

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0336.12
(267 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
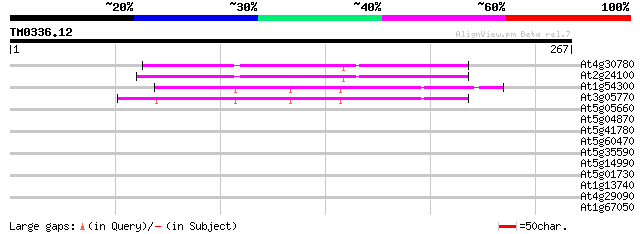
Score E
Sequences producing significant alignments: (bits) Value
At4g30780 unknown protein 115 3e-26
At2g24100 Unknown protein 108 3e-24
At1g54300 hypothetical protein 94 8e-20
At3g05770 unknown protein 91 5e-19
At5g05660 putative protein 32 0.35
At5g04870 calcium-dependent protein kinase 30 1.0
At5g41780 putative protein 29 2.3
At5g60470 putative zinc finger protein 28 5.1
At5g35590 multicatalytic endopeptidase complex alpha subunit-like 28 6.6
At5g14990 putative protein 28 6.6
At5g01730 putative protein 28 6.6
At1g13740 unknown protein (At1g13740) 28 6.6
At4g29090 putative protein 27 8.7
At1g67050 unknown protein 27 8.7
>At4g30780 unknown protein
Length = 589
Score = 115 bits (287), Expect = 3e-26
Identities = 61/156 (39%), Positives = 89/156 (56%), Gaps = 4/156 (2%)
Query: 64 KFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRI 123
K + F A LL +G++ + LV KC FA+ KL+WE+L+ + K K E+ W I
Sbjct: 135 KLKASNFPASLLKIGQWEYKSRYEGDLVAKCYFAKHKLVWEVLE--QGLKSKIEIQWSDI 192
Query: 124 SAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGHAMTYRIHHLE 182
A++A E+ L + L QP+F +E N Q HT W+ + DFT G A R H L+
Sbjct: 193 MALKANCPEDGPGTLTLVLARQPLFFRETNPQPRKHTLWQATS-DFTDGQASMNRQHFLQ 251
Query: 183 FASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
A G + +H+E +++CD RL LS+QP +DSPYF
Sbjct: 252 CAQGIMNKHFEKLVQCDHRLFHLSRQPEIAIDSPYF 287
>At2g24100 Unknown protein
Length = 463
Score = 108 bits (270), Expect = 3e-24
Identities = 58/159 (36%), Positives = 90/159 (56%), Gaps = 4/159 (2%)
Query: 61 TTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPW 120
T K + F A +L +G++ + LV KC FA+ KL+WE+L+ + K K E+ W
Sbjct: 102 TVEKLKASNFPATILRIGQWEYKSRYEGDLVAKCYFAKHKLVWEVLE--QGLKSKIEIQW 159
Query: 121 DRISAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGHAMTYRIH 179
I A++A + E++ L I L +P+F +E N Q HT W+ + DFT G A R H
Sbjct: 160 SDIMALKANLPEDEPGTLTIVLARRPLFFRETNPQPRKHTLWQATS-DFTDGQASMNRQH 218
Query: 180 HLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
L+ G + +H+E +++CD RL LS+QP L +P+F
Sbjct: 219 FLQCPPGIMNKHFEKLVQCDHRLFCLSRQPEINLAAPFF 257
>At1g54300 hypothetical protein
Length = 314
Score = 94.0 bits (232), Expect = 8e-20
Identities = 61/173 (35%), Positives = 90/173 (51%), Gaps = 10/173 (5%)
Query: 70 FYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEIL-----DSSKNKKFKAEVPWDRIS 124
F + +G + + + +V K FA++KLIWE L ++ K K E+ W+ +S
Sbjct: 3 FPISTIRIGGWVVVAKNPDDIVAKFYFAKKKLIWEFLFGEPETNTLRLKRKIEIQWNDVS 62
Query: 125 AIRAFIEE-NKTEILEIELDEQPIFHKEINSQS-THTTWEISDHDFTGGHAMTYRIHHLE 182
+ I ++T IL+IEL ++P F E N Q+ HT W+ DHDFTG HA YR H L
Sbjct: 63 SFEESISSRDETGILKIELKKRPTFFIETNPQAGKHTQWKQLDHDFTGDHASNYRRHTLH 122
