

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0342b.6
(427 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
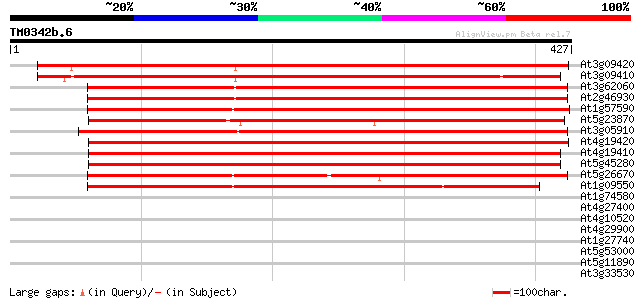
Sequences producing significant alignments: (bits) Value
At3g09420 putative pectinacetylesterase 513 e-145
At3g09410 putative pectinacetylesterase 477 e-135
At3g62060 pectinacetylesterase precursor-like protein 429 e-120
At2g46930 putative pectinesterase 421 e-118
At1g57590 unknown protein 415 e-116
At5g23870 pectinacetylesterase 412 e-115
At3g05910 putative pectinacetylesterase 407 e-114
At4g19420 pectinacetylesterase like protein 401 e-112
At4g19410 putative pectinacetylesterase protein 390 e-109
At5g45280 pectin acetylesterase 373 e-104
At5g26670 pectin acetylesterase precursor - like protein 372 e-103
At1g09550 putative pectinacetylesterase precursor 335 4e-92
At1g74580 hypothetical protein 31 1.1
At4g27400 unknown protein (At4g27400) 31 1.5
At4g10520 putative subtilisin-like protease 30 2.5
At4g29900 Ca2+-transporting ATPase - like protein 29 5.7
At1g27740 putative bHLH transcription factor (AtbHLH054) 29 5.7
At5g53000 PP2A regulatory subunit (TAP46) 28 7.4
At5g11890 unknown protein 28 7.4
At3g33530 unknown protein 28 7.4
>At3g09420 putative pectinacetylesterase
Length = 427
Score = 513 bits (1320), Expect = e-145
Identities = 243/415 (58%), Positives = 308/415 (73%), Gaps = 11/415 (2%)
Query: 22 RKLTTKDYAIAAFAFFLLLS-LAFF--------SQLHHHHHSHSLSNLIPFTPLPNAILR 72
RK D+ +A+ L++ L+FF + S S+L+ A R
Sbjct: 13 RKWAKSDWLVASIGCVLIVFFLSFFFDPTSDSVPSVDRSRPIISPSDLVKLKLSSVAKER 72
Query: 73 GAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTALGSSKYMETVVP 132
GA CLDGS PG+HF +G GSGS+SWL+HLEGGGWCN+++SCS R T LGSS Y E V
Sbjct: 73 GAFCLDGSLPGYHFHEGSGSGSQSWLVHLEGGGWCNTVASCSARALTKLGSSNYFEQEVA 132
Query: 133 FSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESE--KGSGLFFRGQIIWEAIMDEL 190
F G+LSSD SQNP+FFNWNKV IRYCDGASF+G PE+E G+ LFFRGQ+IWEAI+DEL
Sbjct: 133 FQGVLSSDPSQNPEFFNWNKVAIRYCDGASFSGRPEAEFKNGTRLFFRGQLIWEAIIDEL 192
Query: 191 LSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFFLDEKDIRGNNT 250
LS+GMS AKQA+L+GCSAGGLA+L+HCD FR LPK+ VKC++D G+FL+ D+ GN T
Sbjct: 193 LSMGMSDAKQAILTGCSAGGLASLIHCDYFRDHLPKDAAVKCVSDGGYFLNVPDVLGNPT 252
Query: 251 MRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNI 310
MRSFY+DVV LQGV KSL ++C+AK EPSKC+FP E LKNI+TPVFLV+ AYDFWQI+++
Sbjct: 253 MRSFYHDVVNLQGVEKSLDQKCVAKTEPSKCMFPQEFLKNIRTPVFLVNPAYDFWQIQHV 312
Query: 311 LVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIH 370
LVP +DP W C+LNI C+A I +L+ FRSS++ A+ EF Q K GMF++SC+ H
Sbjct: 313 LVPTSADPDKSWAKCRLNIKECDAEQIKVLHGFRSSMMTAIGEFHQNKDGGMFIDSCYAH 372
Query: 371 CQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDFT 425
CQT TW+S SP+I+NKTIAESV DW+F+R+ + LIDCPYPCNP+C N++FT
Sbjct: 373 CQTVMSVTWHSLTSPRIENKTIAESVGDWYFNRKPVKLIDCPYPCNPSCYNMNFT 427
