

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0358.18
(152 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
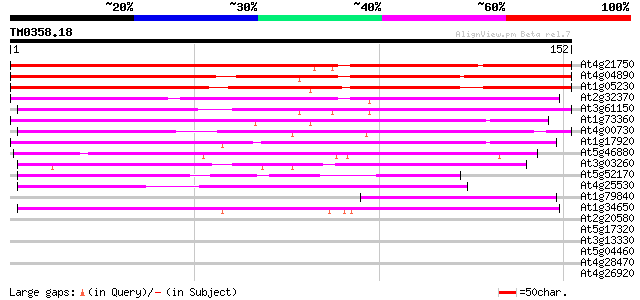
Score E
Sequences producing significant alignments: (bits) Value
At4g21750 L1 specific homeobox gene ATML1/ovule-specific homeobo... 159 7e-40
At4g04890 putative homeotic protein 150 3e-37
At1g05230 homeobox like protein 135 9e-33
At2g32370 putative homeodomain transcription factor 86 6e-18
At3g61150 homeobox protein 86 8e-18
At1g73360 putative homeobox protein 83 7e-17
At4g00730 homeodomain protein AHDP 82 9e-17
At1g17920 unknown protein 80 3e-16
At5g46880 homeobox protein 72 1e-13
At3g03260 unknown protein 52 2e-07
At5g52170 homeodomain transcription factor-like 50 4e-07
At4g25530 homeodomain-containing transcription factor FWA 47 4e-06
At1g79840 homeobox protein (GLABRA2) 42 2e-04
At1g34650 hypothetical protein 40 6e-04
At2g20580 putative 26S proteasome regulatory subunit S2 31 0.30
At5g17320 homeobox protein 30 0.39
At3g13330 unknown protein 30 0.39
At5g04460 putative protein 30 0.66
At4g28470 putative protein 29 0.86
At4g26920 putative homeodomain protein 28 1.5
>At4g21750 L1 specific homeobox gene ATML1/ovule-specific homeobox
protein A20
Length = 762
Score = 159 bits (401), Expect = 7e-40
Identities = 90/166 (54%), Positives = 112/166 (67%), Gaps = 18/166 (10%)
Query: 1 ATCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYN 60
ATC CGG ++GEMS+DEQ +++ENARLRE I+RIS I AKY GK + SS Q +
Sbjct: 150 ATCPNCGGPAAIGEMSFDEQHLRIENARLREEIDRISAIAAKYVGKPLMANSSSFPQLSS 209
Query: 61 QMPSSSRAFDLGVGNYGGDGN-------------DLLRS-SLPPILADADKPIIVEVAVA 106
SR+ DL VGN+G + N D+LRS S+P ++ADKP+IVE+AVA
Sbjct: 210 SHHIPSRSLDLEVGNFGNNNNSHTGFVGEMFGSSDILRSVSIP---SEADKPMIVELAVA 266
Query: 107 AMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGSKPFG 152
AMEELVR+A+ G PLWV S+N +VE LNEEEY R FPRG G KP G
Sbjct: 267 AMEELVRMAQTGDPLWVSSDN-SVEILNEEEYFRTFPRGIGPKPIG 311
>At4g04890 putative homeotic protein
Length = 743
Score = 150 bits (379), Expect = 3e-37
Identities = 86/162 (53%), Positives = 109/162 (67%), Gaps = 19/162 (11%)
Query: 1 ATCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYN 60
ATC CGG ++GEMS+DEQ +++ENARLRE I+RIS I AKY GK S + L+
Sbjct: 150 ATCPNCGGPAAIGEMSFDEQHLRIENARLREEIDRISAIAAKYVGKPLGSSFAPLA---- 205
Query: 61 QMPSSSRAFDLGVGNYG----------GDGNDLLRSSLPPILADADKPIIVEVAVAAMEE 110
+ + SR+ DL VGN+G G G+ L S+P ++ DKPIIVE+AVAAMEE
Sbjct: 206 -IHAPSRSLDLEVGNFGNQTGFVGEMYGTGDILRSVSIP---SETDKPIIVELAVAAMEE 261
Query: 111 LVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGSKPFG 152
LVR+A+ G PLW LS +++VE LNEEEY R FPRG G KP G
Sbjct: 262 LVRMAQTGDPLW-LSTDNSVEILNEEEYFRTFPRGIGPKPLG 302
>At1g05230 homeobox like protein
Length = 721
Score = 135 bits (340), Expect = 9e-33
