

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0381.8
(460 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
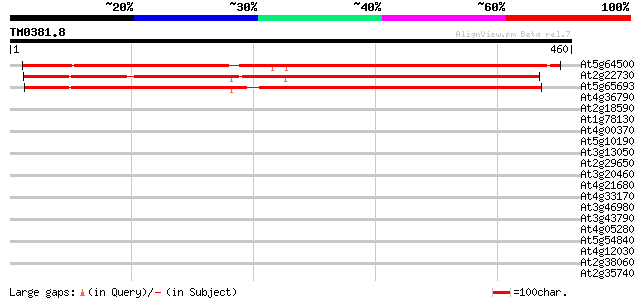
Score E
Sequences producing significant alignments: (bits) Value
At5g64500 unknown protein 565 e-161
At2g22730 hypothetical protein 542 e-154
At5g65693 unknown protein 524 e-149
At4g36790 unknown protein 40 0.002
At2g18590 hypothetical protein 40 0.003
At1g78130 transporter like protein 37 0.030
At4g00370 unknown protein 36 0.039
At5g10190 unknown protein 33 0.25
At3g13050 putative transporter 33 0.25
At2g29650 Na+-dependent inorganic phosphate cotransporter like p... 33 0.33
At3g20460 sugar transporter, putative 32 0.96
At4g21680 peptide transporter - like protein 30 2.8
At4g33170 putative protein 30 3.6
At3g46980 unknown protein 30 3.6
At3g43790 transporter-like protein 30 3.6
At4g05280 putative protein 29 6.2
At5g54840 SGP1 monomeric G-protein (emb|CAB54517.1) 28 8.1
At4g12030 Sodium Bile acid symporter (SBF) like protein 28 8.1
At2g38060 putative Na+-dependent inorganic phosphate cotransporter 28 8.1
At2g35740 putative sugar transporter 28 8.1
>At5g64500 unknown protein
Length = 484
Score = 565 bits (1455), Expect = e-161
Identities = 284/462 (61%), Positives = 344/462 (73%), Gaps = 30/462 (6%)
Query: 11 PSWFTPKRLLLIFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGSGIQGDFNLNN 70
PSWFTPK+LL +FCV+N INY+DRGAIASNG+NG+ +CT SG C++GSGIQGDFNL+N
Sbjct: 30 PSWFTPKKLLFVFCVVNLINYIDRGAIASNGINGSRGSCTS-SGTCSSGSGIQGDFNLSN 88
Query: 71 FQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITIC 130
F+DGVLSSAFMVGLL+ASPIFASLAKS +PFRLIGVGLS+WT AV GCG+SFDFWSITIC
Sbjct: 89 FEDGVLSSAFMVGLLVASPIFASLAKSVNPFRLIGVGLSIWTLAVIGCGLSFDFWSITIC 148
Query: 131 RMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTALGYVYGFAPLESKKTI 190
RM VGVGEASF+SLAAPFIDDNAP QK+AWLA FYMCIP G A GYVYG +
Sbjct: 149 RMFVGVGEASFVSLAAPFIDDNAPHDQKSAWLAVFYMCIPTGYAFGYVYG-------GVV 201
Query: 191 SSIETNVSEHGDDGVLAQDQAFIR-----------GTRSTSKLRNQ----------FTRF 229
S+ + + +L A + T K R F+
Sbjct: 202 GSVLPWRAAFWGEAILMLPFAVLGFVIKPLHLKGFAPDDTGKPRTDNLNVLPVGYGFSAV 261
Query: 230 SKDMQELLYDQVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFGGITIVCGI 289
KD++ LL D+VY+ N+LGYI+YNFV+GAYSYWGPKAGY+IY M N D++FGG+T+VCGI
Sbjct: 262 MKDLKLLLVDKVYVTNILGYIAYNFVLGAYSYWGPKAGYNIYKMENADMIFGGVTVVCGI 321
Query: 290 LGTLGGGLILDRMTSTISNAFKLLSGATLLGAIFCLVAFLFKNLSGFIVFFSIGELLIFA 349
+GTL GG+ILD M +TISNAFK+LS +T +GAIFC AF FK++ F+ F++GELL+FA
Sbjct: 322 VGTLSGGVILDYMDATISNAFKVLSVSTFIGAIFCFAAFCFKSMYAFLALFAVGELLVFA 381
Query: 350 TQAPVNYVSLRCVKPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQDRINDWRKSALCLTS 409
TQ PVN++ L CVKPSLRPL+MA+STVSIHIFGDVPS+PLVGVLQD +N+WR ++L LT
Sbjct: 382 TQGPVNFIVLHCVKPSLRPLAMAMSTVSIHIFGDVPSSPLVGVLQDYVNNWRVTSLVLTF 441
Query: 410 VFFLAAGIWFIGIFLRTVDLSTEDDEDQSATSLRGKMKPLLE 451
V F AA IW IGIFL +VD ED E + T PLL+
Sbjct: 442 VLFPAAAIWSIGIFLNSVDRYNEDSEPDAVTR-ESTAAPLLQ 482
>At2g22730 hypothetical protein
Length = 507
Score = 542 bits (1396), Expect = e-154
Identities = 280/456 (61%), Positives = 331/456 (72%), Gaps = 41/456 (8%)
Query: 12 SWFTPKRLLLIFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGSGIQGDFNLNNF 71
S +P LL+IFC+IN +NY+DRGAIASNGVNG+ +C +D G CT +GIQG FNL+NF
Sbjct: 44 SSLSPVWLLVIFCIINLLNYMDRGAIASNGVNGSTRSC-NDKGKCTLATGIQGHFNLSNF 102
Query: 72 QDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICR 131
+DGVLSS+FMVGLLIASPIFASLAK RLIGVGL+VWT AV GCG SF FW I +CR
Sbjct: 103 EDGVLSSSFMVGLLIASPIFASLAK-----RLIGVGLTVWTIAVLGCGSSFAFWFIVLCR 157
Query: 132 MLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTALGYVYG----------- 180
M VGVGEASFISLAAPFIDDNAP QK AWL FYMCIP+G ALGYVYG
Sbjct: 158 MFVGVGEASFISLAAPFIDDNAPQEQKAAWLGLFYMCIPSGVALGYVYGGYVGKHFSWRY 217
Query: 181 ------------------FAPLESKKTISSIETNVSEHGDDGVLAQDQAFIRGTRSTSKL 222
PL+ K S N + D + DQ + S S
Sbjct: 218 AFWGEAVLMAPFAVLGFLMKPLQLKG--SETLKNNNRLQVDNEIEHDQFEVSIETSKSSY 275
Query: 223 RN----QFTRFSKDMQELLYDQVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDL 278
N FT F+KDM+ L ++V+++NVLGY+SYNFVIGAYSYWGPKAGY+IY M N D+
Sbjct: 276 ANAVFKSFTGFAKDMKVLYKEKVFVVNVLGYVSYNFVIGAYSYWGPKAGYNIYKMKNADM 335
Query: 279 LFGGITIVCGILGTLGGGLILDRMTSTISNAFKLLSGATLLGAIFCLVAFLFKNLSGFIV 338
+FG +TI+CGI+GTL GG ILDR+T+TI NAFKLLSGAT LGA+FC AF K+L GFI
Sbjct: 336 IFGAVTIICGIVGTLSGGFILDRVTATIPNAFKLLSGATFLGAVFCFTAFTLKSLYGFIA 395
Query: 339 FFSIGELLIFATQAPVNYVSLRCVKPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQDRIN 398
F++GELL+FATQAPVNYV L CVKPSLRPLSMAISTV+IHIFGDVPS+PLVG++QD IN
Sbjct: 396 LFALGELLVFATQAPVNYVCLHCVKPSLRPLSMAISTVAIHIFGDVPSSPLVGIVQDHIN 455
Query: 399 DWRKSALCLTSVFFLAAGIWFIGIFLRTVDLSTEDD 434
WRK+ L LTS+ FLAA IWFIGIF+ +VD +++
Sbjct: 456 SWRKTTLILTSILFLAAAIWFIGIFINSVDRFNQEE 491
>At5g65693 unknown protein
Length = 492
Score = 524 bits (1350), Expect = e-149
Identities = 265/462 (57%), Positives = 324/462 (69%), Gaps = 48/462 (10%)
Query: 13 WFTPKRLLLIFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGSGIQGDFNLNNFQ 72
+ TP R + I C+IN INYVDRG IASNGVNG+ C D GVC+AG+GIQG+FNL NF+