>At3g09410 putative pectinacetylesterase
Length = 409
Score = 477 bits (1228), Expect = e-135
Identities = 235/403 (58%), Positives = 295/403 (72%), Gaps = 8/403 (1%)
Query: 22 RKLTTKDYAIAAFAFFLLL---SLAFFSQLHHHHHSHSLSNLIPFTPLPNAILRGAICLD 78
R + D+ +A+ L++ SL+F S S S+L+ A RGA CLD
Sbjct: 10 RTWSKSDWLLASIGIVLIVYSFSLSFNST-SDSIPSVDRSDLVKLKLSSKAKERGAFCLD 68
Query: 79 GSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTALGSSKYMETVVPFSGILS 138
GS PG+HF KG GSGS SWLL+LEGGG C +I SCS R T LGSS + E VPF G+LS
Sbjct: 69 GSLPGYHFHKGSGSGSNSWLLYLEGGGGCRTIESCSARAMTRLGSSNFFEHEVPFFGVLS 128
Query: 139 SDRSQNPDFFNWNKVMIRYCDGASFAGHPESE--KGSGLFFRGQIIWEAIMDELLSIGMS 196
SD SQNPDFFNWN+VMIRYCDGA F+GHPE+E + LFFRGQ+IWEAIMDELLS+GMS
Sbjct: 129 SDPSQNPDFFNWNRVMIRYCDGACFSGHPEAEFKNETRLFFRGQLIWEAIMDELLSMGMS 188
Query: 197 KAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFFLDEKDIRGNNTMRSFYN 256
AK+A+L+GCSAGGL+TL+HCD FR LPK+ TVKC++D G+ L+ D+ GN TM SF++
Sbjct: 189 HAKRAMLTGCSAGGLSTLIHCDYFRDHLPKDATVKCVSDGGYILNVLDVLGNPTMGSFFH 248
Query: 257 DVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNILVPEGS 316
DVV LQ V KSL + C+AKMEPSKC+FP E LKNI+TPVFLV++AYD+WQI+N LVP+
Sbjct: 249 DVVTLQSVDKSLDQNCVAKMEPSKCMFPQESLKNIRTPVFLVNTAYDYWQIQNGLVPDSP 308
Query: 317 DPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRG 376
D WK C+LNI C+A + +L+ FRSSL+ A+ EF K+ GMF+NSC HCQ
Sbjct: 309 DLDERWKICRLNIQECDAAQMKVLHGFRSSLIDAIGEFHVNKEGGMFINSCNSHCQI--R 366
Query: 377 ETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
E+W+S S +I+NKTIAESV DW+F+R+ + LIDCPYPCN +C
Sbjct: 367 ESWHSATSTRIENKTIAESVGDWYFNRKPVKLIDCPYPCNASC 409
>At3g62060 pectinacetylesterase precursor-like protein
Length = 419
Score = 429 bits (1102), Expect = e-120
Identities = 188/366 (51%), Positives = 262/366 (71%), Gaps = 2/366 (0%)
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
+IP T + A +GA+CLDG+ PG+H +GFGSG+ SWL+ LEGGGWCN+ SC RK +
Sbjct: 54 MIPLTLIHGADSKGAVCLDGTLPGYHLDRGFGSGANSWLIQLEGGGWCNNHRSCVYRKTS 113
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSSK+ME + F+GILS+ +NPDFFNWN++ +RYCDGASF+G + E S LF+RG
Sbjct: 114 RRGSSKFMEKALAFTGILSNRSEENPDFFNWNRIKLRYCDGASFSGDSQDES-SQLFYRG 172
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
Q IW+ M+E LS+GM +A QALLSGCSAGGLA+++HCD+FR++LP T VKCL+DAG F
Sbjct: 173 QRIWQVAMEEFLSLGMKQANQALLSGCSAGGLASILHCDEFRELLPSSTKVKCLSDAGMF 232
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
LD D+ G +++R+ + VV +Q + K L C ++P+ C FP ++ +IKTP+FL++
Sbjct: 233 LDSVDVSGGHSLRNMFQGVVTVQNLQKDLSSTCTNHLDPTSCFFPQNLVSDIKTPMFLLN 292
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+ L P +DP G WK+CK + CN++ I FR+ +L A+N F Q
Sbjct: 293 TAYDSWQIQESLAPPTADPGGIWKACKSDHSRCNSSQIQFFQEFRNQMLFAVNSFSNSDQ 352
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDR-EVINLIDCPYPCNPT 418
