

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.1
(573 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
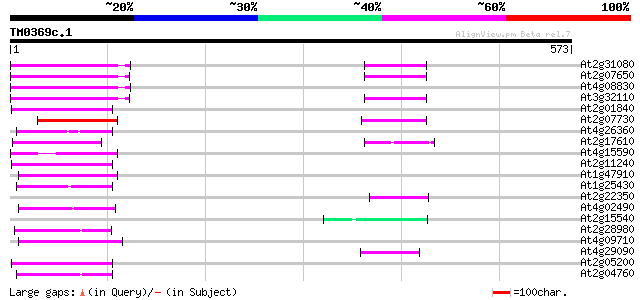
Score E
Sequences producing significant alignments: (bits) Value
At2g31080 putative non-LTR retroelement reverse transcriptase 74 2e-13
At2g07650 putative non-LTR retrolelement reverse transcriptase 69 5e-12
At4g08830 putative protein 69 9e-12
At3g32110 non-LTR reverse transcriptase, putative 65 8e-11
At2g01840 putative non-LTR retroelement reverse transcriptase 61 1e-09
At2g07730 putative non-LTR retroelement reverse transcriptase 54 2e-07
At4g26360 putative protein 54 3e-07
At2g17610 putative non-LTR retroelement reverse transcriptase 52 7e-07
At4g15590 reverse transcriptase like protein 50 3e-06
At2g11240 pseudogene 49 8e-06
At1g47910 reverse transcriptase, putative 49 8e-06
At1g25430 hypothetical protein 49 1e-05
At2g22350 putative non-LTR retroelement reverse transcriptase 48 2e-05
At4g02490 47 2e-05
At2g15540 putative non-LTR retroelement reverse transcriptase 47 2e-05
At2g28980 putative non-LTR retroelement reverse transcriptase 47 3e-05
At4g09710 RNA-directed DNA polymerase -like protein 47 4e-05
At4g29090 putative protein 46 5e-05
At2g05200 putative non-LTR retroelement reverse transcriptase 46 5e-05
At2g04760 putative non-LTR retroelement reverse transcriptase 45 8e-05
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 74.3 bits (181), Expect = 2e-13
Identities = 44/123 (35%), Positives = 73/123 (58%), Gaps = 6/123 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I ++ G +SH+ D ++LF S Q+++I + L+ F EASG KV+L+KS++F S+
Sbjct: 530 IAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHN 589
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + +Q +S GI LGKYL +P L+ R+ KE F ++++ ++ L G K
Sbjct: 590 VSREMEQLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLA------GWK 643
Query: 121 GRA 123
GR+
Sbjct: 644 GRS 646
Score = 51.2 bits (121), Expect = 2e-06
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG-- 420
IWK E+V F+WL ++ + TN++R HL + C C + E + H LR CP
Sbjct: 906 IWKLITPERVRVFIWLVSQNVIMTNVERVRRHLSENAICSVCNGAEETILHVLRDCPAME 965
Query: 421 SVWVR 425
+W R
Sbjct: 966 PIWRR 970
>At2g07650 putative non-LTR retrolelement reverse transcriptase
Length = 732
Score = 69.3 bits (168), Expect = 5e-12
Identities = 43/122 (35%), Positives = 70/122 (57%), Gaps = 6/122 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I +++ G +SH+ D ++LF S Q+++I + L+ F ASG KV+L+KS++F S
Sbjct: 192 ISLSQGGPKLSHICFADDLILFAEASVAQIRVIRRVLERFCVASGQKVSLEKSKIFFSEN 251
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + + +S GI+ LGKYL +P L+ R+ K+ F I++K + L G K
Sbjct: 252 VSRDLGKLISDESGISSTRELGKYLGMPVLQRRINKDTFGDILEKLTTRLA------GWK 305
Query: 121 GR 122
GR
Sbjct: 306 GR 307
Score = 57.4 bits (137), Expect = 2e-08
Identities = 25/65 (38%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG-- 420
+W TA E+V FLWL + A+ TN +R+ HL C C + E + H LR CP
Sbjct: 568 VWSVTAPERVRLFLWLVAQQAIMTNQERHRRHLSTTTICQVCKGAKESIIHVLRDCPAME 627
Query: 421 SVWVR 425
+W+R
Sbjct: 628 GIWLR 632
>At4g08830 putative protein
Length = 947
Score = 68.6 bits (166), Expect = 9e-12
