

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
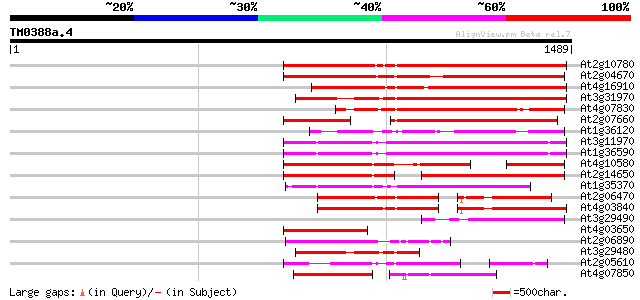
Query= TM0388a.4
(1489 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 812 0.0
At2g04670 putative retroelement pol polyprotein 701 0.0
At4g16910 retrotransposon like protein 683 0.0
At3g31970 hypothetical protein 640 0.0
At4g07830 putative reverse transcriptase 511 e-144
At2g07660 putative retroelement pol polyprotein 486 e-137
At1g36120 putative reverse transcriptase gb|AAD22339.1 474 e-133
At3g11970 hypothetical protein 451 e-126
At1g36590 hypothetical protein 446 e-125
At4g10580 putative reverse-transcriptase -like protein 398 e-110
At2g14650 putative retroelement pol polyprotein 393 e-109
At1g35370 hypothetical protein 372 e-103
At2g06470 putative retroelement pol polyprotein 329 6e-90
At4g03840 putative transposon protein 322 1e-87
At3g29490 hypothetical protein 290 6e-78
At4g03650 putative reverse transcriptase 288 2e-77
At2g06890 putative retroelement integrase 287 4e-77
At3g29480 hypothetical protein 285 1e-76
At2g05610 putative retroelement pol polyprotein 231 3e-60
At4g07850 putative polyprotein 207 5e-53
>At2g10780 pseudogene
Length = 1611
Score = 812 bits (2097), Expect = 0.0
Identities = 409/757 (54%), Positives = 535/757 (70%), Gaps = 13/757 (1%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + EL+A SKC FW V FLGHVIS QGV+VDP KI SI W +P+ A++IRSF+
Sbjct: 834 ERLREHELFAKLSKCSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFL 893
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GLAGYYRRFV +A + PLT+LT K+ F W+++CE+SF E+K LT A +L LP+EGE
Sbjct: 894 GLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGE 953
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
PY VY DAS GLGCV+MQ+ VIAYASRQL+ HEKNYPTHDLE+AA+V+ LKIWR YLY
Sbjct: 954 PYTVYTDASIVGLGCVLMQKGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLY 1013
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
G+ IY+DHKSLKY+F Q +LN+RQRRWME + DY D+ HPGK N VADALSR+
Sbjct: 1014 GAKVQIYTDHKSLKYIFTQPELNLRQRRWMELVADYNLDIAYHPGKANQVADALSRRRSE 1073
Query: 967 VSSLMVKELELLEAF*DL---SLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEW 1023
V + +++L+ L +L ++ P L G + LL I+ QE DE + W
Sbjct: 1074 VEAER-SQVDLVNMMGTLHVNALSKEVEP--LGLGAAD-QADLLSRIRLAQERDEEIKGW 1129
Query: 1024 RKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDL 1083
Q E+ ++ + GRVCVPND ++ IL E H+S+ SIHPG NKMY+DL
Sbjct: 1130 ----AQNNKTEYQTSNNGTIVVNGRVCVPNDRALKEEILREAHQSKFSIHPGSNKMYRDL 1185
Query: 1084 KLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPL 1143
K ++ W GMKK VA +V+ C TC K EHQ P+G LQ+L +PEWKWD I+MDFVT LP
Sbjct: 1186 KRYYHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPIPEWKWDHITMDFVTGLPT 1245
Query: 1144 -TRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRF 1202
+ + +A+WVVVDRLTK+AHF+ IS E +AE +I +IVRLHG+P SIVSDRD+RF
Sbjct: 1246 GIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIVRLHGIPVSIVSDRDTRF 1305
Query: 1203 TSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIE 1262
TS+FW A Q+ALGT++ LS+AY+PQTD Q+ERTIQ+LED+LRACVLD G+W++ L L+E
Sbjct: 1306 TSKFWKAFQKALGTRVNLSTAYHPQTDEQSERTIQTLEDMLRACVLDWGGNWEKYLRLVE 1365
Query: 1263 FTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRI 1322
F YNNSF ASIGM+PYEALYGR C+TPLCW GE + GP +V +TTE +K ++ K++
Sbjct: 1366 FAYNNSFQASIGMSPYEALYGRACRTPLCWTPVGERRLFGPTIVDETTERMKFLKIKLKE 1425
Query: 1323 S*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPV 1382
+ RQKSYA+ RRKELEFQ GD V+L+ G GR KKL+P+++GPY++ ERVG V
Sbjct: 1426 AQDRQKSYANKRRKELEFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAV 1485
Query: 1383 AYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKR 1442
AY++ LPP L+ H+V HVSQLRK ++D +E LK+++TV P++I+DR TK
Sbjct: 1486 AYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDIPPGLKENMTVEAWPVRIMDRMTKG 1545
Query: 1443 LRNK*VSLVKVVWN-QATGDATWELEDKMRESHPDLF 1478
R K L+KV+WN + + TWE E+KM+ + P+ F
Sbjct: 1546 TRGKARDLLKVLWNCRGREEYTWETENKMKANFPEWF 1582
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 701 bits (1809), Expect = 0.0
Identities = 357/746 (47%), Positives = 490/746 (64%), Gaps = 40/746 (5%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + ++L+A SKC FW E+ FLGH++S++GV+VDP KIE+I W +P A++IRSF+
Sbjct: 692 EKLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFL 751
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
L GYYRRFVK +A + P+T+LT K+ PF W+ +CEE F +K+ LT+ +LALP+ G+
Sbjct: 752 RLTGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGQ 811
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
PY VY DAS GLGCV+MQ KVIAYASRQL+ HE NYPTHDLE+A +++ALKIWR YLY
Sbjct: 812 PYMVYTDASRVGLGCVLMQRGKVIAYASRQLRKHEGNYPTHDLEMAVVIFALKIWRSYLY 871
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
G +++DHKSLKY+F+Q +LN+RQ RWME + DY+ ++ HPGK NVVADALSRK +
Sbjct: 872 GGKVQVFTDHKSLKYIFNQPELNLRQMRWMELVADYDLEIAYHPGKANVVADALSRKRVG 931
Query: 967 VSSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKL 1026
+ E L+ L L V +A L V + LL + QE DEGL+
Sbjct: 932 AAPGQSVE-ALVSEIGALRLCV-VAREPLGLEAVD-RADLLTRARLAQEKDEGLI----A 984
Query: 1027 VTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLH 1086
++ + E+ ++ + GRVCVP D +RR IL E H S SIHPG KMY+DLK +
Sbjct: 985 ASKAEGSEYQFAANGTIFVYGRVCVPKDEELRREILSEAHASMFSIHPGATKMYRDLKRY 1044
Query: 1087 FWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRR 1146
+ W GMK+ VA +V+ C C K EHQ P
Sbjct: 1045 YQWVGMKRDVANWVAECDVCQLVKAEHQVP------------------------------ 1074
Query: 1147 RFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRF 1206
DAIWV++DRLTK+AHF+ I LA+ ++++IV+LHGVP SIVSDRDS+FT F
Sbjct: 1075 --DAIWVIMDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTFAF 1132
Query: 1207 WGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYN 1266
W A Q +GTK+++S+AY+PQTDGQ+ERTIQ+LED+LR CVLD G W + L L+EF YN
Sbjct: 1133 WRAFQAKMGTKVQMSTAYHPQTDGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYN 1192
Query: 1267 NSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SR 1326
NS+ ASIGMAP+EALYGR C TPL W Q E + G + VQ+TTE ++ ++ M+ + +R