G+++NSCF HCQT R +TW++ +SP++ K +AESV DW+FDR + + IDCPYPC+ T
Sbjct: 353 NGLYINSCFAHCQTERQDTWFAQDSPQLNGKRVAESVGDWYFDRAKNVKAIDCPYPCDTT 412
Query: 419 CRNLDF 424
C NL F
Sbjct: 413 CHNLIF 418
>At2g46930 putative pectinesterase
Length = 416
Score = 421 bits (1083), Expect = e-118
Identities = 185/365 (50%), Positives = 255/365 (69%), Gaps = 1/365 (0%)
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++P T + A +GA+CLDG+ PG+H G GSG+ WL+ LEGGGWCN+ SC RK T
Sbjct: 52 MVPLTLIQAAASKGAVCLDGTLPGYHLHPGSGSGANRWLIQLEGGGWCNTRRSCIFRKTT 111
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSS +ME V+ F+GILS+ ++NPDFFNWN+V +RYCDGASF G + E S L++RG
Sbjct: 112 RRGSSNHMEKVLAFTGILSNKSNENPDFFNWNRVKLRYCDGASFTGDSQDES-SQLYYRG 170
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
Q IW + M+ELLS GM KA+QALLSGCSAGGLA+++HCD F+++ P TTVKCL+DAG F
Sbjct: 171 QRIWHSAMEELLSKGMQKAEQALLSGCSAGGLASILHCDQFKELFPGTTTVKCLSDAGMF 230
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
+D D+ G +++R + VV +Q + K L C ++P+ C FP ++ IKTP+FL++
Sbjct: 231 MDAVDVSGGHSLRKMFQGVVTVQNLQKELSTACTKHLDPTSCFFPQNLVSGIKTPMFLLN 290
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQ++ L P D SG WK+CK + +CN++ I FR+ ++ A+ F
Sbjct: 291 AAYDAWQVQESLAPPSVDLSGSWKACKSDHSHCNSSQIQFFQDFRTHMVDAVKSFATSTH 350
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
G+F+NSCF HCQ+ R +TWY+P+SP + KT+AESV DW+FDR + IDCPYPC+ TC
Sbjct: 351 NGVFINSCFAHCQSERQDTWYAPDSPTLHGKTVAESVGDWYFDRTTVKAIDCPYPCDKTC 410
Query: 420 RNLDF 424
NL F
Sbjct: 411 HNLIF 415
>At1g57590 unknown protein
Length = 423
Score = 415 bits (1067), Expect = e-116
Identities = 184/367 (50%), Positives = 258/367 (70%), Gaps = 1/367 (0%)
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++ T + +A +GA+CLDGS PG+H +GFGSG+ +WL+ LEGGGWC++I +C RK T
Sbjct: 58 MVGLTLIQSAAAKGAVCLDGSLPGYHLHRGFGSGANNWLVQLEGGGWCDTIRNCVYRKTT 117
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSS YME +PF+GILS + NPDF+NWN+V +RYCDG SF+G E+ K + L FRG
Sbjct: 118 RRGSSSYMEKEIPFTGILSDKAADNPDFYNWNRVKVRYCDGGSFSGDSEN-KAAQLQFRG 176
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
+ IW A M++L++ GM +AKQALLSGCSAGGLA ++ CDDF ++ P T VKCL+DAGFF
Sbjct: 177 KRIWLAAMEDLMAKGMRQAKQALLSGCSAGGLAVILRCDDFGKLFPPSTRVKCLSDAGFF 236
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
LD D+ G ++R Y VV+LQ + +L + C+ ++ P+ C FP ++ +KTP+F+++
Sbjct: 237 LDAIDVSGGRSLRRLYAGVVRLQNLQTNLPQYCVNRLNPTSCFFPQNLINQVKTPLFILN 296
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+ L P+ +DPSG W C+LN C+A+ I L FR+ ++ + F +
Sbjct: 297 AAYDSWQIQESLAPKSADPSGSWNDCRLNYAKCSASQIQFLQGFRTRMVNLVKGFAMPSK 356
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
G+F+NSCF HCQT R +TW++ NSP I+NK IA +V DW+F+R LIDC YPC+ TC
Sbjct: 357 NGVFLNSCFAHCQTERHDTWFAQNSPAIKNKGIAVAVGDWYFERGGAKLIDCAYPCDKTC 416
Query: 420 RNLDFTR 426
NL F R
Sbjct: 417 HNLVFRR 423
>At5g23870 pectinacetylesterase
