

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0348.12
(180 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
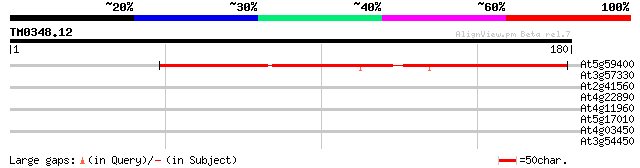
At5g59400 unknown protein 120 3e-28
At3g57330 Ca2+-transporting ATPase-like protein 28 2.7
At2g41560 putative Ca2+-ATPase 28 2.7
At4g22890 unknown protein 27 5.9
At4g11960 unknown protein 27 5.9
At5g17010 sugar transporter - like protein 27 7.7
At4g03450 hypothetical protein 27 7.7
At3g54450 oligopeptide transporter -like protein 27 7.7
>At5g59400 unknown protein
Length = 299
Score = 120 bits (302), Expect = 3e-28
Identities = 67/136 (49%), Positives = 85/136 (62%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + S L S+ + N I+ VLG GYP+AS+ V+V++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRVLKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>At3g57330 Ca2+-transporting ATPase-like protein
Length = 1025
Score = 28.1 bits (61), Expect = 2.7
Identities = 12/37 (32%), Positives = 21/37 (56%)
Query: 135 NIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGA 171
N++ +++ FV SAP+ +Q LW N ++ GA
Sbjct: 812 NVVALIINFVSACITGSAPLTAVQLLWVNMIMDTLGA 848
>At2g41560 putative Ca2+-ATPase
Length = 1030
Score = 28.1 bits (61), Expect = 2.7
Identities = 12/37 (32%), Positives = 21/37 (56%)
Query: 135 NIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGA 171
N++ +++ FV SAP+ +Q LW N ++ GA
Sbjct: 815 NVVALIINFVSACITGSAPLTAVQLLWVNMIMDTLGA 851
>At4g22890 unknown protein
Length = 324
Score = 26.9 bits (58), Expect = 5.9
Identities = 9/15 (60%), Positives = 11/15 (73%)
Query: 164 DLVALRGACPNCGEE 178
D + L+G CPNCG E
Sbjct: 264 DFLILKGPCPNCGTE 278
>At4g11960 unknown protein
Length = 313
Score = 26.9 bits (58), Expect = 5.9
Identities = 9/15 (60%), Positives = 11/15 (73%)
Query: 164 DLVALRGACPNCGEE 178
D + L+G CPNCG E
Sbjct: 253 DFLILKGPCPNCGTE 267
>At5g17010 sugar transporter - like protein
Length = 503
Score = 26.6 bits (57), Expect = 7.7
Identities = 31/105 (29%), Positives = 43/105 (40%), Gaps = 16/105 (15%)
Query: 68 TYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGL 127
T AD GRR L A L+L + L P +Y+V I + +G + H +
Sbjct: 113 TIADVIGRRKELIL----AALLYLVGALVTALAP-TYSVLIIGRVIYGVSVGLAMHAAPM 167
Query: 128 GFS----------IIVNNIIFIVLGFVIGYPLASAPVKVIQGLWR 162
+ ++ FIVLG V GY + S V V G WR
Sbjct: 168 YIAETAPSPIRGQLVSLKEFFIVLGMVGGYGIGSLTVNVHSG-WR 211
>At4g03450 hypothetical protein
Length = 641
Score = 26.6 bits (57), Expect = 7.7
Identities = 20/60 (33%), Positives = 30/60 (49%), Gaps = 4/60 (6%)
Query: 101 PLSYAVGIAYQNAFGSGL-SHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQG 159
P + ++ +AF G+ + HNP L SII +IIF+ FV+ Y LA + G
Sbjct: 544 PALFVSLVSMSSAFFCGVVATTKHNPWLFDSIIFISIIFL---FVVAYLLAPYAIPQFPG 600
>At3g54450 oligopeptide transporter -like protein
Length = 488
Score = 26.6 bits (57), Expect = 7.7
Identities = 10/22 (45%), Positives = 16/22 (72%)
Query: 127 LGFSIIVNNIIFIVLGFVIGYP 148
LGFSII +++ ++ F+IG P
Sbjct: 120 LGFSIIAGSVVIAIVIFLIGIP 141
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.340 0.149 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,873,515
Number of Sequences: 26719
Number of extensions: 144695
Number of successful extensions: 421
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 411
Number of HSP's gapped (non-prelim): 10
length of query: 180
length of database: 11,318,596
effective HSP length: 93
effective length of query: 87
effective length of database: 8,833,729
effective search space: 768534423
effective search space used: 768534423
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0348.12