Sbjct: 1193 NSYQASIGMAPFEALYGRPCWTPLRWTQVEERSIYGADYVQETTERIRVLKLNMKEAQAR 1252
Query: 1327 QKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRI 1386
Q+SYAD RR+ELEF+ GD V+L++ + G R+I KL+P+++GP++I ERVGPVAYR+
Sbjct: 1253 QRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSILETKLSPRYMGPFRIVERVGPVAYRL 1312
Query: 1387 ALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK 1446
LP + H V HV LRK + D VL L+ ++T+ P+++++R K LR K
Sbjct: 1313 ELPDVMRAFHKVFHVLMLRKCLHKDDEVLVKIPEDLQPNMTLEARPVRVLERRIKELRRK 1372
Query: 1447 *VSLVKVVWN-QATGDATWELEDKMR 1471
+ L+KV+W+ + TWE E +M+
Sbjct: 1373 KIPLIKVLWDCDGVTEETWEPEARMK 1398
>At4g16910 retrotransposon like protein
Length = 687
Score = 683 bits (1762), Expect = 0.0
Identities = 347/681 (50%), Positives = 467/681 (67%), Gaps = 19/681 (2%)
Query: 802 LTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGC 861
+ P+TQLT K+ F W+E+CE SF E+K LT A +L LP+EGEPY VY DAS GLGC
Sbjct: 1 MAQPMTQLTGKDTAFNWSEECERSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGC 60
Query: 862 VMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKY 921
V++Q+ VIAYASRQL+ HEKNYPT+DLE+AA+V+ALKIWR YLYG+ I++DHKSLKY
Sbjct: 61 VLIQKGSVIAYASRQLRKHEKNYPTNDLEMAAVVFALKIWRSYLYGAKVQIFTDHKSLKY 120
Query: 922 LFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSR--KAMHVSSLMVKELELLE 979
+F Q +LN+RQRRW++ + DY+ ++ HPGK N V DALSR + V + ++
Sbjct: 121 IFTQPELNLRQRRWIKLVADYDLNIAYHPGKANQVVDALSRLRTVVEAERNQVNLVNMMG 180
Query: 980 AF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIGS 1039
+L ++ P L + LL I+S QE DE + W Q E+ +
Sbjct: 181 TLHLNALSKEVEPLGLR---AANQADLLSRIRSAQERDEEIKGW----AQNNKTEYQSSN 233
Query: 1040 DNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEY 1099
+ + GRVC PND ++ IL E H+S+ SIHPG NKMY+DLK ++ W GMKK VA +
Sbjct: 234 NGTIVVNGRVCGPNDKALKEEILKEAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARW 293
Query: 1100 VSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPL-TRRRFDAIWVVVDRL 1158
V AK EHQ P+G LQ+L +PEWKWD I MDFVT LP + + +A+WVVVDRL
Sbjct: 294 V--------AKEEHQVPSGMLQNLPIPEWKWDHIMMDFVTGLPTGIKSKHNAVWVVVDRL 345
Query: 1159 TKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKL 1218
TK+AHF+ IS E +AE +I +IVRLHG+P SIVSDRD+RFTS+FW Q+ LGT++
Sbjct: 346 TKSAHFMAISDKDAAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKPFQKVLGTRV 405
Query: 1219 RLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPY 1278
LS+AY+PQTDGQ+ERTIQ+LED+LRACVLD G+W++ L L+EF YNNSF ASIGM+PY
Sbjct: 406 NLSTAYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPY 465
Query: 1279 EALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRKEL 1338
EALYGR +TPLCW GE + GP +V +TT+++K ++ K++ + RQKSYA+ RRKEL
Sbjct: 466 EALYGRAGRTPLCWTPVGERRLFGPAVVDETTKKMKFLKIKLKEAQDRQKSYANKRRKEL 525
Query: 1339 EFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDV 1398
EFQ GD V+L+ G GR KKL P+++GPY++ ERVG VAY++ LPP L H+V
Sbjct: 526 EFQVGDLVYLKAMTYKGAGRFTSRKKLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFHNV 585
Query: 1399 LHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWN-Q 1457
HVSQLRK +++ +E LK+++TV P++I+D+ K R K + L+K++WN
Sbjct: 586 FHVSQLRKCLSEQEESMEDVPPGLKENMTVEAWPVRIMDQMKKGTRGKSMDLLKILWNCG 645
Query: 1458 ATGDATWELEDKMRESHPDLF 1478
+ TWE E KM+ + P+ F
Sbjct: 646 GREEYTWETETKMKANFPEWF 666
>At3g31970 hypothetical protein
Length = 1329
Score = 640 bits (1652), Expect = 0.0
Identities = 336/723 (46%), Positives = 455/723 (62%), Gaps = 60/723 (8%)
Query: 759 GVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAW 818
GV+VD KIE+I W +P A++IRSF+GLAGYY+RFVK +A + P+T+LT K+ PF W
Sbjct: 658 GVSVDLEKIEAIRDWPRPTNATEIRSFLGLAGYYKRFVKVFASMAQPMTKLTGKDVPFVW 717
Query: 819 TEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQLK 878
+ +C+E F +K+ LT+ +LALP+ GEPY VY DAS GL CV+MQ KV
Sbjct: 718 SPECDEGFMSLKEMLTSTPVLALPEHGEPYMVYTDASRVGLDCVLMQRGKV--------- 768
Query: 879 IHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEF 938
++DHKSLKY+F Q +LN+RQRRWME
Sbjct: 769 ----------------------------------FTDHKSLKYIFTQPELNLRQRRWMEL 794
Query: 939 LQDYEFDLQ*HPGKTNVVADALSRKAMHVS-SLMVKELELLEAF*DLSLDVKIAPGKLSF 997
+ DY+ ++ H GK NVVADALSRK + S +V E+ L +
Sbjct: 795 VADYDLEIAYHSGKANVVADALSRKRVGGSVEALVSEIGALRL---------CVMAQEPL 845
Query: 998 GMVTVS-SGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDAT 1056
G+ V + LL ++ QE DEGL+ ++ E+ ++ + GRVCVP D
Sbjct: 846 GLEAVDRADLLTRVRLAQEKDEGLIA----ASKADGSEYQFAANGTILVHGRVCVPKDEE 901
Query: 1057 MRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKP 1116
+RR IL E H S SIHP KMY+DLK ++ W GMK+ VA +V+ C C K EHQ P
Sbjct: 902 LRREILSEAHASMFSIHPRATKMYRDLKRYYQWVGMKRDVANWVTECDVCQLVKAEHQVP 961
Query: 1117 AGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKL 1176
G LQSL + EWKWD I+MDFV LP++R + DAIWV+VDRLTK+AHF+ I L
Sbjct: 962 GGLLQSLPILEWKWDFITMDFVVGLPVSRTK-DAIWVIVDRLTKSAHFLAIRKTDGAVLL 1020
Query: 1177 AEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTI 1236
A+ ++++IV LHGVP SIVSDRDS+FTS FW A Q +GTK+++S+AY+PQTDGQ+ERTI
Sbjct: 1021 AKKYVSEIVELHGVPVSIVSDRDSKFTSAFWRAFQGEMGTKVQMSTAYHPQTDGQSERTI 1080
Query: 1237 QSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDG 1296
Q+LED+LR CVLD G W + L L+EF YNNS+ ASI MAP+EALYGR C+TPLCW Q G
Sbjct: 1081 QTLEDMLRMCVLDRGGHWADHLSLVEFAYNNSYQASIRMAPFEALYGRPCRTPLCWTQVG 1140
Query: 1297 EHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGV 1356
E + G + V +TTE ++ ++ M+ + RQ+SYAD RR+ELEF+ GD V+L++ + G
Sbjct: 1141 ERSIYGADYVLETTERIRVLKLNMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGP 1200
Query: 1357 GRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLE 1416
R+I KLTP+++GP++I ERVGPVAYR+ LP + H V HVS LRK + D VL
Sbjct: 1201 NRSISETKLTPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEVLA 1260
Query: 1417 PDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWN-QATGDATWELEDKMRESHP 1475
L+ ++T+ P++I++R K LR K + L+KV+WN + TWE E +M+ S
Sbjct: 1261 KILEDLQPNMTLEARPVRILERRIKELRRKKIPLIKVLWNCDGVTEETWEPEARMKASFK 1320
Query: 1476 DLF 1478
F
Sbjct: 1321 KWF 1323
>At4g07830 putative reverse transcriptase
Length = 611
Score = 511 bits (1316), Expect = e-144
