

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369b.1
(266 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
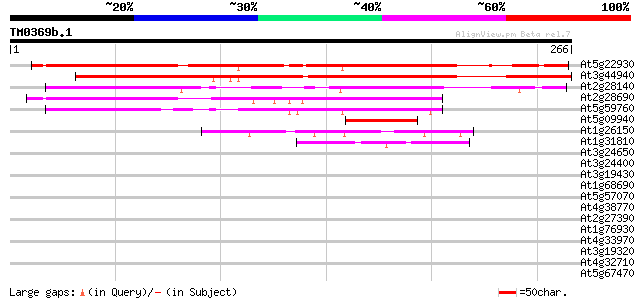
Score E
Sequences producing significant alignments: (bits) Value
At5g22930 putative protein 233 8e-62
At3g44940 putative protein 207 5e-54
At2g28140 unknown protein 186 1e-47
At2g28690 unknown protein 115 2e-26
At5g59760 putative protein 101 4e-22
At5g09940 putative protein 49 4e-06
At1g26150 Pto kinase interactor, putative 45 5e-05
At1g31810 hypothetical protein 42 4e-04
At3g24650 abscisic acid-insensitive protein 3 39 0.002
At3g24400 protein kinase, putative 39 0.003
At3g19430 putative late embryogenesis abundant protein 39 0.003
At1g68690 protein kinase, putative 39 0.003
At5g57070 putative protein 39 0.004
At4g38770 extensin - like protein 39 0.004
At2g27390 pseudogene 39 0.004
At1g76930 extensin 39 0.004
At4g33970 extensin-like protein 38 0.005
At3g19320 unknown protein 38 0.005
At4g32710 putative protein kinase 38 0.006
At5g67470 formin-like protein 37 0.008
>At5g22930 putative protein
Length = 248
Score = 233 bits (594), Expect = 8e-62
Identities = 138/260 (53%), Positives = 172/260 (66%), Gaps = 39/260 (15%)
Query: 11 CAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERDEAQ 70
C Y F ++ EE+RQSL+YTTLEL+QT+ EE+RKRD+QL++LKD+L TI+ERDEA
Sbjct: 22 CFYDFL-QTTEEIRQSLLYTTLELDQTKMFAYEEIRKRDEQLIHLKDILTKTIKERDEAL 80
Query: 71 EKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEP--RRGIDSNNGLSSSDCEESIVS 128
EKCQRL+ + H QQQ+ P SG SSIEDE + + SN SSSDCEESI+S
Sbjct: 81 EKCQRLMFDS----HSLQQQKHMTPPLSGASSIEDEQVQPQQLRSNKSFSSSDCEESIMS 136
Query: 129 SPVFDNMPQPQFPLPLPSVAQSMIELTP---EKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
D++ P P L V+ + I + P +KPLPEKGKLLQAV+KAGPLLQTLLLAGP
Sbjct: 137 PT--DHIMNPP-PCQLEEVSGTEIIMDPLLQDKPLPEKGKLLQAVIKAGPLLQTLLLAGP 193
Query: 186 LPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQCGRVSRK 245
LPQWRHPPPPL+SFEIPPVT+ P + NG CG+ +RK
Sbjct: 194 LPQWRHPPPPLKSFEIPPVTVQCPIVN---------------NG---------CGKFNRK 229
Query: 246 RIFCDGSDSPSETKFQRVVL 265
R+F DG S SE K+Q+V+L
Sbjct: 230 RVFSDG--SYSEAKYQKVLL 247
>At3g44940 putative protein
Length = 220
Score = 207 bits (527), Expect = 5e-54
Identities = 123/245 (50%), Positives = 152/245 (61%), Gaps = 35/245 (14%)
Query: 32 LELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERDEAQEKCQRLLLEKFLFQHQQQQQQ 91
+ELEQT+ EELRKRD+QL++L+D+L T++ERDEA EK LLL L Q +QQQ Q
Sbjct: 1 MELEQTKLVAHEELRKRDEQLIHLEDVLTKTLKERDEALEKYNHLLLNNLLLQQKQQQNQ 60
Query: 92 QQAD-----PNSGVSSI-EDEP----RRGIDSNNGLSSSDCEESIVSSPVFDNMPQPQFP 141
Q P SG SSI EDE + ++SN SSSD EESI+S V D + Q
Sbjct: 61 NQKQELVTPPLSGASSIIEDEQVQPQQPQLNSNKSFSSSDTEESIMSPSVIDPVTMNQ-- 118
Query: 142 LPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEI 201
S + + L P+KPLPEKGKLLQAV+KAGPLLQTLLLAGPLPQWRHPPPPLE+ EI
Sbjct: 119 QIEVSGDEMLATLLPDKPLPEKGKLLQAVIKAGPLLQTLLLAGPLPQWRHPPPPLETSEI 178
Query: 202 PPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQCGRVSRKRIFCDGSDSPSETKFQ 261
PPVTIP P Q ++ CG ++KR F ++ SETK+Q
Sbjct: 179 PPVTIPLPQFQ-----------------------NNGCGNSNKKRAFSISDETYSETKYQ 215
Query: 262 RVVLH 266
+V+LH
Sbjct: 216 KVLLH 220
>At2g28140 unknown protein
Length = 211
Score = 186 bits (471), Expect = 1e-47
Identities = 120/250 (48%), Positives = 148/250 (59%), Gaps = 63/250 (25%)
Query: 18 KSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERDEAQEKCQRLL 77
K++EE+R SL++TT+ELEQTR EEL +DDQ++ LKDLLN I+E+DEAQE+ +R+L
Sbjct: 21 KTIEELRHSLLHTTMELEQTRMVASEELIAKDDQIMQLKDLLNKAIKEKDEAQERYKRIL 80
Query: 78 LEK-FLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSSDCEESIVSSPVFDNMP 136
L++ FL QHQ ++Q DPN NG SSD EESIVSS
Sbjct: 81 LDQNFLLQHQTDEEQ---DPNI----------------NGFHSSDGEESIVSS------- 114
Query: 137 QPQFPLPLPSVAQSMIELT-PEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
F P+ IEL PE LPEKGKLL+AV+KAGPLLQTLLLAG LPQWRHPPP
Sbjct: 115 ---FEPPVE------IELDFPEMTLPEKGKLLKAVVKAGPLLQTLLLAGQLPQWRHPPPQ 165
Query: 196 LESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQCG-RVSRKRIFCDGSDS 254
LESFEIPPV I S + CG ++RKR+ C DS
Sbjct: 166 LESFEIPPVII----------------------AEGSPLSQDSCGNNLNRKRVHC---DS 200
Query: 255 PSETKFQRVV 264
ETK+QR++
Sbjct: 201 KRETKYQRLL 210
