

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0378b.2
(736 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
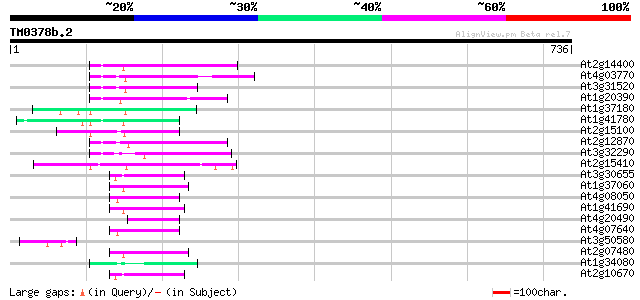
Score E
Sequences producing significant alignments: (bits) Value
At2g14400 putative retroelement pol polyprotein 79 7e-15
At4g03770 hypothetical protein 74 3e-13
At3g31520 hypothetical protein 70 4e-12
At1g20390 hypothetical protein 70 4e-12
At1g37180 hypothetical protein 69 7e-12
At1g41780 hypothetical protein 66 6e-11
At2g15100 putative retroelement pol polyprotein 66 8e-11
At2g12870 putative retroelement pol polyprotein 65 1e-10
At3g32290 hypothetical protein 57 5e-08
At2g15410 putative retroelement pol polyprotein 57 5e-08
At3g30655 hypothetical protein, 3' partial 51 3e-06
At1g37060 Athila retroelment ORF 1, putative 50 3e-06
At4g08050 50 6e-06
At1g41690 hypothetical protein 49 1e-05
At4g20490 putative protein 47 5e-05
At4g07640 putative athila transposon protein 47 5e-05
At3g50580 proline-rich protein 46 7e-05
At2g07480 F9A16.15 45 1e-04
At1g34080 putative protein 45 1e-04
At2g10670 pseudogene 45 1e-04
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 79.3 bits (194), Expect = 7e-15
Identities = 53/201 (26%), Positives = 100/201 (49%), Gaps = 7/201 (3%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVII----AASDAVKCRMFPS 160
PF+ + S+ I D+ K L L+SY+G DPK +L F + A D C++F
Sbjct: 144 PFTPQITSLRIRDSRK-LNLESYNGLEDPKGYLAAFLIAAGRVDLNEADEDVRYCKLFSE 202
Query: 161 TFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYM 220
A+ WFT L GSISNF + S FL Q+S ++ ++ DL+N+ Q E+L+ ++
Sbjct: 203 NLCGQALMWFTQLEPGSISNFNELSVVFLKQYSILMDKSISDTDLWNLSQGPNETLRAFI 262
Query: 221 ARYSAASVKVEDEEPRACALAFKNGL-LPGGLNSKLTRKPARSMEEMRARASTYI-LDEE 278
++ K+ ++ A + GL L ++++ RA+ ++ +++E
Sbjct: 263 TKFKYVLSKLSRISQQSALSALRKGLWYDSRFKEDLILHKLDTIQDALFRANNWMEVEDE 322
Query: 279 DDAFKRKRAKLEKGDTSPKQR 299
++F ++ + + T P ++
Sbjct: 323 KESFAKRDKQAKPAVTFPPKK 343
>At4g03770 hypothetical protein
Length = 464
Score = 73.9 bits (180), Expect = 3e-13
Identities = 62/227 (27%), Positives = 99/227 (43%), Gaps = 32/227 (14%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAAS-------DAVKCRM 157
PF+ + + I D K L ++ ++G SDPK HL F ++ +A + DA C +
Sbjct: 218 PFTHMISNAIISDPGK-LRIEYFNGSSDPKGHLKLF---IISVARAKFRPEERDAGLCHL 273
Query: 158 FPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLK 217
F K A+ WFT L S+ +F++ S+ FL Q+S + + DL+++ QQ E L+
Sbjct: 274 FVEHLKGPALNWFTRLKGNSVDSFQELSTLFLKQYSVLIDPSTSDADLWSLSQQPNEPLR 333
Query: 218 EYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDE 277
+++A++ + KVE A K L +RAR S L
Sbjct: 334 DFLAKFRSTLAKVEGINDLAALSTLKKALC------------------LRARLSAKFLAR 375