Identities = 277/610 (45%), Positives = 381/610 (62%), Gaps = 50/610 (8%)
Query: 864 MQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLF 923
MQ KVIAY SRQLK HE NYPTHDLE+AA+ ++DHKSLKY+F
Sbjct: 1 MQRGKVIAYGSRQLKKHEGNYPTHDLEMAAV------------------FTDHKSLKYIF 42
Query: 924 DQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHVSSLMVKELELLEAF*D 983
Q +LN+R RRWM+ + DY+ ++ HPGK NVV DALSRK + + E ++E
Sbjct: 43 TQLELNLRLRRWMKLVADYDLEIAYHPGKANVVTDALSRKRVGAAPGQSVETLVIE---- 98
Query: 984 LSLDVKIAPGKLSFGMVTVS-SGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIGSDNI 1042
+ A + G+ V + LL ++ QE DEGL+ ++ + E+ ++
Sbjct: 99 IGALRLCAVAREPLGLEAVDQTDLLSRVQLAQEKDEGLIA----ASKAEGFEYQFAANGT 154
Query: 1043 LRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVST 1102
+ GRVCVP D +R+ IL E H S SIHPG KMY+DLK ++ W GMK+ V +V
Sbjct: 155 ILVHGRVCVPKDKELRQEILSEAHASMFSIHPGATKMYRDLKRYYQWVGMKRDVGNWVEE 214
Query: 1103 CLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTA 1162
C C K+EHQ LQSL +PEWKWD I+MDFV LP++R + DAIWV+VDRLTK+A
Sbjct: 215 CDVCQLVKIEHQVSGSLLQSLPIPEWKWDFITMDFVVGLPVSRTK-DAIWVIVDRLTKSA 273
Query: 1163 HFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSS 1222
HF+ I LA+ ++++IV+LHGVP SIVSDRDS+FTS FW A Q +GTK+++S+
Sbjct: 274 HFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMST 333
Query: 1223 AYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALY 1282
AY+PQTDGQ+ERTIQ+LED+LR CVLD +G W + L L+EF YNNS+ ASIGMAP+E LY
Sbjct: 334 AYHPQTDGQSERTIQTLEDMLRMCVLDWRGHWADHLSLVEFAYNNSYQASIGMAPFEVLY 393
Query: 1283 GRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRKELEFQA 1342
GR C+T LCW Q GE + G + VQ+ TE ++ ++ M+ + +RQ+SYAD RRKELEF+
Sbjct: 394 GRPCRT-LCWTQVGERSIYGADYVQEITERIRVLKLNMKEAQNRQRSYADKRRKELEFEV 452
Query: 1343 GDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVS 1402
GD V + G +S+ +RVGPVA+R+ L + H V HVS
Sbjct: 453 GDSV-------SQDGHVARSE-------------QRVGPVAFRLELSDVMRAFHKVFHVS 492
Query: 1403 QLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWN-QATGD 1461
LRK + D VL L+ ++T+ P+++++R K LR K + L+KV+ N +
Sbjct: 493 MLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLRNCDGVTE 552
Query: 1462 ATWELEDKMR 1471
TWE E +++
Sbjct: 553 ETWEPEARLK 562
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 486 bits (1250), Expect = e-137
Identities = 240/446 (53%), Positives = 320/446 (70%), Gaps = 5/446 (1%)
Query: 1010 IKSKQETDEGLLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSR 1069
I+ QE DE + W T E+ ++ + GRVCVPND ++ IL E H+S+
Sbjct: 508 IRLAQERDEEIKGW----TLNNKTEYQTSNNGTIVVNGRVCVPNDRALKEEILREAHQSK 563
Query: 1070 LSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWK 1129
SIHPG NKMY+DLK ++ W GM+K VA +V+ C TC K EHQ P+G LQ+L + EWK
Sbjct: 564 FSIHPGSNKMYRDLKRYYHWVGMRKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPISEWK 623
Query: 1130 WDSISMDFVTALPL-TRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLH 1188
WD I+MDFVT LP + + +A+WVVVDRLTK+AHF+ IS E +AE +I +I+RLH
Sbjct: 624 WDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIMRLH 683
Query: 1189 GVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVL 1248
G+P SIVSDRD+RFTS+FW A Q+ALGT++ LS+AY+PQTDGQ+ERTIQ+LED+LRACVL
Sbjct: 684 GIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLSTAYHPQTDGQSERTIQTLEDMLRACVL 743
Query: 1249 DHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQ 1308
D G+W++ L LIEF YNNSF ASIGM+PYEALYGR C+TPLCW GE + GP +V +
Sbjct: 744 DWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALYGRACRTPLCWTPVGERRLFGPTIVDE 803
Query: 1309 TTEEVKRIQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPK 1368
TTE +K ++ K++ + RQKSYA+ RRKELEFQ GD V+L+ G GR KKL+P+
Sbjct: 804 TTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVYLKAMTYKGAGRFTSRKKLSPR 863
Query: 1369 FIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTV 1428
++GPY++ ERVG VAY++ LPP L+ H+V HVSQLRKY++D +E LK+++TV
Sbjct: 864 YVGPYKVIERVGAVAYKLDLPPKLNVFHNVFHVSQLRKYLSDQEESVEDIPPGLKENMTV 923
Query: 1429 VMPPIKIVDRSTKRLRNK*VSLVKVV 1454
P++I+DR +K R K L+KV+
Sbjct: 924 EAWPVRIMDRMSKGTRGKSRDLLKVL 949
Score = 231 bits (589), Expect = 2e-60
Identities = 111/179 (62%), Positives = 136/179 (75%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + EL+A SK FW V FLGHVIS QGV+VDP KI SI W +P+ A++IRSF+
Sbjct: 329 ERLREHELFAKLSKFSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFL 388
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GLAGYYRRFV +A + PLT+LT K+ F W+++CE+SF E+K L A +L LP+EGE
Sbjct: 389 GLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGE 448
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYL 905
PY VY DAS GLGCV+MQ+ VIAYASRQL+ HEKNYPTHDLE+AA+V+ALKIWR YL
Sbjct: 449 PYTVYTDASIVGLGCVLMQKGSVIAYASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYL 507
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 474 bits (1219), Expect = e-133
Identities = 269/676 (39%), Positives = 371/676 (54%), Gaps = 145/676 (21%)
Query: 797 KDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASH 856
K +A + P+T+LT+K+ PF W+ +CEE F+
Sbjct: 691 KGFASMAQPMTKLTEKDIPFVWSPECEEGFRG---------------------------- 722
Query: 857 NGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDH 916
KVI YASRQL+ HE NYPTHDLE+AAI++ALKIW YLYG +++DH
Sbjct: 723 -----------KVIVYASRQLRKHEGNYPTHDLEMAAIIFALKIWGSYLYGGKVQVFTDH 771
Query: 917 KSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHVSSLMVKELE 976
KSLK +F Q +LN+RQRRWME + DY+ ++ HPGKTNVVADALSRK
Sbjct: 772 KSLKDIFTQPELNLRQRRWMELVADYDLEIAYHPGKTNVVADALSRKR------------ 819
Query: 977 LLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFS 1036
V APG+ +V+ L + +++ KL +A
Sbjct: 820 -----------VGAAPGQSVEALVSEIGALRLCVVAREPL--------KLEAVDRA---- 856
Query: 1037 IGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQV 1096
++ + RVC+P D +RR IL E H S SIHPG KMY+DLK H+ W GMK+ V
Sbjct: 857 --ANGTILVHERVCLPKDEELRREILSEAHASMFSIHPGATKMYRDLKRHYQWVGMKRDV 914
Query: 1097 AEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVD 1156
A +V+ C C K EHQ P G LQSL + EWK D LP
Sbjct: 915 ANWVTECDVCQLVKAEHQVPGGLLQSLPISEWK----KTDGAAVLP-------------- 956