Length = 415
Score = 412 bits (1058), Expect = e-115
Identities = 198/369 (53%), Positives = 257/369 (68%), Gaps = 9/369 (2%)
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+ T + +A GA CLDGS P +H +GFG+GS +W+L EGGGWCN I+SC R T
Sbjct: 35 VSMTLVRDAAALGAFCLDGSLPAYHLDRGFGAGSNNWILQFEGGGWCNDIASCVERAKTR 94
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSG---LFF 177
GS++YM V F+G+LS++ SQNPDF+NWNKV +RYCDGASFAG +S+ G+G L+F
Sbjct: 95 RGSTRYMSKTVVFTGVLSNNASQNPDFYNWNKVRLRYCDGASFAG--DSQFGNGTSLLYF 152
Query: 178 RGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAG 237
RGQ IW AI+ +LL G++KA +ALL+GCSAGGL+T +HCD+F LPK +VKC++DAG
Sbjct: 153 RGQRIWNAIILDLLPKGLAKAHKALLTGCSAGGLSTFLHCDNFTSYLPKNASVKCMSDAG 212
Query: 238 FFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKM--EPSKCLFPSEILKNIKTPV 295
FFLD D+ N TMRSFY+ +V LQG+ K+L C EPS C FP +L+ IKTP
Sbjct: 213 FFLDAIDVAANRTMRSFYSQLVSLQGIQKNLDPSCTHAFFPEPSLCFFPQYVLRFIKTPF 272
Query: 296 FLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKAL-NEF 354
F+++SAYD +Q + LVP +D +G W CKLN+ CN + +D L FR +L AL N F
Sbjct: 273 FILNSAYDVFQFHHGLVPPSADQTGRWNRCKLNVTACNPHQLDALQGFRKDMLGALMNFF 332
Query: 355 QQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDR-EVINLIDCPY 413
+ + GMF+NSCF HCQ+ ETW SP SP+I NKTIAE+V DW+F R E I CPY
Sbjct: 333 RNSTRGGMFINSCFDHCQSALEETWLSPTSPRINNKTIAETVGDWYFGRGEEAKEIGCPY 392
Query: 414 PCNPTCRNL 422
PC+ TC NL
Sbjct: 393 PCDKTCHNL 401
>At3g05910 putative pectinacetylesterase
Length = 415
Score = 407 bits (1047), Expect = e-114
Identities = 182/372 (48%), Positives = 252/372 (66%), Gaps = 1/372 (0%)
Query: 53 HSHSLSNLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISS 112
+S +L+ L+ L GA+CLDG+ PG+H +G GSG+ SWL+ LEGGGWCN+I +
Sbjct: 44 YSSNLNPLMVGLTLIRGADSGAVCLDGTLPGYHLHRGHGSGANSWLIQLEGGGWCNNIRT 103
Query: 113 CSRRKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKG 172
C RK T GSS YME + F+GILS +NPDFFNWN+V +RYCDGASF+G +++
Sbjct: 104 CVYRKTTRRGSSNYMEKQLQFTGILSDKAQENPDFFNWNRVKLRYCDGASFSGDGQNQAA 163
Query: 173 SGLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKC 232
L FRG+ IW A +D+L + GM A QALLSGCSAGGLA ++ CD+FR + P T VKC
Sbjct: 164 Q-LQFRGERIWRAAIDDLKANGMRYANQALLSGCSAGGLAAILRCDEFRNLFPGSTKVKC 222
Query: 233 LADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIK 292
L+DAG FLD D+ G T+R+ YN VV+LQ V +L + C ++P+ C FP ++ +K
Sbjct: 223 LSDAGLFLDTADVSGGRTIRNLYNGVVELQSVKNNLPRICTNHLDPTSCFFPQNLISQMK 282
Query: 293 TPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALN 352
TP+F+V++AYD WQI++ + P +DPSG W C+LN C + L FR +L+ +
Sbjct: 283 TPLFIVNAAYDTWQIQSSIAPTSADPSGFWHDCRLNHGKCTPAQLRFLQGFREQMLRVVK 342
Query: 353 EFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCP 412
F +Q G+F+NSCF HCQT R +TW++ +SP I+ K +A +V DW+FDR + L+DCP
Sbjct: 343 GFSMSRQNGLFINSCFAHCQTERQDTWFADDSPVIRKKAVAIAVGDWYFDRAEVKLVDCP 402
