

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0378b.13
(282 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
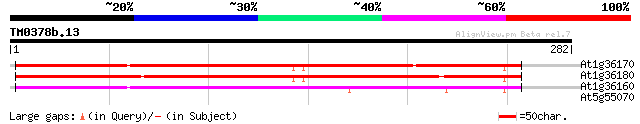
At1g36170 acetyl-CoA carboxylase, putative, 5' partial 200 8e-52
At1g36180 acetyl-CoA carboxylase, putative 199 1e-51
At1g36160 hypothetical protein, 3' partial 194 6e-50
At5g55070 2-oxoglutarate dehydrogenase E2 subunit 29 3.2
>At1g36170 acetyl-CoA carboxylase, putative, 5' partial
Length = 1865
Score = 200 bits (508), Expect = 8e-52
Identities = 139/262 (53%), Positives = 161/262 (61%), Gaps = 10/262 (3%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 258 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 316
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 317 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 376
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + + + + + SPE PFLQW
Sbjct: 377 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVVQDLLNCLDSPELPFLQWQ 436
Query: 180 ECFAVVATRLCKVLATHLPIDLKN*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEKG 239
ECFAV+ATRL K L L + F S + AKLLKG LEAH+SSC +KE+G
Sbjct: 437 ECFAVLATRLPKNLRNMLESKYRE-FESISRNSLTTDFPAKLLKGILEAHLSSCDEKERG 495
Query: 240 AQERLK*P----D*SCEGGRET 257
A ERL P S EGGRE+
Sbjct: 496 ALERLIEPLMSLAKSYEGGRES 517
>At1g36180 acetyl-CoA carboxylase, putative
Length = 2359
Score = 199 bits (506), Expect = 1e-51
Identities = 138/263 (52%), Positives = 163/263 (61%), Gaps = 12/263 (4%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY +EEA GT LLIDGR LLQNDHDP KL L +LVS S
Sbjct: 753 LMQLDGKSHVIYAKEEATGTRLLIDGRTCLLQNDHDPSKLMAETPCKLLRYLVSDNSSID 812
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPL+SPAS +I FK+SEGQ MQA ELIA+LDLDDPSAVRKA+PF
Sbjct: 813 TDT-PYAEVEVMKMCMPLISPASGVIHFKLSEGQAMQAGELIAKLDLDDPSAVRKAKPFR 871
Query: 124 SWALLLQFLVKFNRNVPQ--A*ILH--R*FLLAMSTNMMR*LCKV*SIVYSPEFPFLQW* 179
L + V Q A L+ R L + L + + + SPE PFLQW
Sbjct: 872 GSFPRLGLPTAISGKVHQRCAATLNAARMILAGYDHKVDEVLQDLLNCLDSPELPFLQWQ 931
Query: 180 ECFAVVATRLCKVLATHLPIDLKN-*FPSRGFQAPKFLTFAKLLKGNLEAHISSCPDKEK 238
ECFAV+ATRL K L L + K S+ P F AKLLKG LEAH+SSC +KE+
Sbjct: 932 ECFAVLATRLPKDLRNMLELKYKEFEIISKTSLTPDFP--AKLLKGILEAHLSSCDEKER 989
Query: 239 GAQERLK*P----D*SCEGGRET 257
G+ ERL P S EGGRE+
Sbjct: 990 GSLERLIEPLMSLVKSYEGGRES 1012
>At1g36160 hypothetical protein, 3' partial
Length = 2252
Score = 194 bits (492), Expect = 6e-50
Identities = 137/269 (50%), Positives = 161/269 (58%), Gaps = 16/269 (5%)
Query: 4 LLQLDGNNHVIYLEEEAAGTCLLIDGRAFLLQNDHDP*KLCGFYLFLLWLWLVSKPLSFA 63
L+QLDG +HVIY EEEAAGT LLIDGR LLQNDHDP KL L +L+S S
Sbjct: 640 LMQLDGKSHVIYAEEEAAGTRLLIDGRTCLLQNDHDPSKLMAETPCKLMRYLISDN-SNI 698
Query: 64 E*S*PYAEVEVIKMCMPLLSPASEIIQFKISEGQEMQACELIARLDLDDPSAVRKAEPF* 123
+ PYAEVEV+KMCMPLLSPAS +I FK+SEGQ MQA ELIA LDLDDPSAVRKAEPF
Sbjct: 699 DADTPYAEVEVMKMCMPLLSPASGVIHFKMSEGQAMQAGELIANLDLDDPSAVRKAEPFH 758
Query: 124 SWALLLQFLVKFNRNVPQA*ILHR*FLLAMSTNMMR*LCKV*SIVY-------SPEFPFL 176
L + V Q + + +V ++V SPE PFL
Sbjct: 759 GSFPRLGLPTAISGRVHQRCAATLNAARMILAGYEHKVDEVNTVVQDLLNCLDSPELPFL 818
Query: 177 QW*ECFAVVATRLCKVLATHL-PIDLKN*FPSRGFQAPKFLTF---AKLLKGNLEAHISS 232
QW ECFAV+ATRL K L + L++ + + LT AKLLKG LEAH+SS
Sbjct: 819 QWQECFAVLATRLPKNLRNMVNTCVLESKYREFESISRNSLTTDFPAKLLKGILEAHLSS 878
Query: 233 CPDKEKGAQERLK*P----D*SCEGGRET 257
C +KE+GA ERL P S EGGRE+
Sbjct: 879 CDEKERGALERLIEPLMSLAKSYEGGRES 907
>At5g55070 2-oxoglutarate dehydrogenase E2 subunit
Length = 464
Score = 28.9 bits (63), Expect = 3.2
Identities = 17/54 (31%), Positives = 29/54 (53%), Gaps = 1/54 (1%)
Query: 56 VSKPLSFAE*S*PYAEVEVIKMCMPLLSPASEIIQ-FKISEGQEMQACELIARL 108
+ KP E A++E K+ + + SPAS +IQ F + EG ++ +AR+
Sbjct: 114 LKKPGDRVEADEAIAQIETDKVTIDIASPASGVIQEFLVKEGDTVEPGNKVARI 167
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.359 0.161 0.584
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,482,438
Number of Sequences: 26719
Number of extensions: 202876
Number of successful extensions: 724
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 709
Number of HSP's gapped (non-prelim): 7
length of query: 282
length of database: 11,318,596
effective HSP length: 98
effective length of query: 184
effective length of database: 8,700,134
effective search space: 1600824656
effective search space used: 1600824656
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0378b.13


