

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.1
(728 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
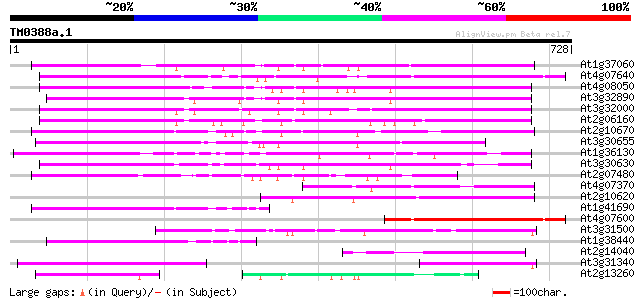
Score E
Sequences producing significant alignments: (bits) Value
At1g37060 Athila retroelment ORF 1, putative 289 4e-78
At4g07640 putative athila transposon protein 288 7e-78
At4g08050 244 1e-64
At3g32890 Athila ORF 1, putative 241 1e-63
At3g32000 unknown protein 239 5e-63
At2g06160 putative Athila retroelement ORF1 protein 233 3e-61
At2g10670 pseudogene 228 7e-60
At3g30655 hypothetical protein, 3' partial 207 2e-53
At1g36130 hypothetical protein 197 2e-50
At3g30630 hypothetical protein 177 2e-44
At2g07480 F9A16.15 175 9e-44
At4g07370 150 2e-36
At2g10620 putative Athila retroelement ORF1 protein 149 5e-36
At1g41690 hypothetical protein 148 1e-35
At4g07600 142 5e-34
At3g31500 hypothetical protein 141 1e-33
At1g38440 hypothetical protein 130 3e-30
At2g14040 putative retroelement pol polyprotein 127 2e-29
At3g31340 Athila ORF 1, putative 125 8e-29
At2g13260 putative Athila retroelement ORF1 protein 123 4e-28
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 289 bits (739), Expect = 4e-78
Identities = 217/734 (29%), Positives = 342/734 (46%), Gaps = 113/734 (15%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NFE++ LI +V N++ G +EDP H++ F R C+ ++ VS D +
Sbjct: 28 GIVPPPVQNNNFEIKSGLIAMVQSNKFHGLPMEDPLDHLDEFDRLCSLTKINRVSEDGFK 87
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A +W S PQGSITSW D + F +F + +L+NDI FTQ+ +E
Sbjct: 88 LRLFPFSLGDKAHQWEKSLPQGSITSWNDCKKAFLAKFFSNSRTARLRNDISGFTQTNNE 147
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
YEAWERFK +CP H + LD AS+G F + G+E
Sbjct: 148 TFYEAWERFKGYQTQCPHHEML-------------------LDTASNGNFLNKDVEDGWE 188
Query: 209 LIEKMA-------------IRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQN 255
++E +A IR + S+++ R + D+L+ ++ + + +
Sbjct: 189 VVENLAQSDGNYNEDYDRSIRTSSDSDEKHRREMKAMNDKLDKLLLMQQKHIHFLGDDET 248
Query: 256 QMKTTKIGSRVAKVEFVTCGG-------------PHDSEECTETRPEEEVKAMGQARNDP 302
++ +V +V G P+ S T ++ Q +N P
Sbjct: 249 LQVQDGETLQLEEVSYVQNQGGYNKGFNNFKQNHPNLSYRSTNVANRQDQVYPSQQQNQP 308
Query: 303 FSNT-YNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQP-------------Q 348
YN G P + QGN +Q+Q P GF Q +QP Q
Sbjct: 309 KPFVPYNQGQGYVPKQQY------QGN-YQQQLPPPGFT-QQQQQPALTTPDSDLKNMLQ 360
Query: 349 ERGEGEGSGSKKSLEELVETFINRTENNYKN----QEAANKNLENQFGQLAKQIAERPQG 404
+ +G+ +G+ + + E N+ + +Y + EA + GQ A + G
Sbjct: 361 QILQGQATGAMDLSKRMAEIH-NKVDCSYNDINIKVEALTSKIRYIEGQTGSTAAPKFTG 419
Query: 405 IFPSDCIPNPKQENASVVATRSGRVMSELKKKTE----------------GEKREEIVGG 448
PS + +E A + RSG+ + + + G E+ +
Sbjct: 420 --PSGKSMSNSKEYAHAITLRSGKELPTKESPNQNTEDSLDQDGEDFCQNGNSAEKAIEE 477
Query: 449 EVV------------------VPIKVREEVDLTPE-TSKIPFPKALAKKSLDKKFSKFVD 489
++ K +E V + P +PFP K + K +
Sbjct: 478 PILHQPTRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEK 537
Query: 490 VFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRK-MPKK 548
K L +++P D L +P K++KD++ R+K V V+++ ECSAI+Q+K +PKK
Sbjct: 538 QLKNLEVTMPLVDCLALIPDSNKYVKDMI--TERIKEVQGMVVLSHECSAIIQQKIIPKK 595
Query: 549 RRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLK 608
DPGSFT+P + +A + LCDLGAS++LMPL + ++L + P +SL +ADRS++
Sbjct: 596 LGDPGSFTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVR 655
Query: 609 TPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPV 668
+G++ED+ V + P DFVVL+M+E+ K PLILGRPFLA RA IDV KG + L +
Sbjct: 656 ISHGLLEDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNL 715
Query: 669 GKE-KVRFSVFNPM 681
G++ K+ F + N M
Sbjct: 716 GRDLKMTFDITNTM 729
>At4g07640 putative athila transposon protein
Length = 866
Score = 288 bits (737), Expect = 7e-78
Identities = 221/710 (31%), Positives = 350/710 (49%), Gaps = 68/710 (9%)
Query: 39 NFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRD 98
NFE++ LI+++ N++ G +EDP H++ F R CN ++ VS D +L LFPFSL D
Sbjct: 38 NFEIKSGLISMIQGNKFYGLPMEDPLDHLDEFDRLCNLTKINGVSADGFKLRLFPFSLGD 97
Query: 99 TAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFK 158
A W + P SI +W+D + F ++F A +L+N+I F+Q T E+ EAWERFK
Sbjct: 98 KAHIWEKNLPHDSIITWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFK 157
Query: 159 KLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAM 218
+CP H T+A ++ Y G+ R LD AS+G F + G+EL++ +A
Sbjct: 158 GYTNQCPHHGFTKASMLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVDNLAQSDG 217
Query: 219 NSSNDRQARRGVLEV-EAYDQLMASNKQLSKQMNEIQ-NQMKTTKIGSRVAKVEFVTCGG 276
N + D R V + ++ D+ K L+ +++ I NQ K + V +F G
Sbjct: 218 NYNED--CDRTVRDTADSDDKHRKEIKALNDKLDRILLNQQK--HVHFLVDDEQFQVQDG 273
Query: 277 PHDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWR--------------QG 322
E EEV + N YN N+PN S+R Q
Sbjct: 274 --------EGNQLEEVSYINN--NQSGYKGYNNFKTNNPNLSYRSTNIANPQDQVYPPQQ 323
Query: 323 SSGQGNGF----QRQFPSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFINRT-ENNY 377
Q F Q P Q FQG QP G+ +++ + + E ++ E+
Sbjct: 324 QQVQNKPFVPYNQGFVPKQQFQGNY--QPPPPPGGKALPTREEPKTVTEDSEDQDGEDLS 381
Query: 378 KNQEAANKNLENQFGQLAKQ----IAERPQGIFPSDCIPNPKQENASVVATRSGRVMSEL 433