>At2g28690 unknown protein
Length = 231
Score = 115 bits (289), Expect = 2e-26
Identities = 77/205 (37%), Positives = 114/205 (55%), Gaps = 24/205 (11%)
Query: 9 LNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERDE 68
L +F+Q +SM+E+RQ L Y++ ELE + EE + +++ NL LL +ERDE
Sbjct: 4 LGSIWFYQ-ESMDELRQKLQYSSFELEAVKAKANEETKLHQEEVKNLLHLLKLARQERDE 62
Query: 69 AQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSN-NGLSSSDCEE-SI 126
A+++ Q+LL K NS ++ +DS +SSS+ ++
Sbjct: 63 AKDQLQKLLTIK---------------TNSSITESNSHGSSPVDSFFEPVSSSEFSNFNM 107
Query: 127 VSSPV----FDNMPQ--PQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTL 180
V P+ F N Q + + V M E+ K LPEKGKLLQ VM++GPLLQTL
Sbjct: 108 VPEPINQVKFKNRQQLVNRNLKKIDPVDALMDEIIKGKALPEKGKLLQTVMESGPLLQTL 167
Query: 181 LLAGPLPQWRHPPPPLESFEIPPVT 205
L+AGPLP+WR+PPP +SF +PP++
Sbjct: 168 LVAGPLPRWRNPPPLQQSFRVPPIS 192
>At5g59760 putative protein
Length = 221
Score = 101 bits (252), Expect = 4e-22
Identities = 74/199 (37%), Positives = 109/199 (54%), Gaps = 33/199 (16%)
Query: 18 KSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERDEAQEKCQRLL 77
+S++E+RQ+L T ELE + E+ R +++ L +LL T +ERDEA+++
Sbjct: 2 QSIDEMRQNLQSTLFELETLKMEANEKSRTHREEVNQLLNLLKFTQQERDEARQQ----- 56
Query: 78 LEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSSDCEESIVSSPV--FDNM 135
L +F+F QQ +PNS R + +N S S S SS + F N+
Sbjct: 57 LSQFIFHTQQ-------NPNS----------RSLTKSNSFSHSQDVSSSSSSELSTFLNI 99
Query: 136 -PQPQFPLPLPSVAQSMIELTP------EKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ 188
PQP L P+ ++ P K PE GKLL+AV++AGPLL+TLLLAGPLP+
Sbjct: 100 HPQPLTNLEDPTAHNHHQQIDPLDVLVMGKAFPETGKLLKAVVEAGPLLETLLLAGPLPK 159
Query: 189 WRHPPPPLES--FEIPPVT 205
W +PPP +S FE+P ++
Sbjct: 160 WINPPPQTQSQRFELPSIS 178
>At5g09940 putative protein
Length = 213
Score = 48.5 bits (114), Expect = 4e-06
Identities = 18/34 (52%), Positives = 27/34 (78%)
Query: 160 LPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPP 193
LPE GK QA+++AG L+ +L + GP+P+WR+PP
Sbjct: 140 LPENGKFQQAIIEAGSLVDSLFITGPIPKWRNPP 173
>At1g26150 Pto kinase interactor, putative
Length = 760
Score = 44.7 bits (104), Expect = 5e-05
Identities = 43/138 (31%), Positives = 60/138 (43%), Gaps = 19/138 (13%)
Query: 92 QQADPNSGVSSIEDEPRRGID---SNNGLSSSDCEESIVSSPVFDNMPQPQFPLP-LPSV 147
Q + P +S EP G +N SS E+ +SSP P+P P P L
Sbjct: 30 QPSFPGDNATSPTREPTNGNPPETTNTPAQSSPPPETPLSSPP----PEPSPPSPSLTGP 85
Query: 148 AQSMIELTPE-KPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP-LESFEIPPVT 205
+ I ++P +P P +A A P+ + P P+ PPPP E+ P+T
Sbjct: 86 PPTTIPVSPPPEPSPPPPLPTEAPPPANPV------SSPPPESSPPPPPPTEAPPTTPIT 139
Query: 206 IPSPPLQ---PPQQPPQL 220
PSPP PP+ PP L
Sbjct: 140 SPSPPTNPPPPPESPPSL 157
Score = 38.5 bits (88), Expect = 0.004
Identities = 38/128 (29%), Positives = 47/128 (36%), Gaps = 12/128 (9%)
Query: 91 QQQADPNSGVSSIEDEPRRGIDSNNGLSSSDCEESIVSSPVFDNMPQPQFPLPLPSVAQS 150
Q P + +SS EP S G + VS P P+P P PLP+ A
Sbjct: 59 QSSPPPETPLSSPPPEPSPPSPSLTGPPPTTIP---VSPP-----PEPSPPPPLPTEAPP 110
Query: 151 MIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPP 210
P PE +A P T + P P PPPP +P PS P
Sbjct: 111 PANPVSSPP-PESSPPPPPPTEAPP---TTPITSPSPPTNPPPPPESPPSLPAPDPPSNP 166
Query: 211 LQPPQQPP 218
L PP+ P
Sbjct: 167 LPPPKLVP 174
Score = 32.7 bits (73), Expect = 0.21
Identities = 31/97 (31%), Positives = 37/97 (37%), Gaps = 5/97 (5%)
Query: 127 VSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLP--EKGKLLQAV-MKAGPLLQTLLLA 183
VSSP ++ P P P P P P P E L A + PL L+
Sbjct: 115 VSSPPPESSPPPPPPTEAPPTTPITSPSPPTNPPPPPESPPSLPAPDPPSNPLPPPKLVP 174
Query: 184 GPLPQWRH-PPPPLESFEIPPVTIPSPPL-QPPQQPP 218
RH P PP PP +PSPP + P PP
Sbjct: 175 PSHSPPRHLPSPPASEIPPPPRHLPSPPASERPSTPP 211
Score = 30.4 bits (67), Expect = 1.0
Identities = 31/94 (32%), Positives = 41/94 (42%), Gaps = 10/94 (10%)
Query: 127 VSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPL 186
++SP +P PQ P + E T P PE ++ P +T L + P
Sbjct: 18 LASPPLMALPPPQPSFPGDNATSPTREPTNGNP-PET---TNTPAQSSPPPETPL-SSPP 72
Query: 187 PQWRHPPPPLESFE-IPPVTIP-SPPLQPPQQPP 218
P+ P PP S PP TIP SPP +P PP
Sbjct: 73 PE---PSPPSPSLTGPPPTTIPVSPPPEPSPPPP 103
>At1g31810 hypothetical protein
Length = 1201
Score = 41.6 bits (96), Expect = 4e-04
Identities = 30/85 (35%), Positives = 36/85 (42%), Gaps = 7/85 (8%)