Query: 278 E-DDAFKRKRAKLEKGDTSPKQ--R*KEEFQIGGKAEGKEQGAFTRE 321
E K + EKG T E+ +EG ++G RE
Sbjct: 376 EIGGGLTIKDLEAEKGKTEQVNAVANPEQAAPAANSEGPKRGRGNRE 422
>At3g31520 hypothetical protein
Length = 528
Score = 70.1 bits (170), Expect = 4e-12
Identities = 45/149 (30%), Positives = 76/149 (50%), Gaps = 11/149 (7%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAAS-------DAVKCRM 157
PF+ + + I D K L ++ ++G SDPK HL F ++ +A + DA C +
Sbjct: 226 PFTHRISNAIISDPGK-LRIEYFNGSSDPKRHLKSF---IISVARTKFRPEERDAGLCHL 281
Query: 158 FPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLK 217
F K A+ WF+ L S+ +F++ S+ FL Q+S + + DL+++ QQ E L+
Sbjct: 282 FVEHLKGPALCWFSRLEGNSMDSFQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLR 341
Query: 218 EYMARYSAASVKVEDEEPRACALAFKNGL 246
+++A+ + KVE A A K L
Sbjct: 342 DFLAKLRSTLAKVEGINDVAALSALKKAL 370
>At1g20390 hypothetical protein
Length = 1791
Score = 70.1 bits (170), Expect = 4e-12
Identities = 55/189 (29%), Positives = 92/189 (48%), Gaps = 12/189 (6%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKM----VIIAASDAVKCRMFPS 160
PF+ + +V+I K + L+SY+G +DPK+ L FN + + I DA +C++F
Sbjct: 156 PFTRRITNVSIRGAQK-IKLESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIE 214
Query: 161 TFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYM 220
A WF+ L SI +F +S FL ++ + DL++I Q ESL+ ++
Sbjct: 215 HLTGPAHNWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFV 274
Query: 221 ARYS--AASVKVEDEEPRACALAFKNGL-LPGGLNSKLTRKPARSMEEMRARASTYI-LD 276
R+ ++ V DE A +A +N + +T ++E+ RAS +I L+
Sbjct: 275 DRFKLVVTNITVPDE---AAIVALRNAVWYDSRFRDDITLHAPSTLEDALHRASRFIELE 331
Query: 277 EEDDAFKRK 285
EE RK
Sbjct: 332 EEKLILARK 340
>At1g37180 hypothetical protein
Length = 661
Score = 69.3 bits (168), Expect = 7e-12
Identities = 56/240 (23%), Positives = 96/240 (39%), Gaps = 25/240 (10%)
Query: 30 PPPSPSQVGSLERSPENSPAPEQQPAVTQEQWRHLM--------RNIGNIQQRNEHLQAQ 81
PPPS + ++P S +P AV+ E + I + +Q + ++
Sbjct: 203 PPPSVEVGTTSTQNPTTSSSPAGFAAVSNETIARTLVPLDPAIRLPISDPRQEPSAVNSE 262
Query: 82 LDFYRRE---------QRDDGSREADSVAEFR---PFSEDVESVAIPDNMKTLVLDSYSG 129
L + + Q + E + V E PF+ + + I + + L Y+G
Sbjct: 263 LQSLQDQIWAMNAKVHQATTSAPEVEKVIEATRRTPFTPRISKLRIRE-FRDFKLPVYNG 321
Query: 130 DSDPKDHLLYFNTKMVIIAAS----DAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFS 185
DP +HL F + + DA C++F A+ WFT L GSI NF+ S
Sbjct: 322 KGDPNEHLTSFQVIVRRVPLEPYEEDAGLCKLFSENLSGPALTWFTQLEEGSIDNFKQLS 381
Query: 186 SKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNG 245
+ F+ Q+ +T L+N+ Q + L+ Y+ + V++ A KNG
Sbjct: 382 TAFIKQYGYFIKSDITEAHLWNLSQSADDPLRTYIIVFKEIMVQIPSLSDSTAQFALKNG 441
>At1g41780 hypothetical protein
