

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.2
(282 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
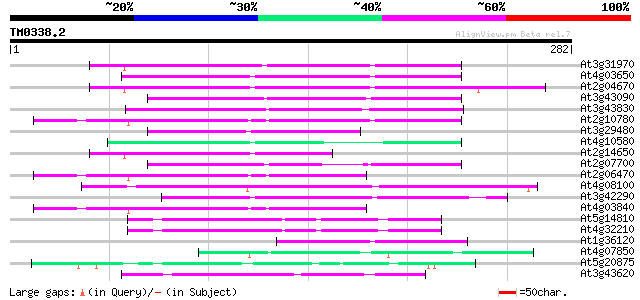
Score E
Sequences producing significant alignments: (bits) Value
At3g31970 hypothetical protein 99 2e-21
At4g03650 putative reverse transcriptase 96 2e-20
At2g04670 putative retroelement pol polyprotein 96 3e-20
At3g43090 putative protein 82 3e-16
At3g43830 putative protein 80 9e-16
At2g10780 pseudogene 77 1e-14
At3g29480 hypothetical protein 75 4e-14
At4g10580 putative reverse-transcriptase -like protein 74 1e-13
At2g14650 putative retroelement pol polyprotein 71 7e-13
At2g07700 putative retroelement pol polyprotein 65 5e-11
At2g06470 putative retroelement pol polyprotein 64 1e-10
At4g08100 putative polyprotein 63 2e-10
At3g42290 putative protein 62 5e-10
At4g03840 putative transposon protein 61 8e-10
At5g14810 putative protein 49 2e-06
At4g32210 putative protein 49 2e-06
At1g36120 putative reverse transcriptase gb|AAD22339.1 49 4e-06
At4g07850 putative polyprotein 44 1e-04
At5g20875 unknown protein 42 3e-04
At3g43620 putative protein 42 4e-04
>At3g31970 hypothetical protein
Length = 1329
Score = 99.4 bits (246), Expect = 2e-21
Identities = 65/192 (33%), Positives = 89/192 (45%), Gaps = 9/192 (4%)
Query: 41 ARELHRLQREAALDQN-----RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQT 95
A + +Q+ AA+ + R + +R F GGT P+EAD W +E F +
Sbjct: 124 APRVAEVQQRAAVAEEVPSYLRMMEQLQRIGTRYFFGGTSPEEADSWKSRVEHNFGSSRC 183
Query: 96 SEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQ 155
+V LA + L GDA WWR H ++W F F KYFP A D E++
Sbjct: 184 PAEYRVDLAVHFLEGDAHLWWRSVTARRRQAH--MSWADFVAEFNAKYFPQEALDRMEAR 241
Query: 156 FLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMR 215
FL L QG ++ EY + L + R D+ +RF+RGLR D+ R L
Sbjct: 242 FLELTQGERSVREYEREFNRLLVYAG--RGMQDDQAQMRRFLRGLRPDLRVRCRVLQYAT 299
Query: 216 FQALVEKATEVE 227
ALVE A EVE
Sbjct: 300 KAALVETAAEVE 311
>At4g03650 putative reverse transcriptase
Length = 839
Score = 96.3 bits (238), Expect = 2e-20
Identities = 58/171 (33%), Positives = 81/171 (46%), Gaps = 4/171 (2%)
Query: 57 RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWW 116
R + +R F+GGT P+EAD W ++E+ F + +V L + L GDA WW
Sbjct: 143 RMMKQLQRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDAHLWW 202
Query: 117 RGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESL 176
R +++W F F KYFP A D E++FL L QG ++ EY K L
Sbjct: 203 RSVTA--RRRQADMSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRL 260
Query: 177 AKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
+ R D+ +RF+RGLR ++ R ALVE A EVE
Sbjct: 261 LVYAG--RGMEDDQAQMRRFLRGLRPNLRVRCRVSQYATKAALVETAAEVE 309
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 95.5 bits (236), Expect = 3e-20
Identities = 69/238 (28%), Positives = 103/238 (42%), Gaps = 13/238 (5%)
Query: 41 ARELHRLQREAALDQN-----RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQT 95
A + +Q+ AA+ + R + +R F+ GT P+EAD W +++ F +
Sbjct: 104 APRMAEVQQRAAVAEEVPSYLRMMEQLQRIGTWYFSSGTSPEEADSWRSRVQRNFGSSRC 163
Query: 96 SEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQ 155
+V LA + L GDA WWR +++W F F KYFP A D E++
Sbjct: 164 PAEYRVDLAVHFLEGDAHLWWRSVTA--RRRQADMSWADFVAEFNAKYFPQEALDRMEAR 221
Query: 156 FLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMR 215