Sbjct: 23 FLTPGRFVTILCIINLINYVDRGVIASNGVNGSSKVC-DAKGVCSAGTGIQGEFNLTNFE 81
Query: 73 DGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRM 132
DG+LSSAFMVGLL+ASPIFA L+K +PF+LIGVGL+VWT AV GCG S++FW I + RM
Sbjct: 82 DGLLSSAFMVGLLVASPIFAGLSKRFNPFKLIGVGLTVWTIAVIGCGFSYNFWMIAVFRM 141
Query: 133 LVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTALGYVYG------------ 180
VGVGEASFISLAAP+IDD+APVA+K WL FYMCIPAG ALGYV+G
Sbjct: 142 FVGVGEASFISLAAPYIDDSAPVARKNFWLGLFYMCIPAGVALGYVFGGYIGNHLGWRWA 201
Query: 181 --------------------------FAPLESKKTISSIETNVSEHGDDGVLAQDQAFIR 214
FA +SKK +SIET V D +
Sbjct: 202 FYIEAIAMAVFVILSFCIKPPQQLKGFADKDSKKPSTSIET---------VAPTDAEASQ 252
Query: 215 GTRSTSKLRNQFTRFSKDMQELLYDQVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMS 274
T K +N F KD++ L ++V+I+NVLGYI+YNFVIGAYSYWGPKAG+ IY M
Sbjct: 253 IKTKTPKSKNLVVLFGKDLKALFSEKVFIVNVLGYITYNFVIGAYSYWGPKAGFGIYKMK 312
Query: 275 NPDLLFGGITIVCGILGTLGGGLILDRMTSTISNAFKLLSGATLLGAIFCLVAFLFKNLS 334
N D++FGG+TI+CGI+GTLGG +LDR+ +T+SN FKLL+ +TLLGA FC AFL KN+
Sbjct: 313 NADMIFGGLTIICGIIGTLGGSYVLDRINATLSNTFKLLAASTLLGAAFCFTAFLMKNMY 372
Query: 335 GFIVFFSIGELLIFATQAPVNYVSLRCVKPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQ 394
FI F++GE+LIFA QAPVN+V L CV+P+LRPLSMA STV IHI GDVPS+PL G +Q
Sbjct: 373 AFIALFAVGEILIFAPQAPVNFVCLHCVRPNLRPLSMASSTVLIHILGDVPSSPLYGKMQ 432
Query: 395 DRINDWRKSALCLTSVFFLAAGIWFIGIFLRTVDLSTEDDED 436
D + +WRKS L +TS+ FLAA IW IGIF+ +VD S E ED
Sbjct: 433 DHLKNWRKSTLIITSILFLAAIIWGIGIFMNSVDRSNEVSED 474
>At4g36790 unknown protein
Length = 489
Score = 40.4 bits (93), Expect = 0.002
Identities = 64/353 (18%), Positives = 123/353 (34%), Gaps = 63/353 (17%)
Query: 86 IASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRMLVGVGEASFISLA 145
+ASP+ L ++ ++ +G W + A G S F + + R + G G A I
Sbjct: 93 LASPLAGVLVITYDRPIVLAIGTFCWALSTAAVGASSYFIQVALWRAVNGFGLAIVIPAL 152
Query: 146 APFIDDNAPVAQKTA---------------------------------WLATFYMCIPAG 172
FI D+ + A W F M
Sbjct: 153 QSFIADSYKDGARGAGFGMLNLIGTIGGIGGGVVATVMAGSEFWGIPGWRCAFIMMAALS 212
Query: 173 TALGYVYGFAPLESKKTISSIETNVSEHGDDGVLAQDQAFIRGTRSTSKLRNQFTRFSKD 232
+G + ++ +K I E + + V A + S
Sbjct: 213 AVIGLLVFLFVVDPRKNIEREELMAHKMNSNSVWNDSLAAAKSVVKVST----------- 261
Query: 233 MQELLYDQVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFGGITIVCGILGT 292
+ + ++G + ++ ++ W G+ H LL G+ G +GT
Sbjct: 262 -----FQIIVAQGIIGSFPWTAMV-FFTMWFELIGFD--HNQTAALL--GVFATGGAIGT 311
Query: 293 LGGGLILDRMTSTISNAFKLLSG--ATLLGAIFCLVAF--LFKNLSGFIVF----FSIGE 344
L GG+I D+M+ N+ +++ + +G F ++ + ++ S + +F F +G
