

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0335b.5
(1648 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
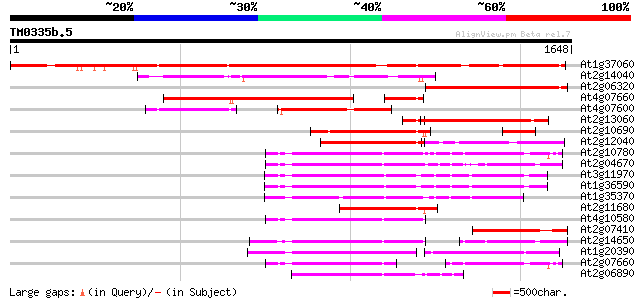
Sequences producing significant alignments: (bits) Value
At1g37060 Athila retroelment ORF 1, putative 1404 0.0
At2g14040 putative retroelement pol polyprotein 724 0.0
At2g06320 putative retroelement pol polyprotein 545 e-154
At4g07660 putative athila transposon protein 535 e-151
At4g07600 456 e-128
At2g13060 pseudogene 446 e-125
At2g10690 putative retroelement pol polyprotein 442 e-123
At2g12040 T10J7.2 389 e-108
At2g10780 pseudogene 380 e-105
At2g04670 putative retroelement pol polyprotein 371 e-102
At3g11970 hypothetical protein 354 2e-97
At1g36590 hypothetical protein 348 1e-95
At1g35370 hypothetical protein 336 8e-92
At2g11680 putative retroelement pol polyprotein 327 4e-89
At4g10580 putative reverse-transcriptase -like protein 309 1e-83
At2g07410 putative retroelement pol polyprotein 290 4e-78
At2g14650 putative retroelement pol polyprotein 290 5e-78
At1g20390 hypothetical protein 273 6e-73
At2g07660 putative retroelement pol polyprotein 263 5e-70
At2g06890 putative retroelement integrase 256 8e-68
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 1404 bits (3633), Expect = 0.0
Identities = 777/1737 (44%), Positives = 1060/1737 (60%), Gaps = 188/1737 (10%)
Query: 2 EDPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAE 61
EDP H+++F R C+ ++ VS + +L LFPFSL D A +W S PQGS+T+W D +
Sbjct: 60 EDPLDHLDEFDRLCSLTKINRVSEDGFKLRLFPFSLGDKAHQWEKSLPQGSITSWNDCKK 119
Query: 62 KFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYD 121
F +FF S +L+NDI FTQ+ +E YEAWE FK +CP H +
Sbjct: 120 AFLAKFFSNSRTARLRNDISGFTQTNNETFYEAWERFKGYQTQCPHHEML---------- 169
Query: 122 GLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAMR--AMNSENERQIRRSSFEVETYDK 179
LD AS+G F + G++++E +A N + +R IR SS E + +
Sbjct: 170 ---------LDTASNGNFLNKDVEDGWEVVENLAQSDGNYNEDYDRSIRTSSDSDEKHRR 220
Query: 180 -LIASNKKLSEEVAEMQKHI-----------QDTKSIGAKVTS----------------- 210
+ A N KL + + QKHI QD +++ + S
Sbjct: 221 EMKAMNDKLDKLLLMQQKHIHFLGDDETLQVQDGETLQLEEVSYVQNQGGYNKGFNNFKQ 280
Query: 211 ----LECKFCGESHDSNQCTVNDDVSKVRALTP---GRGAPTSEQ------------GPL 251
L + ++ +Q + ++ + P G+G +Q G
Sbjct: 281 NHPNLSYRSTNVANRQDQVYPSQQQNQPKPFVPYNQGQGYVPKQQYQGNYQQQLPPPGFT 340
Query: 252 DQPQIVVKTVGKKSLEELIERFID----------------HTSSNYKSHDMAIK--SLES 293
Q Q T L+ ++++ + H + +D+ IK +L S
Sbjct: 341 QQQQQPALTTPDSDLKNMLQQILQGQATGAMDLSKRMAEIHNKVDCSYNDINIKVEALTS 400
Query: 294 QLGQLAKQMSENHQGKFS--SETLTLTEQENTAVVSTRSGRVL---HEPKEKVEGEKNEE 348
++ + Q KF+ S +E ++ RSG+ L P + E +++
Sbjct: 401 KIRYIEGQTGSTAAPKFTGPSGKSMSNSKEYAHAITLRSGKELPTKESPNQNTEDSLDQD 460
Query: 349 AEVERDEGVMEERA------REPTRTRV------------------VNTHTPELRKIPFP 384
E G E+A +PTR V P +PFP
Sbjct: 461 GEDFCQNGNSAEKAIEEPILHQPTRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFP 520
Query: 385 KALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETIL 444
K + K + + K L + +P + L +P K++KD++++R + EV ++
Sbjct: 521 GRFKKVMIQKYKALLEKQLKNLEVTMPLVDCLALIPDSNKYVKDMITER--IKEVQGMVV 578
Query: 445 MTEECSAILQRK-MPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLG 503
++ ECSAI+Q+K +PKK DPGSFT+P + L + LCDLGAS++LM L + K+L
Sbjct: 579 LSHECSAIIQQKIIPKKLGDPGSFTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFN 638
Query: 504 EVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLA 563
+ P ISL +ADRS++ +G++ED+ V + P DFV+L+M+E+ K PLILGRPFLA
Sbjct: 639 KYKPCNISLILADRSVRISHGLLEDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLA 698
Query: 564 TGEAEIKVAKGTLTLKVGED-EVLFNIFDSLKHRADE-EVFRCEVVDELVCEEFVRISIK 621
A I V KG + L +G D ++ F+I +++K E +F E +D L + +
Sbjct: 699 RARAIIDVKKGKIDLNLGRDLKMTFDITNTMKKPTIEGNIFWIEEMDMLADKMLEELGET 758
Query: 622 DPLETTIMEGLEIEDQNLERELDSLY---HEVNATLSQLESVSSLATKSIWKEELTRDEE 678
D L++ + + + D +LE L Y H T + S +SL S + L D+
Sbjct: 759 DHLQSALTKDSKEGDLHLE-TLGGEYILDHITRPTAHSVYSTTSLDHNSSSEANLVSDDW 817
Query: 679 IPIE-EKSELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWTID 737
I+ K +LK LP L+Y +L PVI+N LT + L+ L ++ A+G+++D
