

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0349b.2
(400 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
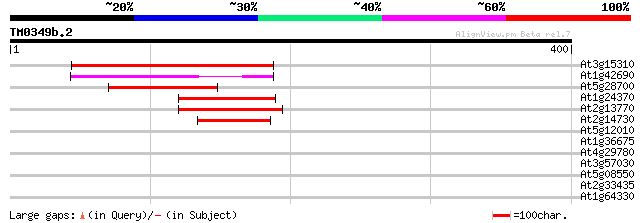
Score E
Sequences producing significant alignments: (bits) Value
At3g15310 unknown protein 129 4e-30
At1g42690 unknown protein 97 2e-20
At5g28700 putative protein 80 2e-15
At1g24370 hypothetical protein 80 2e-15
At2g13770 hypothetical protein 80 3e-15
At2g14730 hypothetical protein 57 2e-08
At5g12010 putative protein 37 0.015
At1g36675 putative protein 35 0.095
At4g29780 unknown protein 34 0.16
At3g57030 unknown protein 29 5.2
At5g08550 putative protein 28 6.8
At2g33435 RRM-containing protein 28 6.8
At1g64330 unknown protein 28 8.9
>At3g15310 unknown protein
Length = 415
Score = 129 bits (323), Expect = 4e-30
Identities = 66/144 (45%), Positives = 92/144 (63%)
Query: 45 RGRKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQ 104
R R + R + L +DYF + + FRRR+RM+K +FLRIV LSS +F
Sbjct: 43 RKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRMKKPLFLRIVDRLSSELMFFQH 102
Query: 105 RVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQ 164
R DA + G SP+ KCT +R+LA G A+DAVDEY+++GETT + CL F KGII +
Sbjct: 103 RRDATGRFGHSPIQKCTAAIRLLAYGYASDAVDEYLRMGETTAMSCLENFTKGIISFFGD 162
Query: 165 EYLRAPTQEDLQRILHVNEMWGVP 188
EYLRAPT +L+R+L++ ++ G P
Sbjct: 163 EYLRAPTATNLRRLLNIGKIRGFP 186
>At1g42690 unknown protein
Length = 333
Score = 97.1 bits (240), Expect = 2e-20
Identities = 57/146 (39%), Positives = 77/146 (52%), Gaps = 31/146 (21%)
Query: 44 PRGRK-HLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYF 102
PR +K ++ R+ + +L++DYF PTY IFRRR+RM K +F+RIV S+ YF
Sbjct: 37 PRKKKLYIERNREEGHIQLVNDYFTENPTYPPHIFRRRFRMNKSLFMRIVERFSNEVPYF 96
Query: 103 TQRVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLY 162
QR DA ++ G S L K T +RMLA G+AADA
Sbjct: 97 KQRRDATRRLGFSALQKSTAAIRMLAYGIAADA--------------------------- 129
Query: 163 EQEYLRAPTQEDLQRILHVNEMWGVP 188
YLR PT++DL R+LH+ E G P
Sbjct: 130 ---YLRRPTRDDLIRLLHIGEQRGFP 152
>At5g28700 putative protein
Length = 292
Score = 80.5 bits (197), Expect = 2e-15
Identities = 42/78 (53%), Positives = 52/78 (65%)
Query: 71 TYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANG 130
TY IFR R+RM K +F+RIV S+ YF QR DA + S L K T +RMLA G
Sbjct: 33 TYPPHIFRHRFRMNKPLFMRIVERFSNEVPYFKQRRDATGRLDFSALQKSTAAIRMLAYG 92
Query: 131 VAADAVDEYIKIGETTTL 148
+AADAVDEY++IGE+T L
Sbjct: 93 IAADAVDEYLRIGESTLL 110
>At1g24370 hypothetical protein
Length = 413
Score = 80.5 bits (197), Expect = 2e-15
Identities = 36/69 (52%), Positives = 48/69 (69%)
Query: 121 TTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILH 180
T RMLA GV AD+ DEYIKIGE+T LE L+RFC+ I+ ++ YLR+P D+ R+LH
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACRYLRSPDANDVARLLH 61
Query: 181 VNEMWGVPK 189
+ E G P+
Sbjct: 62 IGESRGFPR 70
>At2g13770 hypothetical protein
Length = 244
Score = 79.7 bits (195), Expect = 3e-15
Identities = 37/74 (50%), Positives = 50/74 (67%)
Query: 121 TTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILH 180
T RMLA GV AD+ DEYIKIGE+T LE L+RFC+ I+ ++ YLR+P D+ R+LH
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACCYLRSPDANDVARLLH 61
Query: 181 VNEMWGVPKHDREY 194
+ E G P+ +Y
Sbjct: 62 IGESRGFPEAVADY 75
>At2g14730 hypothetical protein
Length = 117
Score = 56.6 bits (135), Expect = 2e-08
Identities = 24/52 (46%), Positives = 38/52 (72%)
Query: 135 AVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMWG 186
AVD+Y++IGE T + C+ + II L+ +EYLR PT++DL+R+L + E+ G
Sbjct: 19 AVDKYLRIGENTLMSCMIHSVEAIIYLFGKEYLRRPTRQDLKRLLRIGELRG 70
>At5g12010 putative protein
Length = 502
Score = 37.4 bits (85), Expect = 0.015
Identities = 27/107 (25%), Positives = 47/107 (43%), Gaps = 4/107 (3%)