++ A+K E Q +Q E+P + P D + P A+ + +
Sbjct: 382 LEKDQADKPHEQPLDQSLEQPLDLSLEQPLDL-PLDNVTRPTTRPIFPAASATAPKPITV 440
Query: 434 KKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDVFKK 493
K K + V VP P ++PFP K DK + F K+
Sbjct: 441 KNKEK-----------VFVP---------PPYKPELPFPGRHKKALADKYRAMFAKNIKE 480
Query: 494 LHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKM-PKKRRDP 552
+ + IP DAL +P KF+KD++ +R ++ V V+++ CSAI+Q+K+ PKK DP
Sbjct: 481 VELRIPLVDALALIPDSHKFLKDLIVER--IQEVQGMVVLSHGCSAIIQKKIIPKKLSDP 538
Query: 553 GSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYG 612
GSFT+P + +A LCDLGAS++LM L++ +RL + + +SL +ADRS++ P+G
Sbjct: 539 GSFTLPCSLGPLAFNRCLCDLGASVSLMALSVAKRLGFTQYKSSNISLILADRSVRIPHG 598
Query: 613 IVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGKE- 671
++++ + + P DFVVL+M+E+ K PLILGRPFLAT A IDV KG + L +GK+
Sbjct: 599 LLKNFGITIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLSLGKDF 658
Query: 672 KVRFSVFNPMIETNHDNDFVCDVIRSRSKVSEETPKVNSQIDEFHPALIK 721
++ F V + M + + I+ ++++E + ++ D + AL K
Sbjct: 659 RMTFDVKDAMKKPTIEGQLFW--IKEMDQLADELLEELAEEDHLNSALTK 706
>At4g08050
Length = 1428
Score = 244 bits (623), Expect = 1e-64
Identities = 201/686 (29%), Positives = 329/686 (47%), Gaps = 70/686 (10%)
Query: 39 NFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRD 98
NFE++ LI+++ N++ G +EDP H++ F R CN ++ VS D +L LFPFSL D
Sbjct: 38 NFEIKSGLISMIQGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGD 97
Query: 99 TAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFK 158
A W + SIT+W+D + F ++F A +L+N+I F+Q T E+ EAWERFK
Sbjct: 98 KAHIWEKNLSHDSITTWDDYKKAFLSKFFSNARTARLRNEIYGFSQKTGESFCEAWERFK 157
Query: 159 KLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAM 218
+CP H+ T+A ++ Y G+ R LD AS+G F + G+EL+E +A
Sbjct: 158 GYTNQCPHHSFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSDG 217
Query: 219 NSSND-RQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGP 277
N + D + RG + + D+ K L+ +++ I + S+ V F+
Sbjct: 218 NYNEDCDRTVRGTADSD--DKHRKEIKALNDKLDRI--------LLSQQKHVHFLVDDEQ 267
Query: 278 HDSEECTETRPEE-EVKAMGQARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFPS 336
+ ++ + EE + Q +N PF YN G+ F QGN + P
Sbjct: 268 YQVQDGEGNQLEEVYLPQQKQGQNKPFV-LYNQGFVPKQQF--------QGN--YQPPPP 316
Query: 337 QGF---QGQSLRQPQERG--------EGEGSGSKKSLEELVETFINRTENNYKN----QE 381
GF Q Q P + +G+ S S + ++L E ++ + +Y + E
Sbjct: 317 PGFVTQQNQCHAAPDAKMKQMVKQLLQGQASSSMEIAKKLSELH-HKLDCSYNDLNAKVE 375
Query: 382 AANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVAT-------RSGRVMSELK 434
A N + GQ A + + + P I NPK+ + T + V+ +
Sbjct: 376 ALNTKVRYLEGQSASTFSPKVTRL-PGKSIQNPKEYATAHAITICHDRELPTRHVLDLIT 434
Query: 435 KKT---EGEKREEI----------VGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLD 481
+ EGE ++ G + E+ + K P L ++L
Sbjct: 435 RDNDVQEGEASTQVEASVVEFNHSAGSRHLTQSTSEEKAAIIERMVKRFKPTPLPLRALP 494
Query: 482 KKFSK-FVDVFK--------KLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVL 532
F K +++ +K ++ +P + L + K +++++ +R ++ +
Sbjct: 495 WTFRKAWMERYKSVAAKQLDEIEAVMPLMEVLNLILDPHKVVRNLILERIKMYHDSDDES 554
Query: 533 MTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGE 592
A +R + +K DPGSFT+P I A + LCDLGAS+NLMPL+M RL +
Sbjct: 555 DATPSRAADKRIVQEKLEDPGSFTLPCSIGEFAFSDCLCDLGASVNLMPLSMARRLEFIQ 614
Query: 593 VTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLAT 652
P L+L +ADRS + +G+++D+ V ++ P DFVVLDME + K PLILGRPFLA+
Sbjct: 615 YKPCDLTLILADRSSRKHFGMLKDLPVMINGVEVPTDFVVLDMEVEHKDPLILGRPFLAS 674
Query: 653 GRAKIDVDKGHLILPVGKE-KVRFSV 677
A IDV +G + L +GK K++F++
Sbjct: 675 VGAVIDVKEGKIGLNLGKHIKLQFNI 700
>At3g32890 Athila ORF 1, putative
Length = 755
Score = 241 bits (615), Expect = 1e-63
Identities = 193/670 (28%), Positives = 318/670 (46%), Gaps = 59/670 (8%)
Query: 49 LVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQP 108
++ N++ G +EDP H++ F R N ++ VS D +L LFPFSL D A W + P
Sbjct: 1 MIQGNKFHGLPMEDPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLP 60
Query: 109 QGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHN 168
SIT+W+D + F ++F A +L+N+I F+Q T E+ EAWERFK +CP H
Sbjct: 61 HDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTRESFCEAWERFKGYTNQCPHHG 120
Query: 169 LTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNSSNDRQARR 228
T+A ++ Y G+ R LD +S+G F + G+EL+E +A N + D
Sbjct: 121 FTKASLLSTLYRGVLLRIRMLLDTSSNGNFQNKDVEEGWELVENLAQSDGNYNEDCDRN- 179
Query: 229 GVLEVEAYDQ-----LMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFV--TCGGPHDSE 281
E++A + L++ K + +++ Q Q++ + G+++ +V ++ GG
Sbjct: 180 ---EIKALNDKLDMILLSQQKHVHFLVDDKQFQVQYGE-GNQLEEVSYIINNQGGYKGYN 235
Query: 282 ECTETRPEEEVKAMG--------QARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQ 333
P ++ Q +N PF YN G+ F QGN +
Sbjct: 236 NFKTNNPNLSYRSTNVANPPQQQQGQNKPFV-PYNQGFVPKQQF--------QGN--YQP 284
Query: 334 FPSQGFQGQSLRQP-----------QERGEGEGSGSKKSLEELVETFINRTENNYKNQ-- 380
P GF Q + P Q+ E + S S + ++L E ++ + +Y +
Sbjct: 285 PPPPGFTTQQNQGPAAPDAEMKQMVQQLLEVQASSSMEIAKKLSELH-HKLDCSYNDMNA 343
Query: 381 --EAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQ-ENASVVATRSGRVMSELKKKT 437
