

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0372a.5
(392 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
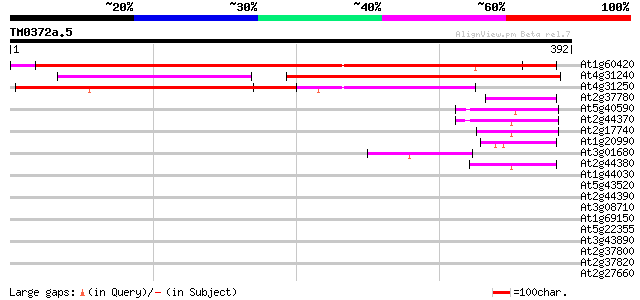
Sequences producing significant alignments: (bits) Value
At1g60420 unknown protein 294 6e-80
At4g31240 predicted protein 278 4e-75
At4g31250 receptor kinase - like protein 179 3e-45
At2g37780 hypothetical protein 47 2e-05
At5g40590 putative protein 45 7e-05
At2g44370 unknown protein 42 4e-04
At2g17740 unknown protein 42 4e-04
At1g20990 hypothetical protein 42 6e-04
At3g01680 unknown protein 42 8e-04
At2g44380 unknown protein 42 8e-04
At1g44030 hypothetical protein 41 0.001
At5g43520 putative protein 40 0.003
At2g44390 unknown protein 40 0.003
At3g08710 thioredoxin-like protein 39 0.004
At1g69150 hypothetical protein 39 0.004
At5g22355 unknown protein 39 0.005
At3g43890 putative protein 39 0.005
At2g37800 hypothetical protein 39 0.005
At2g37820 hypothetical protein 38 0.008
At2g27660 hypothetical protein 38 0.008
>At1g60420 unknown protein
Length = 578
Score = 294 bits (752), Expect = 6e-80
Identities = 140/366 (38%), Positives = 223/366 (60%), Gaps = 3/366 (0%)
Query: 19 ILAAEGAEYLLSCEG-KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGIN 77
+L ++++S +G KVP+SE +GK I L FS R C P+LVE Y L++ +
Sbjct: 179 VLVTPSRDFVISPDGNKVPVSELEGKTIGLLFSVASYRKCTELTPKLVEFYTKLKENKED 238
Query: 78 LEIIFVSFDREEDGFNEHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVE 137
EI+ +S + +E+ FN+ K+ PWLA+PF+ +L + + +P+ + L +
Sbjct: 239 FEIVLISLEDDEESFNQDFKTKPWLALPFNDKSGSKLARHFMLSTLPTLVILGPDGKTRH 298
Query: 138 KNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLAHEGRDFLISGDDRKVPV 197
N+ E I+DYG A+PFT ++ +ELK ++K K E LE LL ++++ D KV V
Sbjct: 299 SNVAEAIDDYGVLAYPFTPEKFQELKELEKAKVEAQTLESLLVSGDLNYVLGKDGAKVLV 358
Query: 198 SELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKEFNVNR 257
S+L GKTI +YF A+W PP RAFT +L +VY +K + FE++ IS+DRD + F+
Sbjct: 359 SDLVGKTILMYFSAHWCPPCRAFTPKLVEVYKQIK-ERNEAFELIFISSDRDQESFDEYY 417
Query: 258 SSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISLNGKFMVTSYGADAFPFTE 317
S PWLA+P+ D + L + + + GIP L +GP G+ ++ + +V ++GADA+PFTE
Sbjct: 418 SQMPWLALPFGDPRKASLAKTFKVGGIPMLAALGPTGQTVTKEARDLVVAHGADAYPFTE 477
Query: 318 SRIRDLEAALRKEGEALPQQVEDVKH-EHVLKLDMAKAYVCDSCKEQGKFWAFSCDVCDY 376
R++++EA + + P++V+ V H EH L+L + Y CD C+E+G W++ CD CD+
Sbjct: 478 ERLKEIEAKYDEIAKDWPKKVKHVLHEEHELELTRVQVYTCDKCEEEGTIWSYHCDECDF 537
Query: 377 DLHPSC 382
DLH C
Sbjct: 538 DLHAKC 543
Score = 232 bits (591), Expect = 3e-61