Sbjct: 312 LMGGIIADKMSRIYPNSGRVMCAQFSAFMGIPFSIILLKVIPQSTSSYSIFSITLFLMGL 371
Query: 345 LLIFATQAPVNYVSLRCVKPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQDRI 397
+ + A + V P R + A F +APLVG+L +++
Sbjct: 372 TITWCGSAVNAPMFAEVVPPRHRTMIYAFDRAFEGSFSSF-AAPLVGILSEKL 423
>At2g18590 hypothetical protein
Length = 433
Score = 40.0 bits (92), Expect = 0.003
Identities = 72/370 (19%), Positives = 137/370 (36%), Gaps = 76/370 (20%)
Query: 74 GVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRML 133
G+LS + +ASP+ A S+ + G W + G+S F +T+
Sbjct: 29 GLLSFIRNIVQGLASPLAGLFAISYDRPTVFAFGSFFWVSSTVATGVSRYFIQVTLGVAF 88
Query: 134 VGVGEASFISLAAPFIDDNAPVAQK---------------------------------TA 160
GVG A + I D+ + + +
Sbjct: 89 NGVGHAIVYPVLQSIIADSFKESSRGFGFGLWNLIGTVGGIGGTVVPTVMAGHDFFGISG 148
Query: 161 WLATFYMCIPAGTALGYVYGF--APLESKKTISSIETNVSEHGDDGVLAQDQAFIRGTRS 218
W F + T +G + F + KKT S I + +H D + + + S
Sbjct: 149 WRCAFILSATLSTIVGILVFFFVSDPREKKTSSVIVHHDDQHERD---ENNGGTMMESPS 205
Query: 219 TSKLRNQFTRFSKDMQELLYDQVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDL 278
+S + + KD+ +L Q+ ++ G + ++ N
Sbjct: 206 SSVWKESWVAI-KDVTKLRTFQIIVLQ--GIVGFDH--------------------NQAA 242
Query: 279 LFGGITIVCGILGTLGGGLILDRMTSTISNAFKLLSG--ATLLGAIFCLV--AFLFKNLS 334
L GI +G+L GG+I D+M+ N+ +L+ + +GA+F +V + ++++
Sbjct: 243 LLNGIFATGQAIGSLVGGIIADKMSRVFPNSGRLICAQFSVFMGAMFSIVLLRMIPQSVN 302
Query: 335 GFIVF----FSIGELLIFATQAPVNYVSLRCVKPSLRPLSMAIS---TVSIHIFGDVPSA 387
F +F F +G + + A + + V R + A V+ FG A
Sbjct: 303 SFYIFLVTLFLMGLTITWCGPAINSPILAEIVPAKHRTMVYAFDRALEVTFSSFG----A 358
Query: 388 PLVGVLQDRI 397
PLVG++ +++
Sbjct: 359 PLVGIMSEKL 368
>At1g78130 transporter like protein
Length = 490
Score = 36.6 bits (83), Expect = 0.030
Identities = 55/267 (20%), Positives = 103/267 (37%), Gaps = 37/267 (13%)
Query: 89 PIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRMLVGVGEASFISLAAPF 148
P+ A +A H+ +I +G +W+ A S F+ + + R L G+G A
Sbjct: 59 PLAAYMAIRHNRAHVIALGAFLWSAATFLVAFSSTFFQVAVSRALNGIGLALVAPAIQSL 118
Query: 149 IDDNAPVAQKTAWLATFYMCIPAGTALGYVYG--FAPLESKKT-----------ISSIET 195
+ D+ A + + G+ LG + APL + S+
Sbjct: 119 VADSTDDANRGTAFGWLQLTANIGSILGGLCSVLIAPLTFMGIPGWRVAFHIVGVISVIV 178
Query: 196 NVSEHGDDGVLAQDQAFIRGTRSTSKLRNQFTRFSKDMQELLYDQVYIINVLGY------ 249
V V A D F++ S F ++++L+ + +I + +
Sbjct: 179 GVLVR----VFANDPHFVKDGVDVSNQPGSRKPFCTEVKDLVREADTVIKIRSFQIIVAQ 234
Query: 250 -ISYNFVIGAYSY---WGPKAGYSIYHMSNPDLLFGGITIVCGILGTLGGGLILDRMTST 305
++ +F A S+ W G+S H L+ G+ + LG L GG + D +++
Sbjct: 235 GVTGSFPWSALSFAPMWLELIGFS--HGKTAFLM--GLFVAASSLGGLFGGKMGDFLSTR 290