Sbjct: 818 SEIKAPKVDLKPLPKGLRYVFLGLNSTYPVIVNDELTADQVNLLITELMKYRKAIGYSLD 877
Query: 738 DIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWV 797
DIKGISP +C H+I LE ++PQRRLNP++K+VV+KEI+KLLDAGVIYPISDS WV
Sbjct: 878 DIKGISPTLCTHRIHLENESYSSIEPQRRLNPNLKEVVKKEILKLLDAGVIYPISDSTWV 937
Query: 798 SPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDR 857
SPV VPKKGG+TVV N +ELIPTR +T R+CI+YR+LN +RK+HFPLPFID ML+R
Sbjct: 938 SPVHCVPKKGGMTVVKNSKDELIPTRTITGHRMCIEYRKLNVASRKEHFPLPFIDHMLER 997
Query: 858 LAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFA 917
LA H YYCFLD YSG+ QI + P DQ KT FTCPYG FAYKRMPFGLCNAPATFQRCM +
Sbjct: 998 LANHPYYCFLDSYSGFFQIPIHPNDQGKTTFTCPYGTFAYKRMPFGLCNAPATFQRCMTS 1057
Query: 918 IFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVL 977
IFSDLIE +E+FMDDFSV+G +F +CL NL VLKRC+ETNLVLNWEKCHFMVR+GIVL
Sbjct: 1058 IFSDLIEEMVEVFMDDFSVYGSSFSSCLLNLCRVLKRCEETNLVLNWEKCHFMVREGIVL 1117
Query: 978 GHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLE 1037
G K+SE+GIEVD+AKI+V+ +L PP +K IRSFLGHAGFYR FIKDFSKLA+P+T LL
Sbjct: 1118 GRKISEEGIEVDKAKIDVMMQLQPPKTVKDIRSFLGHAGFYRIFIKDFSKLARPLTRLLC 1177
Query: 1038 KEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVL 1097
KE F FD+ CL AF+ IKE+L+TAP++ AP+W PFEI+
Sbjct: 1178 KETEFAFDDECLTAFKLIKEALITAPIVQAPNWDFPFEII-------------------- 1217
Query: 1098 YVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQD 1157
+++AQ Y TTEKELL VVFA EKFR Y++G KV ++TDHAALRH++AK+D
Sbjct: 1218 --------TMDDAQVRYATTEKELLAVVFAFEKFRSYLVGSKVTIYTDHAALRHIYAKKD 1269
Query: 1158 SKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVS- 1216
+KPRL+RW+LLLQEFD+EI+D++G +N VADHLSR+ + I + +E+L+A+
Sbjct: 1270 TKPRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMR--IEDEVLIDDSMPEEQLMAIQQ 1327
Query: 1217 -TKEPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCI 1275
++ LPWY N+ V+G P +L+ +KKKF D + WD+P+L+ +C D + R C+
Sbjct: 1328 LNEKKLPWYADHVNYLVSGEEPPNLSSYEKKKFFKDINHFYWDEPYLYTLCKDKIYRTCV 1387
Query: 1276 TEVDFEKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNI 1335
+E + E IL HCHG +YGGHF+ +T +K+LQ+GF+WP++ ++++ F+ CD
Sbjct: 1388 SEDEIEGILLHCHGFAYGGHFATFKTMSKILQAGFWWPSMFKDAQEFISKCD-------- 1439
Query: 1336 SRRNEMPLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKV 1395
E FDVWGIDFMGPFP S+G YILVA+DYVSKWVEA A TND++V
Sbjct: 1440 ------------SFENFDVWGIDFMGPFPSSYGNKYILVAIDYVSKWVEAIASHTNDARV 1487
Query: 1396 VVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEIS 1455
V+ K IF RFGVPR +ISDGG HF N+ FE+LL+K+GVKHK TSGQVEIS
Sbjct: 1488 VLKLFKTIIFPRFGVPRIVISDGGKHFINKGFENLLKKHGVKHK--------TSGQVEIS 1539
Query: 1456 NRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHK 1515
NRE+K ILEK V S+RKDWS KL+D LWAYRTAFKTPIGT+PF+L++GK+CHLPVELE+K
Sbjct: 1540 NREIKAILEKTVGSTRKDWSAKLNDTLWAYRTAFKTPIGTTPFNLLYGKSCHLPVELEYK 1599
Query: 1516 AYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSG 1575
A WA++ LNFD K A EKRL+QLN+L++ RL AYES+ IYKE+TK +HD+KI++R+F G
Sbjct: 1600 AMWAVKLLNFDIKTAEEKRLIQLNDLNKIRLEAYESSKIYKERTKSFHDKKIVSRDFKVG 1659
Query: 1576 QLVLLFNSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVY 1632
VLLFNSRLRLFPGKLKSRWSGPF V V P+GA+ + FTVNGQRLK Y
Sbjct: 1660 DQVLLFNSRLRLFPGKLKSRWSGPFSVTAVRPYGAITLAG--KNGDFTVNGQRLKKY 1714
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 724 bits (1870), Expect = 0.0
Identities = 421/916 (45%), Positives = 548/916 (58%), Gaps = 144/916 (15%)
Query: 376 PELRKIPFPKALVKKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRK 435
P + K+PFP K +K++S F E+ ++L P
Sbjct: 19 PYVPKLPFPGRQRKIQREKEYSLFDEIMRQLQRYSP------------------------ 54
Query: 436 LSEVDETILMTEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLR 495
ECSAILQ +P KR DPGSF +P I T LCDLGA ++LM
Sbjct: 55 ------------ECSAILQNVIPVKREDPGSFVLPSRIGEYTFDRCLCDLGAGVSLMPFS 102
Query: 496 MFKRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPL 555
+ KRL TPT +SL + DRS+ P G+ EDV V+V F P DFVI++++E+ + L
Sbjct: 103 VAKRLGDTNFTPTKMSLVLGDRSISFPVGVAEDVQVRVGNFYIPTDFVIIELDEEPRHRL 162
Query: 556 ILGRPFLATGEAEIKVAKGTLTLKVGEDEVLFNIFDSLKHRADE-EVFRCEVVDELVCEE 614
ILGRPFL A I V K + L++G+ FN+ + E + F +++DELV E