Length = 246
Score = 66.2 bits (160), Expect = 6e-11
Identities = 58/244 (23%), Positives = 97/244 (38%), Gaps = 34/244 (13%)
Query: 10 PVRTTGVQSPSPPPSPPVPPPPPSPSQVGSLERSPENSPAPEQQPAVTQEQWRHLMRNIG 69
P+ T V+ P PP+ P + L + +++ V Q +H
Sbjct: 7 PLTTVDVEGN---PVPPIQVNAPKVNAPAVLAKLRSMMAQLQKKLEVATSQGQHSAEPEQ 63
Query: 70 NIQQRNEHLQAQLDFYRREQRDDG----SREADSVAEFR--------------------- 104
N+ R E Q+ R E + S + DS R
Sbjct: 64 NMTNRVEEADPQVPIPRAESDEADEIKVSDDEDSEENIRWAEDAVSSVPNIDCILEVYHN 123
Query: 105 -PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIA----ASDAVKCRMFP 159
PF+ + + I D K L ++ +SG SDPK HL F + SDA C +F
Sbjct: 124 TPFTRKISNAIISDPEK-LRIEYFSGSSDPKGHLKSFKISVARAKFKPEESDAGLCHLFV 182
Query: 160 STFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEY 219
K + WF+ L S+ +F++ S+ FL Q+S + + DL+++ QQ E L ++
Sbjct: 183 EHLKGPVLDWFSRLEGNSVDSFQELSTLFLKQYSVLIDPGTSDVDLWSLSQQPNEPLSDF 242
Query: 220 MARY 223
+ ++
Sbjct: 243 LTKF 246
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 65.9 bits (159), Expect = 8e-11
Identities = 49/174 (28%), Positives = 77/174 (44%), Gaps = 18/174 (10%)
Query: 62 RHLMRNIGNIQQRNEHLQAQLDFYRRE--QRDDGSREADSVAEFR---PFSEDVESVAIP 116
RH +R + + ++LQ Q+ + Q + E + V E PF+ + + I
Sbjct: 109 RHRLRILSPLNSELQNLQDQIWAMNAKVHQATTSAPEVEKVIEATRRTPFTPRISKLRIR 168
Query: 117 DNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAA--------SDAVKCRMFPSTFKSTAMA 168
+ + L Y+G D K+HL F +IA DA C++F A+
Sbjct: 169 E-FRDFKLPVYNGKGDLKEHLTSFQ----VIAGRVPLEPHEEDAGLCKLFSENLFGLALT 223
Query: 169 WFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMAR 222
WFT L GSI NF+ S+ F+ Q+ N +T L+N Q E L+ Y+ R
Sbjct: 224 WFTQLEEGSIDNFKQLSTAFIKQYEYFINSDITEAHLWNFSQSADEPLRTYIYR 277
>At2g12870 putative retroelement pol polyprotein
Length = 411
Score = 65.1 bits (157), Expect = 1e-10
Identities = 53/191 (27%), Positives = 82/191 (42%), Gaps = 14/191 (7%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK--------CR 156
PF+ + S I D K L +D ++G S PK HL F +I AA D K C
Sbjct: 61 PFTRKISSAIISDPGK-LRIDYFNGSSYPKGHLKSF----IIYAARDKFKPEERDAGLCH 115
Query: 157 MFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESL 216
+F K + F L + +F + S+ FL QFS + +DL+++ QQ E L
Sbjct: 116 LFVEHLKGPVLVLFLRLEGNFVDSFEELSTLFLKQFSVLIDPGTLDSDLWSLSQQPNEPL 175
Query: 217 KEYMARYSAASVKVEDEEPRACALAFKNGL-LPGGLNSKLTRKPARSMEEMRARASTYIL 275
+ + ++ + VE A A K L +L +++ + RAS Y+
Sbjct: 176 MDLLTKFRSTLASVEGIIDVAALSALKEALWYKSEFRKELNLSKPQTICDALHRASDYVS 235
Query: 276 DEEDDAFKRKR 286
EE+ KR
Sbjct: 236 HEEEMELLSKR 246
>At3g32290 hypothetical protein
Length = 612
Score = 56.6 bits (135), Expect = 5e-08