Query: 1157 RLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGT 1216
+ ++++IV+LHGVP SI+S RDS+FTS FW A Q +GT
Sbjct: 957 ---------------------KKYVSEIVKLHGVPVSILSHRDSKFTSAFWRAFQVEMGT 995
Query: 1217 KLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMA 1276
K+++S+AY+PQTDGQ+ERTIQ+LED+L+ CVLD G W + L L++F YNNS+ ASIGMA
Sbjct: 996 KVQMSTAYHPQTDGQSERTIQTLEDMLQMCVLDWGGHWADHLSLVKFAYNNSYQASIGMA 1055
Query: 1277 PYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRK 1336
P+EALYGR C+T LCW Q GE + G + VQ+TTE ++ ++ M+ + RQ+SYAD RR+
Sbjct: 1056 PFEALYGRPCRTLLCWTQVGEKSIYGADYVQETTERIRVLKLNMKEAQDRQRSYADKRRR 1115
Query: 1337 ELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIH 1396
ELEF+ G +I ERVGPVAYR+ LP + H
Sbjct: 1116 ELEFEVGT-----------------------------EIVERVGPVAYRLELPDVMRAFH 1146
Query: 1397 DVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWN 1456
+V HVS LRK + D VL L+ ++T+ P+++++R K +R K + ++KV+W+
Sbjct: 1147 NVFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKEVRRKKIPMIKVLWD 1206
Query: 1457 -QATGDATWELEDKMR 1471
+ TWE E +++
Sbjct: 1207 CDGVTEETWEPEARIK 1222
>At3g11970 hypothetical protein
Length = 1499
Score = 451 bits (1159), Expect = e-126
Identities = 265/753 (35%), Positives = 419/753 (55%), Gaps = 35/753 (4%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + +L+A SKC F + +V++LGH IS+QG+ DP+KI+++ W QP +R F+
Sbjct: 776 EVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLKQLRGFL 835
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GLAGYYRRFV+ + + PL LTK + F WT +++F+++K L A +L+LP +
Sbjct: 836 GLAGYYRRFVRSFGVIAGPLHALTKTDA-FEWTAVAQQAFEDLKAALCQAPVLSLPLFDK 894
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
+ V DA G+G V+MQE +AY SRQLK + + ++ EL A+++A++ WRHYL
Sbjct: 895 QFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLL 954
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
S F I +D +SLKYL +Q+ Q++W+ L ++++++Q GK NVVADALSR
Sbjct: 955 QSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYRQGKENVVADALSR---- 1010
Query: 967 VSSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLE-WRK 1025
V+ E+L M V LL +I++ D L +
Sbjct: 1011 -----VEGSEVLH-----------------MAMTVVECDLLKDIQAGYANDSQLQDIITA 1048
Query: 1026 LVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKL 1085
L + ++ S NILR K ++ VP + ++ IL H S + H G + +Q +K
Sbjct: 1049 LQRDPDSKKYFSWSQNILRRKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKG 1108
Query: 1086 HFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTR 1145
F+W GM K + Y+ +C TC K + G LQ L +P+ W +SMDF+ LP++
Sbjct: 1109 LFYWKGMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSG 1168
Query: 1146 RRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSR 1205
+ I VVVDRL+K AHF+ +S Y +A ++ + +LHG P+SIVSDRD FTS
Sbjct: 1169 GK-TVIMVVVDRLSKAAHFIALSHPYSALTVAHAYLDNVFKLHGCPTSIVSDRDVVFTSE 1227
Query: 1206 FWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTY 1265
FW G L+L+SAY+PQ+DGQTE + LE LR D W + L L E+ Y
Sbjct: 1228 FWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWY 1287
Query: 1266 NNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*S 1325
N ++H+S M P+E +YG+ L + + + +Q+ + + ++ + +
Sbjct: 1288 NTNYHSSSRMTPFEIVYGQVPPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQH 1347
Query: 1326 RQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK-SKKLTPKFIGPYQITERVGPVAY 1384
R K +AD R E EF+ GD+V++++ P ++ ++KL+PK+ GPY+I +R G VAY
Sbjct: 1348 RMKQFADQHRTEREFEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAY 1407
Query: 1385 RIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLR 1444
++ALP + S +H V HVSQL+ + + S + + ++D V P K+V+R +
Sbjct: 1408 KLALPSY-SQVHPVFHVSQLKVLVGNVSTTVHLPSV-MQDVFEKV--PEKVVERKMVNRQ 1463
Query: 1445 NK*VSLVKVVW-NQATGDATWELEDKMRESHPD 1476
K V+ V V W N+ +ATWE ++++ P+
Sbjct: 1464 GKAVTKVLVKWSNEPLEEATWEFLFDLQKTFPE 1496
>At1g36590 hypothetical protein
Length = 1499
Score = 446 bits (1147), Expect = e-125
Identities = 264/753 (35%), Positives = 419/753 (55%), Gaps = 35/753 (4%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + +L+A SKC F + +V++LGH IS+QG+ DP+KI+++ W QP +R F+
Sbjct: 776 EVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLKQLRGFL 835
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GLAGYYRRFV+ + + PL LTK + F WT +++F+++K L A +L+LP +
Sbjct: 836 GLAGYYRRFVRSFGVIAGPLHALTKTDA-FEWTAVAQQAFEDLKAALCQAPVLSLPLFDK 894
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
+ V DA G+G V+MQE +AY SRQLK + + ++ EL A+++A++ WRHYL
Sbjct: 895 QFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLL 954
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
S F I +D +SLKYL +Q+ Q++W+ L ++++++Q GK NVVADALSR
Sbjct: 955 QSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYRQGKENVVADALSR---- 1010
Query: 967 VSSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLE-WRK 1025
V+ E+L M V LL +I++ D L +
Sbjct: 1011 -----VEGSEVLH-----------------MAMTVVECDLLKDIQAGYANDSQLQDIITA 1048
Query: 1026 LVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKL 1085
L + ++ S NILR K ++ VP + ++ IL H S + H G + +Q +K
Sbjct: 1049 LQRDPDSKKYFSWSQNILRRKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKG 1108
Query: 1086 HFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTR 1145
F+ GM K + Y+ +C TC K + G LQ L +P+ W +SMDF+ LP++
Sbjct: 1109 LFYSKGMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSG 1168
Query: 1146 RRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSR 1205
+ I VVVDRL+K AHF+ +S Y +A+ ++ + +LHG P+SIVSDRD FTS
Sbjct: 1169 GK-TVIMVVVDRLSKAAHFIALSHPYSALTVAQAYLDNVFKLHGCPTSIVSDRDVVFTSE 1227
Query: 1206 FWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTY 1265
FW G L+L+SAY+PQ+DGQTE + LE LR D W + L L E+ Y
Sbjct: 1228 FWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWY 1287
Query: 1266 NNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*S 1325
N ++H+S M P+E +YG+ L + + + +Q+ + + ++ + +
Sbjct: 1288 NTNYHSSSRMTPFEIVYGQVPPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQH 1347
Query: 1326 RQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK-SKKLTPKFIGPYQITERVGPVAY 1384