Identities = 73/157 (46%), Positives = 100/157 (63%), Gaps = 18/157 (11%)
Query: 1 ATCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYN 60
A+C CGG T++GEMS+DE ++LENARLRE I+RIS I AKY GK ++Y + +
Sbjct: 152 ASCPNCGGPTAIGEMSFDEHQLRLENARLREEIDRISAIAAKYVGKPVSNYPLM-----S 206
Query: 61 QMPSSSRAFDLGVGNYGGDG-----NDLLRSSLPPILADADKPIIVEVAVAAMEELVRLA 115
P R +L +GN GG+ NDLL+S P ++DKP+I++++VAAMEEL+R+
Sbjct: 207 PPPLPPRPLELAMGNIGGEAYGNNPNDLLKSITAP--TESDKPVIIDLSVAAMEELMRMV 264
Query: 116 RVGHPLWVLSNNHNVETLNEEEYVREFPRGTGSKPFG 152
+V PLW L+EEEY R FPRG G +P G
Sbjct: 265 QVDEPLW------KSLVLDEEEYARTFPRGIGPRPAG 295
>At2g32370 putative homeodomain transcription factor
Length = 721
Score = 86.3 bits (212), Expect = 6e-18
Identities = 55/150 (36%), Positives = 84/150 (55%), Gaps = 7/150 (4%)
Query: 1 ATCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYN 60
A C CGG T++GEM+++E +++ NARL E I+++S K S + + S
Sbjct: 156 ALCPKCGGQTAIGEMTFEEHHLRILNARLTEEIKQLSVTAEKI---SRLTGIPVRSHPRV 212
Query: 61 QMPSSSRAFDLGVGNYGGDGNDLLRSSLPPILADAD-KPIIVEVAVAAMEELVRLARVGH 119
P+ F+ G+G+ G GN ++ P ADA+ KPII+E+A AMEEL+ +A+V
Sbjct: 213 SPPNPPPNFEFGMGSKGNVGNHSRETTGP---ADANTKPIIMELAFGAMEELLVMAQVAE 269
Query: 120 PLWVLSNNHNVETLNEEEYVREFPRGTGSK 149
PLW+ N LN +EY + F G G +
Sbjct: 270 PLWMGGFNGTSLALNLDEYEKTFRTGLGPR 299
>At3g61150 homeobox protein
Length = 808
Score = 85.9 bits (211), Expect = 8e-18
Identities = 54/179 (30%), Positives = 91/179 (50%), Gaps = 38/179 (21%)
Query: 3 CTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYNQM 62
C CGG +GE+S +EQ +++EN+RL++ ++R+ + K+ G+S S+ +
Sbjct: 200 CGNCGGPAVIGEISMEEQHLRIENSRLKDELDRVCALTGKFLGRSNGSH---------HI 250
Query: 63 PSSSRAFDLGVGNYG-------------------------GDGNDLLRS---SLPPILAD 94
P S+ +GVG+ G G G+ L+ + P ++D
Sbjct: 251 PDSALVLGVGVGSGGCNVGGGFTLSSPLLPQASPRFEISNGTGSGLVATVNRQQPVSVSD 310
Query: 95 AD-KPIIVEVAVAAMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGSKPFG 152
D + +++A+AAM+ELV++A+ PLWV S++ E LN+EEY F R G K G
Sbjct: 311 FDQRSRYLDLALAAMDELVKMAQTREPLWVRSSDSGFEVLNQEEYDTSFSRCVGPKQDG 369
>At1g73360 putative homeobox protein
Length = 722
Score = 82.8 bits (203), Expect = 7e-17
Identities = 55/161 (34%), Positives = 83/161 (51%), Gaps = 16/161 (9%)
Query: 1 ATCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYN 60
A C CGG + +DEQ +++ENA LRE +ER+S I +KY G+ + S+L + +
Sbjct: 120 AICPNCGGPPVSEDPYFDEQKLRIENAHLREELERMSTIASKYMGRPISQLSTLHPMHIS 179
Query: 61 QMPSS--------------SRAFDLGVGNYGGDG-NDLLRSSLPPILADADKPIIVEVAV 105
+ S S FDL G+ G N+ L+S ++D DKPI+ +A+
Sbjct: 180 PLDLSMTSLTGCGPFGHGPSLDFDLLPGSSMAVGPNNNLQSQPNLAISDMDKPIMTGIAL 239
Query: 106 AAMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGT 146
AMEEL+RL + PLW ++ + LN Y FPR +
Sbjct: 240 TAMEELLRLLQTNEPLWTRTDGCR-DILNLGSYENVFPRSS 279
>At4g00730 homeodomain protein AHDP
Length = 590
Score = 82.4 bits (202), Expect = 9e-17