EA N + GQ A + G+ P I NPK+ A + R +
Sbjct: 344 KVEALNTKVRYLEGQSASTSTPKVTGL-PGKSIQNPKEYATAHAITICHDRELPTRHVPD 402
Query: 438 EGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSK-FVDVFK---- 492
++ GE E+ + K P L +L F K +++ +K
Sbjct: 403 FITGDSDVQDGEASTQSTSEEKAAIIERMVKRFKPTPLPSSALPWTFRKAWMEKYKSVAS 462
Query: 493 ----KLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKK 548
++ +P + L +P K +++++ +R ++ + A +R + +K
Sbjct: 463 KQLDEIEAVMPLIEVLNLIPDPHKDVRNLILERIKMYHDSDDESDATPSRASDKRIVQEK 522
Query: 549 RRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLK 608
DPGSFT+P I +A + LCDLGAS++LMPL++ RL + P L+L +ADRS +
Sbjct: 523 LEDPGSFTLPCSIGELAFSDCLCDLGASVSLMPLSVARRLEFIQYKPCDLTLILADRSFR 582
Query: 609 TPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPV 668
P+G+++D+ V ++ P DFVVLDME + K PLILGRP LA+ A IDV +G + L +
Sbjct: 583 KPFGMLKDLPVMINGVEVPTDFVVLDMEVEHKDPLILGRPLLASVGAVIDVREGKISLNL 642
Query: 669 GKE-KVRFSV 677
GK K++F +
Sbjct: 643 GKHIKLQFGI 652
>At3g32000 unknown protein
Length = 839
Score = 239 bits (609), Expect = 5e-63
Identities = 193/684 (28%), Positives = 324/684 (47%), Gaps = 62/684 (9%)
Query: 39 NFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRD 98
NFE++ LI+++ N++ G +EDP H++ F R CN ++ VS + +L LFPFSL D
Sbjct: 38 NFEIKSGLISMIQGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSENGFKLRLFPFSLGD 97
Query: 99 TAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFK 158
A W + P SIT+ +D + F ++F +L+N+I F+Q E+ EAWERFK
Sbjct: 98 KALIWEMNLPHDSITTRDDCKKAFLSKFFSNDRTARLRNEISGFSQKIGESFCEAWERFK 157
Query: 159 KLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMA---- 214
+CP H T+A ++ Y G+ R LD AS+G F + G+EL++ +A
Sbjct: 158 DYTNQCPHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVDNLAQSDG 217
Query: 215 ---------IRAMNSSNDRQARRGVLEVEAYDQ-----LMASNKQLSKQMNEIQNQMKTT 260
+R S+D+ + E++A + L++ K + +++ Q Q++
Sbjct: 218 NYNEDCDRTVRGTADSDDKHRK----EIKALNDKLDRILLSQQKHVDFLVDDEQYQVQDG 273
Query: 261 KIGSRVAKVEFV--TCGGPHDSEECTETRPEEEVKAMGQARNDPFSNTYNP---GWRNHP 315
+ G+++ +V ++ GG P ++ A +P Y P +N P
Sbjct: 274 E-GNQLEEVSYINNNQGGYKGYNNFKTNNPNLSYRSTNVA--NPQDQVYPPQQQQGQNKP 330
Query: 316 NFSWRQG--SSGQGNGFQRQFPSQGFQGQSLRQP-----------QERGEGEGSGSKKSL 362
+ QG Q G +Q P GF Q + P Q+ +G+ S S +
Sbjct: 331 FVPYNQGFVPKQQFQGNYQQPPPPGFAPQQNQGPATPDAEMKQMVQQLLQGQASSSMEIA 390
Query: 363 EELVETFINRTENNYKN----QEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNP---- 414
++L E ++ + +Y + EA N + GQ A A + G+ D
Sbjct: 391 KKLSELH-HKLDCSYNDLNAKVEALNTKVRYLEGQSASTSAPKVTGLPGKDSDVQEGEAF 449
Query: 415 KQENASVVATRSGRVMSELKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKA 474
Q SVV L + T EK + ++ + TP S+ P
Sbjct: 450 TQVEVSVVEFNHSAGSRHLIQSTSEEK--------AAIIERMVKRFKPTPLPSR-ALPWT 500
Query: 475 LAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMT 534
K +++ S ++ +P + L +P K +++++ +R ++ +
Sbjct: 501 FRKSWMERYKSVAAKQPDEIEAVMPLMEVLNLIPDPHKDVRNLILERIKMYHDSDDESDA 560
Query: 535 EECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVT 594
A +R + +K DPGSF++P I A + LCDLGA ++LMP ++ RL +
Sbjct: 561 TPSRAADKRIVQEKLEDPGSFSLPCSIGEFAFSDCLCDLGAFVSLMPCSVARRLEFIQYK 620
Query: 595 PTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGR 654
P L+L +ADRS + P+G+++D+ V ++ P DFVVLDME + K PLILGRPFLA+
Sbjct: 621 PCDLTLILADRSSRKPFGMLKDLPVMINGVEVPTDFVVLDMEVEHKDPLILGRPFLASVG 680
Query: 655 AKIDVDKGHLILPVGKE-KVRFSV 677
A IDV +G + L +GK K++F +
Sbjct: 681 AVIDVREGKIGLNLGKHIKLQFGI 704
>At2g06160 putative Athila retroelement ORF1 protein
Length = 750
Score = 233 bits (594), Expect = 3e-61
Identities = 198/716 (27%), Positives = 330/716 (45%), Gaps = 96/716 (13%)
Query: 39 NFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRD 98
NFE++ LI +V N++ G +ED H++ F R C+ ++ VS D + LFPFSL D
Sbjct: 38 NFEIKSGLIAMVQGNKFHGLPMEDSLDHLDEFERLCDLTKINGVSEDGFKFRLFPFSLGD 97
Query: 99 TAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFK 158
W + PQ SIT+W+D + F +F + +L+N+I FTQ +E+ EAWERFK
Sbjct: 98 KTHLWEKTLPQNSITTWDDCKKAFFAKFFSNSRTARLRNEISGFTQKQNESFCEAWERFK 157
Query: 159 KLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMA---- 214
+CP QA ++ Y G+ R LD AS+G F + G+EL+E +A
Sbjct: 158 GYQTKCPHPGFKQASLLSTLYRGVLPKLRMLLDTASNGNFLNKDVEKGWELVENLAQSDG 217
Query: 215 ---------IRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKI--- 262
I + S+D+ R E++A + + Q+ ++ ++ + +I
Sbjct: 218 NYNENYDKSISTSSESDDKHHR----EIKALNDKIDKLLQVQQKHVHFASEHEVFQIQEG 273
Query: 263 -GSRVAKVEFVT------------CGGPHDSEECTE-TRPEEEVKAMGQARNDP---FSN 305
+ A++ +V P+ S T P+++V Q N P S
Sbjct: 274 ENDQCAEISYVQNQDGYNKGYNNYRPNPNLSYRSTNVANPQDQVYPQQQQLNQPKPFVSY 333
Query: 306 TYNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLR--QPQERG---------EGE 354
+ G+ P F G+Q+Q P GF Q + PQE +G+
Sbjct: 334 NQSQGFVPKPQFQ---------GGYQQQQPPPGFTPQQYQASTPQELDIRNMLQKIMQGQ 384
Query: 355 GSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNP 414
G+G ++ ++L + +N + YK + + + + + P + I NP
Sbjct: 385 GAGVMEAAKKLAD--LNIKIDRYKLESLSTRVRYLDGITASPPVTNNPCQV-SGKAIQNP 441
Query: 415 KQ-ENASVVATRSGRVMSELKKKTEGEKREEIVGGEVVVPIKVR-EEVDLTPET----SK 468
K+ A + R R +S T + I GE V I+V E+D +T