R K +AD R E EF+ GD+V++++ P ++ ++KL+PK+ GPY+I +R G VAY
Sbjct: 1348 RMKQFADQHRTEREFEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAY 1407
Query: 1385 RIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLR 1444
++ALP + S +H V HVSQL+ + + S + + ++D V P K+V+R +
Sbjct: 1408 KLALPSY-SQVHPVFHVSQLKVLVGNVSTTVHLPSV-MQDVFEKV--PEKVVERKMVNRQ 1463
Query: 1445 NK*VSLVKVVW-NQATGDATWELEDKMRESHPD 1476
K V+ V V W N+ +ATWE ++++ P+
Sbjct: 1464 GKAVTKVLVKWSNEPLEEATWEFLFDLQKTFPE 1496
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 398 bits (1023), Expect = e-110
Identities = 215/500 (43%), Positives = 300/500 (60%), Gaps = 62/500 (12%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + ++L+A SKC FW E+ FLGH++S++GV+VDP KIE+I W +P A++IRSF+
Sbjct: 666 EKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFL 725
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
G AGYYRRFVK +A + P+T+LT K+ PF W+++CEE F +K+ LT+ +LALP+ G+
Sbjct: 726 GWAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALPEHGQ 785
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
PY VY DAS GLGCV+MQ KVIAYASRQL HE NYPTHDLE+AA+++ALKIWR YLY
Sbjct: 786 PYMVYTDASRVGLGCVLMQHGKVIAYASRQLMKHEGNYPTHDLEMAAVIFALKIWRSYLY 845
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
G +++DHKSLKY+F Q +LN+RQRRWME + DY+ ++ HPGK NVV DALSRK
Sbjct: 846 GGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIAYHPGKANVVVDALSRK--R 903
Query: 967 VSSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVS-SGLLDEIKSKQETDEGLLEWRK 1025
V + + + +E+L + ++ A + G+ V + LL ++ Q+ DEGL
Sbjct: 904 VGAALGQSVEVLVS--EIGALRLCAVAREPLGLEAVDRADLLTRVRLAQKKDEGL----- 956
Query: 1026 LVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKL 1085
++ + + YQ
Sbjct: 957 ------------------------------------------RATKMYRDLKRYYQ---- 970
Query: 1086 HFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTR 1145
W GMK VA +V+ C C K EHQ G LQSL +PEWKWD I+MD V L ++R
Sbjct: 971 ---WVGMKMDVANWVAECDVCQLVKAEHQVLGGMLQSLPIPEWKWDFITMDLVVGLRVSR 1027
Query: 1146 RRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSI--VSDRDSRFT 1203
+ DAIWV+VDRLTK+AHF+ I LA+ +++IV+LHGVP ++ DR +
Sbjct: 1028 TK-DAIWVIVDRLTKSAHFLAIRKTDGAAVLAKKFVSEIVKLHGVPLNMKEAQDRQRSYA 1086
Query: 1204 SRFWGALQQALGTKLRLSSA 1223
+ L+ +G ++ L A
Sbjct: 1087 DKRRRELEFEVGDRVYLKMA 1106
Score = 131 bits (329), Expect = 3e-30
Identities = 63/153 (41%), Positives = 98/153 (63%), Gaps = 1/153 (0%)
Query: 1320 MRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERV 1379
M+ + RQ+SYAD RR+ELEF+ GD V+L++ + G R+I KL+P+++GP++I ERV
Sbjct: 1075 MKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSISETKLSPRYMGPFKIVERV 1134
Query: 1380 GPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRS 1439
PVAYR+ LP + H V HVS LRK + D L L+ ++T+ P+++++R
Sbjct: 1135 EPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEALAKIPEDLQPNMTLEARPVRVLERR 1194
Query: 1440 TKRLRNK*VSLVKVVWN-QATGDATWELEDKMR 1471
K LR K + L+KV+W+ + TWE E +M+
Sbjct: 1195 IKELRQKKIPLIKVLWDCDGVTEETWEPEARMK 1227
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 393 bits (1010), Expect = e-109
Identities = 193/379 (50%), Positives = 267/379 (69%), Gaps = 2/379 (0%)
Query: 1094 KQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWV 1153
+ VA +V+ C C K EHQ P G LQSL +PEWKWD I++DFV LP++R + DAIWV
Sbjct: 941 EDVANWVAECDVCQLVKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLPVSRTK-DAIWV 999
Query: 1154 VVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQA 1213
+VDRLTK+AHF+ I LA+ ++++IV+LHGVP SIVSDRDS+FTS FW A Q
Sbjct: 1000 IVDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAE 1059
Query: 1214 LGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASI 1273
+GTK+++S+AY+PQT GQ+ERTIQ+LED+LR CVLD G W + L L+EF YNNS+ ASI
Sbjct: 1060 MGTKVQMSTAYHPQTYGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPASI 1119
Query: 1274 GMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADN 1333
GMAP+EALY R C+TPLC Q GE + G + VQ+TTE ++ ++ M+ + RQ+SYAD
Sbjct: 1120 GMAPFEALYERPCRTPLCLTQVGERSIYGADYVQETTERIRVLKLNMKEAQDRQRSYADK 1179
Query: 1334 RRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLS 1393
RR+ELEF+ GD V+L++ + G R+I KL+P+++GP++I ERVGPVAYR+ LP +
Sbjct: 1180 RRRELEFEVGDRVYLKMAMLRGPNRSISETKLSPRYMGPFRIVERVGPVAYRLELPDVMR 1239
Query: 1394 HIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKV 1453
H V HVS LRK + D VL L+ ++T+ P+++++R K LR K + L+KV
Sbjct: 1240 AFHKVFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKV 1299
Query: 1454 VWN-QATGDATWELEDKMR 1471
+W+ TWE E +M+
Sbjct: 1300 LWDCDGVTKETWEPEARMK 1318
Score = 316 bits (809), Expect = 7e-86
Identities = 155/296 (52%), Positives = 210/296 (70%), Gaps = 5/296 (1%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + ++L+A SKC FW E+ FLGH++S++GV+VDP KIE+I W P A++IRSF+
Sbjct: 641 EKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTPTNATEIRSFL 700
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GLAGYYRRFVK +A + P+T+LT K+ PF W+ +CEE F +K+ LT+ +LALP+ GE
Sbjct: 701 GLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGE 760
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
PY VY DAS GLGCV+MQ KVIAYASRQL+ HE NYPTHDLE+AA+++ALKIWR YLY
Sbjct: 761 PYMVYTDASGVGLGCVLMQRGKVIAYASRQLRKHEGNYPTHDLEMAAVIFALKIWRSYLY 820
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
G +++DHKSLKY+F Q +LN+RQR+WME + DY+ ++ HPGK NVVADALS K
Sbjct: 821 GGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLEIAYHPGKANVVADALSHK--R 878
Query: 967 VSSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVS-SGLLDEIKSKQETDEGLL 1021
V + + +E L + ++ A + G+ V + LL ++ QE DEGL+
Sbjct: 879 VGAAPGQSVEALVS--EIGALRLCAVAREPLGLEAVDRADLLTRVRLAQEKDEGLI 932
>At1g35370 hypothetical protein
Length = 1447
Score = 372 bits (956), Expect = e-103
Identities = 222/651 (34%), Positives = 355/651 (54%), Gaps = 36/651 (5%)
Query: 733 ELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYY 792
+L+A GSK + LGH IS++ + DP+KI+++ W P +R F+G AGYY
Sbjct: 761 KLFAKGSK--------EHLGHFISAREIETDPAKIQAVKEWPTPTTVKQVRGFLGFAGYY 812
Query: 793 RRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYC 852
