

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.9
(434 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
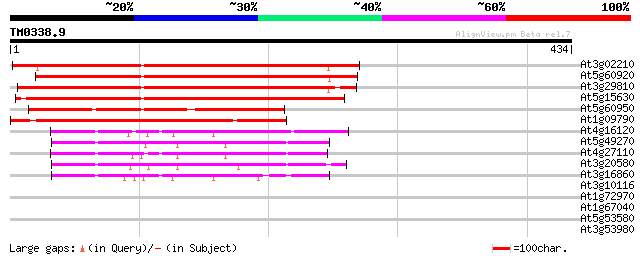
Score E
Sequences producing significant alignments: (bits) Value
At3g02210 predicted GPI-anchored protein 419 e-117
At5g60920 phytochelatin synthetase - like predicted GPI-anchored... 398 e-111
At3g29810 predicted GPI-anchored protein 380 e-106
At5g15630 GPI-anchored protein like 373 e-103
At5g60950 predicted GPI-anchored protein 202 2e-52
At1g09790 putative protein 189 2e-48
At4g16120 COBRA-like 7 (COBL7) predicted GPI-anchored protein 105 5e-23
At5g49270 predicted GPI-anchored protein 102 5e-22
At4g27110 predicted GPI-anchored protein (by homology) 101 9e-22
At3g20580 predicted GPI-anchored protein 96 4e-20
At3g16860 predicted GPI-anchored protein 90 3e-18
At3g10116 putative protein 35 0.081
At1g72970 unknown protein 29 4.4
At1g67040 hypothetical protein 29 4.4
At5g53580 aldo/keto reductase-like protein 28 7.6
At3g53980 unknown protein 28 7.6
>At3g02210 predicted GPI-anchored protein
Length = 452
Score = 419 bits (1076), Expect = e-117
Identities = 188/274 (68%), Positives = 225/274 (81%), Gaps = 7/274 (2%)
Query: 3 SLLLFRFVLPCTLLLLLP--SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQ 60
S + F+F + L+ + SEAYDPLDP+GNIT++WDI++WTGDGYVA VT+ NFQQ
Sbjct: 9 SSIFFKFGISIIFLVSFSGLTPSEAYDPLDPSGNITVKWDIITWTGDGYVATVTVYNFQQ 68
Query: 61 YRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGA 120
YRHIQ PGW+LGW+WAK E+IW M GGQTTEQGDCSKFKG + PHCCKK P VDL+PG+
Sbjct: 69 YRHIQAPGWTLGWSWAKREVIWGMNGGQTTEQGDCSKFKGTI-PHCCKKTPSVVDLLPGS 127
Query: 121 PYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTC 180
PY++Q+ANCC+GGVL S QDPA AV++F++ VG+AGTT K+VR+PKNFT K+PGPGYTC
Sbjct: 128 PYNQQIANCCRGGVLNSWAQDPATAVSAFQLTVGQAGTTNKTVRVPKNFTLKAPGPGYTC 187
Query: 181 GPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPT 240
PA+IVKPTRF+ DKRRVTQA+M+WNVTCTYSQFLA K PTCCVSLS+FYN TIVSCPT
Sbjct: 188 SPAKIVKPTRFIGTDKRRVTQALMTWNVTCTYSQFLAQKTPTCCVSLSSFYNKTIVSCPT 247
Query: 241 CACGC----QSGNCVDPNSPNLTSVISNPGNNGH 270
C+CGC Q GNCVDP P + SVI NPG N +
Sbjct: 248 CSCGCRNTSQPGNCVDPKGPRIASVIPNPGKNAY 281
>At5g60920 phytochelatin synthetase - like predicted GPI-anchored
protein
Length = 456
Score = 398 bits (1023), Expect = e-111
Identities = 172/254 (67%), Positives = 210/254 (81%), Gaps = 6/254 (2%)
Query: 21 SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEI 80
+S+EAYD LDP GNIT++WD++SWT DGYVA VT+ NFQ+YRHIQ PGW+LGW WAK E+
Sbjct: 32 TSTEAYDALDPEGNITMKWDVMSWTPDGYVAVVTMFNFQKYRHIQSPGWTLGWKWAKKEV 91
Query: 81 IWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQ 140
IW M+G QTTEQGDCSK+KG + PHCCKKDP VDL+PG PY++Q+ANCCKGGV+ S Q
Sbjct: 92 IWSMVGAQTTEQGDCSKYKGNI-PHCCKKDPTVVDLLPGTPYNQQIANCCKGGVMNSWVQ 150
Query: 141 DPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRRVT 200
DPA A +SF++ VG AGTT K+VR+P+NFT PGPGYTCGPA+IV+PT+F+ D RR T
