: 609 QKKLSFPSANNICNST-FELLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKS 667
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
+ VF FI V+N+ R+ VRSD+ E F + + GI SCP TP+QN
Sbjct: 668 DVLTVFPAFIDLVENQYDTRVKSVRSDNAKELA---FTEFYKAKGIVSFHSCPETPEQNS 724
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERK++ + +AR ++ ++ M+ +W + V TA ++ NR + NKTP+E+ P
Sbjct: 725 VVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFLINRTPSALLSNKTPFEVLTGKLP 784
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVR 1023
+ S FGC+CY + + HKF +S C+ LGY KG++ + ++ + S +V
Sbjct: 785 DYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKGYKLLDLESNVVHISRNVE 844
Query: 1024 FDDKL----DSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKKSR 1079
F ++L S QS ++ G + P S S + K R
Sbjct: 845 FHEELFPLASSQQSATTASDVFTPMDPLSSGNSITSHLPSPQI-------SPSTQISKRR 897
Query: 1080 ITA--AHPMEL--IMGNKDE--PVRTRSAFRP-SEETLLSLKGLVSLIEPKSIDEALQDK 1132
IT AH + NKD+ P+ + ++ S +L + + + P+S EA K
Sbjct: 898 ITKFPAHLQDYHCYFVNKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKDSK 957
Query: 1133 DWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYS 1192
+W A+++E+ + D W + P +G KWVF K + G + R KAR+VA+GY+
Sbjct: 958 EWCGAIDQEIGAMERTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYT 1017
Query: 1193 QQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGF 1252
Q+EG+DYTETF+P+A++ ++LL+ S + L+Q+D+ +AFLNG + E +Y+ P G+
Sbjct: 1018 QKEGLDYTETFSPVAKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGY 1077
Query: 1253 EDEK----KPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDD 1308
D K P+ V +LKKS+YGLKQA R W+ + S+ LL F + D TLF + +
Sbjct: 1078 ADIKGTSLPPNVVCRLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCIGSE 1137
Query: 1309 ILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQS 1368
+V+ +YVDDI+ S + + ++ ++A F++ +G L YFLG++V +T EG + Q
Sbjct: 1138 FIVLLVYVDDIVIASTTEQAAQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLSQR 1197
Query: 1369 KYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDIL 1428
KY ELL +ML+ + PM P L K D +++YR ++G L+YLT +RPDI
Sbjct: 1198 KYALELLTSADMLDCKPSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDIT 1257
Query: 1429 FSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTE 1488
F+V+ +F S PR HL AV ++L+Y+K T GL Y + L GY DAD+
Sbjct: 1258 FAVNKLCQFSSAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTCPDS 1317
Query: 1489 RKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILES 1548
R+ST+G F+GS+L+SW SK+Q T++ S+AEAEY + A+ S + W+ L ++ S
Sbjct: 1318 RRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRV-HS 1376
Query: 1549 NIPI-YCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
+PI Y D+TAA+ ++ NP+ H R KHIE+ H +R+ + G L
Sbjct: 1377 GVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLDNGQL 1420
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 595 bits (1533), Expect = e-170
Identities = 341/1004 (33%), Positives = 543/1004 (53%), Gaps = 33/1004 (3%)
Query: 605 LKHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYN 664
L +W +DSG + H++ +R +F L V KI G GT+ ++ + N
Sbjct: 426 LSSATWVIDSGATHHVSHDRSLFSSLDTSVLSAVNLPTGPTVKISGVGTLKLNDDILLKN 485
Query: 665 VLLVDGLTHNLLSISQLADK-GYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSEL 723
VL + NL+SIS L D G VIF++ SC I G +L +R N+Y + + +
Sbjct: 486 VLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRMLGQGRRVANLYLLDVGD- 544
Query: 724 EAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
++ +V ++ ++ +WHRRLGHAS++++ +S G K C C K
Sbjct: 545 QSISVNAVVDIS----MWHRRLGHASLQRLDAISDS---LGTTRHKNKGSDFCHVCHLAK 597
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
K+ F N V +LLHID++GP E++ G +Y + IVDD+SR TW+ L K
Sbjct: 598 QRKLSFPTSNKVC-KEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWMYLLKTKS 656
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E VF FI QV+N+ ++ VRSD+ E KF S + GI SCP TP+QN
Sbjct: 657 EVLTVFPAFIQQVENQYKVKVKAVRSDNAPEL---KFTSFYAEKGIVSFHSCPETPEQNS 713
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERK++ + +AR ++ ++ + W + V TA ++ NR + ++NKTPYE+ P
Sbjct: 714 VVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTPYEILTGTAP 773
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVR 1023
FGC+CY + + HKF +S CL LGY KG++ + ++ T+ S +V+
Sbjct: 774 VYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLESNTVFISRNVQ 833
Query: 1024 FDDKL--------DSDQSKLVEKFADLSINV-SDKGKAPEEAEPE-EDEPEEEAGPSDSQ 1073
F +++ KL +S + SD +P + D P + + SQ
Sbjct: 834 FHEEVFPLAKNPGSESSLKLFTPMVPVSSGIISDTTHSPSSLPSQISDLPPQIS----SQ 889
Query: 1074 TLKK--SRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQD 1131
++K + + H + +K T S + S + + + + P + EA
Sbjct: 890 RVRKPPAHLNDYHCNTMQSDHKYPISSTISYSKISPSHMCYINNITKIPIPTNYAEAQDT 949
Query: 1132 KDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGY 1191
K+W A++ E+ K + W + P+ +G KWVF K G++ R KARLVA+GY
Sbjct: 950 KEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVAKGY 1009
Query: 1192 SQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPG 1251
+Q+EG+DYT+TF+P+A++ I+LL+ S + L Q+DV +AFLNG + EE+++ P G
Sbjct: 1010 TQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKIPEG 1069
Query: 1252 FEDEK----KPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKD 1307
+ + K + V +LK+S+YGLKQA R W+++ SS LL F + D TLF K Y
Sbjct: 1070 YAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDG 1129