Query: 306 ISNAFKLL------SGATLLGAIFCLV 326
+ N+ +++ + A L AI LV
Sbjct: 291 LPNSGRIILAQISSASAIPLAAILLLV 317
>At4g00370 unknown protein
Length = 541
Score = 36.2 bits (82), Expect = 0.039
Identities = 30/124 (24%), Positives = 55/124 (44%), Gaps = 14/124 (11%)
Query: 65 DFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFA------VAGC 118
++N ++ G++ S+F G L+ + A ++G G+ W+FA A
Sbjct: 162 EYNWSSATVGLIQSSFFWGYLLTQILGGIWADKFGGKVVLGFGVVWWSFATIMTPIAARL 221
Query: 119 GISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTALGYV 178
G+ F + + R +G+GE + + PV++++ LA Y +G LG V
Sbjct: 222 GLPF----LLVVRAFMGIGEGVAMPAMNNMLSKWIPVSERSRSLALVY----SGMYLGSV 273
Query: 179 YGFA 182
G A
Sbjct: 274 TGLA 277
>At5g10190 unknown protein
Length = 488
Score = 33.5 bits (75), Expect = 0.25
Identities = 29/95 (30%), Positives = 43/95 (44%), Gaps = 10/95 (10%)
Query: 89 PIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRMLVGVGEA----SFISL 144
P+ A L+ H+ +I +G +W A +S F+ + + R L G+G A + SL
Sbjct: 59 PLAAYLSSRHNRAHVIALGAFLWATATFLVAVSTTFFQVAVSRGLNGIGLAIVTPAIQSL 118
Query: 145 AAPFIDD-NAPVAQKTAWLATFYMCIPAGTALGYV 178
A DD N +A WL G+ LGYV
Sbjct: 119 VADSTDDYNRGMA--FGWLG---FTSNIGSILGYV 148
>At3g13050 putative transporter
Length = 500
Score = 33.5 bits (75), Expect = 0.25
Identities = 29/117 (24%), Positives = 53/117 (44%), Gaps = 5/117 (4%)
Query: 59 GSGIQGDFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAG- 117
G +Q +NL+ Q+ +++S G+LI + + ++ H R G ++ VAG
Sbjct: 48 GPAVQSLWNLSARQESLITSVVFAGMLIGAYSWGIVSDKHG--RRKGFIITAVVTFVAGF 105
Query: 118 -CGISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGT 173
S ++ + I R LVG+G LA+ ++ + P + W+ F GT
Sbjct: 106 LSAFSPNYMWLIILRCLVGLGLGGGPVLASWYL-EFIPAPSRGTWMVVFSAFWTVGT 161
>At2g29650 Na+-dependent inorganic phosphate cotransporter like
protein
Length = 512
Score = 33.1 bits (74), Expect = 0.33
Identities = 30/127 (23%), Positives = 53/127 (41%), Gaps = 14/127 (11%)
Query: 62 IQGDFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAV------ 115
+ ++ N G++ S+F G L+ A + R++G G+ W+ A
Sbjct: 130 MSAEYGWNPATVGLIQSSFFWGYLLTQIAGGIWADTVGGKRVLGFGVIWWSIATILTPVA 189
Query: 116 AGCGISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTAL 175
A G+ + + + R +GVGE + + PV +++ LA Y +G L
Sbjct: 190 AKLGLPY----LLVVRAFMGVGEGVAMPAMNNILSKWVPVQERSRSLALVY----SGMYL 241
Query: 176 GYVYGFA 182
G V G A
Sbjct: 242 GSVTGLA 248
>At3g20460 sugar transporter, putative
Length = 488
Score = 31.6 bits (70), Expect = 0.96
Identities = 30/126 (23%), Positives = 52/126 (40%), Gaps = 6/126 (4%)