Sbjct: 151 DPATAASSFQISVGAAGTTNKTVRVPRNFTLMGPGPGYTCGPAKIVRPTKFVTTDTRRTT 210
Query: 201 QAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ-----SGNCVDPNS 255
QAMM+WN+TCTYSQFLA + PTCCVSLS+FYN+TIV CPTCACGCQ SG C+DP++
Sbjct: 211 QAMMTWNITCTYSQFLAQRTPTCCVSLSSFYNETIVGCPTCACGCQNNRTESGACLDPDT 270
Query: 256 PNLTSVISNPGNNG 269
P+L SV+S P G
Sbjct: 271 PHLASVVSPPTKKG 284
>At3g29810 predicted GPI-anchored protein
Length = 441
Score = 380 bits (976), Expect = e-106
Identities = 165/266 (62%), Positives = 210/266 (78%), Gaps = 8/266 (3%)
Query: 7 FRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
F + C+ +++EAYD LDP GNITI+WDI+SWTGDGYVA VT+ NFQQYRHI+
Sbjct: 10 FLLLFLCSWTSFTFTTTEAYDALDPYGNITIKWDIMSWTGDGYVAVVTIFNFQQYRHIEA 69
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGW LGW+W K E+IW M+GGQ TEQGDCSKFKG + PHCCKK P VDL+PG PY++Q+
Sbjct: 70 PGWQLGWSWMKKEVIWSMVGGQATEQGDCSKFKGNI-PHCCKKTPAIVDLLPGTPYNQQI 128
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
+NCC+GGV+++ QDPA A++SF++ VG++GTT +VR P+N T K+PGPGYTCGPA++V
Sbjct: 129 SNCCRGGVISAWAQDPATAISSFQISVGQSGTTNTTVRAPRNITLKAPGPGYTCGPAKLV 188
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC- 245
KP+RF+ DKRR TQ++++WN+TCTYSQFLA K PTCCVSLS FYN+TIV CPTC+CGC
Sbjct: 189 KPSRFISADKRRKTQSLLTWNITCTYSQFLARKTPTCCVSLSAFYNETIVPCPTCSCGCQ 248
Query: 246 ---QSGNCVDPNSPNLTSVISNPGNN 268
Q+G CVD P + SV+ G N
Sbjct: 249 NSSQAGTCVD---PKIASVVPALGKN 271
>At5g15630 GPI-anchored protein like
Length = 431
Score = 373 bits (958), Expect = e-103
Identities = 170/256 (66%), Positives = 202/256 (78%), Gaps = 5/256 (1%)
Query: 5 LLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHI 64
LLF F C ++ ++ AYDPLDP+GNITI+WDI+SWT DGYVA VT+NNFQ YRHI
Sbjct: 3 LLFSF---CFFFFMIIFTATAYDPLDPSGNITIKWDIMSWTADGYVATVTMNNFQIYRHI 59
Query: 65 QEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSE 124
Q PGW+LGWTWAK E+IW M+G QTTEQGDCSKFKG V PHCCKK P VDL+PG PY++
Sbjct: 60 QNPGWTLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGNV-PHCCKKTPTVVDLLPGVPYNQ 118
Query: 125 QVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPAR 184
Q +NCCKGGV+ + QDP+ AV+ F+V G AGTT K+V+LPKNFT PGPGYTCGPA+
Sbjct: 119 QFSNCCKGGVIGAWGQDPSAAVSQFQVSAGLAGTTNKTVKLPKNFTLLGPGPGYTCGPAK 178
Query: 185 IVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACG 244
IV T FL DKRR TQA+M+WNVTCTYSQFLA K P+CCVS S+FYNDTI CP+CACG
Sbjct: 179 IVPSTVFLTTDKRRKTQALMTWNVTCTYSQFLARKHPSCCVSFSSFYNDTITPCPSCACG 238
Query: 245 CQS-GNCVDPNSPNLT 259
C++ +CV +S LT
Sbjct: 239 CENKKSCVKADSKILT 254
>At5g60950 predicted GPI-anchored protein
Length = 204
Score = 202 bits (515), Expect = 2e-52
Identities = 95/199 (47%), Positives = 132/199 (65%), Gaps = 10/199 (5%)
Query: 15 LLLLLPSSSEAYDPLDPN-GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGW 73
LLL+ S + + L N GNIT++WD+L+WT DGYVA VT N+Q+ R I PGW + W
Sbjct: 11 LLLVSFSCLISTEALTSNYGNITVKWDLLNWTPDGYVAVVTAYNYQKQRSI--PGWKMSW 68
Query: 74 TWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGG 133