Identities = 134/362 (37%), Positives = 202/362 (55%), Gaps = 6/362 (1%)
Query: 1 MAGINFEAKHIDSHDVLKILAAEGAEYLLSCEGK-VPLSECDGKVICLFFSANWCRPCRA 59
MA + + D+ D+ +L++ ++L+ +G+ V + GK I L+FSA WC PC+
Sbjct: 1 MAETSKQVNGDDAQDLHSLLSSPARDFLVRNDGEQVKVDSLLGKKIGLYFSAAWCGPCQR 60
Query: 60 FVPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPF-DVNLHRRLIDRY 118
F P+LVE+Y L + + EI+FVS D +E+ F ++ + MPWLAVPF D RL + +
Sbjct: 61 FTPQLVEVYNELSSK-VGFEIVFVSGDEDEESFGDYFRKMPWLAVPFTDSETRDRLDELF 119
Query: 119 QVDQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEEL 178
+V IP+ + + +V +N + I YGADA+PFT ++ +E+K + R R L +
Sbjct: 120 KVRGIPNLVMVDDHGKLVNENGVGVIRSYGADAYPFTPEKMKEIKEDEDRARRGQTLRSV 179
Query: 179 LAHEGRDFLISGDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNC 238
L RDF+IS D KVPVSEL GKTIGL F T +L + Y LK K +
Sbjct: 180 LVTPSRDFVISPDGNKVPVSELEGKTIGLLFSVASYRKCTELTPKLVEFYTKLKENKED- 238
Query: 239 FEIVLISTDRDHKEFNVNRSSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVIS 298
FEIVLIS + D + FN + + PWLA+P+ D++ L R + + +P LV++GPDGK
Sbjct: 239 FEIVLISLEDDEESFNQDFKTKPWLALPFNDKSGSKLARHFMLSTLPTLVILGPDGKTRH 298
Query: 299 LNGKFMVTSYGADAFPFTESRIRDLE--AALRKEGEALPQQVEDVKHEHVLKLDMAKAYV 356
N + YG A+PFT + ++L+ + E + L + +VL D AK V
Sbjct: 299 SNVAEAIDDYGVLAYPFTPEKFQELKELEKAKVEAQTLESLLVSGDLNYVLGKDGAKVLV 358
Query: 357 CD 358
D
Sbjct: 359 SD 360
>At4g31240 predicted protein
Length = 204
Score = 278 bits (711), Expect = 4e-75
Identities = 125/192 (65%), Positives = 161/192 (83%)
Query: 194 KVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKEF 253
+V VS+L GKTIGLYF A+W PP R+FT+QL DVYN L T FE++LISTDRD +EF
Sbjct: 7 QVLVSKLVGKTIGLYFGAHWCPPFRSFTSQLVDVYNELATTDKGSFEVILISTDRDSREF 66
Query: 254 NVNRSSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISLNGKFMVTSYGADAF 313
N+N ++ PWLAIPYEDRTR DLCRI+++K IPALV+IGP+ K ++ N + MV+ YG+ +F
Sbjct: 67 NINMTNMPWLAIPYEDRTRQDLCRIFNVKLIPALVIIGPEEKTVTTNAREMVSLYGSRSF 126
Query: 314 PFTESRIRDLEAALRKEGEALPQQVEDVKHEHVLKLDMAKAYVCDSCKEQGKFWAFSCDV 373
PFTESRI +L+A L+KEG++LP++V+D KHEH LKLDMAKAYVCD CK+QG+FWAFSC+
Sbjct: 127 PFTESRIVELKACLKKEGDSLPRKVKDNKHEHELKLDMAKAYVCDFCKKQGRFWAFSCNA 186
Query: 374 CDYDLHPSCLEK 385
CDYDLHP+C+E+
Sbjct: 187 CDYDLHPTCVEE 198
Score = 94.0 bits (232), Expect = 1e-19
Identities = 50/137 (36%), Positives = 80/137 (57%), Gaps = 1/137 (0%)
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGI-NLEIIFVSFDREEDGF 92
+V +S+ GK I L+F A+WC P R+F +LV++Y L + E+I +S DR+ F
Sbjct: 7 QVLVSKLVGKTIGLYFGAHWCPPFRSFTSQLVDVYNELATTDKGSFEVILISTDRDSREF 66
Query: 93 NEHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEKNLIECIEDYGADAF 152
N ++ +MPWLA+P++ + L + V IP+ + + E+ V N E + YG+ +F
Sbjct: 67 NINMTNMPWLAIPYEDRTRQDLCRIFNVKLIPALVIIGPEEKTVTTNAREMVSLYGSRSF 126
Query: 153 PFTRKRHEELKAIDKRK 169
PFT R ELKA K++
Sbjct: 127 PFTESRIVELKACLKKE 143
>At4g31250 receptor kinase - like protein
Length = 951
Score = 179 bits (453), Expect = 3e-45
Identities = 94/205 (45%), Positives = 132/205 (63%), Gaps = 9/205 (4%)
Query: 5 NFEAKHIDSHDVLKILAAEGAEYLLSCEGKVPLSECDGKVICLFFSANWC---------R 55
+++ K +S D+ ILAAEG E+LLS G+V L + F W R
Sbjct: 747 DYQVKFPESGDLYSILAAEGIEFLLSHSGEVLLLLSRYYIALYFGIIVWSFILCKLTSIR 806
Query: 56 PCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPFDVNLHRRLI 115
PC+ F P L++LYE L+ RG LEIIFVSFD + F EH MPWLAVPF+++L +L
Sbjct: 807 PCKDFTPELIKLYENLQNRGEELEIIFVSFDHDMTSFYEHFWCMPWLAVPFNLSLLNKLR 866
Query: 116 DRYQVDQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANL 175
D+Y + +IPS +PLYS+++ V +++I IEDYG++AFPFT+KR EELKAID KR L
Sbjct: 867 DKYGISRIPSLVPLYSDEISVAEDVIGLIEDYGSEAFPFTKKRKEELKAIDDSKRLGGQL 926
Query: 176 EELLAHEGRDFLISGDDRKVPVSEL 200
E+LL HE R+++++ + KV + L
Sbjct: 927 EKLLTHESRNYVVARNGSKVKRTHL 951
Score = 72.8 bits (177), Expect = 3e-13
Identities = 53/165 (32%), Positives = 83/165 (50%), Gaps = 13/165 (7%)
Query: 171 EEANLEELLAHEGRDFLISGDDRKVPVSELAGKTIGLYF-CAYWS---------PPSRAF 220
E +L +LA EG +FL+S + + L+ I LYF WS P + F
Sbjct: 754 ESGDLYSILAAEGIEFLLSHSGEVLLL--LSRYYIALYFGIIVWSFILCKLTSIRPCKDF 811
Query: 221 TAQLTDVYNNLKATKGNCFEIVLISTDRDHKEFNVNRSSTPWLAIPYEDRTRHDLCRIYD 280
T +L +Y NL+ +G EI+ +S D D F + PWLA+P+ + L Y
Sbjct: 812 TPELIKLYENLQ-NRGEELEIIFVSFDHDMTSFYEHFWCMPWLAVPFNLSLLNKLRDKYG 870
Query: 281 IKGIPALVLIGPDGKVISLNGKFMVTSYGADAFPFTESRIRDLEA 325
I IP+LV + D ++ + ++ YG++AFPFT+ R +L+A
Sbjct: 871 ISRIPSLVPLYSDEISVAEDVIGLIEDYGSEAFPFTKKRKEELKA 915
>At2g37780 hypothetical protein
Length = 286
Score = 46.6 bits (109), Expect = 2e-05
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Query: 333 ALPQQVEDVKHEHVL-KLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
++PQ ++ H H+L +++ Y CD CK G+ + C CDYDLH C
Sbjct: 2 SIPQTIQHFTHNHLLTQVNGIGTYTCDGCKLYGEGRTYRCSDCDYDLHEYC 52
>At5g40590 putative protein
Length = 234
Score = 45.1 bits (105), Expect = 7e-05
Identities = 24/75 (32%), Positives = 38/75 (50%), Gaps = 5/75 (6%)
Query: 312 AFPFTESRIRDLEAALRKEGEALPQQVEDVKHE-HVLKLDMA--KAYVCDSCKEQGKFWA 368
AF T+S D + L K LP++ H+ H L L + Y C++C E G +
Sbjct: 40 AFKCTKSS--DCDYFLHKSCFELPRETNHKSHQPHTLTLIYSPKSTYTCNACGEYGSSFT 97
Query: 369 FSCDVCDYDLHPSCL 383
++C +C YD+H C+
Sbjct: 98 YNCSICQYDVHVGCV 112