Query: 413 YPCNPTCRNLDF 424
YPC+ +C NL F
Sbjct: 403 YPCDKSCHNLVF 414
>At4g19420 pectinacetylesterase like protein
Length = 397
Score = 401 bits (1031), Expect = e-112
Identities = 178/366 (48%), Positives = 253/366 (68%), Gaps = 1/366 (0%)
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+ T + NA+ +GA+CLDGS P +H +G G+G SWL+ LEGGGWCN++++C R T
Sbjct: 25 VNITFVRNAVAKGAVCLDGSPPAYHLDRGSGTGINSWLIQLEGGGWCNNVTNCVSRMHTR 84
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
LGSSK M + FS ILS+ + NPDF+NWN+V +RYCDGASF G E+ + L FRG
Sbjct: 85 LGSSKKMVENLAFSAILSNKKQYNPDFYNWNRVKVRYCDGASFTGDVEAVNPATNLHFRG 144
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
+W A+M ELL+ GM A+ A+LSGCSAGGLA+L+HCD FR +LP T VKCL+DAGFF
Sbjct: 145 ARVWLAVMQELLAKGMINAENAVLSGCSAGGLASLMHCDSFRALLPMGTKVKCLSDAGFF 204
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
L+ +D+ G +++++ DVV L G AK+L + C +++ P+ C FP + + I+TP+F+++
Sbjct: 205 LNTRDVSGVQYIKTYFEDVVTLHGSAKNLPRSCTSRLTPAMCFFPQYVARQIRTPLFILN 264
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+NIL P +DP G W+SC+L+I NC+ + I ++ FR L A+ +
Sbjct: 265 AAYDSWQIKNILAPRAADPYGKWQSCQLDIKNCHPSQIKVMQDFRLEFLSAVIGLGRSSS 324
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
GMF++SC+ HCQT +W+ +SP + TIA++V DW +DR + IDCPYPCNPTC
Sbjct: 325 RGMFIDSCYTHCQTETQTSWFWQDSPILNRTTIAKAVGDWVYDRTLFQKIDCPYPCNPTC 384
Query: 420 RNLDFT 425
+ FT
Sbjct: 385 HHRVFT 390
>At4g19410 putative pectinacetylesterase protein
Length = 391
Score = 390 bits (1002), Expect = e-109
Identities = 168/361 (46%), Positives = 247/361 (67%), Gaps = 2/361 (0%)
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+P T L +A+ +GA+CLDGSAP +HF KGFGSG +W++H+EGGGWC ++SC+ RK T
Sbjct: 24 VPITYLQSAVAKGAVCLDGSAPAYHFDKGFGSGVNNWIVHMEGGGWCTDVASCNERKGTM 83
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
GSSK+M FSGIL +S NPDF+NWN++ +RYCDG+SF G+ E+ + LFFRG
Sbjct: 84 KGSSKFMNKDFGFSGILGGKQSTNPDFYNWNRIKVRYCDGSSFTGNVEAVNPANKLFFRG 143
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
+W A++D+L++ GM A+ A+LSGCSAG LA ++HCD FR ILP+ +VKC++DAG+F
Sbjct: 144 ARVWRAVVDDLMAKGMKNAQNAILSGCSAGALAAILHCDTFRAILPRTASVKCVSDAGYF 203
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
+ KDI G + ++S+Y+ VV L G AKSL C +KM+P C FP ++ +++TP+F+++
Sbjct: 204 IHGKDITGGSYIQSYYSKVVALHGSAKSLPVSCTSKMKPELCFFPQYVVPSMRTPLFVIN 263
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+A+D WQI+N+L P D WK+CKL++ C+A + + FR +++AL+
Sbjct: 264 AAFDSWQIKNVLAPTAVDKGKEWKNCKLDLKKCSAAQLKTVQGFRDQMMRALSPVHSTPS 323
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYP-CNPT 418
G+F++SC HCQ +W P++ N IA++V +WF+ R IDCP P CNPT
Sbjct: 324 RGLFLDSCHAHCQGGSAASWSGDKGPQVANTRIAKAVGNWFYGRSAFQKIDCPSPTCNPT 383
Query: 419 C 419
C
Sbjct: 384 C 384
>At5g45280 pectin acetylesterase
Length = 391
Score = 373 bits (958), Expect = e-104