K E+IW M+G +TT QG CS FKG + P C + P VDL+PG P+++Q+ANCCK G
Sbjct: 69 RGTKKEVIWNMLGAKTTGQGGCSMFKGNI-PQSCVRKPTVVDLLPGTPFNQQIANCCKSG 127
Query: 134 VLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLR 193
VL + ++F++ VG AG + K+ R+P NF F +P Y CGP++ V+PTRF
Sbjct: 128 VLKP------GSESAFQLSVGSAGNSVKTARMPANFMFTAPKQQYICGPSKNVRPTRFTT 181
Query: 194 YDKRRVTQAMMSWNVTCTY 212
DKRR+T A+M+WN+TC +
Sbjct: 182 ADKRRITAALMTWNITCVF 200
>At1g09790 putative protein
Length = 442
Score = 189 bits (481), Expect = 2e-48
Identities = 97/216 (44%), Positives = 132/216 (60%), Gaps = 8/216 (3%)
Query: 1 MDSLLLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQ 60
M +LLL V+ C++L S + YDPLDP G I I+WD+L + + VTL N Q+
Sbjct: 4 MLNLLLVVTVILCSIL----SPTHGYDPLDPFGKIIIKWDLLLSSPGQHHVQVTLENMQE 59
Query: 61 YRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVS-PHCCKKDPVAVDLIPG 119
YRH+++PGW L W W E+IW M G +TTEQG+CS F + PHCC + P VDL+PG
Sbjct: 60 YRHVEKPGWKLSWHWLNQEVIWDMKGAETTEQGNCSAFASSGNLPHCCLERPTIVDLLPG 119
Query: 120 APYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYT 179
A + QVANCC+GGVLTS+ QD AN V++F + VG + + +P NF PGY+
Sbjct: 120 ASLNVQVANCCRGGVLTSMSQDHANHVSAFHMTVGSSPDGPEEFNMPSNFDIGV--PGYS 177
Query: 180 CGPARIVKPTRFLRYDKRRVTQAMMSWN-VTCTYSQ 214
C A V PT+F RR TQA++ + C Y++
Sbjct: 178 CDNATSVSPTKFSTDKGRRKTQALVQQHGRQCVYTR 213
>At4g16120 COBRA-like 7 (COBL7) predicted GPI-anchored protein
Length = 661
Score = 105 bits (262), Expect = 5e-23
Identities = 70/251 (27%), Positives = 114/251 (44%), Gaps = 26/251 (10%)
Query: 32 NGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTT- 90
+G++TI +D++ Y+A VT+ N + W L + W ++E I+ M G +
Sbjct: 216 DGDLTIMYDVVRSYSSNYMAQVTMENHNPLGRLDN--WKLSFDWMRDEFIYTMKGAYPSI 273
Query: 91 -EQGDCSKFKGQVSPH----------CCKKDPVAVDLIPGAPYSEQ----VANCCKGGVL 135
+ DC G + H C + P +DL P Y++ + CC+ G +
Sbjct: 274 VDSSDC--VDGPQAKHYQDLDFSNVLSCARRPTVIDL-PPTKYNDSTFGLIPFCCRNGTI 330
Query: 136 TSLKQDPANAVASFRVVVGRA--GTTTKSVRLPKNFTFKSP-GPGYTCGPARIVKPTRFL 192
DP+ + + F++ V + ++ P+N+ P Y CGP V P++F+
Sbjct: 331 LPRSMDPSKSSSVFQMQVYKMPPDLNISALSPPQNWRINGTLNPDYKCGPPVRVSPSQFV 390
Query: 193 RYDKRRVTQ-AMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCV 251
+ A SW V C +Q A +P CCVS S ++ND+IV C TCACGC S
Sbjct: 391 DPSGLPSNRTAFASWQVVCNITQPKDA-SPRCCVSFSAYFNDSIVPCKTCACGCSSNKAA 449
Query: 252 DPNSPNLTSVI 262
S S++
Sbjct: 450 RACSATAPSLL 460
>At5g49270 predicted GPI-anchored protein
Length = 663
Score = 102 bits (253), Expect = 5e-22
Identities = 66/232 (28%), Positives = 102/232 (43%), Gaps = 20/232 (8%)
Query: 33 GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQ 92
G++TI +D++ Y+ VT+ N + W L + W ++E I +M G T
Sbjct: 216 GDLTIMYDVIRAYDQNYLTEVTMENHNPLGRLDH--WELSFDWMRDEFIQKMQGAYPTVV 273
Query: 93 GDCSKFKGQVS----------PHCCKKDPVAVDLIPGAPYSEQVAN---CCKGGVLTSLK 139
G S C++ P+ +DL P + N CC+ G +
Sbjct: 274 DATKCIFGPQSLIYTGLDFADVLTCERRPIIIDLPPTKKDDSTLGNIPSCCRNGTILPRI 333