Score = 30.8 bits (68), Expect = 1.3
Identities = 16/56 (28%), Positives = 24/56 (42%), Gaps = 9/56 (16%)
Query: 333 ALPQQVEDVKHEHVLKLDMAKAY-------VCDSCKE--QGKFWAFSCDVCDYDLH 379
++P+ V+ H H L L Y CD C++ W++ C CDY H
Sbjct: 113 SMPETVKREDHPHPLTLLYGSPYNQPGLVSKCDVCEDIVPDNLWSYYCKECDYATH 168
Score = 30.0 bits (66), Expect = 2.3
Identities = 19/55 (34%), Positives = 21/55 (37%), Gaps = 6/55 (10%)
Query: 336 QQVEDVKHEHVL---KLDMAKAYVCDSCKEQGKFWAFSCDV---CDYDLHPSCLE 384
Q V H H L K + +C C AF C CDY LH SC E
Sbjct: 5 QSVRHPSHNHPLRGHKCEAKDEIICSGCDLDLVGAAFKCTKSSDCDYFLHKSCFE 59
>At2g44370 unknown protein
Length = 250
Score = 42.4 bits (98), Expect = 4e-04
Identities = 24/77 (31%), Positives = 37/77 (47%), Gaps = 8/77 (10%)
Query: 312 AFPFTESRIRDLEAALRKEGEALPQQVEDVKH-EHVLKL----DMAKAYVCDSCKEQGKF 366
AF T+S + + L K LP++ H +H L L Y CD+C E G
Sbjct: 41 AFKCTKS---ECDYFLHKSCFELPRETRHKAHPDHPLILLYSPPYESTYTCDACGEYGSG 97
Query: 367 WAFSCDVCDYDLHPSCL 383
+ ++C +C YD+H C+
Sbjct: 98 FTYNCSICQYDVHVGCV 114
Score = 33.9 bits (76), Expect = 0.16
Identities = 18/55 (32%), Positives = 24/55 (42%), Gaps = 8/55 (14%)
Query: 333 ALPQQVEDVKHEHVLKLDMAKAYV------CDSCKEQ--GKFWAFSCDVCDYDLH 379
++P+ VE H H L L Y CD C+E W++ C CDY H
Sbjct: 115 SMPESVEREGHAHPLTLLYRSPYQNGLIFNCDVCQETVPDNLWSYYCKECDYGTH 169
Score = 29.3 bits (64), Expect = 3.9
Identities = 18/52 (34%), Positives = 21/52 (39%), Gaps = 5/52 (9%)
Query: 338 VEDVKHEHVL---KLDMAKAYVCDSCKEQGKFWAFSC--DVCDYDLHPSCLE 384
V H H L K + + +C C AF C CDY LH SC E
Sbjct: 8 VRHPSHNHPLRSHKAQVEEEIICSGCDLDLIGAAFKCTKSECDYFLHKSCFE 59
>At2g17740 unknown protein
Length = 248
Score = 42.4 bits (98), Expect = 4e-04
Identities = 20/63 (31%), Positives = 32/63 (50%), Gaps = 6/63 (9%)
Query: 327 LRKEGEALPQQVEDVKH-EHVLKL-----DMAKAYVCDSCKEQGKFWAFSCDVCDYDLHP 380
L K LP+++ H +H L L + Y CD+C E G + ++C +C YD+H
Sbjct: 52 LHKSCFDLPREIRHKSHPDHPLILLYSPQNNNSTYTCDACGEYGSGFTYNCSICQYDVHV 111
Query: 381 SCL 383
C+
Sbjct: 112 GCV 114
Score = 29.6 bits (65), Expect = 3.0
Identities = 17/56 (30%), Positives = 26/56 (46%), Gaps = 10/56 (17%)
Query: 333 ALPQQVEDVKHEHVLKLDMAKA-------YVCDSCKEQ--GKFWAFSCDVCDYDLH 379
++P+ ++ +H H L L + KA + CD C E W + C CDY H
Sbjct: 115 SVPETMKHDEHVHPLAL-IYKAPCPKDHIFTCDVCDETMPHNLWLYYCQKCDYGAH 169
>At1g20990 hypothetical protein
Length = 319
Score = 42.0 bits (97), Expect = 6e-04
Identities = 24/60 (40%), Positives = 32/60 (53%), Gaps = 7/60 (11%)
Query: 330 EGEALPQQV--EDVKH----EHVL-KLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
E A PQ V E++ H +HVL K+++ Y C CKE+G + C CDY LH C