Query: 1308 DILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQ 1367
+ ++V +YVDDI+ S +++ + ++ + F++ +G+L YFLG++V +T G I Q
Sbjct: 1130 EFVIVLVYVDDIVIASTSEAAAAQLTEELDQRFKLRDLGDLKYFLGLEVARTTAGISICQ 1189
Query: 1368 SKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDI 1427
KY ELL+ ML PM P + K+D + YR ++G L+YLT +RPDI
Sbjct: 1190 RKYALELLQSTGMLACKPVSVPMIPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITRPDI 1249
Query: 1428 LFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRT 1487
F+V+ +F S PR THLTA R+L+Y+K T GL Y +S+ L G+ D+D+A +
Sbjct: 1250 TFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVGQGLFYSASSDLTLKGFADSDWASCQD 1309
Query: 1488 ERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILE 1547
R+ST+ F+G +L+SW SK+Q T++ S+AEAEY + A+ + + +W+ L Q
Sbjct: 1310 SRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQASP 1369
Query: 1548 SNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
+Y D+TAAI ++ NP+ H R KHI++ H +R+ + G L
Sbjct: 1370 PVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNGEL 1413
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 567 bits (1460), Expect = e-161
Identities = 341/1002 (34%), Positives = 536/1002 (53%), Gaps = 58/1002 (5%)
Query: 609 SWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSS----PCIYN 664
SWYLD+G S HMTG + F +L G+V FG + + I G G+I + + + +
Sbjct: 279 SWYLDNGASNHMTGNLQWFSKLNEMITGKVRFGDDSRIDIKGKGSIVLITKGGIRKTLTD 338
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKR-KNNIYKIRLSEL 723
V + L N++S+ Q + G DV + +G +L + R +N +YK+ +L
Sbjct: 339 VYFIPDLKSNIISLGQATEAGCDVRMKDDQL-TLHDREGCLLLRATRSRNRLYKV---DL 394
Query: 724 EAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+NVKCL + + + + LV G+ N+ + C +C GK
Sbjct: 395 NVENVKCL------------------QLEAATMVRKELVIGISNIPKEKET-CGSCLLGK 435
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKD 843
+ PF S+ LEL+H DL GP+ + KRY ++++DD++R+ W L K
Sbjct: 436 QARQPFPKATTYRASQVLELVHGDLCGPITQSTTAKKRYILVLIDDHTRYMWSMLLKEKS 495
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F F +V+ E +I R+D GGEF + +F+ GI + P TPQQNG
Sbjct: 496 EAFEKFRDFKTKVEQESGVKIKTFRTDKGGEFVSQEFQDFCAKEGINRHLTAPYTPQQNG 555
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVER+NRTL M R++L+ M + W EAV + YI NR+ R + N+TPYE++K KP
Sbjct: 556 VVERRNRTLLGMTRSILKHMKMPNYLWGEAVRHSTYIINRVGTRSLQNQTPYEVFKQRKP 615
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYN-TDAKTIEESIHV 1022
N+ + FGC+ Y L K D +S + LG SK +R + T+ K I+ +
Sbjct: 616 NVEHLRVFGCIGYAKIEGPHLRKLDDRSKMLVYLGTEPGSKAYRLLDPTNRKIIKWNNSD 675
Query: 1023 RFDDKLDSDQSKLVEKFADLSINVSDK---GKAPEEAE-PEEDEPEEEAGPSDSQTLKKS 1078
+ S + +F + I SD K EE+E E+E E E + +++
Sbjct: 676 SETRDISGTFSLTLGEFGNNGIQESDDIETEKNGEESENSHEEEGENEHNEQEQIDAEET 735
Query: 1079 RITAAHPMELIMGNKDEPVRTRSAFRPS-------------EETLLSLKGLVSLIEPKSI 1125
+ + A P+ + + TR +P+ E+ LL++ EP
Sbjct: 736 QPSHATPLPTLRRS------TRQVGKPNYLDDYVLMAEIEGEQVLLAIND-----EPWDF 784
Query: 1126 DEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKAR 1185
EA + K+W A +EE+ KN WSL+ P VIG KWVF+ K N G + + KAR
Sbjct: 785 KEANKLKEWRDACKEEILSIEKNKTWSLIDLPVRRKVIGLKWVFKIKRNSDGSINKYKAR 844
Query: 1186 LVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVY 1245
LVA+GY Q+ GIDY E FA +AR+E IR++I+ + ++ +H +DVK+AFL+G + E+VY
Sbjct: 845 LVAKGYVQRHGIDYDEVFAHVARIETIRVIIALAASNGWEVHHLDVKTAFLHGELREDVY 904
Query: 1246 VHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTY 1305
V QP GF ++ V+KL K+LYGLKQAPRAW +L+ L E FV+ + +++ +
Sbjct: 905 VTQPEGFTNKDNEGKVYKLHKALYGLKQAPRAWNTKLNKILQELNFVKCSKEPSVYRRQE 964
Query: 1306 KDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYI 1365
+ +L+V IYVDD++ ++ L F K M +FEMS +G+LTY+LGI+V G +
Sbjct: 965 EKKLLIVAIYVDDLLVTGSSLDLILCFKKDMAGKFEMSDLGQLTYYLGIEVLHRKNGIIL 1024
Query: 1366 HQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRP 1425
Q +Y +++++ M PM L K + ++ YR MIG L Y+ +RP
Sbjct: 1025 RQERYAMKIIEEAGMSNCNPVLIPMAAGLELCKAQEEKCITERDYRRMIGCLRYIVHTRP 1084
Query: 1426 DILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGD 1485
D+ + V + +R+ PRE+H A+K++LRYLK T + GL K+ + L GY D+ ++ D
Sbjct: 1085 DLSYCVGVLSRYLQQPRESHGNALKQVLRYLKGTMSHGLYLKRGFKSGLVGYSDSSHSAD 1144
Query: 1486 RTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQL-EDYQ 1544
+ KST+G+ +L ++W S++Q +ALS+ EAE+++A + Q +W++ E
Sbjct: 1145 LDDGKSTAGHIFYLHQCPITWCSQKQQVVALSSCEAEFMAATEAAKQAIWLQDLFAEVCG 1204
Query: 1545 ILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV 1586
+ I DN +AI+L+KN + H R+KHI +YHFIR+ V
Sbjct: 1205 TTSEKVMIRVDNKSAIALTKNLVFHGRSKHIHRRYHFIRECV 1246
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 566 bits (1459), Expect = e-161
Identities = 341/1021 (33%), Positives = 536/1021 (52%), Gaps = 49/1021 (4%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGE-VGFGGNEKGKIVGTGTICVDSSPC---IY 663
+ W+ DS + H+T Q G + V G I TG+ + SS +
Sbjct: 320 KEWHPDSAATAHVTSSTNGLQSATEYEGDDAVLVGDGTYLPITHTGSTTIKSSNGKIPLN 379
Query: 664 NVLLVDGLTHNLLSISQLADK-GYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSE 722
VL+V + +LLS+S+L D V F+ + V+ R+N +Y + E
Sbjct: 380 EVLVVPNIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLQTQKVVTTGPRRNGLYVLENQE 439
Query: 723 LEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKG 782
A + EE VWH RLGHA+ + + L ++ K + +CE CQ G
Sbjct: 440 FVALYSNRQCAATEE--VWHHRLGHANSKALQHLQNSKAIQ---INKSRTSPVCEPCQMG 494
Query: 783 KFTKVPFKAKNVVSTSR---PLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFL 839
