

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.8
(61 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
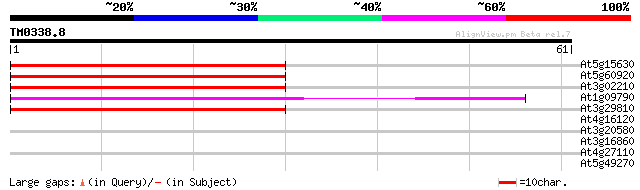
At5g15630 GPI-anchored protein like 67 2e-12
At5g60920 phytochelatin synthetase - like predicted GPI-anchored... 60 2e-10
At3g02210 predicted GPI-anchored protein 56 4e-09
At1g09790 putative protein 54 1e-08
At3g29810 predicted GPI-anchored protein 54 2e-08
At4g16120 COBRA-like 7 (COBL7) predicted GPI-anchored protein 32 0.059
At3g20580 predicted GPI-anchored protein 32 0.078
At3g16860 predicted GPI-anchored protein 32 0.078
At4g27110 predicted GPI-anchored protein (by homology) 31 0.10
At5g49270 predicted GPI-anchored protein 30 0.17
>At5g15630 GPI-anchored protein like
Length = 431
Score = 66.6 bits (161), Expect = 2e-12
Identities = 26/30 (86%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD+ TF+FKQGWAFPRKVYFNGDECM+PPP
Sbjct: 372 KDQKTFTFKQGWAFPRKVYFNGDECMLPPP 401
>At5g60920 phytochelatin synthetase - like predicted GPI-anchored
protein
Length = 456
Score = 60.1 bits (144), Expect = 2e-10
Identities = 21/30 (70%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F++GWAFPR++YFNGD C+MPPP
Sbjct: 394 KDQSTFTFEKGWAFPRRIYFNGDNCVMPPP 423
>At3g02210 predicted GPI-anchored protein
Length = 452
Score = 55.8 bits (133), Expect = 4e-09
Identities = 19/30 (63%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+ + F+F++GWAFPR++YFNGD C+MPPP
Sbjct: 389 KEASAFTFEKGWAFPRRIYFNGDNCVMPPP 418
>At1g09790 putative protein
Length = 442
Score = 54.3 bits (129), Expect = 1e-08
Identities = 25/56 (44%), Positives = 32/56 (56%), Gaps = 12/56 (21%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFCQILLLQA*FPSRHSSSHYSS 56
KD F+F++GWAFPR++ FNGDEC+MP P FP S+H SS
Sbjct: 375 KDMGNFTFREGWAFPRRILFNGDECVMPSPDD------------FPRLPKSAHSSS 418
>At3g29810 predicted GPI-anchored protein
Length = 441
Score = 53.5 bits (127), Expect = 2e-08
Identities = 19/30 (63%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+ F+F++GWAFPR++YFNGD C+MPPP
Sbjct: 380 KNPLEFTFEKGWAFPRRIYFNGDNCVMPPP 409
>At4g16120 COBRA-like 7 (COBL7) predicted GPI-anchored protein
Length = 661
Score = 32.0 bits (71), Expect = 0.059
Identities = 13/27 (48%), Positives = 16/27 (59%)
Query: 11 GWAFPRKVYFNGDECMMPPPTPTHFCQ 37
G FP KV+FNG+EC +P P Q
Sbjct: 613 GDGFPSKVFFNGEECSLPTILPMRSSQ 639
>At3g20580 predicted GPI-anchored protein
Length = 672
Score = 31.6 bits (70), Expect = 0.078
Identities = 12/32 (37%), Positives = 19/32 (58%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTP 32
K+ + +G FP K++FNG+EC +P P
Sbjct: 612 KNIKGLNIPEGDGFPTKLFFNGEECALPKHFP 643
>At3g16860 predicted GPI-anchored protein
Length = 653
Score = 31.6 bits (70), Expect = 0.078
Identities = 18/48 (37%), Positives = 20/48 (41%), Gaps = 13/48 (27%)
Query: 14 FPRKVYFNGDECMMPPPTPTHFCQILLLQA*FPSRHSSSHYSSFLFYL 61
FP KV FNG EC +P PT S H S+FL L
Sbjct: 608 FPTKVLFNGQECSLPSVLPT-------------SNSHRKHVSTFLLIL 642
>At4g27110 predicted GPI-anchored protein (by homology)
Length = 717
Score = 31.2 bits (69), Expect = 0.10
Identities = 12/32 (37%), Positives = 20/32 (62%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTP 32
K+ + + G FP++V+FNG+EC +P P
Sbjct: 651 KNLGSLNIIGGDGFPKRVFFNGEECELPKYFP 682
>At5g49270 predicted GPI-anchored protein
Length = 663
Score = 30.4 bits (67), Expect = 0.17
Identities = 11/19 (57%), Positives = 14/19 (72%)
Query: 14 FPRKVYFNGDECMMPPPTP 32
FP KV FNG+EC++P P
Sbjct: 617 FPAKVIFNGEECLLPDLLP 635
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.335 0.145 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,665,173
Number of Sequences: 26719
Number of extensions: 56767
Number of successful extensions: 181
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 171
Number of HSP's gapped (non-prelim): 10
length of query: 61
length of database: 11,318,596
effective HSP length: 37
effective length of query: 24
effective length of database: 10,329,993
effective search space: 247919832
effective search space used: 247919832
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0338.8


