

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0380.7
(155 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
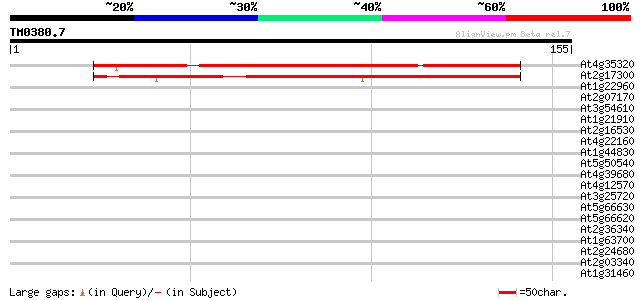
At4g35320 unknown protein 120 2e-28
At2g17300 unknown protein 102 1e-22
At1g22960 putative salt-inducible protein 32 0.10
At2g07170 hypothetical protein 32 0.18
At3g54610 histone acetyltransferase (HAT1) 29 0.88
At1g21910 TINY like protein 29 0.88
At2g16530 unknown protein 29 1.2
At4g22160 putative protein 28 2.0
At1g44830 transcription factor, putative 27 3.4
At5g50540 unknown protein 27 4.4
At4g39680 unknown protein 27 4.4
At4g12570 polyubiquitin-like protein 27 4.4
At3g25720 hypothetical protein 27 5.7
At5g66630 unknown protein 26 7.5
At5g66620 putative protein 26 7.5
At2g36340 hypothetical protein 26 7.5
At1g63700 putative protein kinase 26 7.5
At2g24680 unknown protein 26 9.8
At2g03340 putative WRKY DNA-binding protein 26 9.8
At1g31460 unknown protein 26 9.8
>At4g35320 unknown protein
Length = 147
Score = 120 bits (302), Expect = 2e-28
Identities = 62/122 (50%), Positives = 84/122 (68%), Gaps = 8/122 (6%)
Query: 24 LPVSH----SRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGT 79
LPVS + + D +R+RLSS + S +A+ + FPRSKS+S+MG+ AG+
Sbjct: 16 LPVSQRPNAAATADDEPGLIRRRLSSLSLN---LSNQPAAIAARFPRSKSVSAMGEQAGS 72
Query: 80 SIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPP 139
S+K WW+W W+WILSRKPIF RDLELN E K + GS NRG+ HVF+KLRS+IR F+ P
Sbjct: 73 SVKEWWEWGWSWILSRKPIFIRDLELNKDEAKSI-GSQNRGSIMHVFFKLRSQIRNFMGP 131
Query: 140 TN 141
++
Sbjct: 132 SS 133
>At2g17300 unknown protein
Length = 139
Score = 102 bits (253), Expect = 1e-22
Identities = 56/120 (46%), Positives = 78/120 (64%), Gaps = 11/120 (9%)
Query: 24 LPVSHSRSVDSSEHGL-RKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIK 82
LP+S S + E GL R+RLSS + + S + RSKS+S MG+ G+S+K
Sbjct: 17 LPIS---STTTDEPGLLRRRLSSLSLNLSRNQPSTVS------RSKSVSDMGEQGGSSVK 67
Query: 83 NWWDWTWAWILSRK-PIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPPTN 141
WW+W+W+WIL +K PIF DLE+N ETK G+ RG++ HVF+KLRSEIRR + P++
Sbjct: 68 EWWEWSWSWILLKKLPIFFTDLEVNKNETKSSLGNQQRGSFIHVFFKLRSEIRRLLRPSS 127
>At1g22960 putative salt-inducible protein
Length = 718
Score = 32.3 bits (72), Expect = 0.10
Identities = 25/68 (36%), Positives = 36/68 (52%), Gaps = 17/68 (25%)
Query: 10 RSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTP--AAASASASAVWSSFPRS 67
RSFF+I TT+NN + L + L F+T P AA+S+S+S+ S+ +
Sbjct: 13 RSFFSISTTNNN---------------NNLSRFLFRFSTLPHCAASSSSSSSNLESYYAN 57
Query: 68 KSLSSMGD 75
LSS GD
Sbjct: 58 LILSSHGD 65
>At2g07170 hypothetical protein
Length = 893
Score = 31.6 bits (70), Expect = 0.18
Identities = 14/43 (32%), Positives = 23/43 (52%), Gaps = 4/43 (9%)
Query: 27 SHSRSVDSSEHGLRKRLSSFN----TTPAAASASASAVWSSFP 65
SH++ + ++HGL R S+N T P ++S +W FP
Sbjct: 845 SHTKLQEKADHGLDLRFESYNQFTKTEPEGNKPASSQLWKQFP 887
>At3g54610 histone acetyltransferase (HAT1)
Length = 568
Score = 29.3 bits (64), Expect = 0.88
Identities = 26/75 (34%), Positives = 34/75 (44%), Gaps = 5/75 (6%)
Query: 9 HRSFFTIPTTSNNKQLPV---SHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFP 65