FL L QG ++ EY + L + R D+ +RF+RGLR D+ R
Sbjct: 222 FLELTQGEWSVREYDREFNRLLAYAG--RGMEDDQAQMRRFLRGLRPDLRVQCRVSQYAT 279
Query: 216 FQALVEKATEVELMKNRRM----DRAGTGGPMRSSPRSYQGKGKLQMKKPYQRPTGEG 269
ALVE A EVE R++ T + S GK K+ + P+ G
Sbjct: 280 KAALVETAAEVEEDLQRQVVGVSPAVQTKNTQQQVTPSKGGKPAQGQKRKWDHPSRAG 337
>At3g43090 putative protein
Length = 357
Score = 82.0 bits (201), Expect = 3e-16
Identities = 53/158 (33%), Positives = 72/158 (45%), Gaps = 3/158 (1%)
Query: 70 FTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVE 129
FTGG DP +AD W + +E FE + + + + L +A +WW G G HV
Sbjct: 131 FTGGIDPVKADDWRKLLENNFENARCPVEFQKDIIVHFLREEASHWWDGVLGNTPVQHV- 189
Query: 130 VNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDE 189
+ W FR F K+FP A D E F LRQ + + E +L L++ R E
Sbjct: 190 IFWEDFREEFNRKFFPQEAMDSLEDDFEELRQDTKKVRENERELSHLSRF--SVRAGRGE 247
Query: 190 PYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
M +R +RGLR DI LVEKA +E
Sbjct: 248 QSMIRRLMRGLRPDILTRCASREFFSMIDLVEKAARIE 285
>At3g43830 putative protein
Length = 329
Score = 80.5 bits (197), Expect = 9e-16
Identities = 53/170 (31%), Positives = 81/170 (47%), Gaps = 4/170 (2%)
Query: 59 LNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRG 118
L R D F GGTDP +A W +E F+ ++ LA+ L G+A+ WW
Sbjct: 105 LRFMRDLDIETFNGGTDPVKAYNWRNMLECKFKSMRCPVEFWRDLASCYLRGEAQEWWER 164
Query: 119 ARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAK 178
+ + V+ W+ F+ F +Y P+ D+ E +FL L+QG+ T+ +Y + SL
Sbjct: 165 VKQREQVGCVD-QWSFFKEEFTRRYLPEETIDDLEMKFLRLQQGTKTVRKYEKEFHSLE- 222
Query: 179 HFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVEL 228
RF R + E + +F+ GLR DI LVEKA +E+
Sbjct: 223 --RFERRKRGEHELIHKFISGLRVDIRLCCHVRDFDNMIDLVEKAASLEI 270
>At2g10780 pseudogene
Length = 1611
Score = 76.6 bits (187), Expect = 1e-14
Identities = 61/220 (27%), Positives = 96/220 (42%), Gaps = 13/220 (5%)
Query: 13 QLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRG-----LNDFRRQDP 67
QLA + ++ V DN + +D E+ + + + RG L R
Sbjct: 124 QLATLTKAVIADPVVLKVDNQK----DDDSEVVEVNPQPNHGRKRGDYLSLLEHVSRLGT 179
Query: 68 PKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANH 127
F G TDP AD W +++ F+ + E + +A + L GDA WW +
Sbjct: 180 RHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVE-KRRGDE 238
Query: 128 VEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQV 187
V ++ F F +KYFP A D E +L L QG+ T+ EY + L ++ R+
Sbjct: 239 VR-SFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVG--RELE 295
Query: 188 DEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
+E +RF+RGLR +I + LVE+A +E
Sbjct: 296 EEQAQLRRFIRGLRIEIRNHCLVRTFNSVSELVERAAMIE 335
>At3g29480 hypothetical protein
Length = 718
Score = 75.1 bits (183), Expect = 4e-14
Identities = 38/107 (35%), Positives = 52/107 (48%), Gaps = 2/107 (1%)
Query: 70 FTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVE 129
F+GGT P+EAD W +E+ F + +V LA + L GDA WW+
Sbjct: 51 FSGGTSPEEADSWRSRVERNFGSSRCPVEYRVDLAVHFLEGDAHLWWKSV--TTRRRQAN 108
Query: 130 VNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESL 176
++W F F KYFP A D E+ FL L QG ++ EY + L
Sbjct: 109 MSWADFVAEFNAKYFPQEALDRMEAHFLELTQGERSVREYDREFNRL 155
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 73.6 bits (179), Expect = 1e-13
Identities = 52/178 (29%), Positives = 69/178 (38%), Gaps = 31/178 (17%)
