

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0372b.7
(224 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
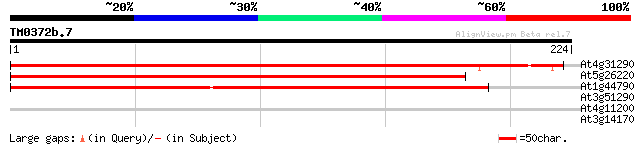
At4g31290 unknown protein 329 6e-91
At5g26220 unknown protein 322 1e-88
At1g44790 unknown protein 189 1e-48
At3g51290 putative protein 27 6.6
At4g11200 putative protein 27 8.6
At3g14170 unknown protein 27 8.6
>At4g31290 unknown protein
Length = 227
Score = 329 bits (844), Expect = 6e-91
Identities = 162/225 (72%), Positives = 183/225 (81%), Gaps = 5/225 (2%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSLVWNPGF YDEKV+G+IK Y+RVFDLACIDHRGTPEHPARTCTLE+ E
Sbjct: 1 MVMWVFGYGSLVWNPGFHYDEKVLGFIKGYKRVFDLACIDHRGTPEHPARTCTLEKAEEA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWG +CVRGGPEKE+L M+YLERRECEYD KT VDFYKE D PA+TGVIVFTSTPD
Sbjct: 61 ICWGTAFCVRGGPEKERLAMEYLERRECEYDLKTSVDFYKEDDPLKPAVTGVIVFTSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KV+NKYYLGPAPL+ MARQIATA+GPCGNNRDYLFLLEKAMH+IGHE+D VIELA+EVRK
Sbjct: 121 KVSNKYYLGPAPLEDMARQIATANGPCGNNRDYLFLLEKAMHDIGHEEDYVIELANEVRK 180
Query: 181 VLGVGN---VLPNDKKLSGPVQIQHSPHAPIPALQLLP-MPEPIA 221
VL + V P + + V + + P A Q+LP PE +A
Sbjct: 181 VLAESSTKKVTPVKESRASRVANKSKNNVP-TAHQILPHHPEAVA 224
>At5g26220 unknown protein
Length = 216
Score = 322 bits (824), Expect = 1e-88
Identities = 147/182 (80%), Positives = 164/182 (89%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSL+WNPGF++DEK+IGYIKDY+RVFDLACIDHRGTPEHPARTCTLE+ G
Sbjct: 1 MVLWVFGYGSLIWNPGFDFDEKLIGYIKDYKRVFDLACIDHRGTPEHPARTCTLEQSTGA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWGA YCVRGGPEKEKL M+YLERRECEYD KTLV+FY E D+S P +TGVIVFTSTPD
Sbjct: 61 ICWGAAYCVRGGPEKEKLAMEYLERRECEYDSKTLVEFYTENDTSTPIVTGVIVFTSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KV+NKYYLGPAPL+ MARQIATA GPCGNNR+YLF LEKAM +I HE++ VIELA+EVRK
Sbjct: 121 KVSNKYYLGPAPLEEMARQIATASGPCGNNREYLFKLEKAMFDIEHEEEYVIELANEVRK 180
Query: 181 VL 182
L
Sbjct: 181 QL 182
>At1g44790 unknown protein
Length = 199
Score = 189 bits (479), Expect = 1e-48
Identities = 91/191 (47%), Positives = 125/191 (64%), Gaps = 1/191 (0%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
M WVFGYGSL+W GF +DE + G+IK YRRVF DHRGTP+ P RT TLE E
Sbjct: 1 MAMWVFGYGSLIWKTGFPFDESLPGFIKGYRRVFHQGSTDHRGTPDFPGRTVTLEAAHEE 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
+C G Y + +K ++ +LE RE +YD K +DF+ + ++S PA+ GV+V+ ++PD
Sbjct: 61 VCCGVAYKITKEEDKRDALL-HLEVREKQYDQKEYLDFFTDSNASEPAVAGVMVYIASPD 119
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
K +N YLGPAPL+ +A+QI A GP G NRDYLF LE+A+ +G +D V +LA++VR
Sbjct: 120 KKSNNNYLGPAPLEDIAKQIVKAKGPSGPNRDYLFNLEEALAQLGFKDKHVTDLANQVRH 179
Query: 181 VLGVGNVLPND 191
+L L D
Sbjct: 180 ILSESEELDID 190
>At3g51290 putative protein
Length = 602
Score = 27.3 bits (59), Expect = 6.6
Identities = 19/73 (26%), Positives = 30/73 (41%), Gaps = 6/73 (8%)
Query: 147 CGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKVLGVGNVLPNDKKLSGPVQIQHSPHA 206
C + YL L KA + + A +R + VG+ L + P+ + H+P +
Sbjct: 17 CKARKRYLKHLVKARQTLS------VSHALYLRSLRAVGSSLVHFSSKETPLHLHHNPPS 70
Query: 207 PIPALQLLPMPEP 219
P P P P P
Sbjct: 71 PSPPPPPPPRPPP 83
>At4g11200 putative protein
Length = 462
Score = 26.9 bits (58), Expect = 8.6
Identities = 26/109 (23%), Positives = 44/109 (39%), Gaps = 20/109 (18%)
Query: 12 VWNPGFEYDEKVIGYIKDYRR--VFDLACIDHRGTPEHPARTCTLEEKEGEICWGAVYCV 69
V+ G ++ + V+ Y+ RR V+D R E C G+ C +YC
Sbjct: 349 VFCSGIDFKKNVVKYVLKTRRNVVYD------RWEKERLGVRCN-----GKGCEWRIYCA 397
Query: 70 RGGPEKEKLVMQYLERRECE-----YDHKT--LVDFYKEGDSSHPALTG 111
P +V Y + +C + KT +V+ + EG P ++G
Sbjct: 398 MENPVGRWMVKTYYPKHQCHPRGRCENIKTPIIVELFMEGIRRDPEMSG 446
>At3g14170 unknown protein
Length = 505
Score = 26.9 bits (58), Expect = 8.6
Identities = 29/106 (27%), Positives = 40/106 (37%), Gaps = 27/106 (25%)
Query: 98 FYKEGDSSHPALTGVIVFTSTPDK---VNNKYYLG------------PAPLDVMARQIAT 142
F K DSSH V S D +NNK +G P P+ V R I+
Sbjct: 52 FIKVSDSSH----STYVSLSNEDNELILNNKLGIGQFFYVDKLDAGTPVPVLVGVRPISG 107
Query: 143 AHGPCGNNRDYLFLL--------EKAMHNIGHEDDMVIELAHEVRK 180
H GN +D + +L E+ HN +D + +RK
Sbjct: 108 RHPFVGNPKDLMQMLVPSETTPREEEYHNQKKKDGARSNIVENIRK 153
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,865,625
Number of Sequences: 26719
Number of extensions: 270992
Number of successful extensions: 550
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 546
Number of HSP's gapped (non-prelim): 6
length of query: 224
length of database: 11,318,596
effective HSP length: 96
effective length of query: 128
effective length of database: 8,753,572
effective search space: 1120457216
effective search space used: 1120457216
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0372b.7