RRFV+++ + PL LTK + F W+ + + +F +K L A +LALP + + V
Sbjct: 813 RRFVRNFGVIAGPLHALTKTDG-FCWSLEAQSAFDTLKAVLCNAPVLALPVFDKQFMVET 871
Query: 853 DASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTI 912
DA G+ V+MQ+ +AY SRQLK + + ++ EL A ++A++ WRHYL S F I
Sbjct: 872 DACGQGIRAVLMQKGHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLLPSHFII 931
Query: 913 YSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHVSSLMV 972
+D +SLKYL +Q+ Q++W+ L ++++++Q GK N+VADALSR + S ++
Sbjct: 932 KTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQYRQGKENLVADALSR--VEGSEVLH 989
Query: 973 KELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKA 1032
L ++E D ++++A S G+L +I S L A
Sbjct: 990 MALSIVEC--DFLKEIQVA---------YESDGVLKDIISA------------LQQHPDA 1026
Query: 1033 PEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGM 1092
+ S +ILR K ++ VPND + +L H S + G + +Q +K F+W GM
Sbjct: 1027 KKHYSWSQDILRRKSKIVVPNDVEITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGM 1086
Query: 1093 KKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIW 1152
K + ++ +C TC K ++ G LQ L +P+ W +SMDF+ LP + + I
Sbjct: 1087 VKDIQAFIRSCGTCQQCKSDNAAYPGLLQPLPIPDKIWCDVSMDFIEGLPNSGGK-SVIM 1145
Query: 1153 VVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQ 1212
VVVDRL+K AHFV ++ Y +A+ + + + HG P+SIVSDRD FTS FW +
Sbjct: 1146 VVVDRLSKAAHFVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFK 1205
Query: 1213 ALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHAS 1272
G +LR+SSAY+PQ+DGQTE + LE+ LR W++ LPL E+ YN ++H+S
Sbjct: 1206 LQGVELRMSSAYHPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSS 1265
Query: 1273 IGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYAD 1332
M P+E +YG+ L + + + +Q+ + ++ + + R K +AD
Sbjct: 1266 SQMTPFELVYGQAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLMRAQHRMKQFAD 1325
Query: 1333 NRRKELEFQAGDHVFLRVTPMTGVGRAIK-SKKLTPKFIGPYQITERVGPV 1382
R E F GD V++++ P ++ ++KL+PK+ GPY+I E+ G V
Sbjct: 1326 QHRTERTFDIGDFVYVKLQPYRQQSVVLRVNQKLSPKYFGPYKIIEKCGEV 1376
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 329 bits (844), Expect = 6e-90
Identities = 167/321 (52%), Positives = 215/321 (66%), Gaps = 7/321 (2%)
Query: 818 WTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQL 877
W CE+SF E+K LT A +L LP+EGEPY VY DAS GLGCV+MQ+ VIAYASRQL
Sbjct: 375 WQRSCEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIAYASRQL 434
Query: 878 KIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWME 937
HEKNYPTHDLE+AA+V+ALKIWR YLYG+ IY+DHKSLKY+F Q +LN+RQRRWME
Sbjct: 435 WKHEKNYPTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFIQPELNLRQRRWME 494
Query: 938 FLQDYEFDLQ*HPGKTNVVADALSRKAMHVSSLMVKELELLEAF*DLSLDVKIAPGKLSF 997
+ DY+ D+ HPGK N VADALSR+ V + +++L+ L ++ ++ S
Sbjct: 495 LVADYDLDIAYHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLHVNA-LSKEVESL 552
Query: 998 GM-VTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDAT 1056
G+ + LL I+ QE DE + W E+ ++ + GRVCVPND
Sbjct: 553 GLGAADQADLLSRIRLAQERDEEIKGW----ALNNKTEYQTSNNGTIVVNGRVCVPNDRA 608
Query: 1057 MRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKP 1116
++ IL E H+S+ SIHPG NKMY+DLK ++ W GMKK VA +V+ C TC K EHQ P
Sbjct: 609 LKEEILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAEHQVP 668
Query: 1117 AGKLQSLDVPEWKWDSISMDF 1137
+G LQ+L +PEWKWD I+MDF
Sbjct: 669 SGLLQNLPIPEWKWDHITMDF 689
Score = 225 bits (573), Expect = 2e-58
Identities = 119/256 (46%), Positives = 166/256 (64%), Gaps = 27/256 (10%)
Query: 1188 HGVPSSIVSD------RDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLED 1241
H VPS ++ + + T F A Q+ALGT++ LS+AY+PQTDGQ+ERTIQ+LED
Sbjct: 665 HQVPSGLLQNLPIPEWKWDHITMDF--AFQKALGTRVNLSTAYHPQTDGQSERTIQTLED 722
Query: 1242 LLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVI 1301
+LRACVLD G+W++ L L ALYGR C+TPLCW GE +
Sbjct: 723 MLRACVLDWGGNWEKYLTL-------------------ALYGRACRTPLCWTPVGERRLF 763
Query: 1302 GPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK 1361
GP +V +TTE +K ++ K++ + RQKSYA+ RRKELEFQ GD V+L+ G GR
Sbjct: 764 GPTIVDETTERMKFLKIKLKEAHDRQKSYANKRRKELEFQVGDLVYLKAMTYKGAGRFTS 823
Query: 1362 SKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDIQ 1421
KKL+P+++GPY++ ERVG VAY++ LPP L+ H+V HVSQLRK ++D +E
Sbjct: 824 RKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDVPPG 883
Query: 1422 LKDDLTVVMPPIKIVD 1437
LK+++TV P++I+D
Sbjct: 884 LKENMTVEAWPVRIMD 899
>At4g03840 putative transposon protein
Length = 973
Score = 322 bits (825), Expect = 1e-87
Identities = 163/322 (50%), Positives = 212/322 (65%), Gaps = 9/322 (2%)
Query: 818 WTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQL 877
W CE++F E+K LT A +L LP+EGEPY VY DAS GL CV+MQ+ VIAYASRQL
Sbjct: 375 WQRSCEKNFLELKAMLTNAPVLVLPEEGEPYTVYTDASVVGLECVLMQKGSVIAYASRQL 434
Query: 878 KIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWME 937
+ HEKNYPTHDLE+AA+V+ALKIWR YLYG+ IY+DHKSLKY+F Q +LN+RQRRWME
Sbjct: 435 RKHEKNYPTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLRQRRWME 494
Query: 938 FLQDYEFDLQ*HPGKTNVVADALSRKAMHVSS--LMVKELELLEAF*DLSLDVKIAPGKL 995
+ DY+ D+ H GK N VADALSR+ V + V + ++ +L ++ P L
Sbjct: 495 LVADYDLDIAYHAGKANQVADALSRRRSEVEAERSQVDLVNMMGTLHVNALSKEVEP--L 552
Query: 996 SFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDA 1055
G + LL I+ QE DE + W E+ ++ + GRVCVPN+
Sbjct: 553 GLGAAD-QANLLSRIRLAQERDEEIKGW----ALNNKTEYQTSNNGTIVVNGRVCVPNNR 607
Query: 1056 TMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQK 1115
++ IL E H+S+ SIHPG NK+Y+DLK ++ W GMKK VA +V+ C TC K EHQ
Sbjct: 608 ALKEEILREAHQSKFSIHPGSNKIYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAEHQV 667
Query: 1116 PAGKLQSLDVPEWKWDSISMDF 1137
P+G LQ+L +PEWKWD I+MDF
Sbjct: 668 PSGLLQNLPIPEWKWDHITMDF 689
Score = 249 bits (637), Expect = 6e-66
Identities = 134/299 (44%), Positives = 188/299 (62%), Gaps = 27/299 (9%)
Query: 1188 HGVPSSIVSDR-------DSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLE 1240
H VPS ++ + D FW A Q+ALGT++ LS+AY+PQTDGQ+ERTIQ+LE
Sbjct: 665 HQVPSGLLQNLPIPEWKWDHITMDFFWKAFQKALGTRVNLSTAYHPQTDGQSERTIQTLE 724