Sbjct: 163 ILGRPFLNIVAALIDVRKSKINLRIGDIVQEFNMERIMSKPTTECQTFWVDIMDELVNEL 222
Query: 615 FVRISIKDPLETTIMEGLEIEDQNLERELDSLYHEVNATLSQLESVSSLATKSIWKEELT 674
++ +DPL+T + + E E L E +++ SS TK + EL
Sbjct: 223 LAELNTEDPLQTVLTKE--------ESEFGYL-GEATTRFARILDSSSPMTKVVAFAELG 273
Query: 675 RDEEIPIEE-------------------KSELKSLPSSLKYAYLEEGENKPVILNSVLTP 715
+E +E+ K ELK LP+ L+ A+L VI+N L
Sbjct: 274 DNE---VEKALVVSSPKDCDDWSELNAPKMELKPLPARLRVAFLGPNSTYLVIINPELNN 330
Query: 716 LKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMKDVV 775
++ LL LR ++ ALG+++DDI GISP +CMH I LE V+ QRRLN +++DVV
Sbjct: 331 VESALLLCELRKYRKALGYSLDDITGISPTLCMHMIHLEGESITSVEHQRRLNSNLRDVV 390
Query: 776 RKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYR 835
+KEI+KLLDAG+IYPISDS WVSPV VVPKKGG+ V+ NE NELIPTR VT R+CIDYR
Sbjct: 391 KKEIMKLLDAGIIYPISDSTWVSPVHVVPKKGGVIVIKNEKNELIPTRTVTGHRMCIDYR 450
Query: 836 RLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVF 895
+LNS TRKD+FPL FIDQML+RL+ YYCFLDGY G+ QI + P+DQEKT FTCPYG F
Sbjct: 451 KLNSATRKDNFPLSFIDQMLERLSNQPYYCFLDGYLGFFQILIHPDDQEKTTFTCPYGTF 510
Query: 896 AYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRC 955
AY+RMPFGLCNAPATFQ CM IFSD+IE +E+F+DDF LVL+RC
Sbjct: 511 AYRRMPFGLCNAPATFQHCMKYIFSDMIEDFMEVFIDDF---------------LVLQRC 555
Query: 956 QETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHA 1015
++ +LVLNWEK HFMVRDGIVLGHK+SEKG+EVDRAKIE++ FLGHA
Sbjct: 556 EDKHLVLNWEKSHFMVRDGIVLGHKISEKGVEVDRAKIEIMR-------------FLGHA 602
Query: 1016 GFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFE 1075
GFYRRFIKDFSK+A+P T LL KE FD+ CL+AF +KESLV+A ++ P+W LPFE
Sbjct: 603 GFYRRFIKDFSKIARPCTQLLCKEQKSEFDDTCLQAFNIVKESLVSALIVQPPEWELPFE 662
Query: 1076 IMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYI 1135
++CDASD +GAVL Q+K++ L+ IYYASR L+ AQ
Sbjct: 663 VICDASDYVVGAVLGQRKDKKLHAIYYASRTLDGAQ------------------------ 698
Query: 1136 LGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRL-- 1193
VIVHT HAALR+L +K+D+KPRL+RW+LLLQ+FDLEI D++G +N VAD+LSRL
Sbjct: 699 ----VIVHTVHAALRYLLSKKDAKPRLLRWILLLQDFDLEIKDKKGIENEVADNLSRLRV 754
Query: 1194 --EGGACSPIP------IQEEFSDE---------KLLAVSTKEP-LPWYVHFANFRVAGL 1235
E +P I++ + DE +L+ + T E LPWY FAN+ +
Sbjct: 755 QEEVLMRDILPGENLASIEDCYMDEVGRLRVSTLELMTLHTGESNLPWYADFANYLSCEV 814
Query: 1236 IPHDLTWQQKKKFLHD 1251
P D T KKK L +
Sbjct: 815 PPPDFTGYFKKKLLKE 830
>At2g06320 putative retroelement pol polyprotein
Length = 466
Score = 545 bits (1403), Expect = e-154
Identities = 255/417 (61%), Positives = 319/417 (76%), Gaps = 2/417 (0%)
Query: 1222 PWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFE 1281
PWY N+ G+ P +LT ++K F D Y WD+P+L+ +C D + R ++E + E
Sbjct: 38 PWYADHVNYLACGIEPPNLTSYERKNFFRDIHHYYWDEPYLYTLCKDKIYRSYVSEDEVE 97
Query: 1282 KILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEM 1341
IL HCH S+YGGHF+ +T +K+LQ+GF+WPT+ ++++ FV CD CQR GNISRRNEM
Sbjct: 98 GILLHCHDSAYGGHFATFKTVSKILQAGFWWPTMFKDAQEFVSKCDSCQRKGNISRRNEM 157
Query: 1342 PLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLK 1401
P ILE+++FDVWGIDFMGPFP S+G YILV VDYVSKWVEA A TND+KVV+ K
Sbjct: 158 PQNPILEVDIFDVWGIDFMGPFPSSYGNKYILVVVDYVSKWVEAIASPTNDAKVVLKLFK 217
Query: 1402 KNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKR 1461
IF RFGV +ISDGG HF N+ FE+LL+K+GVKHKV+TPYHPQTSGQVEISNRE+K
Sbjct: 218 TIIFPRFGVSWVVISDGGKHFINKVFENLLKKHGVKHKVATPYHPQTSGQVEISNREIKT 277
Query: 1462 ILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIR 1521
ILEK V +RKDWS KLDDALWAY+TAFKTPIGT+PF+L+ K+CHL VELE+KA WA++
Sbjct: 278 ILEKTVGITRKDWSTKLDDALWAYKTAFKTPIGTTPFNLLCVKSCHLHVELEYKAMWAVK 337
Query: 1522 KLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLF 1581
LNFD K A EKRL+QL+ELDE RL AYES+ IYKE+TK +HD+KI+ ++F G VLLF
Sbjct: 338 LLNFDIKTAEEKRLIQLSELDEIRLEAYESSKIYKERTKLFHDKKIITKDFQVGDQVLLF 397
Query: 1582 NSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
NSRL++FPGKLKSRWSGPF + V P+GAV + +FTVNGQRLK Y ++L
Sbjct: 398 NSRLKIFPGKLKSRWSGPFCITEVRPYGAVTLAG--KSGVFTVNGQRLKKYLANQIL 452
>At4g07660 putative athila transposon protein
Length = 724
Score = 535 bits (1377), Expect = e-151