Query: 50 EAALDQNRGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLL 109
E L R + +R D F+GGT P+EAD W + + F + +V LA + L
Sbjct: 132 EEVLSYLRMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLE 191
Query: 110 GDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEY 169
GDA WWR +++W F F KYFP A D Q +
Sbjct: 192 GDAHLWWRSVTA--RRRQTDMSWADFVAEFKAKYFPQEALDPYAGQGME----------- 238
Query: 170 AAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
D+ +RF+RGLR D+ R ALVE A EVE
Sbjct: 239 ------------------DDQAQMRRFLRGLRPDLRVRCRVSQYATKAALVETAAEVE 278
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 70.9 bits (172), Expect = 7e-13
Identities = 40/127 (31%), Positives = 59/127 (45%), Gaps = 7/127 (5%)
Query: 41 ARELHRLQREAALDQN-----RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQT 95
A + +Q+ AA+ + R + +R F+GGT P+ AD W +E+ F +
Sbjct: 122 APRVAEVQQRAAVAEEVPSYLRMMEQLQRIGTGYFSGGTSPEVADSWRSRVERNFGSSRC 181
Query: 96 SEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQ 155
++ LA + L GDA WWR +++W F F KYFP A D E+
Sbjct: 182 PAEYRIDLAVHFLEGDAHLWWRSVTA--RRRQADISWADFVAEFNAKYFPQEALDRMEAH 239
Query: 156 FLTLRQG 162
FL L QG
Sbjct: 240 FLELTQG 246
>At2g07700 putative retroelement pol polyprotein
Length = 543
Score = 64.7 bits (156), Expect = 5e-11
Identities = 45/158 (28%), Positives = 69/158 (43%), Gaps = 23/158 (14%)
Query: 70 FTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVE 129
F G + AD W++++E FE+ + + K +A L DA W + +
Sbjct: 53 FKGEHNTVVADKWLRDLEMNFEISRCPDNFKRQIAVNFLDKDARIWRESVTARNQGQMI- 111
Query: 130 VNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDE 189
+W +FR F +KYFP ARD E QF+ K + + Q DE
Sbjct: 112 -SWEAFRREFEKKYFPPEARDRLEQQFM--------------------KRY-LYNGQDDE 149
Query: 190 PYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
M +RF+RGLR +I ++ + L E+A VE
Sbjct: 150 ALMVRRFLRGLRLEIWGRLQAVKYDSVNELTERADNVE 187
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 63.5 bits (153), Expect = 1e-10
Identities = 49/172 (28%), Positives = 75/172 (43%), Gaps = 11/172 (6%)
Query: 13 QLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRG-----LNDFRRQDP 67
QLA + +V VQ DN + +D E+ + + + RG L R
Sbjct: 89 QLATLTKAVVADPVVQRVDNQK----DDDSEVVEVNPQPNHGRKRGDYLSLLEHVSRLGT 144
Query: 68 PKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANH 127
F G TDP AD W +++ F+ + E + +A + L GDA WW +
Sbjct: 145 RHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVE-KRRGDE 203
Query: 128 VEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKH 179
V ++ F F +KYFP A D E +L L QG+ T+ EY + L ++
Sbjct: 204 VR-SFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At4g08100 putative polyprotein
Length = 1054
Score = 63.2 bits (152), Expect = 2e-10
Identities = 57/235 (24%), Positives = 95/235 (40%), Gaps = 13/235 (5%)
Query: 37 AAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTS 96
++ ++ HR +R D GL + P F G +PD W Q+IE +F +
Sbjct: 70 SSHSSQRRHRRERPPPRDPLGGL----KLKIPAFHGTDNPDTYLEWEQKIELVFLCQECL 125
Query: 97 EGAKVGLATYLLLGDAEYWWRG--ARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERES 154
+ KV +A A WW ++ WN + +++ P E
Sbjct: 126 QSNKVKIAATKFYNYALSWWDQLVTSRRRTRDYPIKTWNQLKFVMRKRFVPSYYHRELHQ 185
Query: 155 QFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIM 214
+ L QGS T+ EY ++E+L Q D + RF+ GL +I+D + +