Query: 183 FASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYFGPPVLFKNIRDYNTVQH 235
F G L ++ E ++ D KL + PFP +S YF F+N N+ H
Sbjct: 123 FPPGVLQKNLEKLV-TDSFWSKLYEVPFPVHESRYFDSG--FENNSSRNSHSH 172
>At3g05770 unknown protein
Length = 410
Score = 91.3 bits (225), Expect = 5e-19
Identities = 60/179 (33%), Positives = 93/179 (51%), Gaps = 13/179 (7%)
Query: 52 QKSKKQNSSTTLKFAPA-----TFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEIL 106
Q+++ + ++TL +P F + +G F+ + +V K FA++KL+WE L
Sbjct: 53 QQTENSSKTSTLPKSPEKLKAMNFPISTIKIGDCVFVAKNPDDIVAKFYFAKKKLLWEFL 112
Query: 107 -----DSSKNKKFKAEVPWDRISAIRAFIEE-NKTEILEIELDEQPIFHKEINSQS-THT 159
+ K K E+ W+ +S+ I ++T IL+IEL ++P F E N Q+ HT
Sbjct: 113 FGEPVANMPRLKSKIEIQWNDVSSFEESINSRDETGILKIELKKRPTFFTETNPQAGKHT 172
Query: 160 TWEISDHDFTGGHAMTYRIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
W+ D+DFTG A YR H L F G L ++ E +L D KL + PFP +S YF
Sbjct: 173 QWKQLDYDFTGDQASYYRRHTLHFPPGVLQKNLEKLL-TDSFWSKLYKVPFPVHESLYF 230
>At5g05660 putative protein
Length = 820
Score = 32.0 bits (71), Expect = 0.35
Identities = 20/56 (35%), Positives = 24/56 (42%), Gaps = 4/56 (7%)
Query: 170 GGHAMTYRIHHLEFASGDLGQHYENMLKCDERLMK----LSQQPFPRLDSPYFGPP 221
G H +Y H L+ S L + E+ KCD R K Q P PR P PP
Sbjct: 514 GNHYCSYFCHALDIRSSSLDKRSESCEKCDLRCQKERTPRCQHPCPRRCHPEDCPP 569
>At5g04870 calcium-dependent protein kinase
Length = 610
Score = 30.4 bits (67), Expect = 1.0
Identities = 30/102 (29%), Positives = 43/102 (41%), Gaps = 7/102 (6%)
Query: 1 MNSKPPFGLELSSSTMNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSS 60
+ SKP E S T E +P PK P +KR +SA E + ++ ++ K+ S
Sbjct: 93 LESKPETKQETKSETKPESKPD-PPAKPKKPKHMKRVSSAGLRTESVLQRKTENFKEFYS 151
Query: 61 TTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLI 102
K F L V K TT + K A+RKL+
Sbjct: 152 LGRKLGQGQFGTTFLCVEK-----TTGKEFACK-SIAKRKLL 187
>At5g41780 putative protein
Length = 537
Score = 29.3 bits (64), Expect = 2.3
Identities = 33/140 (23%), Positives = 57/140 (40%), Gaps = 15/140 (10%)
Query: 2 NSKPPFGLELSSST--MNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNS 59
N K +EL T ++E + L+ + + K +KE+E +W K+QK
Sbjct: 165 NQKRELEMELVKKTNQVSETQMRLKRLEEETEKRAKAEMKIVKEKEALWNKVQK------ 218
Query: 60 STTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVP 119
L+ TF K + T NQ + + +I EI D SK +++ +
Sbjct: 219 ---LEAGVDTFRKKRKEFNEEMKSKITENQKL----HTKIAVIDEIEDKSKKLEYQVKEQ 271
Query: 120 WDRISAIRAFIEENKTEILE 139
D I + I++ K + E
Sbjct: 272 EDIIQRLSMEIKDQKKLLKE 291
>At5g60470 putative zinc finger protein
Length = 392
Score = 28.1 bits (61), Expect = 5.1
Identities = 15/68 (22%), Positives = 28/68 (41%)
Query: 93 KCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEI 152
K F + ++L +K F P S I + N +++ +LD+ ++
Sbjct: 121 KDSFISHRSFCDVLAEESSKFFSVPSPLAANSTIATVTDTNNPILIQSQLDQSSTGTADL 180
Query: 153 NSQSTHTT 160
N + HTT
Sbjct: 181 NVNNNHTT 188
>At5g35590 multicatalytic endopeptidase complex alpha subunit-like