Identities = 290/596 (48%), Positives = 386/596 (64%), Gaps = 38/596 (6%)
Query: 451 AILQRKM-PKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLGEVTPTM 509
AI Q+K+ PKK DPGSFT+P + L LCDLGA ++LM L + KRL +
Sbjct: 3 AITQKKIVPKKLIDPGSFTLPCSLGPLAFKRCLCDLGALVSLMPLSVAKRLGFTQYKSCN 62
Query: 510 ISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLATGEAEI 569
ISL +ADRS++ P+ + E++ +++ P DFV+L+M+E+ K PLILGRPFLAT A
Sbjct: 63 ISLILADRSVRIPHSLFENLPIRIGAVDIPTDFVVLEMDEEPKDPLILGRPFLATAGAMN 122
Query: 570 KVAKGTLTLKVGE-DEVLFNIFDSLKHRADE-EVFRCEVVDELVCEEFVRISIKDPLETT 627
V KG + L +G+ + F++ D++K + ++F E +D+L E + +D L +
Sbjct: 123 DVKKGKIDLNLGKYCRMTFDVKDAMKKPTIKGQLFWIEEIDQLADELLEERAEEDHLYSA 182
Query: 628 IMEG-----LEIEDQNLERELDS--------LYHEVNA---------------------- 652
+ + L +E ++ LDS + E+N
Sbjct: 183 LTKRGEDGFLHLETLGYQKLLDSHKAMEESEPFEELNGPETEVMVMSEEGSTQVQPAHSR 242
Query: 653 TLSQLESVSSLATKSIWKEELTRDEEIPIEEKSELKSLPSSLKYAYLEEGENKPVILNSV 712
T S S S+++ + D K +LKSLP L+Y + PVI+N
Sbjct: 243 TYSTNNSTSTISNSGELIIPTSDDWSELKAPKVDLKSLPKGLRYVFFGPNSTYPVIINVE 302
Query: 713 LTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKPIVQPQRRLNPSMK 772
L + LL LR +K A+G+++ DIK ISP++C H+I LE ++PQRRLN ++K
Sbjct: 303 LNDNEVNLLLSELRKYKRAIGYSLSDIKRISPSLCNHRIHLENESYSSIEPQRRLNLNLK 362
Query: 773 DVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCI 832
+VV+KEI+KLLDAGVIYPISDS WV PV VPKKGG+TVV NE +ELIPTR +T RVCI
Sbjct: 363 EVVKKEILKLLDAGVIYPISDSTWVFPVHCVPKKGGMTVVKNEKDELIPTRTITGHRVCI 422
Query: 833 DYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPY 892
DYR+LN+ +RKDHFPLPF +QML+ LA H Y CFLDGYSG+ QI + P DQEKT FTCPY
Sbjct: 423 DYRKLNAASRKDHFPLPFTNQMLEGLANHLYNCFLDGYSGFFQIPIHPNDQEKTTFTCPY 482
Query: 893 GVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVL 952
G FAYKRMPFGLCNAP TFQRCM +IFSDLIE +E+FMDDFSV+GP+F +CL NL VL
Sbjct: 483 GTFAYKRMPFGLCNAPTTFQRCMTSIFSDLIEKMVEVFMDDFSVYGPSFSSCLLNLGRVL 542
Query: 953 KRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGI 1008
+ +ETNLVLNWEKC+FMV++GIVLGHK+SEKGIEVD+ KI+V+ +L PP +K I
Sbjct: 543 TKWEETNLVLNWEKCYFMVKEGIVLGHKISEKGIEVDKEKIKVMMQLQPPKTVKDI 598
Score = 136 bits (343), Expect = 9e-32
Identities = 68/116 (58%), Positives = 90/116 (76%), Gaps = 2/116 (1%)
Query: 1100 IYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSK 1159
IYYASR L+EAQ Y TTEKELL VVFA +KFR Y++ KV V+TDHAALRH++AK+D+K
Sbjct: 598 IYYASRTLDEAQGRYATTEKELLVVVFAFKKFRSYLVESKVTVYTDHAALRHMYAKKDTK 657
Query: 1160 PRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAV 1215
PRL+R +LLLQEFD+EI++++G +N ADHLSR+ PI I + +E+L+ V
Sbjct: 658 PRLLRGILLLQEFDMEIVEKKGIENGAADHLSRMR--IEEPILIDDSMPEEQLMVV 711
>At4g07600
Length = 630
Score = 456 bits (1174), Expect = e-128
Identities = 222/341 (65%), Positives = 263/341 (77%), Gaps = 26/341 (7%)
Query: 788 IYPISDSEW-------VSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCIDYRRLNSV 840
I P SD +W VSPVQ +PKKGGITV+ NE +ELIP R +T R+CIDYR LN+
Sbjct: 309 IIPTSD-DWSKLKAPKVSPVQCIPKKGGITVIKNEKDELIPIRTITGLRMCIDYRNLNAA 367
Query: 841 TRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYGVFAYKRM 900
+R DHFPLPF DQML+RLA H YYCFLDGYSG+ QI + P D EKT FTCPYG FAY+RM
Sbjct: 368 SRNDHFPLPFTDQMLERLANHPYYCFLDGYSGFFQIPIHPNDHEKTTFTCPYGTFAYERM 427
Query: 901 PFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNL 960
PFGLCNAPATFQRCM +IFSD+IE +E+FMDDFSV+GP+F +CL NL VL RC+ETNL
Sbjct: 428 PFGLCNAPATFQRCMTSIFSDIIEEMVEVFMDDFSVYGPSFSSCLLNLGRVLTRCEETNL 487
Query: 961 VLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRR 1020
VLNWEKCHFMV++GI+LGHK+SEKGIEVD+ FLGHAGFYRR
Sbjct: 488 VLNWEKCHFMVKEGIMLGHKISEKGIEVDKG------------------CFLGHAGFYRR 529
Query: 1021 FIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDA 1080
FIKDFSK+A+P+T LL KE F FDE+C ++F +IKE+LV+APV+ A +W PFEIMCDA
Sbjct: 530 FIKDFSKIARPLTRLLCKETEFEFDEDCPKSFHTIKEALVSAPVVRATNWDYPFEIMCDA 589
Query: 1081 SDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKEL 1121
D A+GAVL QK ++ L+VIYYASR L+EAQ Y TTEKEL
Sbjct: 590 LDYAVGAVLGQKIDKKLHVIYYASRTLDEAQGRYATTEKEL 630
Score = 149 bits (377), Expect = 1e-35