Identities = 50/162 (30%), Positives = 85/162 (51%), Gaps = 26/162 (16%)
Query: 3 CTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYNQM 62
CT CGG LG++S +E +++ENARL++ ++R+ + K+ G ++N
Sbjct: 12 CTNCGGPAMLGDVSLEEHHLRIENARLKDELDRVCNLTGKFLG-----------HHHNHH 60
Query: 63 PSSSRAFDLGVGN----------YGGDGNDLLRSSLPPILADA--DKPIIVEVAVAAMEE 110
+SS +G N +GG G L + + K +++E+A+ AM+E
Sbjct: 61 YNSSLELAVGTNNNGGHFAFPPDFGGGGGCLPPQQQQSTVINGIDQKSVLLELALTAMDE 120
Query: 111 LVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGSKPFG 152
LV+LA+ PLWV S + + LN++EY+R F + +KP G
Sbjct: 121 LVKLAQSEEPLWVKSLDGERDELNQDEYMRTF---SSTKPTG 159
>At1g17920 unknown protein
Length = 687
Score = 80.5 bits (197), Expect = 3e-16
Identities = 51/155 (32%), Positives = 82/155 (52%), Gaps = 10/155 (6%)
Query: 1 ATCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLS---- 56
A C +CG + +DEQ +++ENA+LR+ +ER+S I AK+ G+ + LL+
Sbjct: 109 AICPSCGDSPVNEDSYFDEQKLRIENAQLRDELERVSSIAAKFLGRPISHLPPLLNPMHV 168
Query: 57 ---QNYNQMPSSSRAFDLGVGNYGGDGNDLLRSSLPPILADADKPIIVEVAVAAMEELVR 113
+ ++ PS FDL G+ L S +L++ DK ++ +AV AMEEL+R
Sbjct: 169 SPLELFHTGPSLD--FDLLPGSCSSMSVPSLPSQPNLVLSEMDKSLMTNIAVTAMEELLR 226
Query: 114 LARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGS 148
L + PLW+ ++ + LN E Y F R + S
Sbjct: 227 LLQTNEPLWIKTDGCR-DVLNLENYENMFTRSSTS 260
>At5g46880 homeobox protein
Length = 783
Score = 72.0 bits (175), Expect = 1e-13
Identities = 50/171 (29%), Positives = 83/171 (48%), Gaps = 31/171 (18%)
Query: 2 TCTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSY--SSLLSQNY 59
+C +CGG T LG++ ++E + +EN RLRE ++R+ I ++Y G+ S S L
Sbjct: 200 SCPSCGGPTVLGDIPFNE--IHIENCRLREELDRLCCIASRYTGRPMQSMPPSQPLINPS 257
Query: 60 NQMPSSSRAFDLGVGNYGGDGNDLLRSS---LPP--------------------ILADAD 96
+P + +L + Y G+ + + LPP +LAD +
Sbjct: 258 PMLPHHQPSLELDMSVYAGNFPEQSCTDMMMLPPQDTACFFPDQTANNNNNNNMLLADEE 317
Query: 97 KPIIVEVAVAAMEELVRLARVGHPLWVLSNNHNVE----TLNEEEYVREFP 143
K I +E AV+ ++EL ++ PLW+ + + LNEEEY+R FP
Sbjct: 318 KVIAMEFAVSCVQELTKMCDTEEPLWIKKKSDKIGGEILCLNEEEYMRLFP 368
>At3g03260 unknown protein
Length = 699
Score = 51.6 bits (122), Expect = 2e-07
Identities = 43/147 (29%), Positives = 67/147 (45%), Gaps = 17/147 (11%)
Query: 3 CTTCGGLT-SLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYNQ 61
C CGG E ++ Q ++ ENARL++ +RIS V ++ T SL
Sbjct: 113 CPACGGPPFGREERGHNLQKLRFENARLKDHRDRISNFVDQHKPNEPTVEDSLA-----Y 167
Query: 62 MPSSSR-AFDLGVGN-------YGGDGNDLLRSSLPPILADADKPIIVEVAVAAMEELVR 113
+PS R ++ + GN YG +++ P LA+ D ++ E+A +A+EEL R
Sbjct: 168 VPSLDRISYGINGGNMYEPSSSYGPPNFQIIQ---PRPLAETDMSLLSEIAASAVEELKR 224
Query: 114 LARVGHPLWVLSNNHNVETLNEEEYVR 140
L WV S ++ E Y R
Sbjct: 225 LFLAEEQFWVKSCIDETYVIDTESYER 251
>At5g52170 homeodomain transcription factor-like
Length = 682
Score = 50.4 bits (119), Expect = 4e-07