Sbjct: 442 KEYATAHAIIIRHDRELSTQHVSTSNTEDSVIQEGEASVQIEVSVAEIDYAAQTFYQAQS 501
Query: 469 IPFPKALAKKSLDKKFS-----------KFVDVFKKLHISI------------PFADALE 505
KA + + K+F F +K ++ S+ P + L+
Sbjct: 502 DLEEKATIIERMVKRFKPAPSPSLALPWTFRKAWKSIYESLAEKQLDEMESVMPLVEVLK 561
Query: 506 QMPIYAKFMKDILHKRRRL---KGVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIE 562
+ K +++++ +R + +E V+M + A +R + +K DPGSFT+P I
Sbjct: 562 LISDPHKDVRNLILERLNIYQDSDDEEDVIMHQ---AAGERIIQEKLEDPGSFTLPCSIS 618
Query: 563 GMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVD 622
+ LCDLGAS++LMPL++ +L + ++L +ADR+ + P+ I+EDV V ++
Sbjct: 619 QLTFSNCLCDLGASVSLMPLSVARKLRFVQYKSCDITLILADRTSRRPFSILEDVPVMIN 678
Query: 623 KYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGK-EKVRFSV 677
P DFVVL+M+E K PLILGR FLA+ A IDV G + L +G+ +K++F +
Sbjct: 679 GVEVPTDFVVLEMDEKSKDPLILGRLFLASVGAVIDVRFGKIDLNLGRHDKLQFDI 734
>At2g10670 pseudogene
Length = 929
Score = 228 bits (582), Expect = 7e-60
Identities = 197/687 (28%), Positives = 310/687 (44%), Gaps = 76/687 (11%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G ++ P NF+++ LI +V N++ G +EDP H++ R C ++ VS D +
Sbjct: 60 GIVLRPVQNNNFKIKSGLIAMVQGNKFHGLPMEDPLDHLDELERLCGLTKINGVSEDGFK 119
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A W + P+ SIT+W+D F +F + +L+N+I FT +E
Sbjct: 120 LRLFPFSLGDKAHLWEKTLPKNSITTWDDCKMAFLAKFFSNSRTARLRNEISGFTLKQNE 179
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
+ EAWERFK +CP H +QA + Y G+ R LD AS+G F + G+E
Sbjct: 180 SFCEAWERFKGYQTKCPHHGFSQASLLNTLYRGVLPKIRMLLDTASNGNFLKKDIEEGWE 239
Query: 209 LIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNE-IQNQMKTTKIGSRVA 267
L+E +A ++ + N+ R D+ K L+ ++++ IQ Q K
Sbjct: 240 LVENLA-QSGGNYNEDYGRSIYTSSNTDDKHHREMKALNDKLDKIIQMQQK--------- 289
Query: 268 KVEFVTCGGPH-----DSEECTETR------PEEEVKAMGQARNDPFSNTYNPGWRNHPN 316
V F++ P ++++C E R P A PF YN P
Sbjct: 290 HVHFISEDEPFQVQEGENDQCAEIRYVHNQGPSLPSAAAAAEPTKPFV-PYNQSLGFVPK 348
Query: 317 FSWRQGSSGQGNGFQRQFPSQGF--QGQSLRQPQERG---------EGEGSGSKKSLEEL 365
++ G+Q+Q P GF Q PQ +G+ +G+ + ++L
Sbjct: 349 QQFQ-------GGYQQQQPPPGFTPHQQQAHAPQNSDIMTVLQQLIQGQATGAMEIAKKL 401
Query: 366 --VETFINR--TENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQ-ENAS 420
V ++R E N K E+ + + G LA P I NPKQ A
Sbjct: 402 SKVNNKVDRQGVELNSK-FESMSTRMRYVEGILASPSVNNNPCQLPGKAIQNPKQYSTAH 460
Query: 421 VVATRSGRVMSELKKKTEGEKREEIVGGE-----VVVPIKVREEVDLTPETSKIPFPKAL 475
+ R + T + I+ GE V+ EE LT + ++ P A
Sbjct: 461 AITITHDRELPTRYVPTSNTEDSVILEGEDFYQDDVLADNPIEEPILTSQPTRPQAPPA- 519
Query: 476 AKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTE 535
F+K + K I +P +F K ++ K R L +
Sbjct: 520 ------TPFAKKTEAAKTKDIDFVPPPYKPLLPFPGRFKKVLVAKYRALLEKHIKDIPLV 573
Query: 536 ECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTP 595
+C A++ E + + LCDLGAS++LMPL++ ++L P
Sbjct: 574 DCLALIHD----------------EHKQLTFNNCLCDLGASVSLMPLSVVKKLGFVHYKP 617
Query: 596 TMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRA 655
L+L +ADR+ KTP+G++EDV V ++ P DFVVL+M+ + K PLILGRPFLA+ A
Sbjct: 618 CDLTLILADRTSKTPFGLLEDVPVMINGVEVPTDFVVLEMDGESKDPLILGRPFLASVGA 677
Query: 656 KIDVDKGHLILPVGKE-KVRFSVFNPM 681
IDV +G + L +G++ K++F + + M
Sbjct: 678 VIDVKQGKINLNLGEDVKMKFDIRDAM 704
>At3g30655 hypothetical protein, 3' partial
Length = 660
Score = 207 bits (526), Expect = 2e-53
Identities = 179/644 (27%), Positives = 286/644 (43%), Gaps = 77/644 (11%)
Query: 34 PDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFP 93
P NFE++ LI +V N++ G +EDP H++ F R ++ VS D +L LFP
Sbjct: 33 PVQNNNFEIKSGLIAMVQGNKFHGLPMEDPLDHLDEFERLYGLTKINGVSEDGFKLRLFP 92
Query: 94 FSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEA 153
FSL D A W + PQ SIT+W+D + F +F + +L+N+I FTQ +E+ EA
Sbjct: 93 FSLGDKAHLWEKTLPQNSITAWDDCKKAFLAKFFSNSRTARLRNEISGFTQKQNESFGEA 152
Query: 154 WERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKM 213
WERFK +CP H QA ++ Y G+ R LD AS+G F + G+EL+E
Sbjct: 153 WERFKGYQTKCPHHGFKQASLLSTLYRGVLPKIRILLDTASNGNFLKKDVEEGWELVENF 212
Query: 214 AIR--AMNSSNDRQARRGVLEVEAYD-QLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVE 270
A N DR R + +D ++ A N +L K +Q Q K V
Sbjct: 213 AQSDGNYNEDYDRSIRTSSDSNDKHDREMKALNDKLEK---IVQMQQK---------HVH 260
Query: 271 FVTCGGPHDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWRQG-------- 322
F++ P +E + + E Q + N Y P +PN S+R
Sbjct: 261 FISEDEPFPVQEGEKDQWAEISYVQNQGGYNKGYNYYRP----NPNLSYRSTNVANPQDQ 316
Query: 323 ------------------SSGQG--------NGFQRQFPSQGFQGQSLRQP--------- 347
+ QG G+Q+Q P GF Q P
Sbjct: 317 VYAQQQQQQNQPKPFVPYNQNQGFIPKQQFQGGYQQQQPQPGFTPQQQHAPAPQNSDIMI 376
Query: 348 --QERGEGEGSGSKKSLEEL--VETFINRTENNYKNQ-EAANKNLENQFGQLAKQIAERP 402
Q+ +G+ +G+ + ++L V I+R N+ E+ + + G LA +
Sbjct: 377 MLQQLIQGQATGAMEIAKKLCEVNNKIDRQGVELNNKSESMSTRMRYVEGILASPSVNKN 436
Query: 403 QGIFPSDCIPNPKQ-ENASVVATRSGRVMSELKKKTEGEKREEIVGGE-----VVVPIKV 456
P + I NPK+ + R + T + I+ GE V+
Sbjct: 437 PCQLPGNAIQNPKEYATVHAITITHDRELPTRYVPTSSTEDSVILHGEDFYQDDVLANNT 496
Query: 457 REEVDLTPETSKIPFPKA--LAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFM 514
EE LT ++++ A +K K V V +PF +++ + AK+