Identities = 43/123 (34%), Positives = 71/123 (56%), Gaps = 6/123 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I +++ G ISH+ D ++LF S Q+++I + L+ F ASG KV+L KS++F S
Sbjct: 344 IGLSQGGPKISHICFADDLILFAEASVSQIRVIRRILETFCIASGQKVSLDKSKIFFSKN 403
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + ++ +S GI LGKYL +P L+ R+ K+ F ++++ +S L G K
Sbjct: 404 VSRDLEKLISKESGIKSTRELGKYLGMPILQRRINKDTFGEVLERVSSRLA------GWK 457
Query: 121 GRA 123
GR+
Sbjct: 458 GRS 460
Score = 30.0 bits (66), Expect = 3.6
Identities = 17/65 (26%), Positives = 24/65 (36%), Gaps = 18/65 (27%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP--G 420
+W+ A E+V FLW H+ C C E + H L+ CP
Sbjct: 696 LWRVVALERVKTFLW----------------HIGDTSVCQVCKGGDETILHVLKDCPSIA 739
Query: 421 SVWVR 425
+W R
Sbjct: 740 GIWRR 744
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 65.5 bits (158), Expect = 8e-11
Identities = 42/122 (34%), Positives = 69/122 (56%), Gaps = 6/122 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I I++ G +SH+ D ++LF S Q++++ L++F ASG KV+L+KS+++ S
Sbjct: 1206 IKISQSGPRLSHICFADDLILFAEASIDQIRVLRGVLEKFCGASGQKVSLEKSKIYFSKN 1265
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + + +S G+ LGKYL +P L+ R+ KE F +I + +S L G K
Sbjct: 1266 VLRDLGKRISEESGMKATKDLGKYLGVPILQKRINKETFGEVIKRFSSRLA------GWK 1319
Query: 121 GR 122
GR
Sbjct: 1320 GR 1321
Score = 56.6 bits (135), Expect = 4e-08
Identities = 26/65 (40%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP--G 420
+WK A E+V FLWL A+ TN +R HL D C C +E + H LR CP
Sbjct: 1582 VWKVQAPERVRVFLWLVVNQAIMTNSERKRRHLCDSDVCQVCRGGIESILHVLRDCPAMS 1641
Query: 421 SVWVR 425
+W R
Sbjct: 1642 GIWDR 1646
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 61.2 bits (147), Expect = 1e-09
Identities = 36/103 (34%), Positives = 52/103 (49%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
ITK+ + ISHL D L F SN ++ + K++ EASG K+N KS + +
Sbjct: 1031 ITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIFGQKIP 1090
Query: 63 QRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
++Q L ++GI GKYL LP+ R E F I+ K
Sbjct: 1091 TMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTK 1133
Score = 32.0 bits (71), Expect = 0.96
Identities = 17/69 (24%), Positives = 30/69 (42%), Gaps = 2/69 (2%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGS- 421
IW+ K+ F+W AL T + + ++ C RC + E + H + C +
Sbjct: 1388 IWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQ 1447
Query: 422 -VWVRSRFA 429
VW + F+
Sbjct: 1448 VVWRSANFS 1456
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 54.3 bits (129), Expect = 2e-07
Identities = 25/68 (36%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Query: 360 MKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
+K IWK A E+V F+WL + TN++R HL C C + E + H LR CP
Sbjct: 644 LKQIWKLVAPERVRVFIWLVSHMVIMTNVERVRRHLSDIATCSVCNGADESILHVLRDCP 703
Query: 420 G--SVWVR 425
+W R
Sbjct: 704 AMTPIWQR 711
Score = 53.1 bits (126), Expect = 4e-07
Identities = 30/82 (36%), Positives = 50/82 (60%)
Query: 29 QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQELSIIVGITRASYLGKYLDLP 88
Q+ +I + L++F EASG V+L+KS++ SN V + ++ +S GI LGKYL +P
Sbjct: 390 QILIIRRVLEQFCEASGQNVSLEKSKIVFSNNVSRDMERLISGESGIGCTRELGKYLGMP 449
Query: 89 QLKARVTKEHFAPIIDKGNSHL 110
L+ R+ KE +++ +S L
Sbjct: 450 ILQKRMNKETSGEVLEHVSSSL 471