K +++PF ++S SR PL+ +H DL+GP S G +Y I VDDYSR++W L
Sbjct: 495 KSSRLPF----LISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWFYPL 550
Query: 840 TRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTP 899
K E +VF +F V+N+ +I +SD GGEF ++K ++ +GI H SCP TP
Sbjct: 551 HNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCPYTP 610
Query: 900 QQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWK 959
QQNG+ ERK+R L E+ +ML + + FW E+ TA YI NR+ + N +PYE
Sbjct: 611 QQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPYEALF 670
Query: 960 NIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEES 1019
KP+ S FG CY +KFD +S +C+ LGY+ + KG+R + + S
Sbjct: 671 GEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKVYIS 730
Query: 1020 IHVRFDDK---LDSDQSKLVEKFADLSINVSDKGKAPEEAEP------------------ 1058
+V F++ LV +++ + K E + P
Sbjct: 731 RNVIFNESELPFKEKYQSLVPQYSTPLLQAWQHNKISEISVPAAPVQLFSKPIDLNTYAG 790
Query: 1059 -------EEDEPEEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLL 1111
+ EP SD + + AA+ E ++ + R+++ +
Sbjct: 791 SQVTEQLTDPEPTSNNEGSDEEVNPVAEEIAAN-QEQVINSHAMTTRSKAGIQKPNTRYA 849
Query: 1112 SLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRN 1171
+ ++ EPK++ A++ W A+ EE+N+ WSLV + ++++ +KWVF+
Sbjct: 850 LITSRMNTAEPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNILSSKWVFKT 909
Query: 1172 KLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDV 1231
KL+ G + + KARLVA+G+ Q+EG+DY ETF+P+ R IRL++ S + + Q+DV
Sbjct: 910 KLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKGWPIKQLDV 969
Query: 1232 KSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEF 1291
+AFL+G + E V+++QP GF D +KP HV +L K++YGLKQAPRAW++ S+FLL+ F
Sbjct: 970 SNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGF 1029
Query: 1292 VRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYF 1351
V K D +LF IL + +YVDDI+ ++QSL ++ + ++ F M +G YF
Sbjct: 1030 VCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYF 1089
Query: 1352 LGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYR 1411
LGIQ++ G ++HQ+ Y ++L++ M + TP+ L+ ++ +R
Sbjct: 1090 LGIQIEDYANGLFLHQTAYATDILQQAGMSDCNPMPTPLPQQ--LDNLNSELFAEPTYFR 1147
Query: 1412 GMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSE 1471
+ G L YLT +RPDI F+V+ + P + +KRILRY+K T +GL K+ S
Sbjct: 1148 SLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMGLPIKRNST 1207
Query: 1472 YKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICST 1531
LS Y D+D+AG + R+ST+G C LGSNL+SW +KRQ T++ S+ EAEY + +
Sbjct: 1208 LTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAR 1267
Query: 1532 QKLWMKHQLEDYQILE-SNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGV 1590
+ W+ L D I + +YCDN +A+ LS NP LH+R+KH + YH+IR+ V G+
Sbjct: 1268 EITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGL 1327
Query: 1591 L 1591
+
Sbjct: 1328 I 1328
>At1g70010 hypothetical protein
Length = 1315
Score = 562 bits (1449), Expect = e-160
Identities = 334/968 (34%), Positives = 511/968 (52%), Gaps = 35/968 (3%)
Query: 651 TGTICVDSSPCIYNVLLVDGLTHNLLSISQLADK-GYDVIFNQKSCRAVSQIDGSVLFNS 709
+G++ + + +VL + NLLS+S L G + F++ SC ++
Sbjct: 314 SGSVHLGRHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATRELMVGMG 373
Query: 710 KRKNNIYKIRLSELEAQNVKCLLSVNE--EQWVWHRRLGHASMRKISQLSKLNLVRGLPN 767
K+ N+Y + L L ++V +WH+RLGH S++K+ +S L P
Sbjct: 374 KQVANLYIVDLDSLSHPGTDSSITVASVTSHDLWHKRLGHPSVQKLQPMSSL---LSFPK 430
Query: 768 LKFASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIV 827
K +D C C K +PF + N S SRP +L+HID +GP ++ G RY + IV
Sbjct: 431 QKNNTDFHCRVCHISKQKHLPFVSHNNKS-SRPFDLIHIDTWGPFSVQTHDGYRYFLTIV 489
Query: 828 DDYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSY 887
DDYSR TWV L K + V TF+ V+N+ I VRSD+ E F + S
Sbjct: 490 DDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELN---FTQFYHSK 546
Query: 888 GIAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVR 947
GI SCP TPQQN VVERK++ + +AR++ ++ + +W + + TA Y+ NR+
Sbjct: 547 GIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRLPAP 606
Query: 948 PILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFR 1007
+ +K P+E+ P + FGC+CY + HKF ++ C +GY KG++
Sbjct: 607 ILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGFKGYK 666
Query: 1008 FYNTDAKTIEESIHVRFDDKL----DSDQSKLVEKF-ADLS------------INVSDKG 1050
+ + +I S HV F ++L SD S+ + F DL+ +N SD
Sbjct: 667 LLDLETHSIIVSRHVVFHEELFPFLGSDLSQEEQNFFPDLNPTPPMQRQSSDHVNPSDSS 726
Query: 1051 KAPE---EAEPEEDEPEEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSE 1107
+ E A P + PE S + K + + + ++ E + S R ++
Sbjct: 727 SSVEILPSANPTNNVPEPSVQTSHRKAKKPAYLQDYYCHSVVSSTPHEIRKFLSYDRIND 786
Query: 1108 ETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKW 1167
L L L EP + EA + + W AM E + W + P IG +W
Sbjct: 787 PYLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRW 846
Query: 1168 VFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLH 1227
+F+ K N G V R KARLVAQGY+Q+EGIDY ETF+P+A+L +++LL+ + + L
Sbjct: 847 IFKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVAKLNSVKLLLGVAARFKLSLT 906
Query: 1228 QMDVKSAFLNGYISEEVYVHQPPGFE----DEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1283
Q+D+ +AFLNG + EE+Y+ P G+ D P+ V +LKKSLYGLKQA R WY + S
Sbjct: 907 QLDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFS 966
Query: 1284 SFLLENEFVRGKVDTTLFCKTYKDDILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMS 1343