Query: 137 QPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLL---QTLLLAGPLPQWRHPP 193
QP P P P + S +P +P P L + PL Q + P P PP
Sbjct: 524 QPPPPPPPPPLFTSTTSFSPSQPPPPPP--LPSFSNRDPLTTLHQPINKTPPPPP--PPP 579
Query: 194 PPLESFEIPPVTIPSPPLQPPQQPP 218
PPL S IPP PP +PP PP
Sbjct: 580 PPLPSRSIPPPLAQPPPPRPPPPPP 604
Score = 34.3 bits (77), Expect = 0.071
Identities = 30/86 (34%), Positives = 33/86 (37%), Gaps = 13/86 (15%)
Query: 137 QPQFPLPLPSVAQ--------SMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ 188
QP P PLPS + I TP P P L + PL Q P P
Sbjct: 545 QPPPPPPLPSFSNRDPLTTLHQPINKTPPPPPPPPPPLPSRSIPP-PLAQPPPPRPPPP- 602
Query: 189 WRHPPPPLESFEIPPVTIPSPPLQPP 214
PPPP S IP + P PP PP
Sbjct: 603 ---PPPPPSSRSIPSPSAPPPPPPPP 625
Score = 32.7 bits (73), Expect = 0.21
Identities = 32/115 (27%), Positives = 40/115 (33%), Gaps = 28/115 (24%)
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P P PLP S+ + + P +P P + P + P P PPPP
Sbjct: 576 PPPPPPLPSRSIPPPLAQPPPPRPPPPPPPPPSSRSIPSP-------SAPPP----PPPP 624
Query: 196 LESF---------EIPPVTIPSPPLQ--------PPQQPPQLLHQESFFNGSSST 233
SF + PP P PP + PP PP H S G ST
Sbjct: 625 PPSFGSTGNKRQAQPPPPPPPPPPTRIPAAKCAPPPPPPPPTSHSGSIRVGPPST 679
Score = 30.8 bits (68), Expect = 0.78
Identities = 28/89 (31%), Positives = 31/89 (34%), Gaps = 14/89 (15%)
Query: 141 PLPLPSVAQSM---IELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLE 197
PL LPS S + L P P P L + P + P P PPPP
Sbjct: 464 PLNLPSDPPSSGDHVTLLPPPPPPPPPPLFTSTTSFSP-------SQPPP----PPPPPP 512
Query: 198 SFEIPPVTIPSPPLQPPQQPPQLLHQESF 226
F PS P PP PP SF
Sbjct: 513 LFMSTTSFSPSQPPPPPPPPPLFTSTTSF 541
Score = 29.3 bits (64), Expect = 2.3
Identities = 28/99 (28%), Positives = 32/99 (32%), Gaps = 25/99 (25%)
Query: 137 QPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPL 196
QP P P P + S +P +P P PL + P PPPPL
Sbjct: 503 QPPPPPPPPPLFMSTTSFSPSQPPPPP--------PPPPLFTSTTSFSPSQP--PPPPPL 552
Query: 197 ESFEI---------------PPVTIPSPPLQPPQQPPQL 220
SF PP P PPL PP L
Sbjct: 553 PSFSNRDPLTTLHQPINKTPPPPPPPPPPLPSRSIPPPL 591
Score = 28.9 bits (63), Expect = 3.0
Identities = 29/99 (29%), Positives = 36/99 (36%), Gaps = 37/99 (37%)
Query: 134 NMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPP 193
N P+P P PLP + + + P P P PL +T P P PP
Sbjct: 694 NAPKPPAPPPLPP-SSTRLGAPPPPPPP-------------PLSKT-----PAP----PP 730
Query: 194 PPLESFEIPP--------VTIPSPPL------QPPQQPP 218
PPL +PP + PPL PP PP
Sbjct: 731 PPLSKTPVPPPPPGLGRGTSSGPPPLGAKGSNAPPPPPP 769
Score = 28.5 bits (62), Expect = 3.9
Identities = 25/78 (32%), Positives = 29/78 (37%), Gaps = 17/78 (21%)
Query: 141 PLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFE 200
P P P ++ I P+ P P P T L A P P PPPPL
Sbjct: 681 PPPPPPPPKANISNAPKPPAPPPL----------PPSSTRLGAPPPP----PPPPLSKTP 726
Query: 201 IPPVTIPSPPLQPPQQPP 218
PP P P + P PP
Sbjct: 727 APP---PPPLSKTPVPPP 741
>At3g24650 abscisic acid-insensitive protein 3
Length = 720
Score = 39.3 bits (90), Expect = 0.002
Identities = 55/256 (21%), Positives = 97/256 (37%), Gaps = 55/256 (21%)
Query: 27 LVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERD------------EAQEKCQ 74
++ ++LE + AV E K + + ++ +DL I++ EA ++
Sbjct: 235 MMNSSLEQDDDLAAVFLEWLKNNKETVSAEDLRKVKIKKATIESAARRLGGGKEAMKQLL 294
Query: 75 RLLLEKFLFQHQQQQQQQQADPN-SGVSSIEDEPRRGIDSNNGL----SSSDC------- 122
+L+LE H Q+++ N S S + +P + + NN S C
Sbjct: 295 KLILEWVQTNHLQRRRTTTTTTNLSYQQSFQQDPFQNPNPNNNNLIPPSDQTCFSPSTWV 354
Query: 123 -----EESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLL 177
+++ VS P F MP P +P P P ++E P P P +
Sbjct: 355 PPPPQQQAFVSDPGFGYMPAPNYP-PQPEFL-PLLESPPSWPPPPQ-------------- 398
Query: 178 QTLLLAGPLPQWRHPPPPLESFEI--PPVTIPS---PPLQPPQQPPQLLHQESFFNGSSS 232
+GP+P + P PP + P P Q P P + + SS
Sbjct: 399 -----SGPMPHQQFPMPPTSQYNQFGDPTGFNGYNMNPYQYPYVPAGQMRDQRLLRLCSS 453
Query: 233 TSTHSQCGRVSRKRIF 248
+ ++ R++R+R F
Sbjct: 454 ATKEARKKRMARQRRF 469
>At3g24400 protein kinase, putative
Length = 694
Score = 38.9 bits (89), Expect = 0.003
Identities = 31/88 (35%), Positives = 36/88 (40%), Gaps = 15/88 (17%)
Query: 136 PQPQFPLP--LPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPP 193
P P P P PS LTP P A+ + PL + L P P PP
Sbjct: 85 PSPTTPSPPLTPSPTTPSPPLTPSPP--------PAITPSPPLTPSPL--PPSPTTPSPP 134