Sbjct: 186 RLRNLVQGSKTVEEYFLEMETLMLRADL---QEDGEAVMSRFMGGLNREIQDRLETQHYV 242
Query: 215 RFQALVEKATEVELMKNRRMDRAG-TGGPMRSSPRSYQGKGKLQMK---KPYQRP 265
+ ++ KA E R+ R+ T S SYQ + K + KP+ +P
Sbjct: 243 ELEEMLHKAVMFEQQIKRKNARSSHTKTNYSSGKPSYQKEEKFGYQNDSKPFVKP 297
>At3g42290 putative protein
Length = 462
Score = 61.6 bits (148), Expect = 5e-10
Identities = 48/174 (27%), Positives = 72/174 (40%), Gaps = 12/174 (6%)
Query: 77 DEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFR 136
++AD W +++ F + +V LA + L DA WW+ +++W F
Sbjct: 161 EKADSWRSRVKRNFSSSRCLMEYRVDLAVHFLEEDAHLWWKSVTA--RRRQADMSWADFV 218
Query: 137 VAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRF 196
V F KYF D E +FL L ++ EY + + + + D +RF
Sbjct: 219 VEFNAKYFLQDELDRMEVRFLELTLVERSVWEYDREFNRVLVYAGW--GMKDGQAELRRF 276
Query: 197 VRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSY 250
+RGLR D+ R ALVE A EV K G PM+ RS+
Sbjct: 277 MRGLRPDLRVRCRMSQYALKAALVETAAEVAPRKG--------GKPMQGKKRSW 322
>At4g03840 putative transposon protein
Length = 973
Score = 60.8 bits (146), Expect = 8e-10
Identities = 48/172 (27%), Positives = 74/172 (42%), Gaps = 11/172 (6%)
Query: 13 QLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRG-----LNDFRRQDP 67
QLA + +V VQ DN + +D E+ + + + RG L R
Sbjct: 89 QLATLTKAIVANPVVQRVDNQK----DDELEVVEVNPQPNHGRKRGDYLSLLEHVSRLGT 144
Query: 68 PKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANH 127
F G TDP AD W +++ F+ + E + +A + L DA WW +
Sbjct: 145 RHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLERDAHNWWLTVE-KRRGDE 203
Query: 128 VEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKH 179
V ++ F F +KYFP A D E +L L QG+ T+ EY + L ++
Sbjct: 204 VR-SFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At5g14810 putative protein
Length = 716
Score = 49.3 bits (116), Expect = 2e-06
Identities = 43/158 (27%), Positives = 69/158 (43%), Gaps = 12/158 (7%)
Query: 60 NDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGA 119
+DF R + +F G D W+ +IE+ F + +T E KVG A+ A +
Sbjct: 101 SDFARLEHLRFNG----DRITEWLFQIEQFFLIDRTPEELKVGFASLHFDDTAATLHQSI 156
Query: 120 RGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKH 179
M HV +W S+++ L+ + + D LT Q + I EY A+ E ++
Sbjct: 157 VQSMWWKHVRHDWWSYKL-LLQVRYDEHVNDSIAK--LTQLQETEGIEEYHARFELISTR 213
Query: 180 FRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQ 217
F D Y+ ++ GLR D + +VR G Q
Sbjct: 214 LNFAED-----YLVSVYLAGLRTDTQLNVRMFGPQTIQ 246
>At4g32210 putative protein
Length = 1086
Score = 49.3 bits (116), Expect = 2e-06
Identities = 43/158 (27%), Positives = 69/158 (43%), Gaps = 12/158 (7%)
Query: 60 NDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGA 119
+DF R + +F G D W+ +IE+ F + +T E KVG A+ A +
Sbjct: 101 SDFARLEHLRFNG----DRITEWLFQIEQFFLIDRTPEELKVGFASLHFDDTAATLHQSI 156
Query: 120 RGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKH 179
M HV +W S+++ L+ + + D LT Q + I EY A+ E ++
Sbjct: 157 VQSMWWKHVRHDWWSYKL-LLQVRYDEHVNDSIAK--LTQLQETEGIEEYHARFELISTR 213
Query: 180 FRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQ 217
F D Y+ ++ GLR D + +VR G Q
Sbjct: 214 LNFAED-----YLVSVYLAGLRTDTQLNVRMFGPQTIQ 246
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 48.5 bits (114), Expect = 4e-06
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 2/96 (2%)
Query: 135 FRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCK 194