S LL F++ D T F K L V +Y+DDII S N + M++ F++
Sbjct: 967 STLLGLGFIQSYCDHTCFLKISDGIFLCVLVYIDDIIIASNNDAAVDILKSQMKSFFKLR 1026
Query: 1344 MMGELTYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDASG 1403
+GEL YFLG+++ ++ +G +I Q KY +LL + L + PM P+ + +
Sbjct: 1027 DLGELKYFLGLEIVRSDKGIHISQRKYALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGD 1086
Query: 1404 KVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLG 1463
V YR +IG L+YL +RPDI F+V+ A+F PR+ HL AV +IL+Y+K T G
Sbjct: 1087 FVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQG 1146
Query: 1464 LMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEY 1523
L Y TSE +L Y +ADY R R+STSG C FLG +L+ W+S++Q ++ S+AEAEY
Sbjct: 1147 LFYSATSELQLKVYANADYNSCRDSRRSTSGYCMFLGDSLICWKSRKQDVVSKSSAEAEY 1206
Query: 1524 ISAAICSTQKLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFI 1582
S ++ + + +W+ + L++ Q+ L ++CDN AAI ++ N + H R KHIE H +
Sbjct: 1207 RSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEAAIHIANNHVFHERTKHIESDCHSV 1266
Query: 1583 RDYV*KGV 1590
R+ + KG+
Sbjct: 1267 RERLLKGL 1274
>At1g37110
Length = 1356
Score = 539 bits (1388), Expect = e-153
Identities = 341/1033 (33%), Positives = 535/1033 (51%), Gaps = 76/1033 (7%)
Query: 610 WYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSS----PCIYNV 665
W LDSGC+ HMT R F + K + G + + G GTI +D+ + NV
Sbjct: 308 WILDSGCTSHMTSRRDWFISFQEKGNTTILLGDDHSVESQGQGTIRIDTHGGTIKILENV 367
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIR----LS 721
V L NL+S L GY + R + N +Y + +S
Sbjct: 368 KYVPHLRRNLISTGTLDKLGYRHEGGEGKVRYFK--NNKTALRGSLSNGLYVLDGSTVMS 425
Query: 722 EL---EAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLV--RGLPNLKFASDALC 776
EL E VK L WH RLGH SM + L+ L+ + + L+F C
Sbjct: 426 ELCNAETDKVKTAL--------WHSRLGHMSMNNLKVLAGKGLIDRKEINELEF-----C 472
Query: 777 EACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFG-PVKSESIGGKRYGMIIVDDYSRWTW 835
E C GK KV F S L +H DL+G P + SI GK+Y + I+DD +R W
Sbjct: 473 EHCVMGKSKKVSFNVGKHTSEDA-LSYVHADLWGSPNVTPSISGKQYFLSIIDDKTRKVW 531
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSC 895
+ FL KDE+ F + + V+N+ ++ +R+D+G EF N +F+S +GI +C
Sbjct: 532 LYFLKSKDETFDKFCEWKSLVENQVNKKVKCLRTDNGLEFCNSRFDSYCKEHGIERHRTC 591
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPY 955
TPQQNGV ER NRT+ E R +L ++G+ + FW EA TA Y+ NR I + P
Sbjct: 592 TYTPQQNGVAERMNRTIMEKVRCLLNKSGVEEVFWAEAAATAAYLINRSPASAINHNVPE 651
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKT 1015
E+W N KP + FG + YV + +L ++ K LGY +KG++ + + +
Sbjct: 652 EMWLNRKPGYKHLRKFGSIAYVHQDQGKLKP---RALKGFFLGYPAGTKGYKVWLLEEEK 708
Query: 1016 IEESIHVRFDDKL----------DSDQSKLVE---------KFADLS-----INVSDKGK 1051
S +V F + + D+D E KFA+ S I + +
Sbjct: 709 CVISRNVVFQESVVYRDLKVKEDDTDNLNQKETTSSEVEQNKFAEASGSGGVIQLQSDSE 768
Query: 1052 APEEAEPEEDEPEE-EAGPSDSQTLKKSRITAAH-PMELIMGNKDEPVRTRSAFRPSEET 1109
E E D EE E +T K++ +T + + N + P R F
Sbjct: 769 PITEGEQSSDSEEEVEYSEKTQETPKRTGLTTYKLARDRVRRNINPPTR----FTEESSV 824
Query: 1110 LLSLKGLVSLI--EPKSIDEALQDKD---WILAMEEELNQFSKNDVWSLVKKPESVHVIG 1164
+L + + I EP+S EA++ +D W +A +E++ KN W LV KP+ +IG
Sbjct: 825 TFALVVVENCIVQEPQSYQEAMESQDCEKWDMATHDEMDSLMKNGTWDLVDKPKDRKIIG 884
Query: 1165 TKWVFRNKLNEKG-DVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHN 1223
+W+F+ K G + R KARLVA+GY+Q+EG+DY E FAP+ + +IR+L+S V+ +
Sbjct: 885 CRWLFKLKSGIPGVEPTRFKARLVAKGYTQREGVDYQEIFAPVVKHVSIRILMSLVVDKD 944
Query: 1224 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1283
+ L QMDVK+ FL+G + EE+Y+ QP GF + + V +LKKSLYGLKQ+PR W +R
Sbjct: 945 LELEQMDVKTTFLHGDLEEELYMEQPEGFVSDSSENKVCRLKKSLYGLKQSPRQWNKRFD 1004
Query: 1284 SFLLENEFVRGKVDTTLFCKTYKD-DILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEM 1342
F+ +F+R + D ++ K + D + + +YVDD++ A+++ + + EFEM
Sbjct: 1005 RFMSSQQFIRSEHDACVYVKHVSEHDFIYLLLYVDDMLIAGASKAEINRVKEQLSTEFEM 1064
Query: 1343 SMMGELTYFLGIQVDQTPEGTYIHQSK--YTKELLKKFNMLESTVAKTPM---HPTCILE 1397
MG + LGI + + +G + S+ Y +++L +FNM + + P+ +
Sbjct: 1065 KDMGGASRILGIDIYRDRKGGVLKLSQEIYIRKVLDRFNMSGAKMTNAPVGAHFKLAAVR 1124
Query: 1398 KEDASGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYL 1456
+ED Y +GS++Y + +RPD+ +++ L +R+ S P H AVK ++RYL
Sbjct: 1125 EEDECVDTDVVPYSSAVGSIMYAMLGTRPDLAYAICLISRYMSKPGSMHWEAVKWVMRYL 1184
Query: 1457 KDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIAL 1516
K +L L++ K ++ ++GYCD++YA D R+S SG +G N VSW++ Q +A+
Sbjct: 1185 KGAQDLNLVFTKEKDFTVTGYCDSNYAADLDRRRSISGYVFTIGGNTVSWKASLQPVVAM 1244
Query: 1517 STAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIE 1576
ST EAEYI+ A + + +W+K L+D + + + I+CD+ +AI LSKN + H R KHI+
Sbjct: 1245 STTEAEYIALAEAAKEAMWIKGLLQDMGMQQDKVKIWCDSQSAICLSKNSVYHERTKHID 1304
Query: 1577 VKYHFIRDYV*KG 1589
V++++IRD V G
Sbjct: 1305 VRFNYIRDVVESG 1317
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 530 bits (1364), Expect = e-150
Identities = 332/1020 (32%), Positives = 534/1020 (51%), Gaps = 51/1020 (5%)
Query: 610 WYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIY----NV 665
W +D+GCS HMT +R F++L GG V G K+ G GTI V + + NV
Sbjct: 287 WVMDTGCSYHMTYKREWFEDLNEDAGGSVRMGNKTVSKVRGIGTIRVKNEAGMVVRLTNV 346
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
+ + NLLS+ GY + ++ SVL +R +Y ++ +
Sbjct: 347 RYIPEMDRNLLSLGTFEKSGYSFKLENGTLSIIA--GDSVLLTVRRCYTLYLLQWRPVTE 404
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFT 785
+++ ++ ++ +WHRRLGH S + + L K L L K + CE C GK