Query: 194 PPLESFEIPPVT---IPSPPLQPPQQPP 218
PP S PP+T PS PL+P PP
Sbjct: 135 PPSPSIPSPPLTPSPPPSSPLRPSSPPP 162
Score = 37.7 bits (86), Expect = 0.006
Identities = 32/93 (34%), Positives = 38/93 (40%), Gaps = 14/93 (15%)
Query: 136 PQPQFPLPLPSVAQSM--------IELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLP 187
P PQ PLP+P Q + L P P P L + P T+ P+P
Sbjct: 13 PPPQ-PLPIPPPPQPLPVTPPPPPTALPPALPPPPPPTALPPALPPPPPPTTV---PPIP 68
Query: 188 -QWRHPPPPLESFEIPPV-TIPSPPLQPPQQPP 218
PPPPL +PP T PSPPL P P
Sbjct: 69 PSTPSPPPPLTPSPLPPSPTTPSPPLTPSPTTP 101
Score = 28.9 bits (63), Expect = 3.0
Identities = 26/100 (26%), Positives = 35/100 (35%), Gaps = 9/100 (9%)
Query: 126 IVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
+ SP+ + P P P PS+ + +P P + A P T + P
Sbjct: 119 LTPSPLPPSPTTPSPPPPSPSIPSPPLTPSPPPSSPLRPSSPPPPSPATP--STPPRSPP 176
Query: 186 LPQWRHPPPPLESFEIPPVTIPS-------PPLQPPQQPP 218
P PPP + S PP PS P P PP
Sbjct: 177 PPSTPTPPPRVGSLSPPPPASPSGGRSPSTPSTTPGSSPP 216
Score = 27.7 bits (60), Expect = 6.6
Identities = 15/39 (38%), Positives = 18/39 (45%), Gaps = 2/39 (5%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPS--PPLQPPQQPPQLL 221
P PQ PPP + + P P+ PP PP PP L
Sbjct: 13 PPPQPLPIPPPPQPLPVTPPPPPTALPPALPPPPPPTAL 51
>At3g19430 putative late embryogenesis abundant protein
Length = 550
Score = 38.9 bits (89), Expect = 0.003
Identities = 30/95 (31%), Positives = 37/95 (38%), Gaps = 8/95 (8%)
Query: 127 VSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPL 186
V SP P P P P PSV ++P P P +V P + T + P
Sbjct: 126 VPSPTPPVSPPP--PTPTPSVPSPTPPVSPPPPTPTP-----SVPSPTPPVPTDPMPSPP 178
Query: 187 PQWRHPPP-PLESFEIPPVTIPSPPLQPPQQPPQL 220
P PPP P S PP P+PP PP +
Sbjct: 179 PPVSPPPPTPTPSVPSPPDVTPTPPTPSVPSPPDV 213
Score = 34.3 bits (77), Expect = 0.071
Identities = 25/89 (28%), Positives = 31/89 (34%), Gaps = 4/89 (4%)
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQ----TLLLAGPLPQWRH 191
P P P P P V+ TP P P + P + T + P P
Sbjct: 140 PTPSVPSPTPPVSPPPPTPTPSVPSPTPPVPTDPMPSPPPPVSPPPPTPTPSVPSPPDVT 199
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPPQL 220
P PP S PP P+PP PP +
Sbjct: 200 PTPPTPSVPSPPDVTPTPPTPSVPSPPDV 228
Score = 31.2 bits (69), Expect = 0.60
Identities = 30/112 (26%), Positives = 36/112 (31%), Gaps = 25/112 (22%)
Query: 107 PRRGIDSNNGLSSSDCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKL 166
P G D S D +PV P P P PSV ++P P P
Sbjct: 54 PSPGDDGGGDDSGGDDGGYTPPAPVPPVSPPP----PTPSVPSPTPPVSPPPPTP----- 104
Query: 167 LQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
T + P P PPP P V P+PP+ PP P
Sbjct: 105 ------------TPSVPSPTPPVSPPPPT----PTPSVPSPTPPVSPPPPTP 140
Score = 30.4 bits (67), Expect = 1.0
Identities = 21/85 (24%), Positives = 31/85 (35%), Gaps = 4/85 (4%)
Query: 134 NMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPP 193
++P P P+P + ++P P P + P ++ P P P
Sbjct: 161 SVPSPTPPVPTDPMPSPPPPVSPPPPTPTPSVPSPPDVTPTPPTPSV----PSPPDVTPT 216
Query: 194 PPLESFEIPPVTIPSPPLQPPQQPP 218
PP S PP P+PP P P
Sbjct: 217 PPTPSVPSPPDVTPTPPTPPSVPTP 241
Score = 30.0 bits (66), Expect = 1.3
Identities = 28/99 (28%), Positives = 38/99 (38%), Gaps = 19/99 (19%)
Query: 133 DNMPQPQFPL------PLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPL 186
D MP P P+ P PSV S ++TP P P P + P
Sbjct: 172 DPMPSPPPPVSPPPPTPTPSVP-SPPDVTPTPPTPSV-----------PSPPDVTPTPPT 219
Query: 187 PQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQES 225
P PP + PP ++P+P PP PP +E+
Sbjct: 220 PSVPSPPDVTPTPPTPP-SVPTPSGSPPYVPPPSDEEEA 257
>At1g68690 protein kinase, putative
Length = 708
Score = 38.9 bits (89), Expect = 0.003
Identities = 30/107 (28%), Positives = 44/107 (41%), Gaps = 9/107 (8%)
Query: 118 SSSDCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLL 177
S+S + ++ SP P P PLPS + P +P P L+++ P +
Sbjct: 93 STSPPPQPVIPSPPPSASPPPALVPPLPSSPPPPASVPPPRPSPSPPILVRS---PPPSV 149
Query: 178 QTLLLAGPLPQWRH------PPPPLESFEIPPVTIPSPPLQPPQQPP 218
+ + P P R P PP E P + PSPP + P Q P
Sbjct: 150 RPIQSPPPPPSDRPTQSPPPPSPPSPPSERPTQSPPSPPSERPTQSP 196
Score = 37.7 bits (86), Expect = 0.006
Identities = 30/107 (28%), Positives = 43/107 (40%), Gaps = 6/107 (5%)
Query: 118 SSSDCEESIVSSPVFDNMPQPQFPLPLP----SVAQSMIELTPEKPLPEKGKLLQAVMKA 173
SSS + ++ SP P PQ +P P S +++ P P P +