Identities = 166/361 (45%), Positives = 236/361 (64%), Gaps = 2/361 (0%)
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+P T L +A+ +GA+CLDGSAP +HF KG GSG +W++H+EGGGWC I++C +RK T
Sbjct: 24 VPITYLESAVAKGAVCLDGSAPAYHFDKGSGSGVNNWIVHMEGGGWCTDIATCVQRKSTM 83
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
GSSK M FSGIL +S NPDF+NWN++ +RYCDG+SF G E+ + LFFRG
Sbjct: 84 KGSSKLMNKDFGFSGILGGKQSTNPDFYNWNRIKVRYCDGSSFTGDIEAVDPTHKLFFRG 143
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
+W A++D+L++ GMS A+ A+LSGCSAG LA ++HCD F+ LPK VKC++DAG+F
Sbjct: 144 ARVWRAVIDDLMAKGMSNAQNAILSGCSAGALAAILHCDQFKSTLPKTAKVKCVSDAGYF 203
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
+ KDI G + ++S+Y VV G AKSL C + M+P C FP + K ++TP+F+++
Sbjct: 204 IHGKDITGGSYIQSYYAKVVATHGSAKSLPASCTSSMKPDLCFFPQYVAKTLQTPLFVIN 263
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+A+D WQI+N+L P D S WK+CKL++ C A + + +R +L AL +
Sbjct: 264 AAFDSWQIKNVLAPTSVDKSKAWKTCKLDLKKCTAAQLQTVQGYRDQVLAALAPVRSATT 323
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDC-PYPCNPT 418
G+F++SC HCQ TW P + N +A++V DWFF+R +DC CNPT
Sbjct: 324 NGLFLDSCHAHCQGGSAATWSGDKGPTVANTKMAKAVGDWFFERSTFQNVDCSSLNCNPT 383
Query: 419 C 419
C
Sbjct: 384 C 384
>At5g26670 pectin acetylesterase precursor - like protein
Length = 422
Score = 372 bits (954), Expect = e-103
Identities = 176/373 (47%), Positives = 244/373 (65%), Gaps = 11/373 (2%)
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++ T + A +GA+CLDG+ PG+H +G GSG+ SWL+ LEGGGWC++I +C RK +
Sbjct: 52 MVGLTLIRGAGSKGAVCLDGTLPGYHLHRGHGSGANSWLIQLEGGGWCDNIRNCVYRKKS 111
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSS YME + F+GILS+ +NPDFFNWN+V +RYCDG SF+G ++ K + L FRG
Sbjct: 112 RRGSSNYMEKQIQFTGILSNKAQENPDFFNWNRVKLRYCDGGSFSGDSQN-KAARLQFRG 170
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
+ IW A MD+L + GM AKQALLSGCSAGGLA ++ CD+FR + T VKCL+DAG F
Sbjct: 171 EKIWRAAMDDLKAKGMRNAKQALLSGCSAGGLAVILRCDEFRNLFSGWTRVKCLSDAGLF 230
Query: 240 LDEKDIRGNNTMRSFYN-DVVQLQGVAKSLHKECLAKMEPSK-------CLFPSEILKNI 291
LD + R FY + GV +L C + P+ C FP ++ +
Sbjct: 231 LDT--LVSVIEPRLFYVFKGLMYPGVKNNLPHLCTNHLNPTSVSSSLLSCFFPQNLISQM 288
Query: 292 KTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKAL 351
KTP+F+V++AYD WQI++ + P +DPSG+W C+LN C I L FR+ +L+A+
Sbjct: 289 KTPLFIVNAAYDIWQIQSSIAPPSADPSGYWHECRLNHGRCTPAQIRFLQGFRNQMLRAV 348
Query: 352 NEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDC 411
+ F K+ G+F+NSCF HCQT R +TW++ +SP I K +A +V DW+FDR + LIDC
Sbjct: 349 SGFSNSKKNGLFINSCFAHCQTERQDTWFADDSPVIHKKAVAIAVGDWYFDRAEVKLIDC 408
Query: 412 PYPCNPTCRNLDF 424
PYPC+ +C NL F
Sbjct: 409 PYPCDRSCHNLVF 421
>At1g09550 putative pectinacetylesterase precursor
Length = 363
Score = 335 bits (858), Expect = 4e-92
Identities = 153/344 (44%), Positives = 228/344 (65%), Gaps = 2/344 (0%)