Sbjct: 497 IEEPILTSQSTRPQASPATHFVEKPEAAKTKDTVFVPPPYKPPLPFPGRFKKV-LVAKYR 555
Query: 515 KDILHKRRRLKGVDETVLMTEECSAILQRK-MPKKRRDPGSFTIPVEIEGMADVEALCDL 573
+ + + + V L+ +E +AI ++K +P+K DPGSFT+P I + LCDL
Sbjct: 556 ALLEKQIKDMPLVAYLALIPDEHNAITRQKIVPEKLEDPGSFTLPCSIRQLTFSNCLCDL 615
Query: 574 GASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVEDV 617
GAS+++MPL++ +L + P+ L+L +ADR+ + P+G++EDV
Sbjct: 616 GASVSIMPLSVARKLGFVQYKPSDLTLILADRTSRRPFGLLEDV 659
>At1g36130 hypothetical protein
Length = 914
Score = 197 bits (500), Expect = 2e-50
Identities = 192/707 (27%), Positives = 290/707 (40%), Gaps = 101/707 (14%)
Query: 5 LPELENANRSLRDLTSAAMSYDYPGSIVSPDGTGN-FELRPALINLVSQNQYGGGALEDP 63
+P R+L D + Y +IV P N +EL+P LV Q+ + G + E P
Sbjct: 65 IPPAVERRRTLGDFNRPDLFYANRSAIVPPPLQRNDYELKPGYFALVGQHPFHGLSHEQP 124
Query: 64 HAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFT 123
H+ERF + + VS D + LFP+SL A+ WL GS+ +W + F
Sbjct: 125 MDHIERFEDLVLSIKANGVSEDYLLCKLFPYSLAGEADSWLKQLKAGSLKTWRSIKIAFL 184
Query: 124 TRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQ 183
F A +L+N + TFTQ E AW RFK+ R CP H
Sbjct: 185 NNFYDDAKSEELRNKLSTFTQGPAEAFKAAWVRFKEYQRDCPHHGF-------------- 230
Query: 184 YSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASN 243
S LDAAS+G F+ P LIE +A +NS+ + R + MA
Sbjct: 231 --SEMALDAASNGNFNTHYPADATTLIENLA--CINSTKNADFERKKIAGAISGNQMA-- 284
Query: 244 KQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGPHDSEECTETRPEEEVKAMGQARNDPF 303
+++ +M+ + N + K V F EE E PE E D
Sbjct: 285 -EVNAKMDSVHNLLTGKK------HVHFAA------EEETIEPEPESEEGVFYIVGRDTG 331
Query: 304 SNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGEGEGSGSKKSLE 363
S +H S G G + P Q R+ R G+ +S
Sbjct: 332 S------LGSHMATSVATGLQG------TRVPPTTLQSLLSRRLFLRAAFRGTMILESQT 379
Query: 364 ELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAE------RPQGIFPSDCIPNPKQE 417
+LV F + + Y N NL + +L Q+A+ R +G + NP++
Sbjct: 380 KLVVEFNGKFDGVYTNLNGKIDNLSSHLKKLDVQVAQTAQSIKRQEGFLLGNPDANPRKS 439
Query: 418 NASVVATRSGRVMSELKKKTEGEKR-EEIVGGEVVVPIKVREEVDLTPE---TSKIPFPK 473
+++ V EL + E E E+ + +V + PE T +P+P+
Sbjct: 440 CNAILIREGDDVWEELDTEDELELAVTEMCDRHWIRTAPCEVQVPMQPEPVYTPHVPYPR 499
Query: 474 ALAKKSLDKKFSKFVDVFKKLHISIPFADALE-QMPIYAKFMKDILHKRRRLKGVDETVL 532
K + +K + +K+ ++P DALE +++K ++ L DE L
Sbjct: 500 RRRSKQ-EIHAAKCTTIMEKILYTLP-KDALETSSASLNRYVKRLVDNGISL---DEAKL 554
Query: 533 MTEECSAILQRKMPKKRR----------------------DPGSFTIPVEIEGMADVEAL 570
+T + SAI+ K K++ DPGSF + I +L
Sbjct: 555 LTRDISAIMLPKAKKEKAHRVDIAEYIRTITPGQTTEKLPDPGSFVLDCSISTSRFSRSL 614
Query: 571 CDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDF 630
CDLG+SINLMP ++ ERL + PT ++L AD S + P GI+EDV V
Sbjct: 615 CDLGSSINLMPKSVAERLGMTHYRPTRITLLFADCSKRIPEGILEDVPV----------- 663
Query: 631 VVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGKEKVRFSV 677
++ K PLILGR FLAT A+ DV +G + L V ++ F +
Sbjct: 664 ------KEPKDPLILGRAFLATTGARFDVKRGRISLKVCDSEMEFGM 704
>At3g30630 hypothetical protein
Length = 785
Score = 177 bits (448), Expect = 2e-44
Identities = 168/666 (25%), Positives = 287/666 (42%), Gaps = 80/666 (12%)
Query: 39 NFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRD 98
NF+++ LI+++ N++ G +ED H++ F R CN +V VS D +L LFPF L D
Sbjct: 38 NFKIKSGLISMIQGNKFHGLPMEDLLDHLDEFDRLCNLTKVNGVSEDGFKLRLFPFFLGD 97
Query: 99 TAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFK 158
A W + P SI +W D + F +F A +L N+I +F+Q T E+ EAWERFK
Sbjct: 98 KAHIWEKNMPHDSIITWVDCKKAFLAKFFSNAKTARLINEISSFSQKTGESFCEAWERFK 157
Query: 159 KLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAM 218
+CP H +A ++ Y G+ R LD AS+ F + G+ELIE +A
Sbjct: 158 GYTNQCPHHGFKKASLLSTLYRGVLPRIRMLLDTASNENFQNKDVEEGWELIENLA---- 213
Query: 219 NSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGPH 278
SND + + +++ K +N+ Q+++ + S+ V F+
Sbjct: 214 -QSNDNYNKDCERTIRGTTDSDDKHRKEIKALNDKQDKI----LLSQQKHVHFLV----D 264
Query: 279 DSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFP--- 335
D + + +++ + N YN N+PN S+R S+ N + +P
Sbjct: 265 DEQYQVQDGEGNQLEEVSYINNQSGYKGYNNFKTNNPNLSYR--STNIANPQDQVYPPQR 322
Query: 336 SQGFQGQSLRQP-------QERGEGEGSGSKKSLEE--LVETFINRTENNYKN----QEA 382
Q G +QP Q +G + K + + L E + + Y + EA
Sbjct: 323 QQAISGNYQQQPPPRFAPQQHQGPPAPDANMKQMAQQLLQEQASSSMDCRYNDLNAKVEA 382
Query: 383 ANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQ-ENASVVATRSGRVMSELKKKTEGEK 441
N + GQ A A + G+ P I N K+ + + R + K +
Sbjct: 383 LNTKVRYLEGQSASTSAPKVTGL-PGKPIQNQKEYATVNAITICHDRELPTRHVKVSVVE 441
Query: 442 REEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSK-FVDVFK-------- 492
G ++ E+V + +K P L ++L KF K +++ +K
Sbjct: 442 FNHSAGSHHLIQSTSEEKVAIIERMAKRFKPTPLPSRALPWKFRKAWMERYKSVAEKQLN 501
Query: 493 KLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRRDP 552
++ + +P + L + K +++++ +R ++ + A +R + +K DP
Sbjct: 502 EIEVVMPLMEVLNVIHDPHKDVRNLILERIKMYHDSDDECDANPSRAADKRIVQEKLEDP 561
Query: 553 GSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYG 612
GS T+P I +A LCDLGAS++LMPL++ + + L+ A
Sbjct: 562 GSCTLPCSIGELAFSNCLCDLGASVSLMPLSVAR-------SGSAYKLRCA--------- 605