Sbjct: 55 EFPASPQLVQGEEMVHFGHPQHVLVKVELPDIYTCAGCKEEGAGVRYVCQECDYQLHEFC 114
Score = 32.7 bits (73), Expect = 0.35
Identities = 10/26 (38%), Positives = 13/26 (49%)
Query: 357 CDSCKEQGKFWAFSCDVCDYDLHPSC 382
CD C K + F C C + +HP C
Sbjct: 148 CDVCGRSPKGYTFRCKACSFQMHPGC 173
Score = 28.1 bits (61), Expect = 8.7
Identities = 11/29 (37%), Positives = 15/29 (50%), Gaps = 1/29 (3%)
Query: 355 YVCDSCKEQGKFW-AFSCDVCDYDLHPSC 382
++C CK + + C VCDY LH C
Sbjct: 209 FLCGECKRGKRTGRVYRCTVCDYHLHAVC 237
>At3g01680 unknown protein
Length = 740
Score = 41.6 bits (96), Expect = 8e-04
Identities = 24/77 (31%), Positives = 34/77 (43%), Gaps = 4/77 (5%)
Query: 251 KEFNVNRSSTPWLAIPYEDRTRHDLCRI----YDIKGIPALVLIGPDGKVISLNGKFMVT 306
K+F R PW ++ + + P LV+I P G SLN M+
Sbjct: 420 KKFEDLRDPMPWYSVDSPKLIERHVVEFMRGRWHFMNKPILVVIDPQGNEASLNALHMIW 479
Query: 307 SYGADAFPFTESRIRDL 323
+G +AFPFT SR +L
Sbjct: 480 IWGTEAFPFTRSREEEL 496
Score = 35.4 bits (80), Expect = 0.054
Identities = 21/75 (28%), Positives = 34/75 (45%), Gaps = 4/75 (5%)
Query: 92 FNEHLKSMPWLAVPFDVNLHRRLID----RYQVDQIPSFIPLYSEDLIVEKNLIECIEDY 147
F + MPW +V + R +++ R+ P + + + N + I +
Sbjct: 422 FEDLRDPMPWYSVDSPKLIERHVVEFMRGRWHFMNKPILVVIDPQGNEASLNALHMIWIW 481
Query: 148 GADAFPFTRKRHEEL 162
G +AFPFTR R EEL
Sbjct: 482 GTEAFPFTRSREEEL 496
>At2g44380 unknown protein
Length = 247
Score = 41.6 bits (96), Expect = 8e-04
Identities = 21/66 (31%), Positives = 31/66 (46%), Gaps = 5/66 (7%)
Query: 322 DLEAALRKEGEALPQQVEDVKH-EHVLKL----DMAKAYVCDSCKEQGKFWAFSCDVCDY 376
D + L K LP++ H H L L ++Y CD+C E G + ++C C Y
Sbjct: 52 DCDYFLHKSCFDLPRETNHKSHPNHSLTLLHSPPYGQSYTCDACGEYGSGFTYNCSECQY 111
Query: 377 DLHPSC 382
D+H C
Sbjct: 112 DVHVGC 117
Score = 37.4 bits (85), Expect = 0.014
Identities = 21/58 (36%), Positives = 28/58 (48%), Gaps = 13/58 (22%)
Query: 334 LPQQVEDVKHEHVLKL----------DMAKAYVCDSCKEQ--GKFWAFSCDVCDYDLH 379
+P+ VE HEH L L D AK ++CD C+E+ W + C CDY H
Sbjct: 120 IPETVEREDHEHPLTLLYNTPCKGREDGAK-FICDVCEEKMSENLWVYYCKECDYGTH 176
Score = 31.6 bits (70), Expect = 0.79
Identities = 18/52 (34%), Positives = 24/52 (45%), Gaps = 5/52 (9%)
Query: 338 VEDVKHEHVLKLDMAK---AYVCDSCKEQGKFWAFSC--DVCDYDLHPSCLE 384
V H H L++ A+ VC C+ + AF C CDY LH SC +
Sbjct: 12 VRHASHNHPLRVFKARDEDEVVCSGCELELTGQAFKCMKSDCDYFLHKSCFD 63
>At1g44030 hypothetical protein
Length = 597
Score = 40.8 bits (94), Expect = 0.001
Identities = 36/132 (27%), Positives = 53/132 (39%), Gaps = 32/132 (24%)
Query: 283 GIPALVLIGPDGKVISLNGKFMVTSY-----------------------GADAFPFTESR 319
G+ L+ + P KV + GKF VT G DA F
Sbjct: 274 GLEQLISLSPQTKVKLVQGKFHVTGKIERTSKEKGRCQPKNKHRLYLESGEDASHFL--- 330