Sbjct: 405 ESLS-VVKRQDDTILWHRRLGHMSQKNMDLLLKKGL---LDKKKVSKLETCEDCIYGKAK 460
Query: 786 KVPFKAKNVVSTSRPLELLHIDLFG-PVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDE 844
++ F T LE +H DL+G P S+G +Y + +DDY+R + FL KDE
Sbjct: 461 RIGFNLAQH-DTREKLEYVHSDLWGAPSVPFSLGKCQYFISFIDDYTRKVRIYFLKTKDE 519
Query: 845 SHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGV 904
+ F + V+N+ RI +R+D+G EF N F+ GI +C TPQQNGV
Sbjct: 520 AFDKFVEWANLVENQTDKRIKTLRTDNGLEFCNRSFDEFCSQKGILWHRTCAYTPQQNGV 579
Query: 905 VERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPN 964
ER NRTL E R+ML ++G+ K FW EA +T + N+ + + P + W P
Sbjct: 580 AERMNRTLMEKVRSMLSDSGLPKKFWAEATHTTAILINKTPSSALNYEVPDKRWSGKSPI 639
Query: 965 ISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRF 1024
SY FGC+ +V +T D K + ++ K +L+GY KG++ + + K S +V F
Sbjct: 640 YSYLRRFGCIAFV-HTDDG--KLNPRAKKGILVGYPIGVKGYKIWLLEEKKCVVSRNVIF 696
Query: 1025 DDKL---DSDQSKLVEK---------FADLSIN----VSDKGKAP--EEAEPEEDEPEEE 1066
+ D QSK EK + DL ++ ++ G P E P P
Sbjct: 697 QENASYKDMMQSKDAEKDENEAPPSSYLDLDLDHEEVITSGGDDPIVEAQSPFNPSPATT 756
Query: 1067 AGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSE---ETLLSLKGLVSLIEPK 1123
S+ + I + +L+ +R F + E L + + IEP
Sbjct: 757 QTYSEGVNSETDIIQSPLSYQLVRDRDRRTIRAPVRFDDEDYLAEALYTTEDSGE-IEPA 815
Query: 1124 SIDEALQDKDWI---LAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKG-DV 1179
EA + +W LAM EE+ KN W++VK+P+ VIG++W+++ KL G +
Sbjct: 816 DYSEAKRSMNWNKWKLAMNEEMESQIKNHTWTVVKRPQHQKVIGSRWIYKFKLGIPGVEE 875
Query: 1180 VRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGY 1239
R KARLVA+GY+Q++GIDY E FAP+ + +IR+L+S ++ L Q+DVK+AFL+G
Sbjct: 876 GRFKARLVAKGYAQRKGIDYHEIFAPVVKHVSIRILMSIVAQEDLELEQLDVKTAFLHGE 935
Query: 1240 ISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTT 1299
+ E++Y+ P G+E+ K D V L KSLYGLKQAP+ W E+ ++++ E F+R D+
Sbjct: 936 LKEKIYMVPPEGYEEMFKEDEVCLLNKSLYGLKQAPKQWNEKFNAYMSEIGFIRSLYDSC 995
Query: 1300 LFCKTYKDDILV-VQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQ 1358
+ K D V + +YVDD++ + N+ + + + F+M +G LG+++ +
Sbjct: 996 AYIKELSDGSRVYLLLYVDDMLVAAKNKEDISQLKEELSQRFDMKDLGAAKRILGMEIIR 1055
Query: 1359 TPEGT--YIHQSKYTKELLKKFNMLESTVAKTPMHP-----TCILEKEDASGKVCQKL-Y 1410
E ++ Q+ Y ++L+ +NM ES TP+ +EK++ + + Y
Sbjct: 1056 NREENTLWLSQNGYLNKILETYNMAESKHVVTPLGAHLKMRAATVEKQEQDEDYMKSIPY 1115
Query: 1411 RGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKT 1469
+GS++Y + +RPD+ + V + +R+ S P H VK +LRY+K + L YK++
Sbjct: 1116 SSAVGSIMYAMIGTRPDLAYPVGIISRYMSQPAREHWLGVKWVLRYIKGSLGTKLQYKRS 1175
Query: 1470 SEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAIC 1529
S++K+ GYCDAD+A + R+S +G LG + +SW+S +Q +ALST EAEY+S
Sbjct: 1176 SDFKVVGYCDADHAACKDRRRSITGLVFTLGGSTISWKSGQQRVVALSTTEAEYMSLTEA 1235
Query: 1530 STQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KG 1589
+ +WMK L+++ + ++ I+CD+ +AI+LSKN + H R KHI+V+Y +IRD + G
Sbjct: 1236 VKEAVWMKGLLKEFGYEQKSVEIFCDSQSAIALSKNNVHHERTKHIDVRYQYIRDIIANG 1295
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 528 bits (1360), Expect = e-149
Identities = 337/1027 (32%), Positives = 530/1027 (50%), Gaps = 57/1027 (5%)
Query: 610 WYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSP----CIYNV 665
W LD+GCS HMT + F++ K G V G + + G G+I + +S + +V
Sbjct: 285 WVLDTGCSFHMTPRKDWFKDFKELSSGYVKMGNDTYSPVKGIGSIKIRNSDGSQVILTDV 344
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
+ +T NL+S+ L D+G + V S + ++++ +Y + E
Sbjct: 345 RYMPNMTRNLISLGTLEDRGCWFKSQDGILKIVKGC--STILKGQKRDTLYILDGVTEEG 402
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRG--LPNLKFASDALCEACQKGK 783
++ V +E +WH RLGH S + + L K +R + L+F CE C GK
Sbjct: 403 ESHSSA-EVKDETALWHSRLGHMSQKGMEILVKKGCLRREVIKELEF-----CEDCVYGK 456
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFG-PVKSESIGGKRYGMIIVDDYSRWTWVKFLTRK 842
+V F V T L +H DL+G P S+G +Y + VDDYSR W+ FL +K
Sbjct: 457 QHRVSFAPAQHV-TKEKLAYVHSDLWGSPHNPASLGNSQYFISFVDDYSRKVWIYFLRKK 515
Query: 843 DESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQN 902
DE+ F + V+N+ ++ ++R+D+G E+ N FE GI +C TPQQN
Sbjct: 516 DEAFEKFVEWKKMVENQSDRKVKKLRTDNGLEYCNHYFEKFCKEEGIVRHKTCAYTPQQN 575
Query: 903 GVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIK 962
G+ ER NRT+ + R+ML +GM K FW EA +TA Y+ NR I P E W
Sbjct: 576 GIAERLNRTIMDKVRSMLSRSGMEKKFWAEAASTAVYLINRSPSTAINFDLPEEKWTGAL 635
Query: 963 PNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHV 1022
P++S FGC+ Y+ + +L+ +S K + Y E KG++ + + K S +V
Sbjct: 636 PDLSSLRKFGCLAYIHADQGKLNP---RSKKGIFTSYPEGVKGYKVWVLEDKKCVISRNV 692
Query: 1023 RF-------DDKLDSDQSKLVEKFADLSIN-------VSDKGKA------PEEAEPEEDE 1062
F D K DS + DL +N +D+G A P EA +
Sbjct: 693 IFREQVMFKDLKGDSQNTISESDLEDLRVNPDMNDQEFTDQGGATQDNSNPSEATTSHNP 752
Query: 1063 PEEEAGPSDSQTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVS---- 1118
S + ++ A + +D RT A E+ + S
Sbjct: 753 VLNSPTHSQDEESEEEDSDAVEDLSTYQLVRDRVRRTIKANPKYNESNMVGFAYYSEDDG 812
Query: 1119 LIEPKSIDEALQDKDWI---LAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNE 1175
EPKS EAL D DW AM+EE+ SKN W LV KPE V +IG +WVF K
Sbjct: 813 KPEPKSYQEALLDPDWEKWNAAMKEEMVSMSKNHTWDLVTKPEKVKLIGCRWVFTRKAGI 872
Query: 1176 KG-DVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
G + R ARLVA+G++Q+EG+DY E F+P+ + +IR L+S V++N+ L QMDVK+A
Sbjct: 873 PGVEAPRFIARLVAKGFTQKEGVDYNEIFSPVVKHVSIRYLLSMVVHYNMELQQMDVKTA 932
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRG 1294