Identities = 33/120 (27%), Positives = 59/120 (48%), Gaps = 23/120 (19%)
Query: 3 CTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYNQM 62
C CGG GE+S+++ +++ENA+L+E ++RI + ++ G S + L Q N
Sbjct: 147 CIDCGGAVIPGEVSFEQHQLRIENAKLKEELDRICALANRFIGGSIS-----LEQPSNGG 201
Query: 63 PSSSRAFDLGVGNYGGDGNDLLRSSLPPILADADKPIIVEVAVAAMEELVRLARVGHPLW 122
S L +G+ G L+ +++A+ AM+EL++LA + LW
Sbjct: 202 IGSQH---LPIGHCVSGGTSLM---------------FMDLAMEAMDELLKLAELETSLW 243
>At4g25530 homeodomain-containing transcription factor FWA
Length = 686
Score = 47.0 bits (110), Expect = 4e-06
Identities = 31/122 (25%), Positives = 52/122 (42%), Gaps = 14/122 (11%)
Query: 3 CTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYNQM 62
C CG T+ G+ Y+ Q + ENA L I++ + S LS +M
Sbjct: 130 CNICGKATNCGDTEYEVQKLMAENANLEREIDQFN--------------SRYLSHPKQRM 175
Query: 63 PSSSRAFDLGVGNYGGDGNDLLRSSLPPILADADKPIIVEVAVAAMEELVRLARVGHPLW 122
S+S N G + +L S ++ + I + +A+ A+ EL+ L V P W
Sbjct: 176 VSTSEQAPSSSSNPGINATPVLDFSGGTRTSEKETSIFLNLAITALRELITLGEVDCPFW 235
Query: 123 VL 124
++
Sbjct: 236 MI 237
>At1g79840 homeobox protein (GLABRA2)
Length = 747
Score = 41.6 bits (96), Expect = 2e-04
Identities = 20/53 (37%), Positives = 31/53 (57%)
Query: 96 DKPIIVEVAVAAMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGS 148
+K I E++ A EL ++A G P+W+ S E LN +EY++EFP+ S
Sbjct: 253 EKSRIAEISNRATLELQKMATSGEPMWLRSVETGREILNYDEYLKEFPQAQAS 305
>At1g34650 hypothetical protein
Length = 708
Score = 39.7 bits (91), Expect = 6e-04
Identities = 39/169 (23%), Positives = 69/169 (40%), Gaps = 22/169 (13%)
Query: 3 CTTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLS----QN 58
C CGG E + Q ++ +N L+ ER+S + K+ G S S +L
Sbjct: 106 CPPCGGPHGKEEQLCNLQKLRTKNVILKTEYERLSSYLTKHGGYSIPSVDALPDLHGPST 165
Query: 59 YNQMPSSSRAFDLGVGNYGGDGNDLLR----------SSLP-PI-------LADADKPII 100
Y ++ A N+ + LLR + LP P+ L+ +K +
Sbjct: 166 YGSTSNNRPASYGSSSNHLPQQSSLLRRPFTRELINTTPLPKPVLLQHFQQLSQLEKNRM 225
Query: 101 VEVAVAAMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFPRGTGSK 149
E+A A+ E++ L ++ H +W+ S ++ Y R F + + K
Sbjct: 226 FEIAKNAVAEVMSLIQMEHSMWIKSTIDGRAIIDPGNYKRYFTKNSHLK 274
>At2g20580 putative 26S proteasome regulatory subunit S2
Length = 891
Score = 30.8 bits (68), Expect = 0.30
Identities = 17/39 (43%), Positives = 26/39 (66%), Gaps = 2/39 (5%)
Query: 71 LGVG-NYGGDGNDLLRSSLPPILADADKPIIVEVAVAAM 108
+G+G +Y G ND +R+ L PIL DA P+ V +A A++
Sbjct: 488 MGLGISYAGSQNDQIRNKLSPILNDAKAPLDV-IAFASL 525
>At5g17320 homeobox protein
Length = 718
Score = 30.4 bits (67), Expect = 0.39
Identities = 43/182 (23%), Positives = 73/182 (39%), Gaps = 41/182 (22%)
Query: 3 CTTCGGLTSLGEMSYDEQLMKL--ENARLREAIERISGIVAKYAGKS-----TTSY---- 51
C CGG G L KL +NA L++ ER+S + +Y G S T Y
Sbjct: 116 CPPCGG-RGPGREDQLRHLQKLRAQNAYLKDEYERVSNYLKQYGGHSMHNVEATPYLHGP 174