Identities = 52/191 (27%), Positives = 84/191 (43%), Gaps = 26/191 (13%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
PF+ + + I D K L ++ ++G SDPK HL K II S
Sbjct: 164 PFTHMISNAIISDPGK-LRIEYFNGSSDPKGHL-----KSFII----------------S 201
Query: 165 TAMAWFTTLPR---GSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMA 221
A A F R S+ +F++ S+ FL Q+S + + DL+++ QQ E L++++A
Sbjct: 202 VARAKFRQEERDAGNSVDSFQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLA 261
Query: 222 RYSAASVKVEDEEPRACALAFKNGL-LPGGLNSKLTRKPARSMEEMRARASTYILDEEDD 280
++ + KVE A A K L +L ++ + RAS Y+ EE+
Sbjct: 262 KFRSTLAKVEGINDVAALSALKKALWYKSEFRKELNLSKPLTIRDALHRASDYVSHEEEM 321
Query: 281 AFKRKRAKLEK 291
KR + K
Sbjct: 322 ELVAKRHEPSK 332
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 56.6 bits (135), Expect = 5e-08
Identities = 73/291 (25%), Positives = 120/291 (41%), Gaps = 28/291 (9%)
Query: 32 PSPSQVGSL-ERSPENSPAPEQQPAVTQEQWRHLMRNIGNIQQRNEHLQAQL-DFYRREQ 89
P+ VGS+ E + EN+P + +P V R + G + + L+ L D +
Sbjct: 158 PASIVVGSIDEHTRENAPNRDNRPNVDTRARRTEDDDTGARDRELQQLRTTLKDTSSKMH 217
Query: 90 RDDGSR-EADSVAEFR---PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMV 145
R S E D V E PF+ + V I ++ + L +Y G DP+ L + +
Sbjct: 218 RAVSSTPEVDKVLEETQRSPFTSCISDVRIR-HISKIKLANYEGLVDPRPFLTSVSIAIG 276
Query: 146 IIAASD----AVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVT 201
SD A C++F A+ WF+ L SI + ++ FL + + +
Sbjct: 277 RAHFSDEDRDAGSCQLFVKHLSGAALTWFSRLEANSIDSVHALTTSFLKNYGVFMEKGAS 336
Query: 202 INDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTR--KP 259
DL+ + Q ESL+ ++ R+ V + A A A +N L G +S+ T KP
Sbjct: 337 NVDLWTMAQTAKESLRSFIGRFKEIVTSVATPDDAAIA-ALRNALWHGD-SSRCTTLCKP 394
Query: 260 ARSMEEMRA-------RASTYILDEEDDAFKRKRAKLEK------GDTSPK 297
+A + + + DA K KL+K GDT+P+
Sbjct: 395 FHRQGSAKAPIAKEKPKEGHHESRQHYDADYAKEEKLKKGTSYYVGDTAPR 445
>At3g30655 hypothetical protein, 3' partial
Length = 660
Score = 50.8 bits (120), Expect = 3e-06
Identities = 33/103 (32%), Positives = 48/103 (46%), Gaps = 6/103 (5%)
Query: 132 DPKDHL-----LYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSS 186
DP DHL LY TK+ ++ D K R+FP + A W TLP+ SI+ + D
Sbjct: 61 DPLDHLDEFERLYGLTKINGVS-EDGFKLRLFPFSLGDKAHLWEKTLPQNSITAWDDCKK 119
Query: 187 KFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVK 229
FL +F +N N++ Q++ ES E R+ K
Sbjct: 120 AFLAKFFSNSRTARLRNEISGFTQKQNESFGEAWERFKGYQTK 162
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 50.4 bits (119), Expect = 3e-06
Identities = 30/107 (28%), Positives = 46/107 (42%), Gaps = 4/107 (3%)
Query: 132 DPKDHLLYFNTKMVII----AASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSK 187
DP DHL F+ + + D K R+FP + A W +LP+GSI+++ D