H S S + Q P S S SV SS H +++L++ AAAS + SSFP
Sbjct: 4 HSSHLNAANRSRSSQTPSPSHSASASVTSSLH--KRKLAATTAANAAASEDHAPPSSSFP 61
Query: 66 RSKSLSSMGDYAGTS 80
S + D A TS
Sbjct: 62 PSSFSADTRDGALTS 76
>At1g21910 TINY like protein
Length = 230
Score = 29.3 bits (64), Expect = 0.88
Identities = 26/127 (20%), Positives = 52/127 (40%), Gaps = 20/127 (15%)
Query: 26 VSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNWW 85
V R + +S LSS ++ +++S+S+S+ SS S + Y G +++W
Sbjct: 2 VKQERKIQTSSTKKEMPLSSSPSSSSSSSSSSSS--SSCKNKNKKSKIKKYKGVRMRSWG 59
Query: 86 DWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLR------SEIRRFIPP 139
W ++ +Q+T++ GS++ Y + + P
Sbjct: 60 SW------------VSEIRAPNQKTRIWLGSYSTAEAAARAYDVALLCLKGPQANLNFPT 107
Query: 140 TNSHHHL 146
++S HHL
Sbjct: 108 SSSSHHL 114
>At2g16530 unknown protein
Length = 343
Score = 28.9 bits (63), Expect = 1.2
Identities = 17/60 (28%), Positives = 25/60 (41%), Gaps = 14/60 (23%)
Query: 88 TWAWILSRKPIFARDLELNDQETKLLGGS--------------HNRGTWRHVFYKLRSEI 133
TW + P+ + + +L+D ++L GGS H WR VF L EI
Sbjct: 84 TWMYACKMAPLSSEEFQLSDIASRLAGGSDVFSVHKSNMTPVEHRFKVWRAVFLLLLMEI 143
>At4g22160 putative protein
Length = 401
Score = 28.1 bits (61), Expect = 2.0
Identities = 16/46 (34%), Positives = 23/46 (49%)
Query: 27 SHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSS 72
S S + +S G +SS +TT A SA +F R+KS+ S
Sbjct: 31 SDSETESNSSSGTESNMSSDSTTSAGVSAMIRVFLVNFQRAKSVCS 76
>At1g44830 transcription factor, putative
Length = 211
Score = 27.3 bits (59), Expect = 3.4
Identities = 13/45 (28%), Positives = 21/45 (45%)
Query: 43 LSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNWWDW 87
+ + TP S+ +S+ SS S S M Y G +++W W
Sbjct: 2 VKTLQKTPKRMSSPSSSSSSSSSTSSSSIRMKKYKGVRMRSWGSW 46
>At5g50540 unknown protein
Length = 113
Score = 26.9 bits (58), Expect = 4.4
Identities = 13/27 (48%), Positives = 17/27 (62%), Gaps = 1/27 (3%)
Query: 62 SSFPRSKSLSSMGDYAGTSIKNW-WDW 87
SSFPRS L++ D G S+ +W DW
Sbjct: 4 SSFPRSVILTARSDTKGKSVFSWLTDW 30
>At4g39680 unknown protein
Length = 633
Score = 26.9 bits (58), Expect = 4.4
Identities = 15/52 (28%), Positives = 28/52 (53%)
Query: 19 SNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSL 70
S+ ++LP + +V ++E R+R +S + A + SA ++ PRS L
Sbjct: 376 SDKRKLPANDQEAVGNNEPAKRRRWNSNSIKVPEAQITNSATPTTTPRSTGL 427
>At4g12570 polyubiquitin-like protein
Length = 873
Score = 26.9 bits (58), Expect = 4.4
Identities = 19/73 (26%), Positives = 34/73 (46%), Gaps = 1/73 (1%)
Query: 9 HRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAV-WSSFPRS 67
+RS+ + T N ++ S V +E + + AASA + + W S S
Sbjct: 16 NRSYSAVAGTDNKRKRDEDSSDYVGVAESLEMLKKQEIDADHMAASAQQTLISWRSGENS 75
Query: 68 KSLSSMGDYAGTS 80
+SLSS G+ + ++
Sbjct: 76 RSLSSSGECSSSN 88
>At3g25720 hypothetical protein
Length = 282
Score = 26.6 bits (57), Expect = 5.7
Identities = 14/60 (23%), Positives = 24/60 (39%), Gaps = 11/60 (18%)
Query: 55 ASASAVWSSFPRSKSLSSMGDYAGTSIKNWW--------DWTWAWILSRKPIFARDLELN 106
+ ++W + R L +GD S+ +W W W +L +P+ R L N
Sbjct: 49 SGGGSLWVDWHRYHHLQGLGD---ASVSKFWTSQELVSDSWNWKCLLRLRPLAERFLRCN 105