Query: 1241 DLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLV 1300
D+LRAC LD G+W++ L L ALYGR C+TPLCW GE +
Sbjct: 725 DMLRACALDWGGNWEKYLRL-------------------ALYGRACRTPLCWTPVGERRL 765
Query: 1301 IGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAI 1360
GP +V +TTE +K ++ K++ + RQKSYA+ RRKELEFQ D V+L+ G GR
Sbjct: 766 FGPIIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVEDLVYLKAMTYKGAGRFT 825
Query: 1361 KSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDI 1420
KKL+P+++GPY++ ERVG VAY++ LPP L+ H+V HVSQLRK +++ +E
Sbjct: 826 SRKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSNQEESVEDVPP 885
Query: 1421 QLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWNQATGDA-TWELEDKMRESHPDLF 1478
LK+++TV P++I+DR TK R K L+KV+WN + TWE E+KM+ + + F
Sbjct: 886 GLKENMTVEAWPVQIMDRMTKGTRGKSRDLLKVLWNCGGREQYTWETENKMKANFSEWF 944
>At3g29490 hypothetical protein
Length = 438
Score = 290 bits (741), Expect = 6e-78
Identities = 154/381 (40%), Positives = 222/381 (57%), Gaps = 54/381 (14%)
Query: 1092 MKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAI 1151
MK+ VA +V+ C C K EHQ P G LQSL +PEW
Sbjct: 1 MKRDVANWVAECDVCQLVKAEHQVPGGMLQSLPIPEWNT--------------------- 39
Query: 1152 WVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQ 1211
F+ I LA+ ++++IV+LHGVP+
Sbjct: 40 ------------FLAIRKTDGAAVLAKKYVSEIVKLHGVPAE------------------ 69
Query: 1212 QALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHA 1271
+GTK+++S+ Y+PQTDGQ ERTIQ+LED+LR CVLD G W + L L+EF YNNS+ A
Sbjct: 70 --MGTKVQMSTPYHPQTDGQFERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYQA 127
Query: 1272 SIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYA 1331
IGMAP+EALYGR C+TPLCW Q GE + G + VQ+TTE ++ ++ M+ + RQ SYA
Sbjct: 128 GIGMAPFEALYGRPCRTPLCWTQVGERSIYGADYVQETTERIRVLKLNMKEAQDRQWSYA 187
Query: 1332 DNRRKELEFQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPF 1391
D RR+ELEF+ GD V+L++ + G R+I KL+ +++GP++I ERVGPVAY + LP
Sbjct: 188 DKRRRELEFEVGDRVYLKMAMLRGPNRSISETKLSLRYMGPFRIVERVGPVAYMLELPDV 247
Query: 1392 LSHIHDVLHVSQLRKYMADDSHVLEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLV 1451
+ H V HVS LRK + D VL L+ ++T+ +++++R K L+ K +SL+
Sbjct: 248 MRAFHKVFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARQVRVLERRIKELQRKKISLI 307
Query: 1452 KVVWN-QATGDATWELEDKMR 1471
KV+W+ + TW+ E +M+
Sbjct: 308 KVLWDCDGVTEETWQPEARMK 328
>At4g03650 putative reverse transcriptase
Length = 839
Score = 288 bits (737), Expect = 2e-77
Identities = 128/223 (57%), Positives = 172/223 (76%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + ++L+A SKC FW E+ FLGH++S++GV+VDP KIE+I W +P A++IRSF+
Sbjct: 591 EKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFL 650
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GLAGYYRRF+K +A + P+T+LT K+ PF W+ +CEE F +K+ LT+ +LALP+ GE
Sbjct: 651 GLAGYYRRFIKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGE 710
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
PY VY DAS GLGC +MQ KVIAYASRQL+ HE NYPTHDLE+AA+++ALKIWR YLY
Sbjct: 711 PYMVYTDASGVGLGCALMQRGKVIAYASRQLRKHEGNYPTHDLEMAAVIFALKIWRSYLY 770
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*H 949
G +++DHKSLKY+F Q +LN+RQRRWME + DY+ ++ H
Sbjct: 771 GGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIAYH 813
Score = 38.9 bits (89), Expect = 0.022
Identities = 15/23 (65%), Positives = 22/23 (95%)
Query: 1214 LGTKLRLSSAYNPQTDGQTERTI 1236
+GTK+++S+ Y+PQTDGQ+ERTI
Sbjct: 817 MGTKVQMSTTYHPQTDGQSERTI 839
>At2g06890 putative retroelement integrase
Length = 1215
Score = 287 bits (734), Expect = 4e-77
Identities = 168/437 (38%), Positives = 246/437 (55%), Gaps = 35/437 (8%)
Query: 732 KELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGY 791
+ELYAN KC F + + FLG V+S+ GV VD K+++I W PK ++RSF GLAG+
Sbjct: 599 EELYANLKKCTFCTDNLVFLGFVVSADGVKVDEEKVKAIRDWPSPKTVGEVRSFHGLAGF 658
Query: 792 YRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVY 851
YRRF KD++ + +PLT++ KK+ F W + EE+FQ +K +LT A +L L + + +++
Sbjct: 659 YRRFFKDFSTIVAPLTEVMKKDVGFKWEKAQEEAFQSLKDKLTNAPVLILSEFLKTFEIE 718
Query: 852 CDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFT 911
CDAS G+G V+MQ++K+IA+ S +L NYPT+D EL A+V AL+ W+HYL+ +F
Sbjct: 719 CDASGIGIGAVLMQDQKLIAFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLWPKVFV 778
Query: 912 IYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMHVSSLM 971
I++DH+SLK+L Q+ LN R RW+EF++ + + ++ GK NVVADALS++ +S+L
Sbjct: 779 IHTDHESLKHLKGQQKLNKRHARWVEFIETFAYVIKYKKGKDNVVADALSQRYTLLSTLN 838
Query: 972 VKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETDEGLLEWRKLVTQGK 1031
VK + ++IK ETD E K +
Sbjct: 839 VKLMGF------------------------------EQIKEVYETDHDFQEVYKACEKFA 868
Query: 1032 APEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPG 1091
+ + D L + R+CVPN ++R L + E H L H G+ K + + HF WP
Sbjct: 869 SGRY-FRQDKFLFYENRLCVPN-CSLRDLFVREAHGGGLMGHFGIAKTLEVMTEHFRWPH 926
Query: 1092 MKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAI 1151
MK V C TC AK + Q P G L +P+ W+ ISMDFV LP T + D+I
Sbjct: 927 MKCDVKRICGRCNTCKQAKSKIQ-PNGLYTPLPIPKHPWNDISMDFVMGLPRTGK--DSI 983
Query: 1152 WVVVDRLTKTAHFVPIS 1168
+VV F PIS
Sbjct: 984 FVVYSPFQIVYGFNPIS 1000
Score = 42.4 bits (98), Expect = 0.002
Identities = 39/144 (27%), Positives = 70/144 (48%), Gaps = 24/144 (16%)
Query: 1304 ELVQQTTEEVKR-IQEKMRIS*SRQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIKS 1362
ELVQQ E +R I+EK ++ A+ R+E F+ GD V++ + + +
Sbjct: 1022 ELVQQIHENARRNIEEKTKL----YAKQANKGRREQIFEVGDMVWIHLRKERFPAQ--RK 1075
Query: 1363 KKLTPKFIGPYQITERVGPVAYRIALP--------PF--------LSHIHDVLHVSQL-R 1405
KL P+ GP++I +R+ AY++ L PF + I+D+ H +L R
Sbjct: 1076 SKLMPRIDGPFKIIKRINDNAYQLDLQDEPDLRSNPFQEGGDDMIMDSINDMEHEPELER 1135
Query: 1406 KYMADDSHVLEPDDIQLKDDLTVV 1429
+ +A++ L +D + ++ VV
Sbjct: 1136 ELVAEEGAKLVAEDELVAEEKLVV 1159
>At3g29480 hypothetical protein
Length = 718
Score = 285 bits (730), Expect = 1e-76
Identities = 155/335 (46%), Positives = 207/335 (61%), Gaps = 37/335 (11%)
Query: 759 GVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAW 818
GV+VD KIE+I W +P A++IRSF+GLAGYYRRFVK +A + PLT+LT KN PF W
