

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.6
(143 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
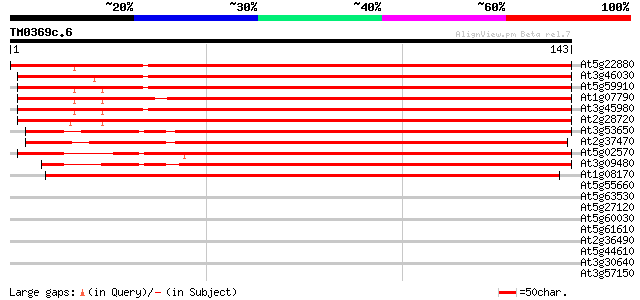
Score E
Sequences producing significant alignments: (bits) Value
At5g22880 histone H2B like protein (emb|CAA69025.1) 241 1e-64
At3g46030 histone H2B -like protein 230 2e-61
At5g59910 histone H2B - like protein 230 2e-61
At1g07790 histone H2B 229 3e-61
At3g45980 histone H2B 229 4e-61
At2g28720 putative histone H2B 219 5e-58
At3g53650 histone H2B - like protein 218 7e-58
At2g37470 putative histone H2B 216 3e-57
At5g02570 putative protein 208 9e-55
At3g09480 putative histone H2B 194 1e-50
At1g08170 hypothetical protein 132 5e-32
At5g55660 putative protein 38 0.002
At5g63530 putative metal-binding protein 36 0.008
At5g27120 SAR DNA-binding protein - like 35 0.014
At5g60030 KED - like protein 33 0.068
At5g61610 putative protein 32 0.089
At2g36490 hypothetical protein 32 0.12
At5g44610 unknown protein 32 0.15
At3g30640 hypothetical protein 32 0.15
At3g57150 putative pseudouridine synthase (NAP57) 31 0.20
>At5g22880 histone H2B like protein (emb|CAA69025.1)
Length = 145
Score = 241 bits (615), Expect = 1e-64
Identities = 130/146 (89%), Positives = 133/146 (91%), Gaps = 4/146 (2%)
Query: 1 MAKGDKKPAEKKPAA---AEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYK 57
MAK DKKPAEKKPA A E AAAAEKKP+A KK+P KE AGDKKKKRSKKNVETYK
Sbjct: 1 MAKADKKPAEKKPAEKTPAAEPAAAAEKKPKAGKKLP-KEPAGAGDKKKKRSKKNVETYK 59
Query: 58 IYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAV 117
IYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLA ESS+LARYNKKPTITSREIQTAV
Sbjct: 60 IYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAGESSKLARYNKKPTITSREIQTAV 119
Query: 118 RLVLPGELAKHAVSEGTKAVTKFTSS 143
RLVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 120 RLVLPGELAKHAVSEGTKAVTKFTSS 145
>At3g46030 histone H2B -like protein
Length = 145
Score = 230 bits (587), Expect = 2e-61
Identities = 122/143 (85%), Positives = 128/143 (89%), Gaps = 3/143 (2%)
Query: 3 KGDKKPAEKKPAAAEEKA--AAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYI 60
K +KKPAEKKP + KA A AEKKP+A KK+P KE GA GDKKKK KK+VETYKIYI
Sbjct: 4 KAEKKPAEKKPVEEKSKAEKAPAEKKPKAGKKLP-KEAGAGGDKKKKMKKKSVETYKIYI 62
Query: 61 FKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLV 120
FKVLKQVHPDIGISSKAMGIMNSFINDIFEKLA ESS+LARYNKKPTITSREIQTAVRLV
Sbjct: 63 FKVLKQVHPDIGISSKAMGIMNSFINDIFEKLASESSKLARYNKKPTITSREIQTAVRLV 122
Query: 121 LPGELAKHAVSEGTKAVTKFTSS 143
LPGELAKHAVSEGTKAVTKFTSS
Sbjct: 123 LPGELAKHAVSEGTKAVTKFTSS 145
>At5g59910 histone H2B - like protein
Length = 150
Score = 230 bits (586), Expect = 2e-61
Identities = 124/148 (83%), Positives = 132/148 (88%), Gaps = 8/148 (5%)
Query: 3 KGDKKPAEKKPAA---AEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
K +KKPAEKKPA+ EEK+ A AEKKP+A KK+P KE GA GDKKKK KK+VET
Sbjct: 4 KAEKKPAEKKPASEKPVEEKSKAEKAPAEKKPKAGKKLP-KEAGAGGDKKKKMKKKSVET 62
Query: 56 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQT 115
YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQE+S+LARYNKKPTITSREIQT