Query: 140 QDPANAVASFRVVVGRAGTTTKSVRL--PKNFTFKSP-GPGYTCGPARIVKPTRFLRYDK 196
DP+ +V+ F + V + L P+N+ K P Y+CGP V PT +
Sbjct: 334 MDPSKSVSVFTMQVAKMPPDFNRSALFPPQNWRIKGTLNPDYSCGPPVRVTPTFYPDPSG 393
Query: 197 RRVTQAMM-SWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS 247
++ SW + C +Q + P CCVS S F+ND+I+ C TCACGC S
Sbjct: 394 MPTNKSSFASWQIVCNITQ-AKTEIPKCCVSFSAFFNDSIIPCNTCACGCVS 444
>At4g27110 predicted GPI-anchored protein (by homology)
Length = 717
Score = 101 bits (251), Expect = 9e-22
Identities = 72/235 (30%), Positives = 104/235 (43%), Gaps = 26/235 (11%)
Query: 32 NGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTE 91
+G++ I +D+L Y+A VT++N + W+L W W + E I M G E
Sbjct: 267 HGDLNIIYDVLQAYASSYMAQVTIDNTSPLGRLDH--WNLTWEWMRGEFIHSMRGAYAAE 324
Query: 92 QG--DCSKFKG----------QVSPHCCKKDPVAVDLIPGAPYSEQVAN----CCKGGVL 135
+ +C K QV+ C+K P+ DL P + V CCK G L
Sbjct: 325 KNTLECLSSKAGQFYGDLDFSQVAN--CQKKPIIKDL-PAERKDDNVTGKLPFCCKNGTL 381
Query: 136 TSLKQDPANAVASFRVVVGRAGTTTKSVRL--PKNFTFKS-PGPGYTCGPARIVKPTRFL 192
DP+ + A F++ V + P+N+ P Y CGP V T F
Sbjct: 382 LPTHMDPSKSKAIFQLQVYKVPPDQNRTAFFPPRNWKIDGIVNPTYKCGPPIRVDATPFP 441
Query: 193 RYDK-RRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ 246
+ T A+ SW V C ++ +A CCVS S FYND+ + C TCACGC+
Sbjct: 442 DPSGLQATTYAIASWQVICNITK-PKPQAARCCVSFSAFYNDSAIPCNTCACGCK 495
>At3g20580 predicted GPI-anchored protein
Length = 672
Score = 95.9 bits (237), Expect = 4e-20
Identities = 75/245 (30%), Positives = 109/245 (43%), Gaps = 23/245 (9%)
Query: 33 GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQ 92
G++ I +D+L Y+A VT++N + W+L + W + E I M G T ++
Sbjct: 229 GDLNIVYDVLQSFDSNYLAQVTIDNDNPLGRLDR--WNLTFEWMRGEFINTMRGAYTHKK 286
Query: 93 --GDCSKFK-GQVSPHC-------CKKDPVAVDLIPGAPYSEQVAN---CCKGGVLTSLK 139
+C K GQ C++ P DL P CCK G L
Sbjct: 287 DPSECLYSKAGQYYKDLDFSQVMNCQRKPAISDLPPEKKEDNMTGKLPFCCKNGTLLPPI 346
Query: 140 QDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPG---PGYTCGPARIVKPTRFLRYDK 196
DP+ + + F++ V + L +K G P Y CGP V P++F
Sbjct: 347 MDPSKSRSMFQLQVFKLPPDLNRTALYPPQHWKIDGVLNPQYKCGPPVRVDPSQFPDPSG 406
Query: 197 R-RVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCVDPNS 255
VT A+ SW V C ++ A+A CCVS S FYN++ V C TCACGC N +D ++
Sbjct: 407 LLAVTYAISSWQVVCNITK-PKAQASRCCVSFSAFYNNSAVPCNTCACGC---NDIDTDT 462
Query: 256 PNLTS 260
N S
Sbjct: 463 CNANS 467
>At3g16860 predicted GPI-anchored protein
Length = 653
Score = 89.7 bits (221), Expect = 3e-18
Identities = 65/238 (27%), Positives = 107/238 (44%), Gaps = 33/238 (13%)
Query: 33 GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGG--QTT 90
G++TI +D+ Y A VT+ N + W L + W K+E ++ G
Sbjct: 210 GDLTIMYDVTRAYQSSYSAQVTIENHNLLGRLDN--WDLSFMWMKDEFLFSTKGAYPSVV 267
Query: 91 EQGDC-----SKFKGQV---SPHCCKKDPVAVDLIPGAPYSE----QVANCCKGGVLTSL 138
+ DC +K+ + + C + P +DL P Y++ ++ CC+ G +
Sbjct: 268 DSSDCITGPQAKYYKDLDFSNVMSCARRPHIIDL-PLTKYNDTNVGRIPYCCRNGTILPR 326
Query: 139 KQDPANAVASFRVVVGRA--GTTTKSVRLPKNFTFKSP-GPGYTCGPARIVKPTRF---- 191