Sbjct: 78 SSSPPPQPVIPSPPPSTSPPPQPVIPSPPPSASPPPALVPPLPSSPPPPASVPPPRPSPS 137
Query: 174 GPLLQTLLLAGPLPQWRHPPPPLE--SFEIPPVTIPSPPLQPPQQPP 218
P+L P PPPP + + PP + PSPP + P Q P
Sbjct: 138 PPILVRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPSPPSERPTQSP 184
Score = 33.1 bits (74), Expect = 0.16
Identities = 29/93 (31%), Positives = 37/93 (39%), Gaps = 7/93 (7%)
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTL--LLAGPLP 187
PV + P P P P S + + P P P Q V+ + P + L PLP
Sbjct: 62 PVANGAPPPPLPKPPESSSPPPQPVIPSPP-PSTSPPPQPVIPSPPPSASPPPALVPPLP 120
Query: 188 QWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQL 220
PPPP PP PSPP+ PP +
Sbjct: 121 S--SPPPPASV--PPPRPSPSPPILVRSPPPSV 149
Score = 32.0 bits (71), Expect = 0.35
Identities = 14/31 (45%), Positives = 16/31 (51%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQ 215
P P+ P PP S PP T SPP PP+
Sbjct: 215 PPPEDTKPQPPRRSPNSPPPTFSSPPRSPPE 245
Score = 31.6 bits (70), Expect = 0.46
Identities = 30/96 (31%), Positives = 38/96 (39%), Gaps = 16/96 (16%)
Query: 127 VSSPVFDNMPQP-QFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
V+SP+ + P P + P P P V S + P P PL + + P
Sbjct: 35 VTSPLPPSAPPPNRAPPPPPPVTTSPPPVANGAPPP-------------PLPKPPESSSP 81
Query: 186 LPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLL 221
PQ P PP + P IPSPP P PP L
Sbjct: 82 PPQPVIPSPPPSTSPPPQPVIPSPP--PSASPPPAL 115
Score = 30.8 bits (68), Expect = 0.78
Identities = 29/111 (26%), Positives = 39/111 (35%), Gaps = 17/111 (15%)
Query: 128 SSPVFDNMPQP-----QFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLL 182
S P+ P P Q P P PS + P P P + Q+ T
Sbjct: 137 SPPILVRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPSPPSERPTQSPPSPPSERPTQSP 196
Query: 183 AGPLP--------QWRHPPPPLESFEIPPVTIPSPP----LQPPQQPPQLL 221
P P PPPP ++ PP P+ P PP+ PP++L
Sbjct: 197 PPPSPPSPPSDRPSQSPPPPPEDTKPQPPRRSPNSPPPTFSSPPRSPPEIL 247
Score = 30.4 bits (67), Expect = 1.0
Identities = 19/51 (37%), Positives = 23/51 (44%), Gaps = 6/51 (11%)
Query: 175 PLLQTLLLAGPLPQWRHPPPPLESFEIPPVT--IPSPPLQPPQQ----PPQ 219
P+ L + P P PPPP + PPV P PPL P + PPQ
Sbjct: 34 PVTSPLPPSAPPPNRAPPPPPPVTTSPPPVANGAPPPPLPKPPESSSPPPQ 84
Score = 30.0 bits (66), Expect = 1.3
Identities = 17/38 (44%), Positives = 19/38 (49%), Gaps = 6/38 (15%)
Query: 187 PQWRHPPPPLESFEI----PPVTIPSPPLQPP--QQPP 218
P PPPPL + PPVT P PP PP + PP
Sbjct: 14 PPVTSPPPPLNNATSPATPPPVTSPLPPSAPPPNRAPP 51
>At5g57070 putative protein
Length = 607
Score = 38.5 bits (88), Expect = 0.004
Identities = 51/217 (23%), Positives = 84/217 (38%), Gaps = 42/217 (19%)
Query: 43 EELRKRDDQLLNLKDLLNNTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVS- 101
++ R +D L+ T E +E++ K + ++ F+ + QQ A P
Sbjct: 201 DKYRSQDSSSYQQFQNLSKTEIEEEESEPK--EIQIDTFVVKPSSPPQQPPATPPPPPPP 258
Query: 102 ---SIEDEPRRGIDSNNGLSSSDCEESIVSSPV-----FDNMPQPQFPLPLPSVAQSMIE 153
+ +PRR ++ + + D +E+ S F P P P P P Q +I
Sbjct: 259 PPVEVPQKPRR---THRSVRNRDLQENAKRSETKFKRTFQPPPSPP-PPPPPPPPQPLIA 314
Query: 154 LTPEKP---------------------LPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHP 192
TP + L +GK + + K+ + + + P+ +
Sbjct: 315 ATPPRKQGTLQRRKSNAAKEIKMVFASLYNQGKKKKKLQKSKR--KERIESSPMVEDVTE 372
Query: 193 PPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNG 229
PP +S IPP PSPP PP PP L +S F G
Sbjct: 373 PPQYQSL-IPP---PSPPPPPPPPPPPLRSSQSVFYG 405
>At4g38770 extensin - like protein
Length = 448
Score = 38.5 bits (88), Expect = 0.004
Identities = 31/114 (27%), Positives = 50/114 (43%), Gaps = 23/114 (20%)
Query: 130 PVFDNMPQPQFPLPLPSVAQS-MIELTP---EKPLPEKGKLLQ-----AVMKAGPLLQT- 179
PV+D P+ + P P+P +EL P +KP P K ++ V K P ++
Sbjct: 212 PVYDPPPKKEVPPPVPVYKPPPKVELPPPIPKKPCPPKPPKIEHPPPVPVYKPPPKIEKP 271
Query: 180 --LLLAGPLPQWRHPPP-----------PLESFEIPPVTIPSPPLQPPQQPPQL 220
+ + P P+ HPPP P + + PPV + PP + P P ++
Sbjct: 272 PPVPVYKPPPKIEHPPPVPVHKLPKKPCPPKKVDPPPVPVHKPPTKKPCPPKKV 325
Score = 33.5 bits (75), Expect = 0.12
Identities = 27/100 (27%), Positives = 43/100 (43%), Gaps = 18/100 (18%)
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PV+ P+ + P P+P + P+KP P K V P + P P
Sbjct: 275 PVYKPPPKIEHPPPVP------VHKLPKKPCPPKKVDPPPVPVHKPPTKK-----PCPPK 323
Query: 190 RHPPPPLESFEIPP-VTIPSPPLQPP------QQPPQLLH 222
+ PPP+ + PP + IP P ++ P + PP++ H