Sbjct: 63 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQEASKLARYNKKPTITSREIQT 122
Query: 116 AVRLVLPGELAKHAVSEGTKAVTKFTSS 143
AVRLVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 123 AVRLVLPGELAKHAVSEGTKAVTKFTSS 150
>At1g07790 histone H2B
Length = 148
Score = 229 bits (585), Expect = 3e-61
Identities = 126/148 (85%), Positives = 131/148 (88%), Gaps = 10/148 (6%)
Query: 3 KGDKKPAEKKPAA---AEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
+ +KKPAEKK AA EE AA AEKKP+A KK+P KE AGDKKKKRSKKNVET
Sbjct: 4 RAEKKPAEKKTAAERPVEENKAAEKAPAEKKPKAGKKLPPKE---AGDKKKKRSKKNVET 60
Query: 56 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQT 115
YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESS+LARYNKKPTITSREIQT
Sbjct: 61 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSKLARYNKKPTITSREIQT 120
Query: 116 AVRLVLPGELAKHAVSEGTKAVTKFTSS 143
AVRLVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 121 AVRLVLPGELAKHAVSEGTKAVTKFTSS 148
>At3g45980 histone H2B
Length = 150
Score = 229 bits (584), Expect = 4e-61
Identities = 124/148 (83%), Positives = 131/148 (87%), Gaps = 8/148 (5%)
Query: 3 KGDKKPAEKKPAA---AEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
+ +KKPAEKKPAA EEK+ A AEKKP+A KK+P KE GA GDKKKK KK+VET
Sbjct: 4 RAEKKPAEKKPAAEKPVEEKSKAEKAPAEKKPKAGKKLP-KEAGAGGDKKKKMKKKSVET 62
Query: 56 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQT 115
YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLA ESS+LARYNKKPTITSREIQT
Sbjct: 63 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLASESSKLARYNKKPTITSREIQT 122
Query: 116 AVRLVLPGELAKHAVSEGTKAVTKFTSS 143
AVRLVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 123 AVRLVLPGELAKHAVSEGTKAVTKFTSS 150
>At2g28720 putative histone H2B
Length = 151
Score = 219 bits (557), Expect = 5e-58
Identities = 119/148 (80%), Positives = 126/148 (84%), Gaps = 7/148 (4%)
Query: 3 KGDKKPAEKKPA----AAEEKAAA---AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
K KKPAEKKPA A EEK A AEKKP+A KK+P + +KKKKR KK+ ET
Sbjct: 4 KAGKKPAEKKPAEKAPAEEEKVAEKAPAEKKPKAGKKLPKEAVTGGVEKKKKRVKKSTET 63
Query: 56 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQT 115
YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQE+S+LARYNKKPTITSREIQT
Sbjct: 64 YKIYIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQEASKLARYNKKPTITSREIQT 123
Query: 116 AVRLVLPGELAKHAVSEGTKAVTKFTSS 143
AVRLVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 124 AVRLVLPGELAKHAVSEGTKAVTKFTSS 151
>At3g53650 histone H2B - like protein
Length = 138
Score = 218 bits (556), Expect = 7e-58
Identities = 118/139 (84%), Positives = 126/139 (89%), Gaps = 7/139 (5%)
Query: 5 DKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFKVL 64
+KKPA KKPA + A AEK P+AEKKI KEGG+ +KKKK+SKKN+ETYKIYIFKVL
Sbjct: 7 EKKPAGKKPA----EKAPAEKLPKAEKKI-TKEGGS--EKKKKKSKKNIETYKIYIFKVL 59
Query: 65 KQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGE 124
KQVHPDIGIS KAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGE
Sbjct: 60 KQVHPDIGISGKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGE 119