DP + + F++ V + S+ P+++ K P Y CGP V ++F
Sbjct: 327 SMDPEKSKSVFQIEVYKMPPDLNISSITPPQSWQIKGNLNPDYKCGPPLRVSSSQFPDTS 386
Query: 192 -LRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCAC-GCQS 247
L +K A SW V C +Q P CCVS S+++ND+++ C TCAC GC S
Sbjct: 387 GLPSNK----SAFASWQVVCNITQ---PTPPKCCVSFSSYFNDSVIPCKTCACGGCSS 437
>At3g10116 putative protein
Length = 170
Score = 35.0 bits (79), Expect = 0.081
Identities = 17/46 (36%), Positives = 28/46 (59%), Gaps = 2/46 (4%)
Query: 45 TGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTT 90
T GY A VT+ N+ +Y I+ P W L W W +++I+ +G Q++
Sbjct: 4 TRGGYTANVTIFNYLKYHDIEGP-WVLDWIW-EDQILLSTVGVQSS 47
>At1g72970 unknown protein
Length = 594
Score = 29.3 bits (64), Expect = 4.4
Identities = 24/75 (32%), Positives = 36/75 (48%), Gaps = 6/75 (8%)
Query: 183 ARIVKPTRFLRY---DKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCP 239
+++V RFL Y DK+ V + M+S +V + L K S++ F DT+V+
Sbjct: 472 SKVVTSNRFLNYTQCDKQNVHK-MLSLSVKANIN--LRPKQLNDTKSMAQFCKDTVVTIW 528
Query: 240 TCACGCQSGNCVDPN 254
GC G V PN
Sbjct: 529 HYHGGCLVGKVVSPN 543
>At1g67040 hypothetical protein
Length = 826
Score = 29.3 bits (64), Expect = 4.4
Identities = 17/52 (32%), Positives = 24/52 (45%), Gaps = 8/52 (15%)
Query: 217 AAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCVDPNSPNLTSVISNPGNN 268
AAK P YN+++ SC +C G+ VD N V+ + GNN
Sbjct: 252 AAKEPVVSPEFQCGYNNSVASCKSC------GSLVDVNGS--IQVVQDTGNN 295
>At5g53580 aldo/keto reductase-like protein
Length = 365
Score = 28.5 bits (62), Expect = 7.6
Identities = 22/75 (29%), Positives = 31/75 (41%), Gaps = 7/75 (9%)
Query: 5 LLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHI 64
LLFR +LP LLL S A + I W I T V + + RH+
Sbjct: 275 LLFRQILPGLEPLLLALSEIAKKRGKTMPQVAINWCICKGT-------VPIPGIKSVRHV 327
Query: 65 QEPGWSLGWTWAKNE 79
++ +LGW +E
Sbjct: 328 EDNLGALGWKLTNDE 342
>At3g53980 unknown protein
Length = 114
Score = 28.5 bits (62), Expect = 7.6
Identities = 29/103 (28%), Positives = 40/103 (38%), Gaps = 14/103 (13%)
Query: 84 MIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLK---Q 140
+IG G+C G+ SP D A+ L P A ++ + GG T +K Q
Sbjct: 16 LIGTVVDGAGEC----GRSSP-----DNEAMKLAPCAGAAQDANSAVPGGCCTQIKRFSQ 66
Query: 141 DPANAVASFRVVVGRAGTTTKSVRL--PKNFTFKSPGPGYTCG 181
+P A +A V L PK F + GY CG
Sbjct: 67 NPKCLCAILLSDTAKASGVDPEVALTIPKRCNFANRPVGYKCG 109
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.145 0.504
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,006,265
Number of Sequences: 26719
Number of extensions: 417357
Number of successful extensions: 1209
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1162
Number of HSP's gapped (non-prelim): 20
length of query: 434
length of database: 11,318,596
effective HSP length: 102
effective length of query: 332
effective length of database: 8,593,258
effective search space: 2852961656
effective search space used: 2852961656
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0338.9


