

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0377b.5
(372 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
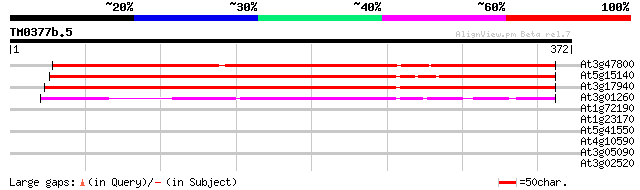
At3g47800 aldose 1-epimerase - like protein 390 e-109
At5g15140 putative aldose 1-epimerase - like protein 383 e-106
At3g17940 aldose 1-epimerase like protein 332 2e-91
At3g01260 putative aldose 1-epimerase 208 3e-54
At1g72190 phosphoglycerate dehydrogenase, putative 30 1.6
At1g23170 unknown protein 29 3.6
At5g41550 disease resistance protein-like 29 4.7
At4g10590 unknown protein 28 8.1
At3g05090 unknown protein 28 8.1
At3g02520 putative 14-3-3 protein 28 8.1
>At3g47800 aldose 1-epimerase - like protein
Length = 358
Score = 390 bits (1002), Expect = e-109
Identities = 190/334 (56%), Positives = 246/334 (72%), Gaps = 6/334 (1%)
Query: 29 DHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGTTA 88
+ I+ ++L +G LS+ +N+GA + SL+LPD+HGK DVVLG+D++ Y ND+TYFG
Sbjct: 29 EKIKTYKLTRGSLSVTFTNYGAVMTSLLLPDRHGKQDDVVLGFDTVDGYKNDTTYFGAIV 88
Query: 89 GRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSY 148
GRVANRIGGA+F LNG YK NEG NTLHGG +GFSDV+W V++YV ITF+Y
Sbjct: 89 GRVANRIGGAKFKLNGHLYKTDPNEGRNTLHGGSKGFSDVIWSVQKYVPTS---HITFTY 145
Query: 149 HSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNI 208
SFDGEEGFPG++ V V+Y+L +N L + M+AK LNKPTP+NL H YWNL +HNSGNI
Sbjct: 146 DSFDGEEGFPGNVTVKVTYMLIGENKLGVKMEAKPLNKPTPINLALHTYWNLHSHNSGNI 205
Query: 209 LGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVL 268
L +Q+ +IT +D LIPTGEI S+ GTPYDFL+P+ +G+RI++LP GYDINYV+
Sbjct: 206 LSHKIQLLAGKITPVDDKLIPTGEITSITGTPYDFLEPREIGSRIHELP--GGYDINYVI 263
Query: 269 DGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCL 328
DG K L K A+V ++ +GR M+L+TN PG+QFYT+N +K GKG VY+ GLCL
Sbjct: 264 DGPIGKHLRK-TAVVTEQVTGRKMELWTNQPGVQFYTSNMMKRVVGKGKAVYEKYGGLCL 322
Query: 329 ESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
E+Q FPDSVNH NFPS IV P + Y HVML +F+
Sbjct: 323 ETQGFPDSVNHKNFPSQIVNPGESYLHVMLFRFT 356
>At5g15140 putative aldose 1-epimerase - like protein
Length = 490
Score = 383 bits (983), Expect = e-106
Identities = 190/336 (56%), Positives = 241/336 (71%), Gaps = 4/336 (1%)
Query: 27 EKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTNDSTYFGT 86
EK+ I L+ELKKG+L++K +NWGA+++SL PDK+GK+ D+VLGYDS+K Y D YFG
Sbjct: 156 EKEKIGLYELKKGNLTVKFTNWGASIISLHFPDKNGKMDDIVLGYDSVKTYKTDKVYFGA 215
Query: 87 TAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITF 146
T GRVANRIG +F LNG YK N+G NTLHGG +GF DV+W V ++ +G KP I F
Sbjct: 216 TVGRVANRIGKGKFKLNGKEYKTSVNDGKNTLHGGKKGFGDVVWAVAKHQYDGKKPHIVF 275
Query: 147 SYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSG 206
++ S DG++GFPG+L VTV+Y L N LS++M+AK +K TPVNL +H+YWNLG HNSG
Sbjct: 276 THTSPDGDQGFPGELSVTVTYKLVKDNELSVVMEAKPKDKATPVNLAHHSYWNLGGHNSG 335
Query: 207 NILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINY 266
+IL E +QI GS T +D LIPTG+I VKGT YDFL+ + + + L GYDINY