Query: 125 LAKHAVSEGTKAVTKFTSS 143
L+KHAVSEGTKAVTKFTSS
Sbjct: 120 LSKHAVSEGTKAVTKFTSS 138
>At2g37470 putative histone H2B
Length = 138
Score = 216 bits (551), Expect = 3e-57
Identities = 115/138 (83%), Positives = 123/138 (88%), Gaps = 6/138 (4%)
Query: 5 DKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFKVL 64
+KKPAEKKPA A AEK P+AEKKI GG+ +KKKK+SKK+VETYKIYIFKVL
Sbjct: 7 EKKPAEKKPAGK----APAEKLPKAEKKISKDAGGS--EKKKKKSKKSVETYKIYIFKVL 60
Query: 65 KQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGE 124
KQVHPD+GIS KAMGIMNSFINDIFEKLAQESS+LARYNKKPTITSREIQTAVRLVLPGE
Sbjct: 61 KQVHPDVGISGKAMGIMNSFINDIFEKLAQESSKLARYNKKPTITSREIQTAVRLVLPGE 120
Query: 125 LAKHAVSEGTKAVTKFTS 142
LAKHAVSEGTKAVTKFTS
Sbjct: 121 LAKHAVSEGTKAVTKFTS 138
>At5g02570 putative protein
Length = 132
Score = 208 bits (529), Expect = 9e-55
Identities = 115/142 (80%), Positives = 121/142 (84%), Gaps = 14/142 (9%)
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGD-KKKKRSKKNVETYKIYIF 61
K +KKPAEK PA P+AEKKI AKEGG + KKKK++KK+ ETYKIYIF
Sbjct: 4 KAEKKPAEKAPA------------PKAEKKI-AKEGGTSEIVKKKKKTKKSTETYKIYIF 50
Query: 62 KVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVL 121
KVLKQVHPDIGIS KAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVL
Sbjct: 51 KVLKQVHPDIGISGKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVL 110
Query: 122 PGELAKHAVSEGTKAVTKFTSS 143
PGELAKHAVSEGTKAVTKFTSS
Sbjct: 111 PGELAKHAVSEGTKAVTKFTSS 132
>At3g09480 putative histone H2B
Length = 126
Score = 194 bits (493), Expect = 1e-50
Identities = 105/135 (77%), Positives = 116/135 (85%), Gaps = 13/135 (9%)
Query: 9 AEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFKVLKQVH 68
AEKKP+ EK P+A+KKI KEGG+ ++KK++KK+ ETYKIY+FKVLKQVH
Sbjct: 5 AEKKPS---------EKAPKADKKI-TKEGGS---ERKKKTKKSTETYKIYLFKVLKQVH 51
Query: 69 PDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGELAKH 128
PDIGIS KAMGIMNSFIND FEK+A ESSRLARYNKKPTITSREIQTAVRLVLPGELAKH
Sbjct: 52 PDIGISGKAMGIMNSFINDTFEKIALESSRLARYNKKPTITSREIQTAVRLVLPGELAKH 111
Query: 129 AVSEGTKAVTKFTSS 143
AVSEGTKAVTKFTSS
Sbjct: 112 AVSEGTKAVTKFTSS 126
>At1g08170 hypothetical protein
Length = 243
Score = 132 bits (333), Expect = 5e-32
Identities = 64/131 (48%), Positives = 94/131 (70%)
Query: 10 EKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFKVLKQVHP 69
E P E A+ +E + K+ K+ KKKKR + Y+ Y++KV+KQVHP
Sbjct: 105 EPSPQPPETPASKSEGTLKKTDKVEKKQENKKKKKKKKRDDLAGDEYRRYVYKVMKQVHP 164
Query: 70 DIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGELAKHA 129
D+GI+SKAM ++N F+ D+FE++AQE++RL+ Y K+ T++SREI+ AVRLVLPGEL++HA
Sbjct: 165 DLGITSKAMTVVNMFMGDMFERIAQEAARLSDYTKRRTLSSREIEAAVRLVLPGELSRHA 224
Query: 130 VSEGTKAVTKF 140
V+EG+KAV+ F
Sbjct: 225 VAEGSKAVSNF 235
>At5g55660 putative protein
Length = 759
Score = 37.7 bits (86), Expect = 0.002
Identities = 34/104 (32%), Positives = 50/104 (47%), Gaps = 16/104 (15%)
Query: 6 KKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFK--- 62
K+ +KPA +E+AAA K EK + K GG DK K+ S + ++T I I K