Sbjct: 324 KVDPPPVPVHKPPPKIVIPPPKIEHPPPVPVYKPPPKIEH 363
Score = 31.6 bits (70), Expect = 0.46
Identities = 28/115 (24%), Positives = 44/115 (37%), Gaps = 23/115 (20%)
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PV + P P+ +P P IE P P+ + ++ P + P+P +
Sbjct: 329 PVPVHKPPPKIVIPPPK-----IEHPPPVPVYKPPPKIEHPPIYIPPIVKKPCPPPVPIY 383
Query: 190 RHP--------PPPLESFEIPPVTIPSPPLQP----------PQQPPQLLHQESF 226
+ P PPP+ ++ P V IP P P P PP+ +H F
Sbjct: 384 KPPVVIPKKPCPPPVPVYKPPVVVIPKKPCPPLPQLPPLPKFPPLPPKYIHHPKF 438
Score = 31.6 bits (70), Expect = 0.46
Identities = 27/93 (29%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTP--EKPLPEKGKLLQAVMKAGPLLQTLLLAGPLP 187
P F P FPLP P +EL P +KP P K V P+ + P
Sbjct: 157 PPFKGFDHP-FPLPPP------LELPPFLKKPCPPKYSPPVEVPPPVPVYEP-------P 202
Query: 188 QWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQL 220
+ PPP+ ++ PP PP+ + PP++
Sbjct: 203 PKKEIPPPVPVYDPPPKKEVPPPVPVYKPPPKV 235
>At2g27390 pseudogene
Length = 134
Score = 38.5 bits (88), Expect = 0.004
Identities = 34/98 (34%), Positives = 41/98 (41%), Gaps = 20/98 (20%)
Query: 138 PQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLE 197
P+ P P P++ PE PLP + +L PL P P R PPP L
Sbjct: 53 PRLPPPFPAL------FPPEPPLPPRFEL------PPPLFP------PPPLPRLPPPLLP 94
Query: 198 SFEIPPVTIPSPPLQPPQQPP--QLLHQESFFNGSSST 233
E PP P PP P + PP L +S NG ST
Sbjct: 95 PPEEPPREPPPPPPPPEEPPPPASCLRTKSPENGIVST 132
>At1g76930 extensin
Length = 373
Score = 38.5 bits (88), Expect = 0.004
Identities = 28/91 (30%), Positives = 41/91 (44%), Gaps = 4/91 (4%)
Query: 130 PVFDNMPQPQFPLPLPSVAQ-SMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ 188
PV+ + P P + P P V S + P P K V K+ P + P P
Sbjct: 85 PVYKSPPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPP--PPVKHYSPPPV 142
Query: 189 WRHPPPPLESFEIPPV-TIPSPPLQPPQQPP 218
++ PPPP++ + PPV P PP++ PP
Sbjct: 143 YKSPPPPVKHYSPPPVYKSPPPPVKYYSPPP 173
Score = 35.8 bits (81), Expect = 0.024
Identities = 28/90 (31%), Positives = 39/90 (43%), Gaps = 10/90 (11%)
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PV P P + P P V +S P P K V K+ P + P P +
Sbjct: 77 PVKYYSPPPVYKSPPPPVYKS-------PPPPVKHYSPPPVYKSPP--PPVKHYSPPPVY 127
Query: 190 RHPPPPLESFEIPPV-TIPSPPLQPPQQPP 218
+ PPPP++ + PPV P PP++ PP
Sbjct: 128 KSPPPPVKHYSPPPVYKSPPPPVKHYSPPP 157
Score = 35.8 bits (81), Expect = 0.024
Identities = 29/94 (30%), Positives = 37/94 (38%), Gaps = 3/94 (3%)
Query: 130 PVFDNMPQPQFPLPLPSVAQ-SMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ 188
PV P P + P P V S + P P K V K+ P + P P
Sbjct: 29 PVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPP--PPVKYYSPPPV 86
Query: 189 WRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
++ PPPP+ PPV SPP PP + H
Sbjct: 87 YKSPPPPVYKSPPPPVKHYSPPPVYKSPPPPVKH 120
Score = 35.4 bits (80), Expect = 0.032
Identities = 28/91 (30%), Positives = 39/91 (42%), Gaps = 4/91 (4%)
Query: 130 PVFDNMPQPQFPLPLPSVAQ-SMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ 188
PV P P + P P V S + P P K V K+ P + P P
Sbjct: 101 PVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPP--PPVKHYSPPPV 158
Query: 189 WRHPPPPLESFEIPPV-TIPSPPLQPPQQPP 218
++ PPPP++ + PPV P PP++ PP
Sbjct: 159 YKSPPPPVKYYSPPPVYKSPPPPVKHYSPPP 189
Score = 34.7 bits (78), Expect = 0.054
Identities = 28/94 (29%), Positives = 38/94 (39%), Gaps = 3/94 (3%)
Query: 130 PVFDNMPQPQFPLPLPSVAQ-SMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ 188
PV P P + P P V S + P P K V K+ P + P P
Sbjct: 165 PVKYYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPP--PPVKHYSPPPV 222
Query: 189 WRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
++ PPPP++ + PPV PP PP + H
Sbjct: 223 YKSPPPPVKYYSPPPVYKSPPPPVHYSPPPVVYH 256
Score = 31.6 bits (70), Expect = 0.46
Identities = 26/93 (27%), Positives = 34/93 (35%), Gaps = 2/93 (2%)
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PV P P + P P V S + P P V+ P + P P
Sbjct: 229 PVKYYSPPPVYKSPPPPVHYSPPPVVYHSPPPPVHYSPPPVVYHSP--PPPVHYSPPPVV 286
Query: 190 RHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
H PPP + PPV SPP PP +++
Sbjct: 287 YHSPPPPVHYSPPPVVYHSPPPPVHYSPPPVVY 319
Score = 29.3 bits (64), Expect = 2.3
Identities = 17/44 (38%), Positives = 21/44 (47%), Gaps = 8/44 (18%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQ------PPQLLH 222