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++ T + A +GA+CLDGS PG+H +G+GSG+ +W++ L+GG WC+SI +C RK +
Sbjct: 16 MVGLTLVQAAAAKGAVCLDGSVPGYHLCRGYGSGANNWIIQLQGGAWCDSIQNCQSRKGS 75
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSS ME + F G+LS+ ++NPDF+NWNKV +RYCDGASF G E+ K + L +RG
Sbjct: 76 GYGSSTLMEKELAFLGLLSNKAAENPDFYNWNKVKVRYCDGASFDGDSEN-KAAQLQYRG 134
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
+ I+ A+M++L+ GM +AKQALLSGCS+GGL+ ++ CDDF + P TTVKC++DAGFF
Sbjct: 135 KRIFLAVMEDLMEKGMRQAKQALLSGCSSGGLSAILRCDDFNNLFPPTTTVKCMSDAGFF 194
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
LD D+ G +++R Y+ VV QG+ +L C + ++P C FP I+ +KTP+F+++
Sbjct: 195 LDAVDVSGGHSLRRMYSGVVNTQGLQNTLPPTCTSHIKPFLCFFPQYIINQVKTPLFILN 254
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
S +D WQI N L P +D SG W +C + C A+ + L F+ S+L AL F + +
Sbjct: 255 SGFDSWQIGNSLAPPSADKSGSWHNCSFSF-RCTASQMHFLEGFKMSMLDALKTFSKFSK 313
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDR 403
G+ + S + HCQ R +TW+ S + K IA +V DW+F+R
Sbjct: 314 NGVLITSGWAHCQAERQDTWFPGYSGAGKAKGIAVAVGDWYFER 357
>At1g74580 hypothetical protein
Length = 763
Score = 31.2 bits (69), Expect = 1.1
Identities = 45/242 (18%), Positives = 91/242 (37%), Gaps = 30/242 (12%)
Query: 123 SSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDG--ASFAGHPESEKGSGLFFRGQ 180
+SK+ E V +++ PD + +N ++ YC G A + F Q
Sbjct: 299 NSKFQEAEVYLGKMVNE--GLEPDSYTYNTLIAGYCKGGMVQLAERIVGDAVFNGFVPDQ 356
Query: 181 IIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFFL 240
+ +++D L G + AL + G+ ++ T +K L++ G L
Sbjct: 357 FTYRSLIDGLCHEGETNRALALFNEALGKGIKP--------NVILYNTLIKGLSNQGMIL 408
Query: 241 DEKDIRGNNTMRSFYNDVVQLQGVAKSLHK-------ECLAKMEPSKCLFPSEILKNIKT 293
+ + + + +V + L K + L K+ SK FP NI
Sbjct: 409 EAAQLANEMSEKGLIPEVQTFNILVNGLCKMGCVSDADGLVKVMISKGYFPDIFTFNI-- 466
Query: 294 PVFLVH------SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSL 347
L+H + +I ++++ G DP + + LN + D++ +++ +
Sbjct: 467 ---LIHGYSTQLKMENALEILDVMLDNGVDPDVYTYNSLLNGLCKTSKFEDVMETYKTMV 523
Query: 348 LK 349
K
Sbjct: 524 EK 525
>At4g27400 unknown protein (At4g27400)
Length = 341
Score = 30.8 bits (68), Expect = 1.5
Identities = 21/63 (33%), Positives = 31/63 (48%), Gaps = 6/63 (9%)
Query: 364 VNSCFIHCQTWRGETWYSPN-SPKIQNKTI---AESVADWF-FDREVINLIDCPYP-CNP 417
V SCF+H + W G+T+ N +P+ Q + I E + F + I +DC P C
Sbjct: 15 VASCFLHVKAWHGQTYCGGNATPRCQLRYIDCPEECPTEMFPNSQNKICWVDCFKPLCEA 74
Query: 418 TCR 420
CR
Sbjct: 75 VCR 77
>At4g10520 putative subtilisin-like protease
Length = 756
Score = 30.0 bits (66), Expect = 2.5
Identities = 22/79 (27%), Positives = 36/79 (44%), Gaps = 15/79 (18%)
Query: 88 KGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTALGSSKYMETVVPFSGILSSDRSQNPDF 147
KGFG W E G N+ C+R+ +G+ +++ +V G++ +R+QNP+
Sbjct: 155 KGFGPIPSRWKGGCESGELFNASIHCNRK---LIGAKYFVDGLVAEFGVV--NRTQNPE- 208