>At4g26360 putative protein
Length = 1141
Score = 53.5 bits (127), Expect = 3e-07
Identities = 31/98 (31%), Positives = 54/98 (54%), Gaps = 2/98 (2%)
Query: 8 IGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQ 67
+ ISHL D V++FF + L I +TL++F SG+KVN KS FC+ ++Q ++
Sbjct: 668 LSISHLMFADDVMIFFDGGSSSLHGICETLEDFASWSGLKVNNDKSHFFCAG-LEQAERN 726
Query: 68 ELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
L+ G + +YL LP + ++ + P+++K
Sbjct: 727 SLA-AYGFPQGCLPIRYLGLPLMCRKLRIAEYEPLLEK 763
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 52.4 bits (124), Expect = 7e-07
Identities = 29/90 (32%), Positives = 46/90 (50%)
Query: 4 TKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQ 63
++ G ISHL L+F + + + LK + +ASG VN QKS + +D
Sbjct: 125 SRNGPAISHLLFAHDSLVFCKATLEECMTLVNVLKLYEKASGQAVNFQKSAILFGKGLDF 184
Query: 64 RKQQELSIIVGITRASYLGKYLDLPQLKAR 93
R ++LS ++GI + G+YL LP+ R
Sbjct: 185 RTSEQLSQLLGIYKTEGFGRYLGLPEFVGR 214
Score = 52.0 bits (123), Expect = 9e-07
Identities = 31/76 (40%), Positives = 41/76 (53%), Gaps = 7/76 (9%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG 420
IWK A K+ F W + +ALPT N+KR L+ D C RCG + EDV H L +C
Sbjct: 501 IWKQNAPPKIKHFWWRSAHNALPTAGNLKR--RRLITDDTCQRCGEASEDVNHLLFQCRV 558
Query: 421 S--VWVRSRFAKALPG 434
S +W ++ K PG
Sbjct: 559 SKEIWEQAHI-KLCPG 573
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 50.1 bits (118), Expect = 3e-06
Identities = 33/110 (30%), Positives = 56/110 (50%), Gaps = 17/110 (15%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I I++ G +SH+ D ++LF S Q KV+L+KS++F SN
Sbjct: 482 ISISQGGPKVSHVCFADDLILFAEASVAQ-----------------KVSLEKSKIFFSNN 524
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHL 110
V + + ++ GI LGKYL +P L+ R+ K+ F ++++ +S L
Sbjct: 525 VSRDLEGLITAETGIGSTRELGKYLGMPVLQKRINKDTFGEVLERVSSRL 574
Score = 40.8 bits (94), Expect = 0.002
Identities = 17/43 (39%), Positives = 24/43 (55%)
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFR 403
K IW A E+V FLWL G+ + TN++R H+ + C R
Sbjct: 822 KRIWGVIAPERVRVFLWLVGQQVIMTNVERVRRHIGDIEVCQR 864
>At2g11240 pseudogene
Length = 1044
Score = 48.9 bits (115), Expect = 8e-06
Identities = 30/103 (29%), Positives = 48/103 (46%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
++K ++HL D + F + LK++ EASG +N KS + S
Sbjct: 495 VSKGNPRVNHLLFADDTIFFCRSDLKSCKTFLCILKKYEEASGQMINKSKSAITFSRKTP 554
Query: 63 QRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
+ E I+GI LGKYL LP++ R ++ F I+D+
Sbjct: 555 DHIKTEAQQILGIQLVGGLGKYLGLPKMFGRKKRDLFNQIVDR 597
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 48.9 bits (115), Expect = 8e-06
Identities = 27/101 (26%), Positives = 50/101 (48%)
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
+SHL D L F + Q +I + LK++ SG ++N KS + + V+ + ++
Sbjct: 460 VSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFGHKVEDSIKADI 519
Query: 70 SIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHL 110
+I+GI +G YL LP+ + F+ + D+ S +
Sbjct: 520 KLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRI 560
Score = 38.9 bits (89), Expect = 0.008
Identities = 22/75 (29%), Positives = 32/75 (42%), Gaps = 3/75 (4%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC--PG 420
IWK K+ FLW +P + ++ C CGAS E + H L +C