Query: 58 AGSGIQGDFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAG 117
A +GI NL+ + + +G L+ + + LA +GV S F +AG
Sbjct: 77 AQTGIMAGLNLSLAEFSFFGAVLTIGGLVGAAMSGKLADVFGRRGALGVSNS---FCMAG 133
Query: 118 ---CGISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTA 174
S WS+ I R+ +GV + +I + AP + + A + + A A
Sbjct: 134 WLMIAFSQATWSLDIGRLFLGVAAGVASYVVPVYIVEIAPKKVRGTFSAINSLVMCASVA 193
Query: 175 LGYVYG 180
+ Y+ G
Sbjct: 194 VTYLLG 199
>At4g21680 peptide transporter - like protein
Length = 589
Score = 30.0 bits (66), Expect = 2.8
Identities = 33/145 (22%), Positives = 64/145 (43%), Gaps = 22/145 (15%)
Query: 282 GITIVCGILGTLGGGLI-LDRMTSTISNAFKLLSGATLLGAIFCLVAFLFKNLSGFIVFF 340
GI +V I+ + G++ + R+ + + +S ++ L + + ++ S VF
Sbjct: 424 GIGLVIAIMAMISAGIVEIHRLKNKEPESATSISSSSTLSIFWQVPQYMLIGASE--VFM 481
Query: 341 SIGELLIFATQAPVNYVSLRCVKPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQ-----D 395
+G+L F +QAP L+ + A+ SI + G+ S+ LV ++ D
Sbjct: 482 YVGQLEFFNSQAPT----------GLKSFASALCMASISL-GNYVSSLLVSIVMKISTTD 530
Query: 396 RINDWRKSAL---CLTSVFFLAAGI 417
++ W L L +FL AG+
Sbjct: 531 DVHGWIPENLNKGHLERFYFLLAGL 555
>At4g33170 putative protein
Length = 990
Score = 29.6 bits (65), Expect = 3.6
Identities = 24/70 (34%), Positives = 28/70 (39%), Gaps = 6/70 (8%)
Query: 104 IGVGLSVWTFAVAGCGISFDFW-SITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWL 162
I G V +A+ G D W S I M V G+ S AA F D+ PV AW
Sbjct: 533 INQGKQVHAYAIKS-GYDLDLWVSSGILDMYVKCGDMS----AAQFAFDSIPVPDDVAWT 587
Query: 163 ATFYMCIPAG 172
CI G
Sbjct: 588 TMISGCIENG 597
>At3g46980 unknown protein
Length = 533
Score = 29.6 bits (65), Expect = 3.6
Identities = 16/67 (23%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Query: 74 GVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFA--VAGCGISFDFWSITICR 131
G++ S+F+ G LI+ +L + ++ G+++W+ A + W++ R
Sbjct: 151 GIVQSSFLWGYLISPIAGGTLVDRYGGKVVMAWGVALWSLATFLTPWAADSSLWALLAAR 210
Query: 132 MLVGVGE 138
+VGV E
Sbjct: 211 AMVGVAE 217
>At3g43790 transporter-like protein
Length = 478
Score = 29.6 bits (65), Expect = 3.6
Identities = 23/67 (34%), Positives = 31/67 (45%), Gaps = 4/67 (5%)
Query: 71 FQDGVLSSAFMVGLLIASPIFASLAKSH--SPFRLIGVGLSVWTFAVAGCGISFDFWSIT 128
F G + S+FM+G + S + LA + P LIG SV F G+S FW
Sbjct: 76 FYAGFVGSSFMIGRALTSIFWGKLADRYGRKPIILIGT-FSVIIFNTL-FGLSTSFWLAI 133
Query: 129 ICRMLVG 135
R L+G
Sbjct: 134 SVRFLLG 140
>At4g05280 putative protein
Length = 1312
Score = 28.9 bits (63), Expect = 6.2
Identities = 15/57 (26%), Positives = 31/57 (54%), Gaps = 2/57 (3%)
Query: 187 KKTISSIETNVSEHGDD--GVLAQDQAFIRGTRSTSKLRNQFTRFSKDMQELLYDQV 241