Sbjct: 415 GVSVDLEKIEAIKDWPRPTNATEIRSFLGLAGYYRRFVKGFASMAPPLTKLTGKNVPFVW 474
Query: 819 TEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQLK 878
+ +CEE F +K+ LT+ +LALP+ GE Y VY DAS GLGCV+MQ KVIAYASRQL+
Sbjct: 475 SPECEEGFVSLKEILTSTPVLALPEHGESYMVYTDASRVGLGCVLMQRGKVIAYASRQLR 534
Query: 879 IHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEF 938
HE NYPTHDLE++A +++DHKSLKY+ Q +LN+RQRRWME
Sbjct: 535 KHEGNYPTHDLEMSA------------------VFTDHKSLKYIITQPELNLRQRRWMEL 576
Query: 939 LQDYEFDLQ*HPGKTNVVADALSRKAMHVS-----SLMVKELELLEAF*DLSLDVKIAPG 993
+ DY+ ++ H GK +VVADALSRK + V+ +V E+ L A
Sbjct: 577 VADYDLEIAYHHGKASVVADALSRKRVGVAPGQSVEALVSEIRALRL---------CAVA 627
Query: 994 KLSFGMVTVSS-GLLDEIKSKQETDEGLLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVP 1052
+ G+ V LL ++ QE DEGL+ ++ + ++ ++ + GRVCV
Sbjct: 628 REPLGLEAVDRVDLLSRVRLAQEKDEGLI----AASKAEGSKYQFAANGTILVHGRVCVL 683
Query: 1053 NDATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHF 1087
ND +RR IL E H S SIHPG KMY+DLK ++
Sbjct: 684 NDEELRREILSEAHASMFSIHPGATKMYRDLKEYY 718
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 231 bits (588), Expect = 3e-60
Identities = 147/472 (31%), Positives = 232/472 (49%), Gaps = 84/472 (17%)
Query: 727 EYCKGKELYANGSKCEFWMEEVKFLGHVISSQGVAVDPSKIESIMSWEQPKIASDIRSFV 786
E + L+A SKC F + V++LGH IS +G+A DP+KI+++ W P + F+
Sbjct: 218 EVMRVHHLFAKMSKCAFAVPRVEYLGHFISGEGIATDPAKIKAVQDWPVPVNLKQLCGFL 277
Query: 787 GLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGE 846
GL GYYRRF
Sbjct: 278 GLTGYYRRF--------------------------------------------------- 286
Query: 847 PYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLY 906
+ V DA +G+G V+MQE +A+ SRQLK + + ++ EL A+++ ++ WRHYL
Sbjct: 287 -FVVEMDACGHGIGAVLMQEGHPLAFISRQLKGKQLHLSIYEKELLAVIFVVRKWRHYLI 345
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKTNVVADALSRKAMH 966
S F I +D +SLKYL +Q+ Q++W+ L ++++++Q GK N+VADALSR
Sbjct: 346 SSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYKQGKENLVADALSR---- 401
Query: 967 VSSLMVKELELLEAF*DLSLDVKIAPGKLSFGMVTVSSGLLDEIKSKQETD---EGLLEW 1023
V+ E+L + V LL EI++ TD +G++
Sbjct: 402 -----VEGSEVLH-----------------MALSVVECDLLKEIQAGYVTDGDIQGII-- 437
Query: 1024 RKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDL 1083
L Q + + S +LR K ++ VPN++ ++ IL H S + H G +Q +
Sbjct: 438 TILQQQADSKKHYTWSQGVLRRKNKIVVPNNSGIKDTILRWLHCSGMGGHSGKEVTHQRV 497
Query: 1084 KLHFWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPL 1143
K F+W M K + ++ +C TC K ++ G LQ L +P+ W +SMDF+ LPL
Sbjct: 498 KGLFYWKSMVKDIQAFIRSCGTCQQCKSDNAASPGLLQPLPIPDRIWSDVSMDFIDGLPL 557
Query: 1144 TRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIV 1195
+ + I VVVDRL+K AHF+ ++ Y +A+ ++ + +LHG PSSIV
Sbjct: 558 SNGK-TVIMVVVDRLSKAAHFIALAHPYSAMTVAQAYLDNVFKLHGCPSSIV 608
Score = 79.7 bits (195), Expect = 1e-14
Identities = 49/156 (31%), Positives = 84/156 (53%), Gaps = 5/156 (3%)
Query: 1275 MAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*SRQKSYADNR 1334
M PYEA+YG+ L + + + +Q+ + ++ + + R K AD
Sbjct: 625 MTPYEAVYGQPPPLHLPYLPGESKVAVVARSMQERESMILFLKFHLMRAQHRMKQLADQH 684
Query: 1335 RKELEFQAGDHVFLRVTPMTGVGRAIKS-KKLTPKFIGPYQITERVGPVAYRIALPPFLS 1393
E EF+ GD+VF+++ P ++S +KL+PK+ GPY++ +R G VAY++ LP S
Sbjct: 685 ITEREFEVGDYVFVKLQPYRQQSVVMRSTQKLSPKYFGPYKVIDRCGEVAYKLQLPA-NS 743
Query: 1394 HIHDVLHVSQLRKY---MADDSHVLEPDDIQLKDDL 1426
+H V HVSQLR + +H+L + L+++L
Sbjct: 744 QVHPVFHVSQLRVLVGTVTTSTHLLRCYLMSLRENL 779
>At4g07850 putative polyprotein
Length = 1138
Score = 207 bits (526), Expect = 5e-53
Identities = 100/210 (47%), Positives = 146/210 (68%)
Query: 754 VISSQGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKN 813
V S+ GV VD K+++I W PK +RSF GLAG+YRRFV+D++ L +PLT++ KKN
Sbjct: 531 VASTDGVKVDEEKVKAIREWPSPKSVGKVRSFHGLAGFYRRFVRDFSTLAAPLTEVIKKN 590
Query: 814 QPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYA 873
F W + E++FQ +K++LT A +L+LP + +++ CDA G+G V+MQ+KK IAY
Sbjct: 591 VGFKWEQAPEDAFQALKEKLTHAPVLSLPDFLKTFEIECDAPGVGIGAVLMQDKKPIAYF 650
Query: 874 SRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQR 933
S +L NYPT+D EL A+V AL+ W+HYL+ F I++DH+SLK+L Q+ LN R
Sbjct: 651 SEKLGGATLNYPTYDKELYALVRALQTWQHYLWPKEFVIHTDHESLKHLKGQQKLNKRHA 710
Query: 934 RWMEFLQDYEFDLQ*HPGKTNVVADALSRK 963
RW+EF++ + + ++ GK NVVADALSR+
Sbjct: 711 RWVEFIETFPYVIKYKKGKDNVVADALSRR 740
Score = 204 bits (518), Expect = 4e-52
Identities = 114/297 (38%), Positives = 170/297 (56%), Gaps = 15/297 (5%)
Query: 1007 LDEIKSKQETDEGLLEWRKLV-TQGKAPEFSIGSDNIL------RCKG------RVCVPN 1053
L +K +Q+ ++ W + + T ++ G DN++ R +G R+C+PN
Sbjct: 696 LKHLKGQQKLNKRHARWVEFIETFPYVIKYKKGKDNVVADALSRRHEGFLFYDNRLCIPN 755
Query: 1054 DATMRRLILDEGHKSRLSIHPGMNKMYQDLKLHFWWPGMKKQVAEYVSTCLTC*NAKVEH 1113
+++R L + E H L H G++K + ++ HF WP MK+ V C TC AK +
Sbjct: 756 -SSLRELFIREAHGGGLMGHFGVSKTLKVMQDHFHWPHMKRDVERMCERCTTCKQAKAKS 814
Query: 1114 QKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKV 1173
Q P G L +P W+ ISMDFV LP TR D+I+VVVDR +K AHF+P
Sbjct: 815 Q-PHGLCTPLPIPLHPWNDISMDFVVGLPRTRTGKDSIFVVVDRFSKMAHFIPCHKTDDA 873
Query: 1174 EKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTE 1233
+A + ++VRLHG+P +IVSDRD++F S FW L LGTKL S+ +PQTDGQTE
Sbjct: 874 MHIANLFFREVVRLHGMPKTIVSDRDTKFLSYFWKTLWSKLGTKLLFSTTCHPQTDGQTE 933
Query: 1234 RTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPL 1290
++L LLRA + + +W++ LP +EF YN+S H++ +P++ +YG TPL
Sbjct: 934 VVNRTLSTLLRALIKKNLKTWEDCLPHVEFAYNHSVHSATKFSPFQIVYGFNPITPL 990
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.342 0.148 0.511
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,725,100
Number of Sequences: 26719
Number of extensions: 1214929
Number of successful extensions: 3797
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 3582
Number of HSP's gapped (non-prelim): 135
length of query: 1489
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1377
effective length of database: 8,326,068
effective search space: 11464995636
effective search space used: 11464995636
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0388a.4