Query: 320 IRDLEAALRKEGEALPQQVEDVKH-EHVLKLDMAKAYV-----CDSCKEQGKFWAFSCDV 373
+D E E P +V+ + H +H L+L + K+ + C C E K + C V
Sbjct: 331 CKDCNGEDHVECEKTPVEVKHLLHPKHSLQLVLQKSSITQTRKCFCCDEDLKLIFYYCTV 390
Query: 374 CDYDLHPSCLEK 385
CDY ++ +C EK
Sbjct: 391 CDYAMNIACAEK 402
>At5g43520 putative protein
Length = 250
Score = 39.7 bits (91), Expect = 0.003
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 6/62 (9%)
Query: 327 LRKEGEALPQQVEDVKHE-HVLKL-----DMAKAYVCDSCKEQGKFWAFSCDVCDYDLHP 380
L K LP + + H+ H L L D Y CD+C + G +++ C C+Y LH
Sbjct: 61 LHKSCFDLPSETNNKSHQDHPLTLLHSPPDDRSVYTCDACDQYGSGFSYHCSNCNYSLHV 120
Query: 381 SC 382
C
Sbjct: 121 GC 122
Score = 32.3 bits (72), Expect = 0.46
Identities = 19/54 (35%), Positives = 24/54 (44%), Gaps = 5/54 (9%)
Query: 336 QQVEDVKHEHVLKLDMAKAY---VCDSCKEQGKFWAFSC--DVCDYDLHPSCLE 384
Q V H H L++ AK C C+ + AF C CDY LH SC +
Sbjct: 14 QSVRHPSHNHPLRVFKAKEEDETACSGCELELTGQAFKCIKSECDYFLHKSCFD 67
Score = 29.6 bits (65), Expect = 3.0
Identities = 16/56 (28%), Positives = 22/56 (38%), Gaps = 10/56 (17%)
Query: 334 LPQQVEDVKHEHVLKLDMAKA--------YVCDSCKE--QGKFWAFSCDVCDYDLH 379
+P+ V+ HEH L L + C +C E W + C CDY H
Sbjct: 125 IPETVDREDHEHPLTLLYCTPCKGREDTYFTCSACDETISEDLWMYYCKDCDYGTH 180
>At2g44390 unknown protein
Length = 209
Score = 39.7 bits (91), Expect = 0.003
Identities = 13/30 (43%), Positives = 18/30 (59%)
Query: 353 KAYVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
+ Y CD+C E G + ++C C YDLH C
Sbjct: 51 ETYTCDACGEYGSGFTYNCSECQYDLHVGC 80
Score = 34.3 bits (77), Expect = 0.12
Identities = 17/57 (29%), Positives = 25/57 (43%), Gaps = 11/57 (19%)
Query: 334 LPQQVEDVKHEHVLKL---------DMAKAYVCDSCKEQ--GKFWAFSCDVCDYDLH 379
+P+ VE HEH L L + ++C+ C+E W + C CDY H
Sbjct: 83 IPETVEREDHEHPLTLLYNTPCKGCEDGAMFICNVCEEDMSENLWVYYCKECDYGTH 139
>At3g08710 thioredoxin-like protein
Length = 140
Score = 39.3 bits (90), Expect = 0.004
Identities = 16/40 (40%), Positives = 24/40 (60%)
Query: 30 SCEGKVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYE 69
S + K+ ++ DGK++ FSA WC PC+ P +EL E
Sbjct: 33 SWDDKLAEADRDGKIVVANFSATWCGPCKIVAPFFIELSE 72
>At1g69150 hypothetical protein
Length = 517
Score = 39.3 bits (90), Expect = 0.004
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Query: 335 PQQVEDVK--HEHVLKLDMAKA-YVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
P +V DV H+H LKL+M K+ + C +C + +++ C CD H +C
Sbjct: 86 PSEVIDVPQHHDHKLKLEMVKSCFSCATCGKDSDGYSYKCHECDLKFHVNC 136
Score = 28.5 bits (62), Expect = 6.7
Identities = 22/74 (29%), Positives = 37/74 (49%), Gaps = 12/74 (16%)