KK IS ++ + E+ +D G ++ + RS +L +Q + +DM++ + DQ+
Sbjct: 382 KKQISEMDCDKEENAEDCFGEPVPERFIVEMRRSFKELEDQMYQMHEDMKDFVRDQI 438
>At5g54840 SGP1 monomeric G-protein (emb|CAB54517.1)
Length = 288
Score = 28.5 bits (62), Expect = 8.1
Identities = 37/171 (21%), Positives = 63/171 (36%), Gaps = 24/171 (14%)
Query: 96 KSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPV 155
++ S + G+ L TF V G ISF W VG E D+ P+
Sbjct: 125 ENQSFLEMTGLNLMDKTFYVQGVTISFSIWD-------VGGDEKR--------SKDHIPI 169
Query: 156 AQKTAWLATFYMCIPAGTALGYVYGFAPLESKKTISSIETNVSEHGDDGVLAQDQ---AF 212
A K A F + + + L V+G+ K ++I + DD V
Sbjct: 170 ACKDAVAILFMFDLTSRSTLNSVFGWYSQARKWNKTAIPILIGTKFDDFVRLPPNLQWTI 229
Query: 213 IRGTRSTSKLRNQFTRFSKDMQELLYDQVY------IINVLGYISYNFVIG 257
+ R+ +K+ N FS + ++++ + N+ I N +G
Sbjct: 230 VTQARAYAKVMNASLFFSSATHNINVNKIFKFILARLFNLPWKIDRNLTLG 280
>At4g12030 Sodium Bile acid symporter (SBF) like protein
Length = 407
Score = 28.5 bits (62), Expect = 8.1
Identities = 21/74 (28%), Positives = 37/74 (49%), Gaps = 2/74 (2%)
Query: 279 LFGGITIVCGILGT-LGGGLILDRMTSTISNAFK-LLSGATLLGAIFCLVAFLFKNLSGF 336
+FG I+ + ++ T + GL+L+R+ +SNA K L T++ C+ A L N+
Sbjct: 252 VFGMISSILQVVITPIAAGLLLNRLFPRLSNAIKPFLPALTVIDMSCCIGAPLALNIDSI 311
Query: 337 IVFFSIGELLIFAT 350
+ F L + T
Sbjct: 312 LSPFGATILFLVIT 325
>At2g38060 putative Na+-dependent inorganic phosphate cotransporter
Length = 561
Score = 28.5 bits (62), Expect = 8.1
Identities = 12/41 (29%), Positives = 25/41 (60%)
Query: 74 GVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFA 114
GV+ S+F+ G + +S I +L + R++ G+++W+ A
Sbjct: 138 GVVQSSFLWGYIFSSVIGGALVDRYGGKRVLAWGVALWSLA 178
>At2g35740 putative sugar transporter
Length = 580
Score = 28.5 bits (62), Expect = 8.1
Identities = 18/59 (30%), Positives = 30/59 (50%), Gaps = 2/59 (3%)
Query: 125 WSITICRMLVGVGEASFISLAAP-FIDDNAPVAQKTAWLATFYMCIPAGTALGYVYGFA 182
W I + R+LVG G S+ +P +I + +P + A ++T + I G L Y+ A
Sbjct: 120 WVIILGRLLVGFG-VGMASMTSPLYISEMSPARIRGALVSTNGLLITGGQFLSYLINLA 177
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,060,732
Number of Sequences: 26719
Number of extensions: 423914
Number of successful extensions: 1010
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 989
Number of HSP's gapped (non-prelim): 27
length of query: 460
length of database: 11,318,596
effective HSP length: 103
effective length of query: 357
effective length of database: 8,566,539
effective search space: 3058254423
effective search space used: 3058254423
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0381.8