Sbjct: 627 KRKKTEKPA--KEQAAAPLKSVSKEKPVIGKRGGKGKDKNKEPSDEELKTAIIDILKGVD 684
Query: 63 --------VLKQVHP--DIGISSKAMGIMNSFINDIFEKLAQES 96
+LK++ +I ++SK I I D KLA E+
Sbjct: 685 FNTATFTDILKRLDAKFNISLASKKSSI-KRMIQDELTKLADEA 727
Score = 27.3 bits (59), Expect = 2.9
Identities = 14/42 (33%), Positives = 20/42 (47%)
Query: 11 KKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKN 52
+KP A + ++K K+ P K AAG KRS K+
Sbjct: 447 EKPHATTDVLVNEKEKGVKRKRTPKKSSPAAGSSSSKRSAKS 488
>At5g63530 putative metal-binding protein
Length = 355
Score = 35.8 bits (81), Expect = 0.008
Identities = 20/50 (40%), Positives = 28/50 (56%), Gaps = 4/50 (8%)
Query: 10 EKKPAAAEEKAAAA----EKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
EKKP AAEEK EKK +KK+ A++ G DKK + + N ++
Sbjct: 5 EKKPEAAEEKKMEEKKPEEKKEGEDKKVDAEKKGEDSDKKPQEGESNKDS 54
>At5g27120 SAR DNA-binding protein - like
Length = 533
Score = 35.0 bits (79), Expect = 0.014
Identities = 20/52 (38%), Positives = 30/52 (57%), Gaps = 4/52 (7%)
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPR----AEKKIPAKEGGAAGDKKKKRSK 50
K +KK E +P AEE A +KK R E ++PAK+ + KKKK+++
Sbjct: 481 KSNKKKTEAEPETAEEPAKKEKKKKRKHEEEETEMPAKKKEKSEKKKKKKTE 532
Score = 29.6 bits (65), Expect = 0.57
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 4/57 (7%)
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAK----EGGAAGDKKKKRSKKNVET 55
K DKK +K E K KK +KK A+ E A +KKKKR + ET
Sbjct: 457 KKDKKKKKKADDEEEAKTEEPSKKKSNKKKTEAEPETAEEPAKKEKKKKRKHEEEET 513
>At5g60030 KED - like protein
Length = 292
Score = 32.7 bits (73), Expect = 0.068
Identities = 14/51 (27%), Positives = 28/51 (54%)
Query: 4 GDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVE 54
G++K +KK ++E+ + E+K + ++K + G KKKR K ++
Sbjct: 241 GERKKEKKKKRKSDEEIVSEERKSKKKRKSDEEMGSEERKSKKKRKLKEID 291
Score = 25.8 bits (55), Expect = 8.3
Identities = 12/45 (26%), Positives = 25/45 (54%)
Query: 10 EKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVE 54
EK+ E++ + E+K +KK + E + ++K K+ +K+ E
Sbjct: 228 EKEKEKLEDEQRSGERKKEKKKKRKSDEEIVSEERKSKKKRKSDE 272
>At5g61610 putative protein
Length = 225
Score = 32.3 bits (72), Expect = 0.089
Identities = 19/50 (38%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Query: 1 MAKGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSK 50
+ KGDK P E+ AEE+ + KP K P K+ A GDK + K
Sbjct: 141 LLKGDK-PVEEDKLPAEEEKPPQKDKPAEGHKPPQKDKPAEGDKPVEEDK 189
Score = 28.5 bits (62), Expect = 1.3
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 5/51 (9%)
Query: 5 DKKPAEKKPAAAE-----EKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSK 50
DK P + KP+ + +K +K P E+K P K+ A G K ++ K
Sbjct: 127 DKLPRKDKPSKEDNLLKGDKPVEEDKLPAEEEKPPQKDKPAEGHKPPQKDK 177
Score = 28.1 bits (61), Expect = 1.7
Identities = 17/43 (39%), Positives = 21/43 (48%), Gaps = 8/43 (18%)
Query: 2 AKGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDK 44
A+G K P + KPA + KP E K P K+ A GDK
Sbjct: 167 AEGHKPPQKDKPAEGD--------KPVEEDKPPQKDKPAEGDK 201
>At2g36490 hypothetical protein
Length = 1207
Score = 32.0 bits (71), Expect = 0.12
Identities = 20/57 (35%), Positives = 26/57 (45%), Gaps = 2/57 (3%)