>At5g66630 unknown protein
Length = 702
Score = 26.2 bits (56), Expect = 7.5
Identities = 12/23 (52%), Positives = 14/23 (60%)
Query: 44 SSFNTTPAAASASASAVWSSFPR 66
SS + TP AASAS WS F +
Sbjct: 635 SSSSRTPPAASASKKGEWSDFDK 657
>At5g66620 putative protein
Length = 644
Score = 26.2 bits (56), Expect = 7.5
Identities = 12/23 (52%), Positives = 14/23 (60%)
Query: 44 SSFNTTPAAASASASAVWSSFPR 66
SS + TP AASAS WS F +
Sbjct: 577 SSSSRTPPAASASKKGEWSDFDK 599
>At2g36340 hypothetical protein
Length = 414
Score = 26.2 bits (56), Expect = 7.5
Identities = 21/80 (26%), Positives = 35/80 (43%), Gaps = 4/80 (5%)
Query: 17 TTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAV-WSSFPRSKSLSSMGD 75
T + N+++ V S R ++ TTP + S+SAS + WS L + D
Sbjct: 18 TLTRNREVEVEAMLSRRKQLRTTTTRTTTTRTTPLSLSSSASKMNWSKNDELVILGGIVD 77
Query: 76 YAG---TSIKNWWDWTWAWI 92
Y S ++ WD + +I
Sbjct: 78 YENETKLSYRSDWDALYRYI 97
>At1g63700 putative protein kinase
Length = 883
Score = 26.2 bits (56), Expect = 7.5
Identities = 36/135 (26%), Positives = 49/135 (35%), Gaps = 37/135 (27%)
Query: 20 NNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGT 79
N + PV+ S S D S H + F P + + + S+ PR + L GT
Sbjct: 168 NGNRTPVNIS-SRDQSMHSNKNSAEMFKPVP-----NKNRILSASPRRRPL-------GT 214
Query: 80 SIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPP 139
+KN I RDL L LL RS +R FIP
Sbjct: 215 HVKNL------------QIPQRDLVLCSAPDSLLSSPS------------RSPMRSFIPD 250
Query: 140 TNSHHHLPRTTPQAD 154
S+H L + P +D
Sbjct: 251 QVSNHGLLISKPYSD 265
>At2g24680 unknown protein
Length = 851
Score = 25.8 bits (55), Expect = 9.8
Identities = 19/96 (19%), Positives = 39/96 (39%), Gaps = 6/96 (6%)
Query: 7 QQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPR 66
++ S P+ N K ++ + S ++ R + TP A + S F +
Sbjct: 710 EKESSICAEPSRGNKKWKATNNRKERRDSSSAIQNRYVTLTLTPEDVRACTLILPSQFMK 769
Query: 67 SKSLSSMG--DYAGTSIKNWWDWTWAWILSRKPIFA 100
+ ++ +G G + K W +A++LS+ A
Sbjct: 770 ANGINKLGKKTLLGQNRKKW----FAYLLSKSGFVA 801
>At2g03340 putative WRKY DNA-binding protein
Length = 513
Score = 25.8 bits (55), Expect = 9.8
Identities = 13/37 (35%), Positives = 23/37 (62%)
Query: 43 LSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGT 79
L+ +PAAA+ +A+AV ++ +SS+GD G+
Sbjct: 66 LAGAMASPAAAAVAAAAVVATAHHQTPVSSVGDGGGS 102
>At1g31460 unknown protein
Length = 301
Score = 25.8 bits (55), Expect = 9.8
Identities = 11/25 (44%), Positives = 19/25 (76%)
Query: 17 TTSNNKQLPVSHSRSVDSSEHGLRK 41
++S N+Q P+SHS+SV+ + G R+
Sbjct: 70 SSSMNRQKPLSHSKSVNVNFSGNRR 94
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.127 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,682,961
Number of Sequences: 26719
Number of extensions: 139900
Number of successful extensions: 508
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 493
Number of HSP's gapped (non-prelim): 21
length of query: 155
length of database: 11,318,596
effective HSP length: 91
effective length of query: 64
effective length of database: 8,887,167
effective search space: 568778688
effective search space used: 568778688
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0380.7