Sbjct: 61 DPLDHLDEFDRLCSLTKINRVSEDGFKLRLFPFSLGDKAHQWEKSLPQGSITSWNDCKKA 120
Query: 188 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEE 234
FL +F +N ND+ Q E+ E R+ + E
Sbjct: 121 FLAKFFSNSRTARLRNDISGFTQTNNETFYEAWERFKGYQTQCPHHE 167
>At4g08050
Length = 1428
Score = 49.7 bits (117), Expect = 6e-06
Identities = 30/96 (31%), Positives = 43/96 (44%), Gaps = 4/96 (4%)
Query: 132 DPKDHLLYF----NTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSK 187
DP DHL F N + + D K R+FP + A W L SI+ + D+
Sbjct: 61 DPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLSHDSITTWDDYKKA 120
Query: 188 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARY 223
FL +F +N N++Y Q+ GES E R+
Sbjct: 121 FLSKFFSNARTARLRNEIYGFSQKTGESFCEAWERF 156
>At1g41690 hypothetical protein
Length = 371
Score = 48.9 bits (115), Expect = 1e-05
Identities = 30/102 (29%), Positives = 44/102 (42%), Gaps = 4/102 (3%)
Query: 132 DPKDHLLYFNTKMVII----AASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSK 187
DP DHL F + + D K R+FP + A W TLP+GSI+ + D
Sbjct: 61 DPLDHLDEFERLCSLTKINGVSEDGFKLRLFPFSLGDKAHLWEKTLPQGSITTWDDCKKA 120
Query: 188 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVK 229
FL +F + N++ Q++ ES E R+ K
Sbjct: 121 FLAKFFSYSRTARLRNEISGFTQKQSESFCEAWERFKGYQTK 162
>At4g20490 putative protein
Length = 853
Score = 46.6 bits (109), Expect = 5e-05
Identities = 21/69 (30%), Positives = 37/69 (53%)
Query: 155 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGE 214
C++F A+ F+ L SI NF S+ FL Q+ + +DL+++ Q+ GE
Sbjct: 222 CQLFVEGLTGNALTCFSRLEANSIDNFTQLSTAFLKQYRVFIQPGASSSDLWSMTQENGE 281
Query: 215 SLKEYMARY 223
+L +Y+ R+
Sbjct: 282 TLNDYLGRF 290
>At4g07640 putative athila transposon protein
Length = 866
Score = 46.6 bits (109), Expect = 5e-05
Identities = 30/96 (31%), Positives = 42/96 (43%), Gaps = 4/96 (4%)
Query: 132 DPKDHLLYF----NTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSK 187
DP DHL F N + ++D K R+FP + A W LP SI + D
Sbjct: 61 DPLDHLDEFDRLCNLTKINGVSADGFKLRLFPFSLGDKAHIWEKNLPHDSIITWDDCKKA 120
Query: 188 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARY 223
FL +F +N N++ Q+ GES E R+
Sbjct: 121 FLSKFFSNARTARLRNEISGFSQKTGESFCEAWERF 156
>At3g50580 proline-rich protein
Length = 265
Score = 46.2 bits (108), Expect = 7e-05
Identities = 30/88 (34%), Positives = 42/88 (47%), Gaps = 16/88 (18%)
Query: 14 TGVQSPSPPPSPPVPPPPP--SPSQVGSLERSPENSP---------APEQQPAVTQEQWR 62
T +SPSPPP+P +PPP P SPS +P+ SP +P P Q W
Sbjct: 121 TPKKSPSPPPTPSLPPPAPKKSPSTPSLPPPTPKKSPPPPPSHHSSSPSNPPHHQQNPWE 180
Query: 63 HLMR---NIGNIQQRNEHLQAQLDFYRR 87
H+ R N+G + +Q ++ FY R
Sbjct: 181 HIERCMINMGPVGMC--RMQMEVSFYTR 206
Score = 35.0 bits (79), Expect = 0.15