Query: 321 RDLEAALRKEGEALPQQVED------VKH-EHVLKLDMAKAYV----CDSCKEQGKFWAF 369
++++ +E + LP +V D V H EH L+LD + ++ C++C +F
Sbjct: 317 KEMKDEPEEEEDILPFKVIDENTIQHVSHKEHNLRLDKSGIFIEERICEACVYPIYHHSF 376
Query: 370 -SCDVCDYDLHPSC 382
SC C + LH SC
Sbjct: 377 YSCMSCSFILHESC 390
>At5g22355 unknown protein
Length = 664
Score = 38.9 bits (89), Expect = 0.005
Identities = 15/42 (35%), Positives = 23/42 (54%)
Query: 343 HEHVLKLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSCLE 384
H H L + K + CD+C ++ + SCD CD+DL C +
Sbjct: 498 HRHPLYYEHKKDHCCDACYKEIDGYLLSCDTCDFDLDLHCTD 539
Score = 29.6 bits (65), Expect = 3.0
Identities = 15/52 (28%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query: 335 PQQVEDVK-HEH-VLKLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSCLE 384
P +E K HEH ++ L ++ C++C QG + C C + +H +C++
Sbjct: 233 PLAIEHHKTHEHRLVLLSRLISFECNACGMQGDRSPYMCVQCGFVVHRTCID 284
>At3g43890 putative protein
Length = 661
Score = 38.9 bits (89), Expect = 0.005
Identities = 17/51 (33%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Query: 335 PQQVEDVK--HEHVLKLDMA-KAYVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
P +V DV H+H LKL+M ++ C +C + G +++ C C+ H +C
Sbjct: 126 PSEVIDVPQHHDHKLKLEMVISSFTCAACGKDGDGYSYKCQECNLTFHVNC 176
Score = 28.1 bits (61), Expect = 8.7
Identities = 15/57 (26%), Positives = 28/57 (48%), Gaps = 7/57 (12%)
Query: 334 LPQQVEDVKHEHVLKL-----DMAKAYVCDSCKEQ--GKFWAFSCDVCDYDLHPSCL 383
LP +V+ +H L L + + + CD C+++ K W ++ C+ LH C+
Sbjct: 534 LPYEVKHSVDQHFLSLCYGEENASGKHWCDICEKEMDPKTWFYTSKDCELTLHTDCV 590
>At2g37800 hypothetical protein
Length = 396
Score = 38.9 bits (89), Expect = 0.005
Identities = 21/57 (36%), Positives = 28/57 (48%), Gaps = 3/57 (5%)
Query: 329 KEGEALPQQ--VEDVKHEHVL-KLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
K + P+Q V+ H H L K+D + CD CK G + C CDY+LH C
Sbjct: 134 KMSSSRPEQLVVQHFTHIHPLTKVDGYGEFTCDGCKTYGFGKTYRCTRCDYNLHDHC 190
Score = 36.2 bits (82), Expect = 0.032
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 2/52 (3%)
Query: 334 LPQQVEDVKHE-HVLKLDM-AKAYVCDSCKEQGKFWAFSCDVCDYDLHPSCL 383
LPQ V V H H+L+L + +C C+ + W + C C D+H C+
Sbjct: 252 LPQHVRHVPHPAHLLELSQWGASSICMVCRGAIRSWRYKCGPCGLDVHMECI 303
Score = 33.1 bits (74), Expect = 0.27
Identities = 10/29 (34%), Positives = 17/29 (58%)
Query: 356 VCDSCKEQGKFWAFSCDVCDYDLHPSCLE 384
+CD C E + + C+ C +D+HP C +
Sbjct: 223 MCDICDESAEGLYYQCEPCGFDVHPLCTQ 251
Score = 32.0 bits (71), Expect = 0.60
Identities = 10/29 (34%), Positives = 16/29 (54%)
Query: 356 VCDSCKEQGKFWAFSCDVCDYDLHPSCLE 384