Query: 613 IVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGKE- 671
++E+ K LIL RPFLA+ A IDV + + L +G
Sbjct: 606 ---------------------RLKEEHKDHLILERPFLASVGAVIDVREVKINLNLGNHI 644
Query: 672 KVRFSV 677
K++F +
Sbjct: 645 KLQFDI 650
>At2g07480 F9A16.15
Length = 1012
Score = 175 bits (443), Expect = 9e-44
Identities = 163/624 (26%), Positives = 269/624 (42%), Gaps = 121/624 (19%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NF+++ LI+++ N++ G +EDP H++ F R C+ ++ VS ++ +
Sbjct: 23 GIVPPPIQNNNFKIKSDLISMIQGNKFHGLPMEDPLDHLDNFDRLCSLTKINGVSEESFK 82
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A W + P SI +W+D + F +F + +L+++I F Q E
Sbjct: 83 LRLFPFSLGDKAHLWEKTLPVKSIDTWDDCKKAFLAKFFSNSRKARLRSEISGFNQKNSE 142
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQY--SSRFGLDAASSGEFDALLPQVG 206
+ EAWERFK +CP H E + Y L+ R LD AS+G G
Sbjct: 143 SFSEAWERFKGYTTQCPHH-----ESLPPQYSILRCLPKIRMLLDTASNGNILNKDVAEG 197
Query: 207 YELIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRV 266
+EL+E +A N + D YD+ +N+ S ++ + ++KT + R+
Sbjct: 198 WELVENLAQSHRNYNED------------YDR---TNRGSSDSEDKHKKEIKT--LNDRI 240
Query: 267 AKVEFVTCGGPH--DSEECTETRPEEE--VKAMGQARNDPFSNTYNPGWRN----HPNFS 318
K+ + EE T+ + E ++ + +N YN G+ N HPN S
Sbjct: 241 DKLVLAQQRNVYYITEEELTQLQDGENLTIEEVSYLQN---QGGYNKGFNNYKPPHPNLS 297
Query: 319 WRQGS-----------------------SGQGNGFQRQFPSQGFQGQSLRQ--------- 346
+R + QG ++ F GF Q +
Sbjct: 298 YRSNNVANPQDQVYPPQNQPTQAKPFVPYNQGYNQKQNFGPPGFTQQPQQPSAQDSEMKT 357
Query: 347 -PQERGEGEGSGSKKSLEELVETFINRTENNYKNQ----EAANKNLENQFGQLAKQIAER 401
PQ+ +G S S ++L E R + +Y + +A N +++ G +A A +
Sbjct: 358 LPQQLVQGHASCSMTMDKKLAE-LTTRIDCSYNDLNIKIDALNTRVKSMEGHIASTSAPK 416
Query: 402 PQGIFPSDCIPNPKQENASVVATRSGRVMSELKKKTEGEKREEIVGGEVVVPIKVREEVD 461
G P + NPK E A ++T T + I GEV+ P + R+E++
Sbjct: 417 HLGQLPGKAVQNPK-EYAHAIST----------VNTSATEDSGIQEGEVLRP-RSRQEIE 464
Query: 462 L--------------------TPETSKIPFPKALA---KKSLDKKFSKFVDVFKKLHISI 498
L +P K FP+ +A + K F+ K+L +
Sbjct: 465 LDFFARLVERAHDPSNPIHIPSPYVPKPAFPERIAQIDEMIFQKHKMMFIKCIKELEEKV 524
Query: 499 PFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRK-MPKKRRDPGSFTI 557
P D ++M + R + + V ++ ECSAI+Q+K +PKK DPGSFT+
Sbjct: 525 PLVDTPKEMIM------------ERPQEAQQIVELSFECSAIIQKKVIPKKLGDPGSFTL 572
Query: 558 PVEIEGMADVEALCDLGASINLMP 581
P + + +LCDLGAS++ P
Sbjct: 573 PCSLAPLVFNNSLCDLGASMDEEP 596
>At4g07370
Length = 531
Score = 150 bits (380), Expect = 2e-36
Identities = 108/326 (33%), Positives = 163/326 (49%), Gaps = 49/326 (15%)
Query: 381 EAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSELKKKTEGE 440
EA N + G+ A + + G+ P I NPK+E V S E E +
Sbjct: 43 EALNSKVRYLEGESASTSSPKVTGL-PGKSIQNPKEERPKSVTEDSAYQDGEDFSLNEDQ 101
Query: 441 KREEIVGGEVVVPIKVREEVDLTPETSK-----------------------IPFPKALAK 477
+ EV+ PI R+ P TS +PFP K
Sbjct: 102 VDKPT---EVLEPILDRDTRPTNPLTSSAALKLVAAKNNEIAFIPPPYKPPLPFPGRHKK 158
Query: 478 KSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEEC 537
+ DK + F KK+ + IP ADAL +P KF+KD++ +R ++ V +T +++ EC
Sbjct: 159 EVEDKYRAMFAKNIKKVELRIPLADALTHIPDSQKFLKDLIMER--IQEVQKTTVLSHEC 216
Query: 538 SAILQRK-MPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPT 596
SAI+Q + +K DPGSFT+P + + + LCDLG S+NLMPL++
Sbjct: 217 SAIIQENDVSEKLGDPGSFTLPCSLGSLTFNKCLCDLGPSVNLMPLSV------------ 264
Query: 597 MLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAK 656
A RS++ P+G++ED+ + + P DFVVL+M+E+ K PLILGRPFLAT
Sbjct: 265 ------AKRSVRLPHGLLEDLPIKIRNVEVPTDFVVLNMDEEPKDPLILGRPFLATVGVI 318
Query: 657 IDVDKGHLILPVGKE-KVRFSVFNPM 681
IDV +G + L +GK +++F + + M
Sbjct: 319 IDVKQGKIDLNLGKNFEMKFDINDAM 344
>At2g10620 putative Athila retroelement ORF1 protein
Length = 451
Score = 149 bits (376), Expect = 5e-36
Identities = 114/376 (30%), Positives = 194/376 (51%), Gaps = 23/376 (6%)
Query: 326 QGNGFQRQFPSQGFQ--GQSLRQPQERGEGEGSGSKKSLEEL-----VETFINRTENNYK 378
+GN Q Q G++++ P+E S K+L + V + K
Sbjct: 52 EGNSASTSAQKQTSQLPGKAIQNPKEYAHAITLRSGKALPTIEAPKTVTVGKRPSRKTSK 111
Query: 379 NQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSELKKKTE 438
N N G AK++ +R G+ ++ + E+ S+ ++ + + + ++
Sbjct: 112 NDRGRTGRERNSKGTKAKEMGKRFGGVIVTEDSVDHDGEDFSLTKDQADKELEQPLDQSL 171
Query: 439 GEKREE-------IVGGEVVVPIKVR--EEVDLTPETSKIPFPKALAKKSLDKKFS-KFV 488
+ + I V P+ V+ E+V + P P KK+L K+ F
Sbjct: 172 EQPLDHFTRPPFPITAPTAVKPVVVKNKEKVFVPPPYKPHPSFPGRHKKALAHKYRVMFA 231
Query: 489 DVFKKLHISIPFADALEQMPIYAKFMKDILHKR-RRLKGVDETVLMTEECSAILQRKM-P 546
K++ + IP DAL + KF+KD++ +R + L+G+ V++++ECSAI+Q+K+
Sbjct: 232 KNIKEVELQIPLVDALALIMDSHKFLKDLIVERIQELQGM---VVLSDECSAIIQKKIIH 288
Query: 547 KKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRS 606
KK DPGSFT+P + +A + LCDLGAS +LMPL++ +RL + +SL +ADRS
Sbjct: 289 KKLSDPGSFTLPCSLGPLAFNKCLCDLGASASLMPLSVTKRLGFTQYKSCNISLILADRS 348
Query: 607 LKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLIL 666
++ P+ +E++ + + P DFVVL+M+E+ K LILGRPFL T A I + KG + L
Sbjct: 349 VRIPHDFLENIPIRIRAVEIPTDFVVLEMDEEPKDHLILGRPFLTTVGAMIYIKKGKINL 408
Query: 667 PVGKE-KVRFSVFNPM 681
+GK+ ++ F V + M
Sbjct: 409 NLGKDFRMTFDVKDTM 424
>At1g41690 hypothetical protein
Length = 371
Score = 148 bits (373), Expect = 1e-35