Query: 148 FNWNKVMIRYCDGASFAGH 166
Y FAGH
Sbjct: 209 ---------YLSPRDFAGH 218
>At4g29900 Ca2+-transporting ATPase - like protein
Length = 1069
Score = 28.9 bits (63), Expect = 5.7
Identities = 13/41 (31%), Positives = 25/41 (60%), Gaps = 5/41 (12%)
Query: 153 VMIRYCDGASFAGHPESEKGSGLFFRGQIIWEAIMDELLSI 193
+++RY F GH ++E+G F G+ +E ++D+L+ I
Sbjct: 391 LVVRY-----FTGHTKNEQGGPQFIGGKTKFEHVLDDLVEI 426
>At1g27740 putative bHLH transcription factor (AtbHLH054)
Length = 258
Score = 28.9 bits (63), Expect = 5.7
Identities = 39/151 (25%), Positives = 56/151 (36%), Gaps = 30/151 (19%)
Query: 40 LSLAFFSQLHHH----------HHSHSLSNLIPFTPLPNAILRGAICLDGSAPGFHFQKG 89
LS + LH H HH+ L ++ PF +P L LD H Q+
Sbjct: 36 LSSLYNGHLHQHQHHNNVLSSDHHAFLLPDMFPFGAMPGGNL--PAMLDSWDQSHHLQE- 92
Query: 90 FGSGSRSWLLHLEGGGWCNSISSCSRRKFTALGSSKYMETVVPFSGILSSDRSQNPDFFN 149
S + LL +E C + S+C + L SK + V S S D N
Sbjct: 93 -TSSLKRKLLDVE--NLCKTNSNCDVTR-QELAKSKKKQRV--------SSESNTVDESN 140
Query: 150 WNKVMIRYCDGASFAGHPESEKGSGLFFRGQ 180
N + DG S + + EK S +G+
Sbjct: 141 TN-----WVDGQSLSNSSDDEKASVTSVKGK 166
>At5g53000 PP2A regulatory subunit (TAP46)
Length = 405
Score = 28.5 bits (62), Expect = 7.4
Identities = 23/86 (26%), Positives = 43/86 (49%), Gaps = 4/86 (4%)
Query: 322 WKSCKLNIHNCNA-NLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWY 380
W S +N+ C A +L+++L R +L A+ E Q + G F + +T + ETW+
Sbjct: 193 WLS-SINLAICKAIDLLEMLKR-EEEMLSAIKERQLKDGEGGFSRDA-LDDRTKKAETWH 249
Query: 381 SPNSPKIQNKTIAESVADWFFDREVI 406
+ +IQ A+ + F ++V+
Sbjct: 250 RDAAARIQYSKPAQPITCATFAQDVL 275
>At5g11890 unknown protein
Length = 287
Score = 28.5 bits (62), Expect = 7.4
Identities = 22/85 (25%), Positives = 40/85 (46%), Gaps = 9/85 (10%)
Query: 5 PSFSITNLRLRTLVVFFRKLTTKDYAIAAFAFFLLLSLAFFSQLHHHHHSHSLSNLIPFT 64
PSFS+ ++R+ + + T+ D +++ + F +L S+ + H S S PFT
Sbjct: 113 PSFSVPSIRISRVNL----TTSSDSSVSHLSSFFNFTL--ISENPNQHLSFSYD---PFT 163
Query: 65 PLPNAILRGAICLDGSAPGFHFQKG 89
N+ G + +G+ P F G
Sbjct: 164 VTVNSAKSGTMLGNGTVPAFFSDNG 188
>At3g33530 unknown protein
Length = 1350
Score = 28.5 bits (62), Expect = 7.4
Identities = 13/36 (36%), Positives = 19/36 (52%)
Query: 332 CNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSC 367
CN + + I + + L AL E QQ + MFV +C
Sbjct: 1247 CNYSYLSIYSGYCKEALAALREVQQPDTVAMFVLAC 1282
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,228,902
Number of Sequences: 26719
Number of extensions: 450128
Number of successful extensions: 1838
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1798
Number of HSP's gapped (non-prelim): 30
length of query: 427
length of database: 11,318,596
effective HSP length: 102
effective length of query: 325
effective length of database: 8,593,258
effective search space: 2792808850
effective search space used: 2792808850
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0342b.6