Identities = 95/276 (34%), Positives = 160/276 (57%), Gaps = 12/276 (4%)
Query: 399 FLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETILMTEECSAILQRKM- 457
F + K++ + IP +AL + KF+KD++ +R + EV ++++ ECSAI+Q+K+
Sbjct: 2 FAKNIKEVELRIPLVDALALILDTHKFLKDLIVER--IQEVQGMVVLSHECSAIIQKKIV 59
Query: 458 PKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLNLGEVTPTMISLQMADR 517
PKK DPGSFT+P + + LCDLGA ++ M L + KRL + ISL +ADR
Sbjct: 60 PKKLSDPGSFTLPCFLGTVAFNRCLCDLGALVSPMPLSIAKRLGFTQYKSCNISLILADR 119
Query: 518 SLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPFLATGEAEIKVAKGTLT 577
S++ +G+++++ +++ DFVIL+M+E+ K PLIL RPFLAT A I V KG +
Sbjct: 120 SVRISHGLLKNLPIRIGAAEISTDFVILEMDEEPKDPLILRRPFLATAGAMIDVKKGKID 179
Query: 578 LKVGED-EVLFNIFDSLKH-RADEEVFRCEVVDELVCEEFVRISIKDPLETTIMEG---- 631
L +G+D + F+I D++K + ++F E +D+L E ++ +D L + + +
Sbjct: 180 LNLGKDFRMTFDIKDAMKKPNIEGQLFWIEEMDQLADELLEELAQEDHLNSALTKTGQDG 239
Query: 632 -LEIEDQNLERELDSLYHEVNATLSQLESVSSLATK 666
L ++ ++ LDS H+ E ++ ATK
Sbjct: 240 FLHLKVLGYQKLLDS--HKAMEESKPFEELNGPATK 273
>At2g13060 pseudogene
Length = 930
Score = 446 bits (1148), Expect = e-125
Identities = 209/374 (55%), Positives = 267/374 (70%), Gaps = 6/374 (1%)
Query: 1208 SDEKLLAVSTKEPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICS 1267
S E + ++ PWY N+ A + P + T KK+FL + + Y WD+P+L+K
Sbjct: 123 STENCAVTAVEKDYPWYADIVNYLAADVEPDNFTDYNKKRFLREIRRYQWDEPYLYKHSY 182
Query: 1268 DGVIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCD 1327
DG+ RRCI + IL HCH SSYGGHF+ +T +KVLQ+ F+WPT+ R+++ F+ CD
Sbjct: 183 DGIYRRCIAATEVPSILSHCHSSSYGGHFATFKTVSKVLQADFWWPTMFRDTQKFISQCD 242
Query: 1328 RCQRTGNISRRNEMPLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASA 1387
CQR G IS+RNEMP LE+E+FD WGIDFMGPFPPS YILV VDYVSKWVEA A
Sbjct: 243 PCQRRGKISKRNEMPPNFRLEVEVFDRWGIDFMGPFPPSNKNLYILVDVDYVSKWVEAIA 302
Query: 1388 LSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQ 1447
NDS VV+ K IF RFGVPR +ISDG HF N+ E LL +YGV+H+V+TPYHPQ
Sbjct: 303 SLKNDSAVVMKLFKSIIFPRFGVPRIVISDGDKHFINKILEKLLLQYGVQHRVATPYHPQ 362
Query: 1448 TSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACH 1507
TSGQVE+SNR++K ILEK V ++K+WS KL DALWAY+TAFKTP+GT+PFHL++GKACH
Sbjct: 363 TSGQVEVSNRQIKEILEKTVGKAKKEWSYKLYDALWAYKTAFKTPLGTTPFHLLYGKACH 422
Query: 1508 LPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKI 1567
LPVELEHKA WA++ +NFD K A E +ELDE R+ AY+++ +YKE+TK +HD+KI
Sbjct: 423 LPVELEHKAAWAVKMMNFDIKSAGE------SELDEIRIHAYDNSKLYKERTKAYHDKKI 476
Query: 1568 LNREFVSGQLVLLF 1581
L R F +L F
Sbjct: 477 LTRTFEPNDQILRF 490
Score = 65.9 bits (159), Expect = 2e-10
Identities = 30/65 (46%), Positives = 45/65 (69%), Gaps = 2/65 (3%)
Query: 1155 KQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLA 1214
K+D+KPRL+ W+LLLQEFD+E+ D++G +N VADHLSR+ +PI + +E +
Sbjct: 3 KKDAKPRLLIWILLLQEFDIEVRDKKGVENGVADHLSRIR--IDDDVPINDFLPEENIYM 60
Query: 1215 VSTKE 1219
+ T E
Sbjct: 61 IDTVE 65
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 442 bits (1136), Expect = e-123
Identities = 222/372 (59%), Positives = 275/372 (73%), Gaps = 24/372 (6%)
Query: 883 QEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFD 942
++KT FTCP G FAYKRM FGLCNAP TFQR M +IFSD IE +E+FMDDFS F
Sbjct: 163 KKKTTFTCPNGTFAYKRMSFGLCNAPGTFQRSMTSIFSDFIEEIMEVFMDDFS----GFS 218
Query: 943 ACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPP 1002
+CL NL VL+RC+ETNLVLNWEKCHFMV +GIVLGHK+S KGIEVD+AKI+V+ +L PP
Sbjct: 219 SCLLNLCRVLERCEETNLVLNWEKCHFMVHEGIVLGHKISGKGIEVDKAKIDVMIQLQPP 278
Query: 1003 TNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTA 1062
+K IRSFLGHA FYRRFIKDFSK+A+P+T LL KE F FDE+CL+AF IKE+LV+A
Sbjct: 279 KTVKDIRSFLGHAEFYRRFIKDFSKIARPLTRLLCKETEFNFDEDCLKAFHLIKETLVSA 338
Query: 1063 PVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELL 1122
P+ AP+W PFEIMCDA D A+GAVL QK + L+VIYYASR ++EAQ Y T EKELL
Sbjct: 339 PIFQAPNWDHPFEIMCDAFDYAVGAVLDQKIDDKLHVIYYASRTMDEAQTRYATIEKELL 398
Query: 1123 GVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGK 1182
VVFA EK R Y++GFKV V+ DHAALRH++AK+++K RL+RW+LLLQEFD+EIID++G
Sbjct: 399 AVVFAFEKIRSYLVGFKVKVYIDHAALRHIYAKKETKSRLLRWILLLQEFDMEIIDKKGV 458