FL+G++ EE+Y+ QP GFE ++ + V LK+SLYGLKQ+PR W R F+ ++ R
Sbjct: 933 FLHGFLEEEIYMAQPEGFEIKRGSNKVCLLKRSLYGLKQSPRQWNLRFDEFMRGIKYTRS 992
Query: 1295 KVDTTLFCKTYKDDILV-VQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLG 1353
D+ ++ K D + + +YVDD++ SAN+S E +++ EFEM +G+ LG
Sbjct: 993 AYDSCVYFKKCNGDTYIYLLLYVDDMLIASANKSEVNELKQLLSREFEMKDLGDAKKILG 1052
Query: 1354 IQVDQTPEGTYI--HQSKYTKELLKKFNMLESTVAKTPMHPTCIL------EKEDASGKV 1405
+++ + + + Q Y K++L+ F M + TP+ L E E+ ++
Sbjct: 1053 MEISRDRDAGLLTLSQEGYVKKVLRSFQMDNAKPVSTPLGIHFKLKAATDKEYEEQFERM 1112
Query: 1406 CQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGL 1464
Y IGS++Y + +RPD+ +S+ + +RF S P + H AVK +LRY++ T L
Sbjct: 1113 KIVPYANTIGSIMYSMIGTRPDLAYSLGVISRFMSKPLKDHWQAVKWVLRYMRGTEKKKL 1172
Query: 1465 MYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYI 1524
++K ++ L GYCD+DY + R+S +G +G N +SW+SK Q +A+S+ EAEY+
Sbjct: 1173 CFRKQEDFLLRGYCDSDYGSNFDTRRSITGYVFTVGGNTISWKSKLQKVVAISSTEAEYM 1232
Query: 1525 SAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRD 1584
+ + LW+K + + + ++ D+ +AI+L+KN + H R KHI+++ HFIRD
Sbjct: 1233 ALTEAVKEALWLKGFAAELGHSQDYVEVHSDSQSAITLAKNSVHHERTKHIDIRLHFIRD 1292
Query: 1585 YV*KGVL 1591
+ G++
Sbjct: 1293 IICAGLI 1299
>At4g17450 retrotransposon like protein
Length = 1433
Score = 513 bits (1322), Expect = e-145
Identities = 312/998 (31%), Positives = 503/998 (50%), Gaps = 50/998 (5%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVDSSPCIYNVLL 667
+ W +DSG + H+T R ++ + V + KI G G I + + ++NVL
Sbjct: 430 RGWVIDSGATHHVTHNRDLYLNFRSLENTFVRLPNDCTVKIAGIGFIQLSDAISLHNVLY 489
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
+ NL+S + + SQ+ + + N+ ++ +
Sbjct: 490 IPEFKFNLIS----------ELTKELMIGRGSQVGNLYVLDFNENNHTVSLKGTTSMCPE 539
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSK-LNL-VRGLPNLKFASDALCEACQKGKFT 785
SV + WH+RLGH + KI LS LNL V+ + +C C K
Sbjct: 540 FSVCSSVVVDSVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQK 599
Query: 786 KVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRWTWVKFLTRKDES 845
+ F+++ + S +L+HID +GP + TW+ L K +
Sbjct: 600 HLSFQSRQNMC-SAAFDLVHIDTWGPFSVPTNDA--------------TWIYLLKNKSDV 644
Query: 846 HAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVV 905
VF FI V + ++ VRSD+ E KF LF ++GI SCP TP+QN VV
Sbjct: 645 LHVFPAFINMVHTQYQTKLKSVRSDNAHEL---KFTDLFAAHGIVAYHSCPETPEQNSVV 701
Query: 906 ERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYELWKNIKPNI 965
ERK++ + +AR +L ++ + FW + V TA ++ NR+ + NK+PYE KNI P
Sbjct: 702 ERKHQHILNVARALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAY 761
Query: 966 SYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTIEESIHVRFD 1025
FGC+CY + + HKF+ ++ C+ LGY KG++ + + + S HV F
Sbjct: 762 ESLKTFGCLCYSSTSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFH 821
Query: 1026 DKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDSQTLKK-SRITAAH 1084
+ + S ++ ++ D + +D P E+ D+ + S A
Sbjct: 822 EDIFPFISSTIKD------DIKDFFPLLQFPARTDDLPLEQTSIIDTHPHQDVSSSKALV 875
Query: 1085 PMELIMGNKDEPVRTRSAFRPSEETL----LSLKGLVSLIEPKSIDEALQDKDWILAMEE 1140
P + + + +P + F T + + + + P+ EA K W AM+E
Sbjct: 876 PFDPLSKRQKKPPKHLQDFHCYNNTTEPFHAFINNITNAVIPQRYSEAKDFKAWCDAMKE 935
Query: 1141 ELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYT 1200
E+ + + WS+V P + IG KWVF K N G + R KARLVA+GY+Q+EG+DY
Sbjct: 936 EIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGLDYE 995
Query: 1201 ETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFED---EKK 1257
ETF+P+A+L ++R+++ + +HQ+D+ +AFLNG + EE+Y+ PPG+ D E
Sbjct: 996 ETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVGEAL 1055
Query: 1258 PDH-VFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILVVQIYV 1316
P H + +L KS+YGLKQA R WY +LS+ L F + D TLF K ++ V +YV
Sbjct: 1056 PPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKYANGVLMGVLVYV 1115
Query: 1317 DDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQVDQTPEGTYIHQSKYTKELLK 1376
DDI+ S + +F+ +++ F++ +G YFLGI++ ++ +G I Q KY ELL
Sbjct: 1116 DDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQRKYILELLS 1175
Query: 1377 KFNMLESTVAKTPMHPTCILEKEDASGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCAR 1436
L S + P+ P+ L KED YR ++G L+YL +RPDI ++V+ +
Sbjct: 1176 TTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVNTLCQ 1235
Query: 1437 FQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNC 1496
F P HL+AV ++LRYLK T GL Y ++ L GY D+D+ R+ + C
Sbjct: 1236 FSHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCVAAYC 1295
Query: 1497 QFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIP---IY 1553
F+G LVSW+SK+Q T+++STAEAE+ + + + + +W+ +D+++ IP +Y
Sbjct: 1296 MFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKV--PFIPPAYLY 1353
Query: 1554 CDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KGVL 1591
CDNTAA+ + N + H R K +E+ + R+ V G L
Sbjct: 1354 CDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFL 1391
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 503 bits (1295), Expect = e-142
Identities = 322/1036 (31%), Positives = 543/1036 (52%), Gaps = 85/1036 (8%)
Query: 603 APLKHQSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTICVD----S 658
A + +W LD+GCS HMT + + K G+V G + ++ G G + + S
Sbjct: 305 AEVTPDTWILDTGCSFHMTCRKDWIIDFKETASGKVRMGNDTYSEVKGIGDVRIKNEDGS 364
Query: 659 SPCIYNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI 718