Query: 3 KGDKKPAEKK--PAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYK 57
K +KP KK P E E KPRA +K +G + K+K +K VE K
Sbjct: 84 KTPEKPKRKKHRPKVRREAKPKREPKPRAPRKSVVTDGQESKTPKRKYVRKKVEVSK 140
>At5g44610 unknown protein
Length = 168
Score = 31.6 bits (70), Expect = 0.15
Identities = 23/59 (38%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 5 DKKPAEKKPAAAEEKAAAAEKKPRAE---KKIPAKEGGAAGDKKKK---RSKKNVETYK 57
++K E KP AAA EKKP E K P +E A +++KK KK VE K
Sbjct: 70 EEKKEEAKPVEVPVLAAAEEKKPAVEEEKKTAPVEEKKPAVEEEKKPAVEEKKPVEEEK 128
Score = 29.3 bits (64), Expect = 0.75
Identities = 28/66 (42%), Positives = 31/66 (46%), Gaps = 15/66 (22%)
Query: 2 AKGDKKPA-----------EKKPAAAEE-KAAAAEKKPRAEKKIPAKEGGAAGDKKKKRS 49
A +KKPA EKKPA EE K A EKKP E+K KE AA + S
Sbjct: 86 AAEEKKPAVEEEKKTAPVEEKKPAVEEEKKPAVEEKKPVEEEK---KEVVAAVPVAETPS 142
Query: 50 KKNVET 55
K ET
Sbjct: 143 TKAPET 148
>At3g30640 hypothetical protein
Length = 661
Score = 31.6 bits (70), Expect = 0.15
Identities = 15/48 (31%), Positives = 26/48 (53%)
Query: 5 DKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKN 52
D K +EKK ++K A + P ++K+ EG + +K ++ KKN
Sbjct: 41 DVKTSEKKKKLVKDKDADVSETPAKKQKVSHSEGVHSREKDAQKKKKN 88
>At3g57150 putative pseudouridine synthase (NAP57)
Length = 565
Score = 31.2 bits (69), Expect = 0.20
Identities = 17/46 (36%), Positives = 28/46 (59%), Gaps = 1/46 (2%)
Query: 7 KPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKN 52
K E + AEEK +++KK + +K+ KE A +KK+K+ KK+
Sbjct: 460 KTKEVEGEEAEEKVKSSKKKKKKDKE-EEKEEEAGSEKKEKKKKKD 504
Score = 31.2 bits (69), Expect = 0.20
Identities = 18/57 (31%), Positives = 28/57 (48%), Gaps = 5/57 (8%)
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKE-----GGAAGDKKKKRSKKNVE 54
K KK +K +E+ A +EKK + +KK +E +KKKK+ K+ E
Sbjct: 474 KSSKKKKKKDKEEEKEEEAGSEKKEKKKKKDKKEEVIEEVASPKSEKKKKKKSKDTE 530
Score = 30.4 bits (67), Expect = 0.34
Identities = 18/53 (33%), Positives = 31/53 (57%), Gaps = 4/53 (7%)
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
K DK+ +++ A +E+K EKK + +KK E A+ +KK+ KK+ +T
Sbjct: 481 KKDKEEEKEEEAGSEKK----EKKKKKDKKEEVIEEVASPKSEKKKKKKSKDT 529
Score = 28.1 bits (61), Expect = 1.7
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 6/56 (10%)
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKK------IPAKEGGAAGDKKKKRSKKN 52
K KK +KK EE A+ +K + +K + A++ AA +KK+ KK+
Sbjct: 497 KEKKKKKDKKEEVIEEVASPKSEKKKKKKSKDTEAAVDAEDESAAEKSEKKKKKKD 552
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.308 0.125 0.326
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,850,937
Number of Sequences: 26719
Number of extensions: 111175
Number of successful extensions: 1009
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 675
Number of HSP's gapped (non-prelim): 304
length of query: 143
length of database: 11,318,596
effective HSP length: 89
effective length of query: 54
effective length of database: 8,940,605
effective search space: 482792670
effective search space used: 482792670
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0369c.6