Query: 72 YDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGV 131
Y + F++ +RM K F I +L+S+ + D A + I + + LA G
Sbjct: 170 YPEEDFKKAFRMSKSTFELICDELNSA----VAKEDTALRNAIPVRQRVAVCIWRLATGE 225
Query: 132 AADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRI 178
V + +G +T + + CK I + +YL+ P E L+ I
Sbjct: 226 PLRLVSKKFGLGISTCHKLVLEVCKAIKDVLMPKYLQWPDDESLRNI 272
>At1g36675 putative protein
Length = 268
Score = 34.7 bits (78), Expect = 0.095
Identities = 15/19 (78%), Positives = 17/19 (88%)
Query: 94 DLSSSDNYFTQRVDAAKKE 112
DLSSSDNYFT+R +A KKE
Sbjct: 175 DLSSSDNYFTKRFEATKKE 193
>At4g29780 unknown protein
Length = 540
Score = 33.9 bits (76), Expect = 0.16
Identities = 23/109 (21%), Positives = 47/109 (43%), Gaps = 4/109 (3%)
Query: 67 LNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRM 126
++ P + + FRR +RM K F I +L ++ + + ++ I + +
Sbjct: 203 VSRPDFPEDEFRREFRMSKSTFNLICEELDTT----VTKKNTMLRDAIPAPKRVGVCVWR 258
Query: 127 LANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDL 175
LA G V E +G +T + + C+ I + +YL P+ ++
Sbjct: 259 LATGAPLRHVSERFGLGISTCHKLVIEVCRAIYDVLMPKYLLWPSDSEI 307
>At3g57030 unknown protein
Length = 374
Score = 28.9 bits (63), Expect = 5.2
Identities = 28/106 (26%), Positives = 43/106 (40%), Gaps = 11/106 (10%)
Query: 82 RMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLA--NGVAADAVDEY 139
R Q+ FL V ++ + + + D + K K T ++ LA NGVA +
Sbjct: 182 RFQRRQFLAAVLNVDKTGRFI--KYDRSSK-------KATVLLQGLAFANGVALSKDRSF 232
Query: 140 IKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMW 185
+ + ETTT + LR + G Q + P D R E W
Sbjct: 233 VLVVETTTCKILRLWLSGPNAGTHQVFAELPGFPDNIRRNSNGEFW 278
>At5g08550 putative protein
Length = 908
Score = 28.5 bits (62), Expect = 6.8
Identities = 23/79 (29%), Positives = 35/79 (44%), Gaps = 8/79 (10%)
Query: 90 RIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLE 149
RIVA LS + V + PL CT T+R + +D+ ETT L
Sbjct: 826 RIVASLSGV--WTGPSVTRTHSRPLQPLVDCTLTLRRILEKRLGSGLDD----AETTGL- 878
Query: 150 CLRRFCKGIIRLYEQEYLR 168
RR + ++ L+E ++ R
Sbjct: 879 -ARRLKRILVELHEHDHAR 896
>At2g33435 RRM-containing protein
Length = 979
Score = 28.5 bits (62), Expect = 6.8
Identities = 32/130 (24%), Positives = 50/130 (37%), Gaps = 19/130 (14%)
Query: 19 KSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGANQ-------RLIDDYFLNEPT 71
K R + + + R + I S+ RG K + DH GA Q ++ DY E
Sbjct: 449 KKVRREERVKDNSRKKEEAI---SSSRGEKPIKEDHVGAAQLLGNDLVEMVSDYHETEKG 505
Query: 72 YDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGV 131
YD R K + +SSS + +D KK+ + T+ + A G
Sbjct: 506 YDRSKKLSREERVKDSSRKKEEAISSSRE---ENLDKQKKD------ESTSNRKRKAEGE 556
Query: 132 AADAVDEYIK 141
+ A E I+
Sbjct: 557 CSTAETESIE 566
>At1g64330 unknown protein
Length = 555
Score = 28.1 bits (61), Expect = 8.9
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 12/64 (18%)
Query: 12 DLEAYLEKSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGAN----------QRL 61
+ EA LE+ ++E +LN+ D + +LE A LS++H N ++L
Sbjct: 241 ETEAELEREKQEKPALLNQINDVQKALLEQEA--AYNTLSQEHKQINGLFEEREATIKKL 298
Query: 62 IDDY 65
DDY
Sbjct: 299 TDDY 302
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.345 0.152 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,224,106
Number of Sequences: 26719
Number of extensions: 314220
Number of successful extensions: 1270
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1259
Number of HSP's gapped (non-prelim): 16
length of query: 400
length of database: 11,318,596
effective HSP length: 101
effective length of query: 299
effective length of database: 8,619,977
effective search space: 2577373123
effective search space used: 2577373123
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0349b.2