Length = 246
Score = 27.7 bits (60), Expect = 6.6
Identities = 19/63 (30%), Positives = 33/63 (52%), Gaps = 1/63 (1%)
Query: 33 GLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVV 92
G K T++ MKE+E + ++K K+N S T T + L +V + +F T + VV
Sbjct: 162 GHKATSAGMKEQEAV-NFLEKKMKENPSFTFDETVQTAISALQSVLQEDFKATEIEVGVV 220
Query: 93 KCQ 95
+ +
Sbjct: 221 RAE 223
>At5g14990 putative protein
Length = 666
Score = 27.7 bits (60), Expect = 6.6
Identities = 22/98 (22%), Positives = 46/98 (46%), Gaps = 9/98 (9%)
Query: 92 VKC----QFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQPI 147
VKC + R+ +EIL ++++F+ E+ W ++ + + E +IE + + +
Sbjct: 483 VKCMDCLENLNREKDYEIL--LEDEEFRQELSWIIVTELLREVSETVENHEKIEANNKRV 540
Query: 148 FHKEINSQSTHTTWEISDHDFTGGHAM---TYRIHHLE 182
+E+N + D DF + T+R+ +LE
Sbjct: 541 IEEEVNRACLEISLLYDDFDFKIQEKLKMVTFRLQNLE 578
>At5g01730 putative protein
Length = 1200
Score = 27.7 bits (60), Expect = 6.6
Identities = 19/50 (38%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 30 PPLGLKRTTSAMKEREKMWR--KMQKSKKQNSSTTLKFAP---ATFYAKL 74
P L TTSA+ K+ + ++++SKK+ S TT+K P T +AKL
Sbjct: 194 PSLLKTHTTSAVVATSKLGKDKRLRQSKKKGSHTTIKETPEDSRTSHAKL 243
>At1g13740 unknown protein (At1g13740)
Length = 348
Score = 27.7 bits (60), Expect = 6.6
Identities = 19/53 (35%), Positives = 29/53 (53%), Gaps = 4/53 (7%)
Query: 4 KPPFGLELSSSTMNEREPHLRNVCPKPP---LGLKRTTSAMKEREKMWRKMQK 53
K P L+ SSS + + P + +P +GL+RTTS E E+ WRK ++
Sbjct: 75 KTPRKLKRSSSVL-DTVPFNDSTVAEPENYTVGLERTTSLPAEMEEEWRKRKE 126
>At4g29090 putative protein
Length = 575
Score = 27.3 bits (59), Expect = 8.7
Identities = 23/69 (33%), Positives = 30/69 (43%), Gaps = 10/69 (14%)
Query: 45 EKMWRKMQKSKKQNS-----STTLKFAPATFYAKLLNVGKYNFIPT---TVNQLVVKCQF 96
+K+W+ K Q+ S +L A A Y L P+ TVN L+ KC F
Sbjct: 255 QKIWKSQTSPKIQHFLWKCLSNSLPVAGALAYRHLSKESACIRCPSCKETVNHLLFKCTF 314
Query: 97 ARRKLIWEI 105
AR L W I
Sbjct: 315 AR--LTWAI 321
>At1g67050 unknown protein
Length = 264
Score = 27.3 bits (59), Expect = 8.7
Identities = 19/64 (29%), Positives = 35/64 (54%), Gaps = 3/64 (4%)
Query: 2 NSKPPFGLELSSSTMNEREPHLRNVCPKPPLGLKRTT--SAMKEREKMWRKMQKSKKQNS 59
N+K +G + SSS +N + R++CP P L +T ++ K+++ RK + K
Sbjct: 133 NTKSFWGFKRSSS-LNCGSTYGRSLCPLPLLNRSNSTGSTSSKQKQSSSRKHNEHVKLQQ 191
Query: 60 STTL 63
S++L
Sbjct: 192 SSSL 195
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,105,867
Number of Sequences: 26719
Number of extensions: 254769
Number of successful extensions: 779
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 766
Number of HSP's gapped (non-prelim): 15
length of query: 267
length of database: 11,318,596
effective HSP length: 98
effective length of query: 169
effective length of database: 8,700,134
effective search space: 1470322646
effective search space used: 1470322646
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0336.12