Sbjct: 336 DILSEEIQILGSGYTPVDGELIPTGKINPVKGTAYDFLQLRPIKDNMKDL--KTGYDINY 393
Query: 267 VLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGL 326
LD GK+K + K+ +V DKKSGR M+L N GLQFYT +K+ KGK G VYQ GL
Sbjct: 394 CLD-GKAKKMRKIVELV-DKKSGRKMELSGNQAGLQFYTGGMLKDVKGKNGAVYQAFGGL 451
Query: 327 CLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
CLE+Q +PD++NHP FPS IV P K YKH ML KFS
Sbjct: 452 CLETQSYPDALNHPKFPSQIVEPGKKYKHTMLFKFS 487
>At3g17940 aldose 1-epimerase like protein
Length = 341
Score = 332 bits (851), Expect = 2e-91
Identities = 167/341 (48%), Positives = 227/341 (65%), Gaps = 4/341 (1%)
Query: 24 AKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND-ST 82
A + K+ +FEL G + +K+SN+G T+ SL +PDK+GKLGDVVLG+DS+ Y +
Sbjct: 2 ADQSKNTPEIFELNNGTMQVKISNYGTTITSLSVPDKNGKLGDVVLGFDSVDPYVKGLAP 61
Query: 83 YFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKP 142
YFG GRVANRI +F+LNG++Y L N+ N+LHGG +GF +W+V + ++G+KP
Sbjct: 62 YFGCIVGRVANRIKEGKFSLNGVNYTLPINKPPNSLHGGNKGFDKKIWEVAGHKRDGEKP 121
Query: 143 RITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGN 202
ITF YHS DGEEG+PG + VT +Y L S ++ + M+A NK TP+NL H YWNL
Sbjct: 122 FITFKYHSADGEEGYPGAVSVTATYTLTSATTMRLDMEAVPENKDTPINLAQHTYWNLAG 181
Query: 203 HNSGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGY 262
H+SGNIL +QI+GS IT +D + +PTGEI VKGTP+DF + + +G I ++ GY
Sbjct: 182 HDSGNILDHKIQIWGSHITPVDEYTVPTGEILPVKGTPFDFTEEKRIGESIGEV--GIGY 239
Query: 263 DINYVLD-GGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQ 321
D NYVLD + K +K AA + D S RV+ L+TN PG+QFYT N V GKG VY
Sbjct: 240 DHNYVLDCPDQEKEGLKHAAKLSDAASSRVLNLWTNVPGMQFYTGNYVNGVVGKGNAVYG 299
Query: 322 PRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
AG+CLE+Q FP+++N NFPS +V + Y H ML +FS
Sbjct: 300 KHAGVCLETQGFPNAINQSNFPSVVVKAGEKYNHTMLFEFS 340
>At3g01260 putative aldose 1-epimerase
Length = 378
Score = 208 bits (530), Expect = 3e-54
Identities = 127/342 (37%), Positives = 188/342 (54%), Gaps = 57/342 (16%)
Query: 21 NGSAKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND 80
N EK +R+ ELKKG+L++K +N GA+++SL+ P+ + +
Sbjct: 91 NDDGDDEKKTLRVHELKKGNLTVKFTNRGASIMSLLFPNINASKKE-------------- 136
Query: 81 STYFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGD 140
K AN+G NT+HGG + SDV+W+V+++ G
Sbjct: 137 ---------------------------KAAANDGKNTIHGGTKESSDVIWRVKKHKHNGK 169
Query: 141 KPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNL 200
KP I F+Y + +G G L VTV+Y L +N L ++M+AKA K TP+NL++ +YWNL
Sbjct: 170 KPYIVFTYTT--SPDGDQGKLEVTVTYKLVGENRLKMVMQAKAKEKTTPLNLVHRSYWNL 227
Query: 201 GNHNSGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTN 260
G HN+ +I E +QI GS T LD +L PTG+I +VKGTP+DF + + + IN+L
Sbjct: 228 GGHNNEDIFSEEIQILGSGYTHLDDNLTPTGKILAVKGTPFDFRQLRPIKDNINEL--KT 285
Query: 261 GYDINYVLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVY 320
GY INY LDG +K ++ ++D KS M+L T+ GL ++K K K G V+
Sbjct: 286 GYGINYCLDGVANK--MRKVVELVDNKSKIKMELSTDQSGL------RLKTTKPKNGSVH 337