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 8/56 (14%)
Query: 9 SPVRTTGVQSPSPPPSPPVPPPPPSPSQVGS--------LERSPENSPAPEQQPAV 56
+P+ +T PPP+P PPPP+P + S +P+ SP+P P++
Sbjct: 79 TPIPSTPSTPSPPPPAPKKSPPPPTPKKSPSPPSLTPFVPHPTPKKSPSPPPTPSL 134
>At2g07480 F9A16.15
Length = 1012
Score = 45.4 bits (106), Expect = 1e-04
Identities = 28/107 (26%), Positives = 46/107 (42%), Gaps = 4/107 (3%)
Query: 132 DPKDHLLYFNTKMVII----AASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSK 187
DP DHL F+ + + ++ K R+FP + A W TLP SI + D
Sbjct: 56 DPLDHLDNFDRLCSLTKINGVSEESFKLRLFPFSLGDKAHLWEKTLPVKSIDTWDDCKKA 115
Query: 188 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEE 234
FL +F +N + +++ Q+ ES E R+ + + E
Sbjct: 116 FLAKFFSNSRKARLRSEISGFNQKNSESFSEAWERFKGYTTQCPHHE 162
>At1g34080 putative protein
Length = 811
Score = 45.4 bits (106), Expect = 1e-04
Identities = 37/142 (26%), Positives = 56/142 (39%), Gaps = 29/142 (20%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
PF+ + + I + + L Y+G DPK++L F +IA +
Sbjct: 193 PFTLRISKLRIRE-FQDFKLPGYNGKGDPKEYLTSFQ----VIAGRE------------- 234
Query: 165 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYS 224
GSI NF+ S+ F+ Q+ VT L+N+ Q E L+ Y +
Sbjct: 235 -----------GSIDNFKQLSTAFIKQYEYFIKSDVTEAQLWNLSQAADEPLRTYNTAFK 283
Query: 225 AASVKVEDEEPRACALAFKNGL 246
V+V A A KNGL
Sbjct: 284 EIMVQVPGLSDSAAQSALKNGL 305
>At2g10670 pseudogene
Length = 929
Score = 45.1 bits (105), Expect = 1e-04
Identities = 31/103 (30%), Positives = 46/103 (44%), Gaps = 6/103 (5%)
Query: 132 DPKDHL-----LYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSS 186
DP DHL L TK+ ++ D K R+FP + A W TLP+ SI+ + D
Sbjct: 93 DPLDHLDELERLCGLTKINGVS-EDGFKLRLFPFSLGDKAHLWEKTLPKNSITTWDDCKM 151
Query: 187 KFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVK 229
FL +F +N N++ ++ ES E R+ K
Sbjct: 152 AFLAKFFSNSRTARLRNEISGFTLKQNESFCEAWERFKGYQTK 194
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.336 0.147 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,330,338
Number of Sequences: 26719
Number of extensions: 716336
Number of successful extensions: 15696
Number of sequences better than 10.0: 318
Number of HSP's better than 10.0 without gapping: 213
Number of HSP's successfully gapped in prelim test: 119
Number of HSP's that attempted gapping in prelim test: 6943
Number of HSP's gapped (non-prelim): 3025
length of query: 736
length of database: 11,318,596
effective HSP length: 107
effective length of query: 629
effective length of database: 8,459,663
effective search space: 5321128027
effective search space used: 5321128027
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0378b.2


