

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0344.6
(242 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
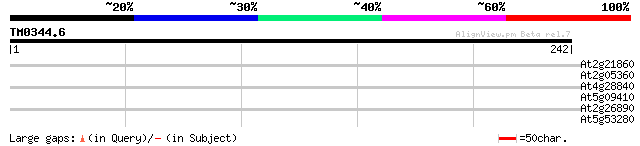
Score E
Sequences producing significant alignments: (bits) Value
At2g21860 unknown protein 30 1.2
At2g05360 hypothetical protein 29 2.0
At4g28840 putative protein 28 4.4
At5g09410 Calmodulin-binding transcription activator 1 (CAMTA1) 28 5.8
At2g26890 unknown protein 27 7.5
At5g53280 unknown protein 27 9.9
>At2g21860 unknown protein
Length = 522
Score = 30.0 bits (66), Expect = 1.2
Identities = 46/202 (22%), Positives = 84/202 (40%), Gaps = 34/202 (16%)
Query: 11 NNSEE------NDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPY 64
NNSE N N++ + S P + N++ T + N D E+ V K+ V +
Sbjct: 145 NNSESVNWIQTNSKNVKNMICFESSPNLM-NRLGGTDVGSVNKDKEVTEVVKT--VGDAW 201
Query: 65 QSISILDSR-----IANQHVRKYGYLHIGLVQVAVKPLT-RLRYCDLPILICLRDKRMLR 118
+ + D R I N ++R L L L+ ++ C IL CL D
Sbjct: 202 ERRNSDDIRFCLLVIINAYIRPVPVLQ-NLRSKGFSTLSCMVKNCGPQILNCLLDPNC-- 258
Query: 119 FHESVVAVLESNIQNGPAYFNCFSSYP--------VCLRDPHVTTCVDLDIKIPNEMFID 170
+++ + + + + + C +SY +C+ H C++LD KIP + +
Sbjct: 259 -RKALQCLNQCSPVDQVCSYRCIASYEGPYFEAFSLCVLQKH--NCLELDAKIPEKPY-- 313
Query: 171 GIPPVSIM--YRFCYKVTDNLF 190
+PP++ C+ ++LF
Sbjct: 314 -VPPMTSFRGKELCHDTAEDLF 334
>At2g05360 hypothetical protein
Length = 358
Score = 29.3 bits (64), Expect = 2.0
Identities = 18/68 (26%), Positives = 33/68 (48%), Gaps = 2/68 (2%)
Query: 72 SRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNI 131
SR+ + L +G+VQV + + RL+ C + CL + +L+ + ++S I
Sbjct: 226 SRLLKEGHTNNNILQVGIVQVGIVQVCRLQICLQEVCFCLYE--LLQTTTLLRLQVKSKI 283
Query: 132 QNGPAYFN 139
GP+ N
Sbjct: 284 LTGPSITN 291
>At4g28840 putative protein
Length = 151
Score = 28.1 bits (61), Expect = 4.4
Identities = 15/56 (26%), Positives = 24/56 (42%)
Query: 12 NSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSI 67
N + ++ L + KIR + Y + F P Y T ++ +QG Y SI
Sbjct: 24 NGSDKPKQPQRGLGVAQLEKIRLHGEYNCNSFNTYPSYHPSTYQEDVRIQGGYPSI 79
>At5g09410 Calmodulin-binding transcription activator 1 (CAMTA1)
Length = 1007
Score = 27.7 bits (60), Expect = 5.8
Identities = 15/61 (24%), Positives = 29/61 (46%), Gaps = 5/61 (8%)
Query: 155 TCVDLDIKIPNEMFIDGI-----PPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEIN 209
+C+ ++++P E+ +DG+ PP + + Y N F VR++ +IN
Sbjct: 448 SCMFGEVEVPAEILVDGVLCCHAPPHTAGHVPFYVTCSNRFACSEVREFDFLSGSTQKIN 507
Query: 210 A 210
A
Sbjct: 508 A 508
>At2g26890 unknown protein
Length = 2535
Score = 27.3 bits (59), Expect = 7.5
Identities = 15/53 (28%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Query: 15 ENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSI 67
E SN+ +++QN K++ Y+ ++PD + EK AVQ Y+ +
Sbjct: 1505 EEISNISKQIQNLDEEKLKRQ--YRKLAMRYHPDKNPEGREKFLAVQKAYECL 1555
>At5g53280 unknown protein
Length = 272
Score = 26.9 bits (58), Expect = 9.9
Identities = 22/80 (27%), Positives = 38/80 (47%), Gaps = 4/80 (5%)
Query: 14 EENDSNLEQKLQNWSIPKIRTN-QVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISILDS 72
++NDS +++ S+ IRT + + + F+ + + EK A+ QS +L
Sbjct: 72 DDNDSTIQEAK---SLNAIRTALENLEDQLEFFHTIHTQQRTEKDVAIARLEQSRILLAM 128
Query: 73 RIANQHVRKYGYLHIGLVQV 92
R+A H + YG L L V
Sbjct: 129 RLAEHHGKNYGVLEEALAFV 148
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,424,109
Number of Sequences: 26719
Number of extensions: 211922
Number of successful extensions: 459
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 458
Number of HSP's gapped (non-prelim): 6
length of query: 242
length of database: 11,318,596
effective HSP length: 96
effective length of query: 146
effective length of database: 8,753,572
effective search space: 1278021512
effective search space used: 1278021512
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0344.6