Query: 1183 DNSVADHLSRLEGGACSPIPIQEEFSDEKLL-------AVSTK-----------EPLPWY 1224
+N VADHLSR+ +PI + +E+L+ + TK E LPWY
Sbjct: 459 ENGVADHLSRMR--IEDSVPIDDTMPEEQLMFYDLVNKSFDTKDMPEEADAVEEEKLPWY 516
Query: 1225 VHFANFRVAGLI 1236
N+ +G +
Sbjct: 517 ADLVNYLTSGQV 528
Score = 135 bits (340), Expect = 2e-31
Identities = 61/98 (62%), Positives = 80/98 (81%)
Query: 1448 TSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACH 1507
TSGQVE+SNR++K IL KVV S++DWS KLD+ LWAYRTA+KTPIG +PF +++GK+CH
Sbjct: 524 TSGQVEVSNRQMKDILTKVVGVSKRDWSTKLDETLWAYRTAYKTPIGRTPFQMLYGKSCH 583
Query: 1508 LPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFR 1545
LPVE+E+KA WA + LN + K A EKR + L+EL+E R
Sbjct: 584 LPVEVEYKAIWATKLLNLEIKGAQEKRPIDLHELEEIR 621
Score = 38.1 bits (87), Expect = 0.043
Identities = 19/49 (38%), Positives = 29/49 (58%)
Query: 684 KSELKSLPSSLKYAYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSAL 732
K +LK LP L+YA+L + PVI+N L+ + +LL LR ++ L
Sbjct: 102 KLDLKPLPKGLRYAFLGPNDTYPVIINDGLSDEQVNQLLNELRKYRGQL 150
>At2g12040 T10J7.2
Length = 976
Score = 389 bits (1000), Expect = e-108
Identities = 184/308 (59%), Positives = 240/308 (77%), Gaps = 2/308 (0%)
Query: 912 QRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMV 971
+R M +IF+D+IE +E+FMDDFSV+G F+ CL NL VL RC+E +LVLNWEKCHF V
Sbjct: 79 ERGMMSIFTDMIEDIMEVFMDDFSVYGSLFEDCLENLYKVLARCEEKHLVLNWEKCHFRV 138
Query: 972 RDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKP 1031
+DGIVLGH++SE GIE DRAKIEV+ L N+K +RSFLGHAGFYRRFIKDFSK+A+P
Sbjct: 139 QDGIVLGHRISEYGIEADRAKIEVMTSLQALDNLKAVRSFLGHAGFYRRFIKDFSKIARP 198
Query: 1032 MTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQ 1091
+T+LL KE F F + C AF+ IK++L++AP++ PDW LPFE+MCDASD A+GAVL Q
Sbjct: 199 LTSLLCKEVKFEFTQECHDAFQQIKQALISAPIVQPPDWDLPFEVMCDASDFAVGAVLGQ 258
Query: 1092 KKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRH 1151
+K++ L+ IYYASR L++A+RNY TTEKE L VVFA EKFR Y++G KVIVHTDHAAL++
Sbjct: 259 RKDKKLHAIYYASRTLDDAKRNYATTEKEFLAVVFAFEKFRSYLVGLKVIVHTDHAALKY 318
Query: 1152 LFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEK 1211
L +D+KPRL+RW+LLLQEFD+E+ D++G + VADHLSR+ +PI + +E
Sbjct: 319 LMQNKDAKPRLLRWILLLQEFDIEVRDKKGVEKCVADHLSRIR--IDDDVPINDFLPEEN 376
Query: 1212 LLAVSTKE 1219
+ + T E
Sbjct: 377 IYMIDTAE 384
Score = 313 bits (803), Expect = 4e-85
Identities = 183/422 (43%), Positives = 237/422 (55%), Gaps = 41/422 (9%)
Query: 1210 EKLLAVSTKEPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDG 1269
E + ++ PWY + A + + KK+FL + + Y WD P+L
Sbjct: 447 ENCAVTALEKDYPWYADIVIYLGADVELDNFIDYNKKRFLREIRRYYWDKPYL------- 499
Query: 1270 VIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRC 1329
I ++F + C+ S S A V SG S + S D C
Sbjct: 500 --PISIVLMEFIGEMHCCNRDSIQSSSSRLLVAYNV--SGC--------SEIHIAS-DPC 546
Query: 1330 QRTGNISRRNEMPLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALS 1389
QR G IS NEMP K ILE+++FD WGI+FMGP PPS YILV VDYVSKWVEA A
Sbjct: 547 QRRGKISNCNEMPQKFILEVKVFDCWGINFMGPIPPSNKNLYILVVVDYVSKWVEAIASP 606
Query: 1390 TNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTS 1449
NDS VV+ K IF FGVPR +ISDGG HF N+ LL +YGV+H+V+TPYHPQTS
Sbjct: 607 KNDSAVVMKLFKCIIFPHFGVPRIVISDGGKHFINKILTKLLLQYGVQHRVATPYHPQTS 666
Query: 1450 GQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLP 1509
GQVE+SNR++K ILEK V AFKTP+GT+PFHL++GKACHLP
Sbjct: 667 GQVEVSNRQIKEILEKTV--------------------AFKTPLGTTPFHLLYGKACHLP 706
Query: 1510 VELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILN 1569
VELEHKA W ++ LNFD K A E+RL+QLNELDE ++ AY+++ +YKE+TK +HD+KIL
Sbjct: 707 VELEHKAAWTVKMLNFDIKSAGERRLIQLNELDEIQIHAYDNSKLYKERTKAYHDKKILT 766
Query: 1570 REFVSGQLVLLFNSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNI-FTVNGQR 1628
R F + F G+ K ++ +F H + E E FT++
Sbjct: 767 RTFEPKDQKKKKTRETKPFKGRGKKSERVGSLLIPIFEHFRIRFEGEEVNTTRFTMDESY 826
Query: 1629 LK 1630
LK
Sbjct: 827 LK 828
Score = 30.4 bits (67), Expect = 8.9
Identities = 15/33 (45%), Positives = 21/33 (63%)
Query: 748 MHKILLEENYKPIVQPQRRLNPSMKDVVRKEII 780
MH+I LE+ K V+ QRRLN K +RK ++
Sbjct: 1 MHRIHLEDESKSSVEHQRRLNLIQKKRLRKRLL 33
>At2g10780 pseudogene
Length = 1611