F F KYFP A D E++FL L +G +++ EY + + L + R D+ +
Sbjct: 157 FVAEFNAKYFPQEALDRMEARFLELTKGELSVREYGREFKRLLVYAG--RGMEDDQAQMR 214
Query: 195 RFVRGLRADIEDSVRPLGIMRFQALVEKATEVELMK 230
RF+RGLR D+ R AL E +L +
Sbjct: 215 RFLRGLRPDLRVRCRVSQYATKAALTAAEVEEDLQR 250
>At4g07850 putative polyprotein
Length = 1138
Score = 43.5 bits (101), Expect = 1e-04
Identities = 43/173 (24%), Positives = 68/173 (38%), Gaps = 15/173 (8%)
Query: 96 SEGAKVGLATYLLLGDAEYWWRGA---RGMMEANHVEVNWNSFRVAFLEKYFPDSARDER 152
+E KV +A A WW + + + +E WN + ++ P E
Sbjct: 91 TEANKVKVAATEFYDYALSWWDQVVTTKRRLGDDSIET-WNQLKNIMKRRFVPSHYHREL 149
Query: 153 ESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDE--PYMCKRFVRGLRADIEDSVRP 210
+ L QG+ T+ EY ++E+L R V E RF+ GL DI D
Sbjct: 150 HQRLRNLVQGNRTVEEYFKEMETL-----MLRADVQEECEATMSRFMGGLNRDILDRFEV 204
Query: 211 LGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQGKGKLQMKKPYQ 263
+ + L KA +M +++ R SS SYQ + K +K Y+
Sbjct: 205 IHYENLEELFHKA----VMFEKQIKRRSAKPSYNSSKPSYQREEKSGFQKEYK 253
>At5g20875 unknown protein
Length = 272
Score = 42.4 bits (98), Expect = 3e-04
Identities = 60/250 (24%), Positives = 99/250 (39%), Gaps = 51/250 (20%)
Query: 12 NQLAEMVATLVQAMTVQTNDNA------------------QRRAAEDAR---ELHRLQRE 50
N +AEM + L+Q T + Q AAED HR+ R
Sbjct: 47 NSMAEMKSALLQQPTATATSESTKVTVDHDLPVAPPPPTKQSTAAEDNSIDTHPHRMSRY 106
Query: 51 AALDQNRGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLG 110
L + + + P F G + W+ E+ FE ++ K+ L + G
Sbjct: 107 IQLMKPK-------IEMPVFEG----PNVNSWLTRAERYFEFGSFTDAEKIQLVYMSVEG 155
Query: 111 DAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYA 170
A W+ +E + V+WN F+ L+++ + ER LTL+Q I Y
Sbjct: 156 RALCWFN-----LENRNPFVDWNDFKARVLQRFGDPRSAMER---LLTLKQVDSVI-SYL 206
Query: 171 AKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVR---PLG---IMRFQALVEKAT 224
+ E L+ D V E FV+GL+ +I++ +R P G I+ +E +
Sbjct: 207 GEFEDLSTQCLDDLDTVLEVV----FVKGLKEEIQEMLRVFQPKGLSDIIMMARRLENSP 262
Query: 225 EVELMKNRRM 234
L+KN ++
Sbjct: 263 FCRLIKNSQV 272
>At3g43620 putative protein
Length = 221
Score = 42.0 bits (97), Expect = 4e-04
Identities = 36/153 (23%), Positives = 67/153 (43%), Gaps = 12/153 (7%)
Query: 57 RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWW 116
R L + D P+F G D W+ +IE+ F + T E KV +A+ W
Sbjct: 61 RRLRRLGKVDFPRFNGDGIKD----WLFQIEQFFLIDHTPEELKVDIASIHFDDIDATWH 116
Query: 117 RGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESL 176
+ + HV +W ++++ +Y + + L L Q + I Y A+ ES+
Sbjct: 117 QSIVQSIMWRHVRHDWWNYKLLLQVRY---NKHVDDSIAKLKLLQETEGIEVYHARFESI 173
Query: 177 AKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVR 209
R ++DE ++ ++ GL+ D + ++R
Sbjct: 174 CT-----RVKLDEDFLVSLYLTGLKTDTQVNIR 201
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.134 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,206,545
Number of Sequences: 26719
Number of extensions: 263505
Number of successful extensions: 711
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 659
Number of HSP's gapped (non-prelim): 45
length of query: 282
length of database: 11,318,596
effective HSP length: 98
effective length of query: 184
effective length of database: 8,700,134
effective search space: 1600824656
effective search space used: 1600824656
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0338.2