+ + +V + ++ NL+S+ L DKG ++K + + D +VL K+++ +Y +
Sbjct: 365 TILLTDVRYIPEMSKNLISLGTLEDKGC-WFESKKGILTIFKNDLTVL-TGKKESTLYFL 422
Query: 719 RLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQL-SKLNLVRGLPNLKFASDALCE 777
+ + L A + +E +WH RLGH + + L SK +L + + + F +
Sbjct: 423 QGTTL-AGEANVIDKEKDETSLWHSRLGHIGAKGLQVLVSKGHLDKNIM-ISFGA----- 475
Query: 778 ACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSE-SIGGKRYGMIIVDDYSRWTWV 836
AK+V T L+ +H DL+G SIG +Y + +DD++R TW+
Sbjct: 476 -------------AKHV--TKDKLDYVHSDLWGSTNVPFSIGKCQYFITFIDDFTRRTWI 520
Query: 837 KFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDFSCP 896
F+ KDE+ + F + Q++N++ ++ + +D+G EF N +F+S G+ +C
Sbjct: 521 YFIRTKDEAFSKFVEWKTQIENQQDKKLKILITDNGLEFCNQEFDSFCRKEGVIRHRTCA 580
Query: 897 RTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKTPYE 956
TPQQNGV ER NRT+ R ML E+G+ K FW EA +TA ++ N+ I P E
Sbjct: 581 YTPQQNGVAERMNRTIMNKVRCMLSESGLGKQFWAEAASTAVFLINKSPSSSIEFDIPEE 640
Query: 957 LWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYNTDAKTI 1016
W P+ FG V Y+ + + +L+ ++ K + LGY + K F+ + + +
Sbjct: 641 KWTGHPPDYKILKKFGSVAYIHSDQGKLNP---RAKKGIFLGYPDGVKRFKVWLLEDRKC 697
Query: 1017 EESIHVRFDDKLDSDQSKLVEKFADLSIN-VSDKGKAPEEAE---------PEEDEPEEE 1066
S + F + + + +L N +S++ K E E +DE + E
Sbjct: 698 VVSRDIVFQEN---------QMYKELQKNDMSEEDKQLTEVERTLIELKNLSADDENQSE 748
Query: 1067 AGPSDSQTLKKSRITAAHPMELIMGNKDEP---------------VRTRSAFRPSEETLL 1111
G + +Q + +A+ ++ + D+ +R F +++L+
Sbjct: 749 GGDNSNQEQASTTRSASKDKQVEETDSDDDCLENYLLARDRIRRQIRAPQRFVEEDDSLV 808
Query: 1112 SLKGLVS----LIEPKSIDEALQDKD---WILAMEEELNQFSKNDVWSLVKKPESVHVIG 1164
++ + EP++ +EA++ + W A EE++ KND W ++ KPE VIG
Sbjct: 809 GFALTMTEDGEVYEPETYEEAMRSPECEKWKQATIEEMDSMKKNDTWDVIDKPEGKRVIG 868
Query: 1165 TKWVFRNKLNEKG-DVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISFSVNHN 1223
KW+F+ K G + R KARLVA+G+SQ+EGIDY E F+P+ + +IR L+S V +
Sbjct: 869 CKWIFKRKAGIPGVEPPRYKARLVAKGFSQREGIDYQEIFSPVVKHVSIRYLLSIVVQFD 928
Query: 1224 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1283
+ L Q+DVK+AFL+G + E + + QP G+EDE + V LKKSLYGLKQ+PR W +R
Sbjct: 929 MELEQLDVKTAFLHGNLDEYILMSQPEGYEDEDSTEKVCLLKKSLYGLKQSPRQWNQRFD 988
Query: 1284 SFLLENEFVRGKVDTTLFCKTYKD-DILVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEM 1342
SF++ + + R K + ++ + D + + +YVDD++ S N+ ++ + + EFEM
Sbjct: 989 SFMINSGYQRSKYNPCVYTQQLNDGSYIYLLLYVDDMLIASQNKDQIQKLKESLNREFEM 1048
Query: 1343 SMMGELTYFLGIQVDQTPEGTYIH--QSKYTKELLKKFNMLESTVAKTPMHPTCIL---- 1396
+G LG+++ + E + QS+Y +L+ F M +S V++TP+ L
Sbjct: 1049 KDLGPARKILGMEITRNREQGILDLSQSEYVAGVLRAFGMDQSKVSQTPLGAHFKLRAAN 1108
Query: 1397 EKEDASGKVCQKL--YRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
EK A KL Y IGS++Y + SRPD+ + V + +RF S P + H AVK ++
Sbjct: 1109 EKTLARDAEYMKLVPYPNAIGSIMYSMIGSRPDLAYPVGVVSRFMSKPSKEHWQAVKWVM 1168
Query: 1454 RYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWESKRQST 1513
RY+K T + L +KK ++++ GYCD+DYA D R+S +G G N +SW+S Q
Sbjct: 1169 RYMKGTQDTCLRFKKDDKFEIRGYCDSDYATDLDRRRSITGFVFTAGGNTISWKSGLQRV 1228
Query: 1514 IALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAK 1573
+ALST EAEY++ A + +W++ + + + + CD+ +AI+LSKN + H R K
Sbjct: 1229 VALSTTEAEYMALAEAVKEAIWLRGLAAEMGFEQDAVEVMCDSQSAIALSKNSVHHERTK 1288
Query: 1574 HIEVKYHFIRDYV*KG 1589
HI+V+YHFIR+ + G
Sbjct: 1289 HIDVRYHFIREKIADG 1304
>At4g03810 putative retrotransposon protein
Length = 964
Score = 478 bits (1229), Expect = e-134
Identities = 306/952 (32%), Positives = 488/952 (51%), Gaps = 60/952 (6%)
Query: 664 NVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNN--------- 714
N V + N++S+S L +G+ K C S + + S +N
Sbjct: 7 NCYYVPAINKNIISVSCLDMEGFHFSIKNKCC---SFDRDDMFYGSAPLDNGLHVLNQSM 63
Query: 715 -IYKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASD 773
IY IR + ++ ++ ++WH RLGH + + I +L L L + + S
Sbjct: 64 PIYNIRTKKFKSNDLN-------PTFLWHCRLGHINEKHIQKLHSDGL---LNSFDYESY 113
Query: 774 ALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKSESIGGKRYGMIIVDDYSRW 833
CE+C GK TK PF + S L L+H D+ GP+ + + G +Y + DD+SR+
Sbjct: 114 ETCESCLLGKMTKAPFTGHSE-RASDLLGLIHTDVCGPMSTSARGNYQYFITFTDDFSRY 172
Query: 834 TWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSYGIAHDF 893
+V + K +S F F +VQN+ I +RSD GGE+ + F GI
Sbjct: 173 GYVYLMKHKSKSFENFKEFQNEVQNQFGKSIKALRSDRGGEYLSQVFSDHLRECGIVSQL 232
Query: 894 SCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVRPILNKT 953
+ P TPQ NGV ER+NRTL +M R+M+ T + FW A+ T+ ++ NR + + KT
Sbjct: 233 TPPGTPQWNGVSERRNRTLLDMVRSMMSHTDLPSPFWGYALETSAFMLNRCPSKSV-EKT 291
Query: 954 PYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFRFYN-TD 1012
PYE+W PN+S+ +GC Y + K KS KC +GY + +KG+ FY+ TD
Sbjct: 292 PYEIWTGKVPNLSFLKIWGCESYA--KRLITDKLGPKSDKCYFVGYPKETKGYYFYHPTD 349
Query: 1013 AKTIEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEAEPEEDEPEEEAGPSDS 1072
K + + + L +F + S K E EP+ D P +
Sbjct: 350 NKVF-----------VVRNGAFLEREFLSKGTSGS-KVLLEEVREPQGDVPTSQEEHQLD 397
Query: 1073 QTLKKSRITAAHPMELIMGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDK 1132
I + ++ EP R R ++ EP S +EAL
Sbjct: 398 LRRVVEPILVEPEVRRSERSRHEPDRFRDWVMDDHALF-----MIESDEPTSYEEALMGP 452
Query: 1133 D---WILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQ 1189
D W+ A + E+ S+N VW+LV P+ V I KW+F+ K++ G++ KA LVA+
Sbjct: 453 DSDKWLEAAKSEMESMSQNKVWTLVDLPDGVKPIECKWIFKKKIDMDGNIQIYKAGLVAK 512