P P H PPP + PPV SPP PP++ PP +H
Sbjct: 298 PPPVVYHSPPPPVHYSPPPVVYHSPP--PPKKHYEYKSPPPPVH 339
Score = 28.5 bits (62), Expect = 3.9
Identities = 12/28 (42%), Positives = 17/28 (59%), Gaps = 1/28 (3%)
Query: 192 PPPPLESFEIPPV-TIPSPPLQPPQQPP 218
PPPP++ + PPV P PP++ PP
Sbjct: 26 PPPPVKHYSPPPVYKSPPPPVKHYSPPP 53
Score = 27.7 bits (60), Expect = 6.6
Identities = 14/38 (36%), Positives = 17/38 (43%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
P P + PPP+ PPV SPP PP + H
Sbjct: 27 PPPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKH 64
>At4g33970 extensin-like protein
Length = 699
Score = 38.1 bits (87), Expect = 0.005
Identities = 28/99 (28%), Positives = 41/99 (41%), Gaps = 5/99 (5%)
Query: 126 IVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVM--KAGPLLQTLLLA 183
+ ++PV P P P+ PS ++ +KP P + +Q K P
Sbjct: 453 VPTTPVQKPSPVPTTPVHEPS---PVLATPVDKPSPVPSRPVQKPQPPKESPQPDDPYDQ 509
Query: 184 GPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
P+ + R PPP + PPV P PP P PP +H
Sbjct: 510 SPVTKRRSPPPAPVNSPPPPVYSPPPPPPPVHSPPPPVH 548
Score = 35.4 bits (80), Expect = 0.032
Identities = 16/34 (47%), Positives = 17/34 (49%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPPP+ S PPV P PP P PP
Sbjct: 536 PPPPVHSPPPPVHSPPPPPVYSPPPPPPPVHSPP 569
Score = 32.7 bits (73), Expect = 0.21
Identities = 23/93 (24%), Positives = 35/93 (36%), Gaps = 8/93 (8%)
Query: 126 IVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
++++PV P P P+ P + + P+ P + + P+ P
Sbjct: 475 VLATPVDKPSPVPSRPVQKPQPPKESPQ--PDDPYDQSPVTKRRSPPPAPV------NSP 526
Query: 186 LPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P PPPP PP + SPP P PP
Sbjct: 527 PPPVYSPPPPPPPVHSPPPPVHSPPPPPVYSPP 559
Score = 32.3 bits (72), Expect = 0.27
Identities = 24/84 (28%), Positives = 34/84 (39%), Gaps = 2/84 (2%)
Query: 141 PLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP--LES 198
P P+PS + E P P+ V K + + P P + PPPP + S
Sbjct: 483 PSPVPSRPVQKPQPPKESPQPDDPYDQSPVTKRRSPPPAPVNSPPPPVYSPPPPPPPVHS 542
Query: 199 FEIPPVTIPSPPLQPPQQPPQLLH 222
P + P PP+ P PP +H
Sbjct: 543 PPPPVHSPPPPPVYSPPPPPPPVH 566
Score = 32.3 bits (72), Expect = 0.27
Identities = 16/35 (45%), Positives = 18/35 (50%), Gaps = 1/35 (2%)
Query: 185 PLPQWRHPPPPLESFEIP-PVTIPSPPLQPPQQPP 218
P P PPPP+ S P PV P PP+ P PP
Sbjct: 582 PPPPVHSPPPPVHSPPPPAPVHSPPPPVHSPPPPP 616
Score = 30.8 bits (68), Expect = 0.78
Identities = 15/43 (34%), Positives = 19/43 (43%), Gaps = 7/43 (16%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQ-------PPQQPPQL 220
P P PPPP + PP PP Q PP +PP++
Sbjct: 605 PPPPVHSPPPPPPVYSPPPPVFSPPPSQSPPVVYSPPPRPPKI 647
Score = 30.4 bits (67), Expect = 1.0
Identities = 30/110 (27%), Positives = 42/110 (37%), Gaps = 16/110 (14%)
Query: 129 SPVFDNMPQPQFPLPLPSVAQSMIELTP-EKPLPEKGKLLQAVMKAGPLLQTLLLAGPLP 187
SPV P P P+ P+ + TP +KP P V K P+ P
Sbjct: 423 SPVHKPTPVPTTPVHKPTP----VPTTPVQKPSPVP---TTPVQKPSPV--------PTT 467
Query: 188 QWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHS 237
P P L + P +PS P+Q PQ P + + ++ S T S
Sbjct: 468 PVHEPSPVLATPVDKPSPVPSRPVQKPQPPKESPQPDDPYDQSPVTKRRS 517
Score = 30.0 bits (66), Expect = 1.3
Identities = 15/33 (45%), Positives = 17/33 (51%), Gaps = 1/33 (3%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQP 217
P P PPPP+ S PPV P PP+ P P
Sbjct: 561 PPPPVHSPPPPVFS-PPPPVYSPPPPVHSPPPP 592
Score = 30.0 bits (66), Expect = 1.3
Identities = 17/38 (44%), Positives = 19/38 (49%), Gaps = 2/38 (5%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
P P PPPP+ S PPV P PP P PP +H
Sbjct: 575 PPPPVYSPPPPVHS-PPPPVHSPPPP-APVHSPPPPVH 610
Score = 29.6 bits (65), Expect = 1.7
Identities = 17/38 (44%), Positives = 20/38 (51%), Gaps = 2/38 (5%)
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
P P PPPP+ S PPV P PP+ P PP +H
Sbjct: 568 PPPPVFSPPPPVYS-PPPPVHSPPPPVHSP-PPPAPVH 603
Score = 28.9 bits (63), Expect = 3.0
Identities = 12/28 (42%), Positives = 13/28 (45%)
Query: 191 HPPPPLESFEIPPVTIPSPPLQPPQQPP 218
H PPP PP + SPP PP P
Sbjct: 594 HSPPPPAPVHSPPPPVHSPPPPPPVYSP 621
Score = 28.5 bits (62), Expect = 3.9
Identities = 30/109 (27%), Positives = 40/109 (36%), Gaps = 9/109 (8%)
Query: 121 DCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEK-PLPEKGKLLQAVMKAGPLLQT 179