+CD C E + + C C +D+HP C +
Sbjct: 35 MCDICDESAEGLYYQCKPCGFDVHPLCTQ 63
>At2g37820 hypothetical protein
Length = 290
Score = 38.1 bits (87), Expect = 0.008
Identities = 16/46 (34%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Query: 338 VEDVKHEHVL-KLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSC 382
V+ H H L + + ++CD CK G + C+ C+YDLH C
Sbjct: 11 VQHFTHIHPLTEFNSIGDFICDGCKTYGSGKTYRCEPCNYDLHEYC 56
Score = 38.1 bits (87), Expect = 0.008
Identities = 17/51 (33%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 334 LPQQVEDVKHE-HVLKLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSCL 383
LPQ V V H H L+ + A C C + W + C++C +D+H C+
Sbjct: 118 LPQHVRHVLHPAHHLEFRPSGASTCMVCHGPCQSWRYRCELCRFDIHMECI 168
Score = 33.1 bits (74), Expect = 0.27
Identities = 10/28 (35%), Positives = 17/28 (60%)
Query: 357 CDSCKEQGKFWAFSCDVCDYDLHPSCLE 384
CD C E + + C +C++D+HP C +
Sbjct: 90 CDICDESVEGLFYRCKICEFDVHPLCTQ 117
>At2g27660 hypothetical protein
Length = 718
Score = 38.1 bits (87), Expect = 0.008
Identities = 21/71 (29%), Positives = 31/71 (43%), Gaps = 6/71 (8%)
Query: 321 RDLEAALRKEGEALPQQVEDVKH-EHVLKLDMAKAYV-----CDSCKEQGKFWAFSCDVC 374
R +L + + Q + H H L L +A Y CD C G +++ C VC
Sbjct: 48 RPCNFSLHESCSKMKQVITHPSHPSHTLSLLVAPVYDGGYFNCDGCGIHGTGFSYQCSVC 107
Query: 375 DYDLHPSCLEK 385
D+D+H C K
Sbjct: 108 DFDIHALCAYK 118
Score = 34.3 bits (77), Expect = 0.12
Identities = 13/36 (36%), Positives = 20/36 (55%), Gaps = 1/36 (2%)
Query: 353 KAYVCDSCKEQGKF-WAFSCDVCDYDLHPSCLEKVN 387
K + CD C++ GK W + C C++D H C+ N
Sbjct: 144 KGFSCDICRKIGKNQWLYRCIPCEFDAHVGCITGPN 179
Score = 29.6 bits (65), Expect = 3.0
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 2/47 (4%)
Query: 338 VEDVKHEHVLKLDMAKAYV-CDSCK-EQGKFWAFSCDVCDYDLHPSC 382
+ H H L+L A + C +CK G +SC C++ LH SC
Sbjct: 12 INHFSHPHRLQLTPATSSPPCSACKLTGGNGRVYSCRPCNFSLHESC 58
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.140 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,354,287
Number of Sequences: 26719
Number of extensions: 419665
Number of successful extensions: 1619
Number of sequences better than 10.0: 135
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 75
Number of HSP's that attempted gapping in prelim test: 1281
Number of HSP's gapped (non-prelim): 412
length of query: 392
length of database: 11,318,596
effective HSP length: 101
effective length of query: 291
effective length of database: 8,619,977
effective search space: 2508413307
effective search space used: 2508413307
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0372a.5