Query: 1190 GYSQQEGIDYTETFAPIARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQP 1249
GY Q GIDY ET++P+A L++IR+L++ + +++ + QMDVK+AFLNG + E VY+ QP
Sbjct: 513 GYKQVHGIDYDETYSPVAMLKSIRILLATAAHYDYEIWQMDVKTAFLNGNLEEHVYMTQP 572
Query: 1250 PGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDI 1309
GF + V KL +S+YGLKQA R+W R + + E +F+R + + ++ KT +
Sbjct: 573 EGFTVPEAARKVCKLHRSIYGLKQASRSWNLRFNEAIKEFDFIRNEEEPCVYKKTSGSAV 632
Query: 1310 LVVQIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELTYFLGIQV--DQTPEGTYIHQ 1367
+ +YVDDI+ + L + + + F M MGE Y LGI++ D+ + + Q
Sbjct: 633 AFLVLYVDDILLLGNDIPLLQSVKTWLGSCFSMKDMGEAAYILGIRIYRDRLNKIIGLSQ 692
Query: 1368 SKYTKELLKKFNMLESTVAKTPMHPTCILEK------EDASGKVCQKLYRGMIGSLLY-L 1420
Y ++L +FNM +S PM L K D ++ + Y IGS++Y +
Sbjct: 693 DTYIDKVLHRFNMHDSKKGFIPMSHGITLSKTQCPSTHDERERMSKIPYASAIGSIMYAM 752
Query: 1421 TASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKDTTNLGLMYKKTSEYKLSGYCDA 1480
+RPD+ ++ + +R+QSDP E+H V+ I +YL+ T + L+Y + E +SGY DA
Sbjct: 753 LYTRPDVACALSMTSRYQSDPGESHWIVVRNIFKYLRRTKDKFLVYGGSEELVVSGYTDA 812
Query: 1481 DYAGDRTERKSTSGNCQFLGSNLVSWESKRQSTIALSTAEAEYISAAICSTQKLWMKHQL 1540
+ D+ + +S SG L VSW+S +QST+A ST EAEYI+A+ + + +W++ +
Sbjct: 813 SFQTDKDDFRSQSGFFFCLNGGAVSWKSTKQSTVADSTTEAEYIAASEAAKEVVWIRKFI 872
Query: 1541 EDYQILES---NIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV*KG 1589
+ ++ S I +YCDN AI+ +K P H ++KHI+ +YH IR+ + +G
Sbjct: 873 TELGVVPSISGPIDLYCDNNGAIAQAKEPKSHQKSKHIQRRYHLIREIIDRG 924
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 454 bits (1168), Expect = e-127
Identities = 282/801 (35%), Positives = 425/801 (52%), Gaps = 59/801 (7%)
Query: 828 DDYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDHGGEFENDKFESLFDSY 887
+D+SR WV FL KDE+ A F+ + V+ + ++ +R+D+G EF N KF+ +
Sbjct: 319 NDWSRKVWVYFLKTKDEAFASFTEWKKMVETQSERKLKHLRTDNGLEFCNHKFDEVCKKE 378
Query: 888 GIAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWTEAVNTACYIQNRISVR 947
GI +C TPQQNGV ER NRT+ R+ML E+G+ K FW +A +TA Y+ NR
Sbjct: 379 GIVRHRTCTYTPQQNGVAERLNRTIMNKVRSMLSESGLDKKFWAKAASTAVYLINRSPSS 438
Query: 948 PILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSERSKGFR 1007
I NK P ELW + PN S FGCV YV + + +L D ++ K + +GY KGFR
Sbjct: 439 SIENKIPEELWTSAVPNFSGLKRFGCVVYVYSQEGKL---DPRAKKGVFVGYPNGVKGFR 495
Query: 1008 FYNTDAKTIEESIHVRF-DDKLDSD---QSKLVEKFADLSINVSDKGKAPEEAEPEEDEP 1063
+ + + S +V F +D + D QS F D + + +EDE
Sbjct: 496 VWMIEEERCSISRNVVFREDVMYKDILNQSTSGMSF-DFPLATNRIPSFECAGNRKEDEI 554
Query: 1064 EEEAGPSDSQTLKKSRITAAHP--------MELIMGNKDEPVRTRSA------FRPSEET 1109
+ G SD T + S + +D+P R + +EE
Sbjct: 555 SVQGGVSDDDTKQSSEESPISTGSSGQNSGQRTYQIARDKPKRQTKIPDKLRDYELNEEV 614
Query: 1110 LLSLKGLVSLI-------EPKSIDEALQDKD---WILAMEEELNQFSKNDVWSLVKKPES 1159
L + G +I EP +ALQD D W+ A++EE+ KN+ W LV + +
Sbjct: 615 LDEIAGYAYMITEDGGNPEPNDYQKALQDSDYKMWLKAVDEEIESLLKNNTWVLVNRDQF 674
Query: 1160 VHVIGTKWVFRNKLNEKG-DVVRNKARLVAQGYSQQEGIDYTETFAPIARLEAIRLLISF 1218
IG KWVF+ K G + R KARLV +GYSQ+EGIDY E F+P+ + +IRLL+S
Sbjct: 675 QKPIGCKWVFKRKSGIVGVEKPRFKARLVVKGYSQKEGIDYQEIFSPVVKHVSIRLLLSM 734
Query: 1219 SVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAW 1278
+ ++ L QMDVK+AFL+GY+ E +Y+ QP G+ ++ PD V LK+SLYGL+Q+PR W
Sbjct: 735 VTHCDMELQQMDVKTAFLHGYLDETIYIEQPEGYVHKRYPDKVCLLKRSLYGLRQSPRQW 794
Query: 1279 YERLSSFLLENEFVRGKVDTTLFCKTYKD-DILVVQIYVDDIIFGSANQSLCKEFSKMMQ 1337
R + F+ + + R K D+ ++ K + + + + +YVDDI+ S ++ + ++
Sbjct: 795 NNRFNEFMQKIGYERSKYDSCVYFKELQSGEYIYLLLYVDDILIASRDKRTVCDLKALLN 854
Query: 1338 AEFEMSMMGELTYFLGIQV--DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPM----- 1390
+EFEM +G+ LG+++ D+ I Q Y ++L F M ++ TPM
Sbjct: 855 SEFEMKDLGDAKKILGMEIVRDRKAGTMSISQEGYLLKVLGNFGMDQAKPVFTPMGAHFK 914
Query: 1391 -HPTCILEKEDASGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTA 1448
P E S + Y+ +GSL+Y + +RPD+ SV L RF S P + H A
Sbjct: 915 LKPATDEEVMRQSEVMRAVPYQSAVGSLMYSMIGTRPDLAHSVGLVCRFMSKPLKEHWQA 974
Query: 1449 VKRILRYLKDTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWES 1508
VK ILRY++ + + L YK E L GYCD+DYA D+ R+STSG
Sbjct: 975 VKWILRYIRGSIDRKLCYKNEGELILEGYCDSDYAADKEGRRSTSG-------------- 1020
Query: 1509 KRQSTIALSTAEAEYISAAICSTQKLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPIL 1568
+ALS+ EAEY++ + + +W+K + + ++ + I+CD+ +AI+L+KN +
Sbjct: 1021 --VKVVALSSTEAEYMALTDGAKEAIWLKGHVSELGFVQKTVNIHCDSQSAIALAKNAVY 1078
Query: 1569 HSRAKHIEVKYHFIRDYV*KG 1589
H R KHI+VKYHFIRD V G
Sbjct: 1079 HERTKHIDVKYHFIRDLVNNG 1099
Score = 33.9 bits (76), Expect = 0.77
Identities = 13/47 (27%), Positives = 22/47 (46%)
Query: 608 QSWYLDSGCSRHMTGERRMFQELKLKPGGEVGFGGNEKGKIVGTGTI 654
+ W +D+GCS HMT + + G+V N ++ G G +
Sbjct: 264 EEWIMDTGCSFHMTPRKEYLMDFVEAKSGKVRMANNSFSEVKGIGKV 310
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.332 0.142 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,299,539
Number of Sequences: 26719
Number of extensions: 1332035
Number of successful extensions: 7290
Number of sequences better than 10.0: 146
Number of HSP's better than 10.0 without gapping: 114
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 6522
Number of HSP's gapped (non-prelim): 317
length of query: 1591
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1478
effective length of database: 8,299,349
effective search space: 12266437822
effective search space used: 12266437822
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0388a.6