Sbjct: 828 IWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPAR 887
Query: 421 SVWVRSRFAKALPGL 435
+W S+ A PG+
Sbjct: 888 QIWALSQIPTA-PGI 901
>At1g25430 hypothetical protein
Length = 1213
Score = 48.5 bits (114), Expect = 1e-05
Identities = 27/98 (27%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Query: 8 IGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQ 67
+ ISHL D V++FF + L I +TL +F SG+KVN KS ++ + + + +
Sbjct: 690 LSISHLMFADDVMIFFDGGSFSLHGICETLDDFASWSGLKVNKDKSHLYLAGL--NQLES 747
Query: 68 ELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
+ G + +YL LP + ++ + P+++K
Sbjct: 748 NANAAYGFPIGTLPIRYLGLPLMNRKLRIAEYEPLLEK 785
Score = 32.0 bits (71), Expect = 0.96
Identities = 27/96 (28%), Positives = 42/96 (43%), Gaps = 11/96 (11%)
Query: 338 WSLHRTIC-----VQPFE*LRHPKFAVMKL---IWKATAAEKVMFFLWLAGRDALPTNMK 389
WS++ +C + +E +R PK V IW A K F +W++ + L T +
Sbjct: 1023 WSVNGFLCQGFSAAKTWEAIR-PKATVKSWASSIWFKGAVPKYAFNMWVSHLNRLLTRQR 1081
Query: 390 RYHSHLVKFDACFRCGASVEDVEHALRRCPGS--VW 423
++ DAC C + E +H L C S VW
Sbjct: 1082 LASWGHIQSDACVLCSFASESRDHLLLICEFSAQVW 1117
>At2g22350 putative non-LTR retroelement reverse transcriptase
Length = 321
Score = 47.8 bits (112), Expect = 2e-05
Identities = 23/60 (38%), Positives = 30/60 (49%)
Query: 368 AAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGSVWVRSR 427
A E+V FLWL + + TN++RY HL C C E + H LR CP + SR
Sbjct: 2 APERVRVFLWLVVQQVIITNVERYRRHLSDTRVCQICQGGEETILHVLRDCPAMAGIWSR 61
>At4g02490
Length = 657
Score = 47.4 bits (111), Expect = 2e-05
Identities = 28/99 (28%), Positives = 53/99 (53%), Gaps = 1/99 (1%)
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
I+HL D +L+F S L I L F + SG+ +NLQK+ + +R + +
Sbjct: 239 ITHLSFADDILVFCDGSLSSLVAILDILDVFKKGSGLGINLQKTALLLDGGNFER-NRIM 297
Query: 70 SIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNS 108
+ +G+++ S +YL +P + ++ K + P++D+ NS
Sbjct: 298 AASLGVSQGSLPVRYLGVPLMSQKMKKHDYQPLVDRINS 336
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 47.4 bits (111), Expect = 2e-05
Identities = 32/109 (29%), Positives = 43/109 (39%), Gaps = 6/109 (5%)
Query: 321 TFYADGGYQVTLERD*RWSLHRTICVQPFE*LRHPKFAVMKL-IWKATAAEKVMFFLWLA 379
+F G Y V L R C PF+ P + ++ +WK K F W
Sbjct: 886 SFTKSGNYTVKSGYWAARDLSRPTCDLPFQ---GPSVSALQAQVWKIKTTRKFKHFEWQC 942
Query: 380 GRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGS--VWVRS 426
L TN + + H+ C RCGA E + H L CP S +W S
Sbjct: 943 LSGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSRQIWALS 991
Score = 33.1 bits (74), Expect = 0.43
Identities = 21/73 (28%), Positives = 34/73 (45%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
I++ SHL D L F Q + I L+ + EASG ++N KS + + VD
Sbjct: 616 ISRASPSTSHLLFADDSLFFCKADPLQGKEIIDILRLYGEASGQQLNPDKSSVMFGHEVD 675
Query: 63 QRKQQELSIIVGI 75
+ + + +GI
Sbjct: 676 NSIRNTIKVSLGI 688
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 47.0 bits (110), Expect = 3e-05
Identities = 27/99 (27%), Positives = 51/99 (51%), Gaps = 1/99 (1%)
Query: 6 EGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRK 65
E IG++HL D +++F ++ + KEF SG++++L+KS ++ + V +
Sbjct: 991 EKIGLTHLCFADDLMVFVDGHQWSIEGVINVFKEFAGRSGLQISLEKSTIYLAGVSASDR 1050
Query: 66 QQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIID 104
Q LS +YL LP L ++T ++P+I+