Score = 380 bits (975), Expect = e-105
Identities = 285/904 (31%), Positives = 453/904 (49%), Gaps = 88/904 (9%)
Query: 751 ILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGIT 810
I LE PI + R+ P+ ++K++ +LLD G I P S S W +PV V KK G
Sbjct: 651 IELEPGTTPISKAPYRMAPAEMAKLKKQLEELLDKGFIRP-SSSPWGAPVLFVKKKDG-- 707
Query: 811 VVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGY 870
+R+CIDYR LN VT K+ +PLP ID+++D+L G Q++ +D
Sbjct: 708 ----------------SFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLA 751
Query: 871 SGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIF 930
SGY+QI + P D KTAF Y F + MPFGL NAPA F + M +F D ++ + IF
Sbjct: 752 SGYHQIPIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIF 811
Query: 931 MDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDR 990
++D V+ +++A +L VL+R +E L KC F R LGH +S++G+ VD
Sbjct: 812 INDILVYSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQRSVGFLGHVISDQGVSVDP 871
Query: 991 AKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLR 1050
KI I++ P N IRSFLG AG+YRRF+ F+ +A+P+T L K+ F + + C +
Sbjct: 872 EKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEK 931
Query: 1051 AFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEA 1110
+F +K L APV+V P+ P+ + DAS + LG VL QK VI YASR L +
Sbjct: 932 SFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGS----VIAYASRQLRKH 987
Query: 1111 QRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQ 1170
++NY T + E+ VVF + +R Y+ G KV ++TDH +L+++F + + R RW+ L+
Sbjct: 988 EKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLRQRRWMELVA 1047
Query: 1171 EFDLEIIDRRGKDNSVADHLSRLEG---GACSPIPIQEEFSDEKLLAVSTKEPLPWYVHF 1227
+++L+I GK N VAD LSR S + + + A+S KE P +
Sbjct: 1048 DYNLDIAYHPGKANQVADALSRRRSEVEAERSQVDLVNMMGTLHVNALS-KEVEPLGLGA 1106
Query: 1228 ANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVI----RRCI--TEVDFE 1281
A+ A L+ Q++ + + K + ++ ++ ++G I R C+ E
Sbjct: 1107 AD--QADLLSRIRLAQERDE---EIKGWAQNNKTEYQTSNNGTIVVNGRVCVPNDRALKE 1161
Query: 1282 KILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEM 1341
+IL H S + H G + L+ ++W + ++ +V C CQ + +++
Sbjct: 1162 EILREAHQSKFSIH-PGSNKMYRDLKRYYHWVGMKKDVARWVAKCPTCQL---VKAEHQV 1217
Query: 1342 P---LKNILEIE-LFDVWGIDFMGPFPPSFGC*Y--ILVAVDYVSKWVEASALSTNDSKV 1395
P L+N+ E +D +DF+ P + + V VD ++K A+S D
Sbjct: 1218 PSGLLQNLPIPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAE 1277
Query: 1396 VVAFLKKNIFTRF-GVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEI 1454
++A + R G+P +I+SD T F ++ +++ + G + +ST YHPQT Q E
Sbjct: 1278 IIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQTDEQSER 1337
Query: 1455 SNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEH 1514
+ + L+ +L V +W + L +AY +F+ IG SP+ ++G+AC P+
Sbjct: 1338 TIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACRTPL---- 1393
Query: 1515 KAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKE---KTKKWHDRKILNRE 1571
W E+RL +DE R KE + K + +++ E
Sbjct: 1394 -----------CWTPVGERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELE 1442
Query: 1572 FVSGQLVLL----------FNSRLRLFPGKLKSRWSGPF-VVKRVFPHGAV--EVENPET 1618
F G LV L F SR +L P R+ GP+ V++RV GAV +++ P
Sbjct: 1443 FQVGDLVYLKAMTYKGAGRFTSRKKLSP-----RYVGPYKVIERV---GAVAYKLDLPPK 1494
Query: 1619 KNIF 1622
N F
Sbjct: 1495 LNAF 1498
Score = 32.0 bits (71), Expect = 3.1
Identities = 38/184 (20%), Positives = 74/184 (39%), Gaps = 20/184 (10%)
Query: 39 DAAEEWLNSQPQ--GSLTTWEDLAEKFTTRFFPRSLLRKLKNDIMTFTQSTDENLYEAWE 96
DA WL + + + ++ D ++F ++FP +L+ + Q + + E E
Sbjct: 223 DAHNWWLTVEKRRGDEVRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQG-NRTVREYDE 281
Query: 97 HFKKLLRKCPQHNLTQAECVAKFYDGLLYSSRFGLDAASSGEFDALPPQAGYDLIEKMAM 156
F +L R + + + +F GL R + F+++ +L+E+ AM
Sbjct: 282 EFNRLRRYVGRELEEEQAQLRRFIRGLRIEIR---NHCLVRTFNSVS-----ELVERAAM 333
Query: 157 RAMNSENERQIRRSSFEVETYDKLIASNKKLSEEVAEMQKHIQDTKSIGAKVTSLECKFC 216
E ER + R + ++KK + + +TKS + EC C
Sbjct: 334 IEEGIEEERYLNREKAPIRNNQSTKPADKKRKFD------KVDNTKS---DAKTGECVTC 384
Query: 217 GESH 220
G++H
Sbjct: 385 GKNH 388
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 371 bits (952), Expect = e-102