Query: 321 QPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
+ +GLCLES ++N+ S I+ P + YKH ML KFS
Sbjct: 338 KAYSGLCLESH----AINYQYRSSQIIEPGETYKHTMLFKFS 375
>At1g72190 phosphoglycerate dehydrogenase, putative
Length = 344
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/53 (30%), Positives = 29/53 (54%)
Query: 168 ILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRI 220
+L +N + I ++ + L +PT L+ + LG N G L + ++ FGSR+
Sbjct: 137 LLKKQNEMQISLRNRLLGEPTGDTLLGKTVFILGYGNIGIELAKRLKPFGSRV 189
>At1g23170 unknown protein
Length = 569
Score = 29.3 bits (64), Expect = 3.6
Identities = 16/49 (32%), Positives = 24/49 (48%)
Query: 226 HLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGKSK 274
+LIP G +++ G + L+ Q G + L D V DGG+SK
Sbjct: 54 NLIPNGTLSNGGGNVFRSLEEQAEGRHLQILAAKKASDTADVSDGGRSK 102
>At5g41550 disease resistance protein-like
Length = 1085
Score = 28.9 bits (63), Expect = 4.7
Identities = 20/64 (31%), Positives = 27/64 (41%), Gaps = 1/64 (1%)
Query: 290 RVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQP-RAGLCLESQVFPDSVNHPNFPSSIVT 348
RV F N L FY K+ E +G + QP +CL + P +H +SI
Sbjct: 822 RVCCSFGNPTILTFYNCLKLDEEARRGIIMQQPVDEYICLPGKEIPAEFSHKAVGNSITI 881
Query: 349 PEKP 352
P P
Sbjct: 882 PLAP 885
>At4g10590 unknown protein
Length = 910
Score = 28.1 bits (61), Expect = 8.1
Identities = 18/67 (26%), Positives = 33/67 (48%), Gaps = 13/67 (19%)
Query: 229 PTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGKSKGLIKLAAIVMDKKS 288
P ++S+K + +V R+NQ+PK +G +L GG+ + +++
Sbjct: 551 PLDSLSSIKDDEH------IVAYRLNQMPKGSGKAKLEILHGGQKRP-------ILESVR 597
Query: 289 GRVMKLF 295
GR +KLF
Sbjct: 598 GRDVKLF 604
>At3g05090 unknown protein
Length = 753
Score = 28.1 bits (61), Expect = 8.1
Identities = 23/100 (23%), Positives = 40/100 (40%), Gaps = 5/100 (5%)
Query: 110 IANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYIL 169
+A + NN + G G +W +E + KP S +G G VT +
Sbjct: 134 VAAKNNNVVASGGLGGEVFIWDIEAALSPVTKPNDANEDSSSNGANG-----PVTSLRTV 188
Query: 170 GSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNIL 209
GS N++S+ PT + + L +++G +L
Sbjct: 189 GSSNNISVQSSPSHGYTPTIAKGHKESVYALAMNDTGTML 228
>At3g02520 putative 14-3-3 protein
Length = 265
Score = 28.1 bits (61), Expect = 8.1
Identities = 13/32 (40%), Positives = 19/32 (58%)
Query: 206 GNILGEVVQIFGSRITVLDSHLIPTGEIASVK 237
G I E+ +I + +LDSHL+PT +A K
Sbjct: 90 GKIETELSKICDGILNLLDSHLVPTASLAESK 121
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,881,641
Number of Sequences: 26719
Number of extensions: 406599
Number of successful extensions: 795
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 775
Number of HSP's gapped (non-prelim): 11
length of query: 372
length of database: 11,318,596
effective HSP length: 101
effective length of query: 271
effective length of database: 8,619,977
effective search space: 2336013767
effective search space used: 2336013767
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0377b.5