Query: 52 ----------SSLLSQNYNQMPSSSRAFDLGVGNYGGDGNDLLRSSLPP----------- 90
+L + N++P S F G Y GN + ++ PP
Sbjct: 175 SNHASTSKNRPALYGTSSNRLPEPSSIFR---GPY-TRGN--MNTTAPPQPRKPLEMQNF 228
Query: 91 -ILADADKPIIVEVAVAAMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFPR-GTGS 148
L+ +K ++E A A+ E++ L ++ +W S+ + ++ Y + F + T
Sbjct: 229 QPLSQLEKIAMLEAAEKAVSEVLSLIQMDDTMWKKSSIDDRLVIDPGLYEKYFTKTNTNG 288
Query: 149 KP 150
+P
Sbjct: 289 RP 290
>At3g13330 unknown protein
Length = 1711
Score = 30.4 bits (67), Expect = 0.39
Identities = 34/142 (23%), Positives = 56/142 (38%), Gaps = 13/142 (9%)
Query: 7 GGLTSLGEMSYDEQLMKLENARLREAIE---RISGIVAKYAGKSTTSYSSLLSQNYNQMP 63
GG S + ++Q + + L AI+ ++ + Y G S Y LL +
Sbjct: 573 GGKGSAETLVSNKQDKQTLSPALEAAIDYQLKVLSVAITYGGSSLLPYKGLLIE------ 626
Query: 64 SSSRAFDLGVGNYGGDGNDLLRSSLPPIL----ADADKPIIVEVAVAAMEELVRLARVGH 119
+ S AF+ G G+ LLRS L ++ D K + A A+EE +
Sbjct: 627 AISSAFNSSSWKVNGAGDHLLRSLLGSLILYYPIDQYKCLSRHPAAPALEEWISTKASSK 686
Query: 120 PLWVLSNNHNVETLNEEEYVRE 141
V + +V T E ++ E
Sbjct: 687 DEQVAHSRWHVPTQEETQFANE 708
>At5g04460 putative protein
Length = 831
Score = 29.6 bits (65), Expect = 0.66
Identities = 25/114 (21%), Positives = 46/114 (39%), Gaps = 5/114 (4%)
Query: 4 TTCGGLTSLGEMSYDEQLMKLENARLREAIERISGIVAKYAGKSTTSYSSLLSQNYNQMP 63
T G+ + DE ++ R+RE + + + + S+ + S LS+N +
Sbjct: 149 TQASGILQMWRELEDEHVLNRARERVRERLRQQRSVESNTNLSSSIASESQLSENNGSLR 208
Query: 64 SSSRA-FDLGV----GNYGGDGNDLLRSSLPPILADADKPIIVEVAVAAMEELV 112
SS + D G N GD N+ P L D ++ + +A M+ +
Sbjct: 209 DSSESENDFGSWSHDRNEHGDNNNTSSREQSPDLGDGERERVRHIARGWMDSRI 262
>At4g28470 putative protein
Length = 1103
Score = 29.3 bits (64), Expect = 0.86
Identities = 23/59 (38%), Positives = 32/59 (53%), Gaps = 5/59 (8%)
Query: 53 SLLSQNYNQMPSSSRA---FDLGVGNYGGDGNDLLRSSLPPILADADKPIIVEVAVAAM 108
+LLS + SS R LG+ Y G ND ++ L PIL DA+ P+ V +A AA+
Sbjct: 512 ALLSGYIDNEDSSVRIGAIMGLGIA-YAGSQNDQIKIRLSPILNDANAPLDV-IAFAAL 568
>At4g26920 putative homeodomain protein
Length = 461
Score = 28.5 bits (62), Expect = 1.5
Identities = 16/41 (39%), Positives = 21/41 (51%), Gaps = 1/41 (2%)
Query: 103 VAVAAMEELVRLARVGHPLWVLSNNHNVETLNEEEYVREFP 143
V + E++ LA + PLW S + TLN E Y R FP
Sbjct: 67 VLACIVNEIIALATLESPLWRRSQREEMLTLN-EYYSRFFP 106
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.132 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,407,578
Number of Sequences: 26719
Number of extensions: 128956
Number of successful extensions: 340
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 295
Number of HSP's gapped (non-prelim): 34
length of query: 152
length of database: 11,318,596
effective HSP length: 90
effective length of query: 62
effective length of database: 8,913,886
effective search space: 552660932
effective search space used: 552660932
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0358.18