D + S PV PQP P P +T + P P + + P
Sbjct: 481 DKPSPVPSRPV--QKPQPPKESPQPDDPYDQSPVTKRRSPPPAPVNSPPPPVYSPPPPPP 538
Query: 180 LLLAGPLPQWRHPPPPLESFEIPPVTIPSPP---LQPP---QQPPQLLH 222
+ + P P PPPP+ S PP + SPP PP PP +H
Sbjct: 539 PVHSPPPPVHSPPPPPVYSPPPPPPPVHSPPPPVFSPPPPVYSPPPPVH 587
Score = 28.1 bits (61), Expect = 5.1
Identities = 18/42 (42%), Positives = 23/42 (53%), Gaps = 4/42 (9%)
Query: 185 PLPQWRH-PPPPLES-FEIPPVTIPSPPL--QPPQQPPQLLH 222
P P H PPPP+ S PPV P PP+ PP Q P +++
Sbjct: 597 PPPAPVHSPPPPVHSPPPPPPVYSPPPPVFSPPPSQSPPVVY 638
Score = 27.3 bits (59), Expect = 8.6
Identities = 13/32 (40%), Positives = 16/32 (49%)
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQ 223
PP P+E E PP P+P PP + HQ
Sbjct: 656 PPAPVEKKETPPAHAPAPSDDEFIIPPFIGHQ 687
>At3g19320 unknown protein
Length = 493
Score = 38.1 bits (87), Expect = 0.005
Identities = 28/83 (33%), Positives = 32/83 (37%), Gaps = 20/83 (24%)
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P+P+ LPLP Q TP P P Q+L P P+ H PPP
Sbjct: 54 PEPEDYLPLPPPPQ-----TPPPPPPP---------------QSLPPPSPSPEPEHYPPP 93
Query: 196 LESFEIPPVTIPSPPLQPPQQPP 218
I P P PL PP PP
Sbjct: 94 PYHHYITPSPPPPRPLPPPPPPP 116
>At4g32710 putative protein kinase
Length = 731
Score = 37.7 bits (86), Expect = 0.006
Identities = 29/85 (34%), Positives = 38/85 (44%), Gaps = 5/85 (5%)
Query: 127 VSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPL 186
+S P D++P P P P ++ L+ PLP L PL + + P
Sbjct: 18 MSLPPADSVPDTSSP-PAPPLSPLPPPLSSPPPLPSPPPLSAPTASPPPLP---VESPPS 73
Query: 187 PQWRHPPPPL-ESFEIPPVTIPSPP 210
P PPPPL ES PP+ PSPP
Sbjct: 74 PPIESPPPPLLESPPPPPLESPSPP 98
Score = 34.3 bits (77), Expect = 0.071
Identities = 26/86 (30%), Positives = 33/86 (38%), Gaps = 10/86 (11%)
Query: 143 PLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQ-------WRHPPP- 194
P P+ + + L P +P+ PL L PLP PPP
Sbjct: 9 PAPATSPPAMSLPPADSVPDTSS--PPAPPLSPLPPPLSSPPPLPSPPPLSAPTASPPPL 66
Query: 195 PLESFEIPPVTIPSPPLQPPQQPPQL 220
P+ES PP+ P PPL PP L
Sbjct: 67 PVESPPSPPIESPPPPLLESPPPPPL 92
Score = 30.4 bits (67), Expect = 1.0
Identities = 30/103 (29%), Positives = 37/103 (35%), Gaps = 23/103 (22%)
Query: 129 SPVFDNMPQP-QFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLP 187
+P +P P P PLPS PLP + + P L L P
Sbjct: 34 APPLSPLPPPLSSPPPLPSPPPLSAPTASPPPLPVESPPSPPIESPPPPL----LESP-- 87
Query: 188 QWRHPPPPLES-----------FEIPPVT-IPSPPLQPPQQPP 218
PPPPLES PP+ +P+ P PP PP
Sbjct: 88 ----PPPPLESPSPPSPHVSAPSGSPPLPFLPAKPSPPPSSPP 126
Score = 27.3 bits (59), Expect = 8.6
Identities = 26/92 (28%), Positives = 30/92 (32%), Gaps = 5/92 (5%)
Query: 129 SPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGK---LLQAVMKAGPLLQTLLLAGP 185
SP ++ P P P P +S +P P L A P P
Sbjct: 73 SPPIESPPPPLLESPPPPPLESPSPPSPHVSAPSGSPPLPFLPAKPSPPPSSPPSETVPP 132
Query: 186 LPQWRHPPPPLESFEIPPVTIPSPPLQPPQQP 217
PP L S PPV SPP PP P
Sbjct: 133 GNTISPPPRSLPSESTPPVNTASPP--PPSPP 162
>At5g67470 formin-like protein
Length = 899
Score = 37.4 bits (85), Expect = 0.008
Identities = 35/133 (26%), Positives = 53/133 (39%), Gaps = 23/133 (17%)
Query: 126 IVSSPVFDNMPQPQFPLPLPSVAQSMIELT--------PEKPLPEKGKLLQAVMKAGPLL 177
++ + +P P P PL + EL + P P QA+ +
Sbjct: 307 VIIPSIKQKLPPPVQPPPLRGLESDEQELPYSQNKPKFSQPPPPPNRAAFQAITQE---- 362
Query: 178 QTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQ--ESFFNGSSSTST 235
P+P R PPPL++ PP P PPL PP PPQ + + ++S +T
Sbjct: 363 -----KSPVPPPRRSPPPLQT---PPPPPPPPPLAPP-PPPQKRPRDFQMLRKVTNSEAT 413
Query: 236 HSQCGRVSRKRIF 248
+ SRK+ F
Sbjct: 414 TNSTTSPSRKQAF 426
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,549,455
Number of Sequences: 26719
Number of extensions: 336250
Number of successful extensions: 4877
Number of sequences better than 10.0: 248
Number of HSP's better than 10.0 without gapping: 112
Number of HSP's successfully gapped in prelim test: 146
Number of HSP's that attempted gapping in prelim test: 2054
Number of HSP's gapped (non-prelim): 1478
length of query: 266
length of database: 11,318,596
effective HSP length: 98
effective length of query: 168
effective length of database: 8,700,134
effective search space: 1461622512
effective search space used: 1461622512
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0369b.1