Sbjct: 1051 VQTLSSF-PFANGQLPVRYLGLPLLTKQMTTADYSPLIE 1088
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 46.6 bits (109), Expect = 4e-05
Identities = 30/108 (27%), Positives = 53/108 (48%), Gaps = 2/108 (1%)
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
++HL D + F + ++ LK++ ASG +NL KS + S+ Q ++ +
Sbjct: 617 VNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSAITFSSKTPQDIKRRV 676
Query: 70 SIIVGITRASYLGKYLDLPQLKARVTKEHFAPIID--KGNSHLGRISF 115
+ + I +GKYL LP+ R ++ F+ I+D + SH I F
Sbjct: 677 KLSLRIDNEGGIGKYLGLPEHFGRRKRDIFSSIVDRIRQRSHSWSIRF 724
Score = 37.0 bits (84), Expect = 0.030
Identities = 22/63 (34%), Positives = 27/63 (41%), Gaps = 1/63 (1%)
Query: 357 FAVMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALR 416
F K IWK + KV FLW A + ALP ++ C RCG E H +
Sbjct: 968 FNWQKNIWKIHTSPKVKHFLWKAMKGALPVGEALSRRNIEAEVTCKRCG-QTESSLHLML 1026
Query: 417 RCP 419
CP
Sbjct: 1027 LCP 1029
>At4g29090 putative protein
Length = 575
Score = 46.2 bits (108), Expect = 5e-05
Identities = 21/60 (35%), Positives = 31/60 (51%)
Query: 359 VMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC 418
+ + IWK+ + K+ FLW ++LP + HL K AC RC + E V H L +C
Sbjct: 253 IYQKIWKSQTSPKIQHFLWKCLSNSLPVAGALAYRHLSKESACIRCPSCKETVNHLLFKC 312
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 46.2 bits (108), Expect = 5e-05
Identities = 27/103 (26%), Positives = 49/103 (47%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
++ G ++HL D + F +++ L + +ASG +N KS + S+
Sbjct: 564 VSINGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTP 623
Query: 63 QRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
+ + ++ I+ I + GKYL LP+ R ++ F IIDK
Sbjct: 624 RSVKGQVKRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDK 666
Score = 40.4 bits (93), Expect = 0.003
Identities = 20/63 (31%), Positives = 27/63 (42%), Gaps = 3/63 (4%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC--PG 420
+WK K+ +W A +ALP ++ H+ AC RCGA E H C
Sbjct: 934 LWKLQTLPKIKHLMWKAAMEALPVGIQLVRRHISPSAACHRCGAP-ESTTHLFFHCEFAA 992
Query: 421 SVW 423
VW
Sbjct: 993 QVW 995
>At2g04760 putative non-LTR retroelement reverse transcriptase
Length = 631
Score = 45.4 bits (106), Expect = 8e-05
Identities = 24/98 (24%), Positives = 49/98 (49%), Gaps = 1/98 (1%)
Query: 8 IGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQ 67
+ ++HL VD +++F ++ + EF SG+ ++L+KS ++ + V + +
Sbjct: 266 LSLTHLCFVDDLMVFIDGQQRSIEGVINIFHEFAGKSGLHISLEKSTLYLAGVSEPNRDH 325
Query: 68 ELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
LS +YL LP + ++T ++P+IDK
Sbjct: 326 ILSAF-SFASGQLPVRYLGLPLMTKQMTTADYSPLIDK 362
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.339 0.147 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,970,193
Number of Sequences: 26719
Number of extensions: 462918
Number of successful extensions: 1341
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 52
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 1213
Number of HSP's gapped (non-prelim): 122
length of query: 573
length of database: 11,318,596
effective HSP length: 105
effective length of query: 468
effective length of database: 8,513,101
effective search space: 3984131268
effective search space used: 3984131268
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0369c.1