Identities = 100/311 (32%), Positives = 156/311 (50%), Gaps = 20/311 (6%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NFE++ +LI +V N++ G +EDP H++ F R C+ ++ VS D +
Sbjct: 28 GIVPPPVQNNNFEIKSSLIAMVQGNKFHGLPMEDPLDHLDEFERLCSLTKINGVSEDGFK 87
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A W + PQGSIT+W+D + F +F + +L+N+I FTQ E
Sbjct: 88 LRLFPFSLGDKAHLWEKTLPQGSITTWDDCKKAFLAKFFSYSRTARLRNEISGFTQKQSE 147
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
+ EAWERFK +CP H QA ++ Y G+ R LD AS+G F + G+E
Sbjct: 148 SFCEAWERFKGYQTKCPHHGFKQASLLSTLYRGVLPKIRMLLDTASNGNFLKKDVEEGWE 207
Query: 209 LIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEI-QNQMKTTKIGSRVA 267
L+EK A ++ + N+ R ++ D+ K L+ ++++I Q Q K
Sbjct: 208 LVEKFA-QSDGNYNEDYDRSIRTSSDSDDKHRRDMKALNDKLDKILQVQQK--------- 257
Query: 268 KVEFVTCGGPHDSEECTETRPEEEVKAMGQARND-PFSNTYNPGWRNHPNFSWRQGSSGQ 326
V ++ P +E E + A +N + YN +R +PN S+R S+
Sbjct: 258 HVHLISEDEPFQVQE-----GENDQCAYSYVQNQGGYDKGYN-NYRPNPNLSYR--STNI 309
Query: 327 GNGFQRQFPSQ 337
N + +P Q
Sbjct: 310 ANPHDQVYPQQ 320
>At4g07600
Length = 630
Score = 142 bits (359), Expect = 5e-34
Identities = 86/237 (36%), Positives = 143/237 (60%), Gaps = 6/237 (2%)
Query: 487 FVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKM- 545
F K++ + IP DAL + KF+KD++ +R ++ V V+++ ECSAI+Q+K+
Sbjct: 2 FAKNIKEVELRIPLVDALALILDTHKFLKDLIVER--IQEVQGMVVLSHECSAIIQKKIV 59
Query: 546 PKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADR 605
PKK DPGSFT+P + +A LCDLGA ++ MPL++ +RL + +SL +ADR
Sbjct: 60 PKKLSDPGSFTLPCFLGTVAFNRCLCDLGALVSPMPLSIAKRLGFTQYKSCNISLILADR 119
Query: 606 SLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLI 665
S++ +G+++++ + + DFV+L+M+E+ K PLIL RPFLAT A IDV KG +
Sbjct: 120 SVRISHGLLKNLPIRIGAAEISTDFVILEMDEEPKDPLILRRPFLATAGAMIDVKKGKID 179
Query: 666 LPVGKE-KVRFSVFNPMIETNHDNDFVCDVIRSRSKVSEETPKVNSQIDEFHPALIK 721
L +GK+ ++ F + + M + N + I ++++E + +Q D + AL K
Sbjct: 180 LNLGKDFRMTFDIKDAMKKPNIEGQLFW--IEEMDQLADELLEELAQEDHLNSALTK 234
>At3g31500 hypothetical protein
Length = 591
Score = 141 bits (356), Expect = 1e-33
Identities = 134/523 (25%), Positives = 234/523 (44%), Gaps = 56/523 (10%)
Query: 190 LDAASSGEFDALLPQVGYELIEKMAIRAMNSS--NDRQARRGVLEVEAYDQLMASNKQLS 247
LD AS+GEF +LIE +A N + DR R G E + + +L A +QL
Sbjct: 3 LDTASNGEFMTKTETEATKLIENLAGSNSNHNVDYDRSNRGGGGESKQFAELSAKVEQLM 62
Query: 248 KQMNEIQNQMKTTKIGSRVAKVEFVTCGGPHDSEECTETRPEEEVKAMGQARNDPFSNTY 307
++ + N + + G + EF G E V G +N +N
Sbjct: 63 RRDQKSVNFCEDSSKG--MVHQEFSGDGSEDLQAEINF------VNGYGNYQNGGSTNVE 114
Query: 308 NPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGEGEGSGS--------- 358
NP + +P + QG Q GFQ + +QGQ + P G G
Sbjct: 115 NPQDQVYPTQAGSQGQKFQPYGFQNK---GNYQGQ-FQPPAGTGHASSFGDNEMKLMMQQ 170
Query: 359 -----KKSLEEL---VETFINRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDC 410
KK+ ++ V++ N + + K LENQ Q+ + RP G
Sbjct: 171 VLEDQKKNAADINVKVDSMYNDLNGKFATLSSHVKTLENQVSQIVSA-SMRPDGTHSGKV 229
Query: 411 IPNPKQENASVVATRSGRVMSELKKKTEGEKRE-EIVGGEVVVPIKVREE-----VDLTP 464
P K++ +++ + EL++ ++ E +V E +V K+ E+ V+ P
Sbjct: 230 KPKGKEQCYAIM------IQEELREIVVAKQVETNVVVVETLVEDKIVEDDEPLSVEPPP 283
Query: 465 ETSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRL 524
K+PFP + K++++F ++ K+L++ +PF + +P Y ++K IL +R +
Sbjct: 284 YVPKLPFPGRERQIQRQKEYARFDEIMKQLYVRLPFLQLVLHVPSYRSYLKYILSNKRSI 343
Query: 525 KGVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTM 584
+ + + EE + +++ + +K T VE+ ++ C + P T+
Sbjct: 344 EEGVKLISKGEEHAQLVESQRQQKE---AQLTNVVEMLAGKEMVNTCSA-----IPPATI 395
Query: 585 FERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLI 644
++L + P+ +SL +ADRS++ P G+ E+V V + P +FVVL+++++ PL
Sbjct: 396 PKKLGITNFKPSRISLILADRSVQFPMGLAENVHARVGNFYIPTNFVVLELDKEPHDPLN 455
Query: 645 LGRPFLATGRAKIDVDKGHLILPVGKEKVRFSV----FNPMIE 683
LGRPFL T A IDV + + L +G + F + NP IE
Sbjct: 456 LGRPFLNTVEAIIDVRRSTINLQIGDYALEFDIKGTRKNPTIE 498
>At1g38440 hypothetical protein
Length = 263
Score = 130 bits (327), Expect = 3e-30
Identities = 86/273 (31%), Positives = 136/273 (49%), Gaps = 14/273 (5%)
Query: 49 LVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQP 108
++ +N++ G +EDP H++ F R N ++ VS D +L LFPFSL D A W + P
Sbjct: 1 MIQENKFHGLPMEDPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLP 60
Query: 109 QGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHN 168
SIT+W+D + F ++F A +L+N+I F+Q T E+ EAWERFK +CP H
Sbjct: 61 HDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTSESFCEAWERFKGYTNQCPHHG 120
Query: 169 LTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNSSND-RQAR 227
T+A ++ Y G+ LDAAS+G F + G+EL+E +A N + D +
Sbjct: 121 FTKASLLSTLYRGVLPCIIMLLDAASNGNFQNKDVEEGWELVENLAQSYGNYNEDCDRTV 180
Query: 228 RGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGPHDSEECTETR 287
RG + SN + K++ + +++ + S+ V F+ ++ E
Sbjct: 181 RGTAD---------SNDKHRKEIKALNDKLDRIFL-SQQKHVHFLVDDKQFQVQD-GEGN 229
Query: 288 PEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWR 320
EEV + N YN N+PN S+R
Sbjct: 230 QLEEVSYIN--NNQGGYKGYNNFKTNNPNLSYR 260
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 127 bits (319), Expect = 2e-29
