

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.4
(197 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
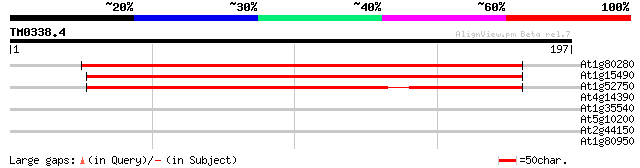
Sequences producing significant alignments: (bits) Value
At1g80280 unknown protein 234 3e-62
At1g15490 unknown protein 219 1e-57
At1g52750 alpha/beta fold family protein 164 3e-41
At4g14390 hypothetical protein 28 4.1
At1g35540 auxin response factor, putative 28 4.1
At5g10200 putative protein 27 5.4
At2g44150 unknown protein 27 7.0
At1g80950 unknown protein 27 9.1
>At1g80280 unknown protein
Length = 647
Score = 234 bits (596), Expect = 3e-62
Identities = 111/155 (71%), Positives = 134/155 (85%)
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSL 85
+ L VSSLPV+V + D+LVP L+S+FTC+ CYG +H RYAFKSSLTDIPLVSL+RS
Sbjct: 26 VSLLVSSLPVLVAIGDVLVPTFLLSSFTCLTCYGFKEHLSRYAFKSSLTDIPLVSLVRSF 85
Query: 86 IILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRL 145
+++CVYS+ DAP LSHGPYLGTV+LCS++S+VLLSVK CVFT NS++ +AS+S RQRL
Sbjct: 86 LVICVYSLSDAPALSHGPYLGTVSLCSVVSVVLLSVKACVFTANSQLNDQASSSPSRQRL 145
Query: 146 HLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
HLKKSWGMPVLFLSSVVF+LGH+VVA RTSCRARR
Sbjct: 146 HLKKSWGMPVLFLSSVVFALGHMVVAYRTSCRARR 180
>At1g15490 unknown protein
Length = 648
Score = 219 bits (557), Expect = 1e-57
Identities = 102/153 (66%), Positives = 128/153 (82%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L VSSLP++V + D+LVP L+S+FTC+ CYG +H RY FK SLTDIP+VS++RS ++
Sbjct: 28 LLVSSLPLLVAIGDVLVPSFLLSSFTCVTCYGAKEHLRRYGFKRSLTDIPIVSVVRSFLV 87
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
+C+Y + D P LSHGPYLGTV+LCS++S++LLSVK C+FTVNS++ EAS S R+RLHL
Sbjct: 88 ICIYLLSDVPALSHGPYLGTVSLCSVVSVLLLSVKACLFTVNSQLNNEASFSPSRKRLHL 147
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
KKSWGMPVLFLSSVVF+LGH VVA RTSCRARR
Sbjct: 148 KKSWGMPVLFLSSVVFALGHTVVAYRTSCRARR 180
>At1g52750 alpha/beta fold family protein
Length = 633
Score = 164 bits (415), Expect = 3e-41
Identities = 82/153 (53%), Positives = 110/153 (71%), Gaps = 7/153 (4%)
Query: 28 LTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
L SSLPV++ V D++VPC+++S+ TC+ C+ +HF +Y+FK+SL D+PLVSL+RSL+I
Sbjct: 30 LLASSLPVLITVADVVVPCLIVSSITCLTCHSPAQHFRQYSFKTSLIDVPLVSLIRSLVI 89
Query: 88 LCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
+C+ +C+ L++G YL TV L S I+LL VKTCVFTVNS +E + + HL
Sbjct: 90 ICLSWLCEDTRLAYGLYLETVMLSSFGGILLLLVKTCVFTVNSTLE-------EAKVYHL 142
Query: 148 KKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
K SWGMPVL LSS VF L H+VVA R SC ARR
Sbjct: 143 KISWGMPVLLLSSAVFGLAHVVVAYRKSCGARR 175
>At4g14390 hypothetical protein
Length = 691
Score = 27.7 bits (60), Expect = 4.1
Identities = 25/82 (30%), Positives = 39/82 (47%), Gaps = 13/82 (15%)
Query: 100 SHGPYLGTVTLCS------ILSIVLLSVKTCVFTVNSRIEAE-ASASLKRQRLH------ 146
S P+LG TL + L + +L++++ V T+ I A+ L R LH
Sbjct: 543 SSAPHLGRATLATNPTLFIFLVLDILAMQSSVATIGILIWAQLGDPVLIRSSLHVALPLL 602
Query: 147 LKKSWGMPVLFLSSVVFSLGHI 168
L MP+ FL VV ++GH+
Sbjct: 603 LFALLCMPLAFLFGVVTAVGHV 624
>At1g35540 auxin response factor, putative
Length = 661
Score = 27.7 bits (60), Expect = 4.1
Identities = 26/106 (24%), Positives = 41/106 (38%), Gaps = 4/106 (3%)
Query: 88 LCVYSICDAPTLSHGPYL---GTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQR 144
LC +CD P L Y G + L S LS+ S+ + +++ + S SL
Sbjct: 31 LCAGPLCDIPKLGEKVYYFPQGHIELVSSLSLYSFSLLSLSLSLSLSLSLSLSLSLSPFS 90
Query: 145 LHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARRSSCFIESTLK 190
L S P+ S+ L H+ + R + C S L+
Sbjct: 91 LFSLFSLS-PLSLSLSLSLLLSHVEASTREELNELQPICDFPSKLQ 135
>At5g10200 putative protein
Length = 621
Score = 27.3 bits (59), Expect = 5.4
Identities = 15/45 (33%), Positives = 25/45 (55%)
Query: 127 TVNSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVA 171
TVNSR + S+ ++ LH+K++ + V + +FS G I A
Sbjct: 373 TVNSRQRLKWEKSMPKEDLHIKQAAALVVKLEGNSLFSSGDIAGA 417
>At2g44150 unknown protein
Length = 363
Score = 26.9 bits (58), Expect = 7.0
Identities = 10/27 (37%), Positives = 17/27 (62%)
Query: 47 VLISNFTCIRCYGITKHFGRYAFKSSL 73
+ +S T RC+G+ +HF Y+ K S+
Sbjct: 317 IRLSRPTSDRCFGLVRHFDEYSRKHSV 343
>At1g80950 unknown protein
Length = 398
Score = 26.6 bits (57), Expect = 9.1
Identities = 14/34 (41%), Positives = 18/34 (52%)
Query: 73 LTDIPLVSLMRSLIILCVYSICDAPTLSHGPYLG 106
+T +PL L+ I+L Y IC TL PY G
Sbjct: 71 VTLVPLRFLLSMSILLLYYLICRVFTLFSAPYRG 104
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.343 0.149 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,663,437
Number of Sequences: 26719
Number of extensions: 131285
Number of successful extensions: 640
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 632
Number of HSP's gapped (non-prelim): 8
length of query: 197
length of database: 11,318,596
effective HSP length: 94
effective length of query: 103
effective length of database: 8,807,010
effective search space: 907122030
effective search space used: 907122030
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0338.4