Identities = 81/238 (34%), Positives = 115/238 (48%), Gaps = 51/238 (21%)
Query: 432 ELKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDVF 491
+L+K+T EK ++ V L P K+PFP K +K++S F ++
Sbjct: 2 KLRKQTLEEKAKKTV---------------LPPYVPKLPFPGRQRKIQREKEYSLFDEIM 46
Query: 492 KKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRRD 551
++L P ECSAILQ +P KR D
Sbjct: 47 RQLQRYSP------------------------------------ECSAILQNVIPVKRED 70
Query: 552 PGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPY 611
PGSF +P I LCDLGA ++LMP ++ +RL TPT +SL + DRS+ P
Sbjct: 71 PGSFVLPSRIGEYTFDRCLCDLGAGVSLMPFSVAKRLGDTNFTPTKMSLVLGDRSISFPV 130
Query: 612 GIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVG 669
G+ EDV V V + P DFV+++++E+ + LILGRPFL A IDV K + L +G
Sbjct: 131 GVAEDVQVRVGNFYIPTDFVIIELDEEPRHRLILGRPFLNIVAALIDVRKSKINLRIG 188
>At3g31340 Athila ORF 1, putative
Length = 781
Score = 125 bits (314), Expect = 8e-29
Identities = 76/247 (30%), Positives = 124/247 (49%), Gaps = 2/247 (0%)
Query: 11 ANRSLRDLTSAAMSYDYPGSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERF 70
A ++LRD + + + + P +FE++ L+ LV QNQ+ + E+ H++ F
Sbjct: 58 ARQTLRDHDAPNIDFFRDRNQHLPINRNDFEIKTGLLALVKQNQFFVHSSENHVYHLDNF 117
Query: 71 IRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRA 130
C R+ V D I+ LF FSL D A WL S ++ SWED F ++ ++
Sbjct: 118 EDICRITRMNGVPDDIIKCRLFIFSLADNAHRWLKSLDPINLRSWEDYKAAFLGQYFTQS 177
Query: 131 LLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGL 190
L+N I +F Q E+ EAWERFK R CP H ++A ++ FY G+ + + L
Sbjct: 178 RTAILRNKISSFQQGGTESFPEAWERFKDYYRECPHHGFSRATLISTFYQGVDKAYKMAL 237
Query: 191 DAASSGEFDALLPQVGYELIEKMAIRAMNSS--NDRQARRGVLEVEAYDQLMASNKQLSK 248
D AS+G+F +LI+ +A +N + DR R G E++ +L +QL +
Sbjct: 238 DTASNGDFMTKTETEATKLIKNLAASNINHNVDYDRSNRGGGGELKQLAELSVKVEQLLR 297
Query: 249 QMNEIQN 255
+ ++ N
Sbjct: 298 RDQKVIN 304
Score = 122 bits (306), Expect = 7e-28
Identities = 68/155 (43%), Positives = 93/155 (59%), Gaps = 4/155 (2%)
Query: 533 MTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGE 592
M CSAI +P+K DPGSF +P I A LCDLGA +NLMPL+M +RL +
Sbjct: 516 MVSTCSAIPPATIPEKLGDPGSFVLPCRIGKSAFERCLCDLGAGVNLMPLSMSKRLGITN 575
Query: 593 VTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLAT 652
P+ +SL +ADRS++ P G+ E+V V V + P DFVVL+++++ PL LGRPFL T
Sbjct: 576 FKPSRISLILADRSVRFPVGLAENVHVRVGDFYIPTDFVVLELDKEPHDPLTLGRPFLNT 635
Query: 653 GRAKIDVDKGHLILPVGKEKVRFSV----FNPMIE 683
A IDV + + L +G + F + NP IE
Sbjct: 636 VGAIIDVRRSTINLQIGDFALEFDMKGTRKNPTIE 670
>At2g13260 putative Athila retroelement ORF1 protein
Length = 622
Score = 123 bits (308), Expect = 4e-28
Identities = 63/170 (37%), Positives = 96/170 (56%), Gaps = 9/170 (5%)
Query: 34 PDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFP 93
P NFE++ +LIN+V +++ G ++EDP H+E+F C+T ++ +S D +L LFP
Sbjct: 51 PVENNNFEIKSSLINMVQTSKFHGLSMEDPLDHLEQFDMLCSTVKINGISEDAFKLRLFP 110
Query: 94 FSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEA 153
FSL D A W + PQ SITSW+ F ++F +L+N+I +FTQ ++E+ EA
Sbjct: 111 FSLGDRARIWEKNLPQRSITSWDQCKRAFLSKFFSTTRTARLRNEISSFTQRSNESFCEA 170
Query: 154 WERFKKLLRRCPQH---------NLTQAEQVAKFYDGLQYSSRFGLDAAS 194
WERFK +CP H NL Q++ A L+ R +D A+
Sbjct: 171 WERFKGYKMQCPHHGFSKESILKNLAQSDTNALLRQILEGQGRGAIDLAT 220
Score = 77.4 bits (189), Expect = 3e-14
Identities = 86/357 (24%), Positives = 138/357 (38%), Gaps = 81/357 (22%)
Query: 303 FSNTYNPGWRNH-PNFSWRQGSS-----GQGNGFQRQFPSQGFQGQSLRQPQERG----- 351
FS T RN +F+ R S + G++ Q P GF +S+ + +
Sbjct: 144 FSTTRTARLRNEISSFTQRSNESFCEAWERFKGYKMQCPHHGFSKESILKNLAQSDTNAL 203
Query: 352 -----EGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIF 406
EG+G G+ L ++ + +N Y A + L + ++ P G
Sbjct: 204 LRQILEGQGRGAI-DLATQMKGMHTKVDNIYGELNAKIERLNVHVYSPSSSTSKHPMGTL 262
Query: 407 PSDCIPNPKQ-------------ENASVVATRS-GRV------MSELKKKTEGEKREEIV 446
P N K+ EN S +R GR+ + E+ K G E +V
Sbjct: 263 PGKSETNHKEFCNAIFINDFDVVENMSYTQSREDGRIDENEKAIEEISKLLYGSNVENLV 322
Query: 447 ------------GGEVV---VPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDVF 491
G +VV V +V+ P +PFP + K+ K FS F
Sbjct: 323 VASDEKAKKSTNGNDVVTKSVEKNAASKVESPPYEPPLPFPGRVLTKAKKKVFSSFKSNM 382
Query: 492 KKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRRD 551
++ +P + L Q+P++ +F++ IL R +L+ D
Sbjct: 383 SRVGAPLPCVENLSQVPLHFQFIQAILENREKLE-------------------------D 417
Query: 552 PGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLK 608
PG FTIP + + +ALCD GAS+N+M L MF G +T + + +SLK
Sbjct: 418 PGKFTIPCSLGDLQLDDALCDSGASVNVMSLAMF----CGTITNEEVKTKKEPKSLK 470
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,606,474
Number of Sequences: 26719
Number of extensions: 752009
Number of successful extensions: 2434
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 2287
Number of HSP's gapped (non-prelim): 123
length of query: 728
length of database: 11,318,596
effective HSP length: 107
effective length of query: 621
effective length of database: 8,459,663
effective search space: 5253450723
effective search space used: 5253450723
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0388a.1