Identities = 281/889 (31%), Positives = 429/889 (47%), Gaps = 93/889 (10%)
Query: 751 ILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGIT 810
I LE P+ + R+ P+ ++K++ LL G I P S S W +PV V KK G
Sbjct: 509 IELEPGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGFIRP-STSPWGAPVLFVKKKDG-- 565
Query: 811 VVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGY 870
+R+CIDYR LN VT K+ +PLP ID++LD+L G + +D
Sbjct: 566 ----------------SFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLT 609
Query: 871 SGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIF 930
SGY+QI +A D KTAF YG F + MPF L NAPA F R M ++F + ++ + IF
Sbjct: 610 SGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVIIF 669
Query: 931 MDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDR 990
+DD V+ + + +L V+++ +E L KC F R+ LGH VS +G+ VD
Sbjct: 670 IDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSVDP 729
Query: 991 AKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLR 1050
KIE I PTN IRSFL G+YRRF+K F+ +A+PMT L K+ PF + C
Sbjct: 730 EKIEAIRDWPRPTNATEIRSFLRLTGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEE 789
Query: 1051 AFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEA 1110
F S+KE L + PV+ P+ P+ + DAS + LG VL Q+ + VI YASR L +
Sbjct: 790 GFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQRGK----VIAYASRQLRKH 845
Query: 1111 QRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQ 1170
+ NY T + E+ V+FA + +R Y+ G KV V TDH +L+++F + + R +RW+ L+
Sbjct: 846 EGNYPTHDLEMAVVIFALKIWRSYLYGGKVQVFTDHKSLKYIFNQPELNLRQMRWMELVA 905
Query: 1171 EFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSD--EKLLAVSTKEPLPWYVHFA 1228
++DLEI GK N VAD LSR GA ++ S+ L V +EPL
Sbjct: 906 DYDLEIAYHPGKANVVADALSRKRVGAAPGQSVEALVSEIGALRLCVVAREPL----GLE 961
Query: 1229 NFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVI----RRCITEVD--FEK 1282
A L+ Q+K + L A + ++ ++G I R C+ + + +
Sbjct: 962 AVDRADLLTRARLAQEKDEGLIAASKAEGSE---YQFAANGTIFVYGRVCVPKDEELRRE 1018
Query: 1283 ILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMP 1342
IL H S + H G + L+ + W + R+ +V CD CQ + +++P
Sbjct: 1019 ILSEAHASMFSIH-PGATKMYRDLKRYYQWVGMKRDVANWVAECDVCQL---VKAEHQVP 1074
Query: 1343 LKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKK 1402
+W V +D ++K A+ D V+A KK
Sbjct: 1075 ---------DAIW------------------VIMDRLTKSAHFLAIRKTDGAAVLA--KK 1105
Query: 1403 ---NIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNREL 1459
I GVP +I+SD + F + + K G K ++ST YHPQT GQ E + + L
Sbjct: 1106 YVSEIVKLHGVPVSIVSDRDSKFTFAFWRAFQAKMGTKVQMSTAYHPQTDGQSERTIQTL 1165
Query: 1460 KRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWA 1519
+ +L V W+ L +AY +++ IG +PF ++G+ C P+ +
Sbjct: 1166 EDMLRMCVLDWGGHWADHLSLVEFAYNNSYQASIGMAPFEALYGRPCWTPLRWTQVEERS 1225
Query: 1520 IRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVL 1579
I ++ + R+L+LN + + + + + D++ EF G V
Sbjct: 1226 IYGADYVQETTERIRVLKLNMKEA------------QARQRSYADKRRRELEFEVGDRVY 1273
Query: 1580 LFNSRLR-----LFPGKLKSRWSGPF-VVKRVFPHGAVEVENPETKNIF 1622
L + LR + KL R+ GPF +V+RV P A +E P+ F
Sbjct: 1274 LKMAMLRGPNRSILETKLSPRYMGPFRIVERVGP-VAYRLELPDVMRAF 1321
>At3g11970 hypothetical protein
Length = 1499
Score = 354 bits (909), Expect = 2e-97
Identities = 267/839 (31%), Positives = 407/839 (47%), Gaps = 70/839 (8%)
Query: 749 HKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGG 808
HKI L E P+ Q R + K+ + K + LL G + S S + SPV +V KK G
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQA-SSSPYASPVVLVKKKDG 649
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
WR+C+DYR LN +T KD FP+P I+ ++D L G + +D
Sbjct: 650 T------------------WRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKID 691
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
+GY+Q+ + P+D +KTAF G F Y MPFGL NAPATFQ M IF + +
Sbjct: 692 LRAGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVL 751
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
+F DD V+ + + +L V + + L KC F V LGH +S +GIE
Sbjct: 752 VFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIET 811
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
D AKI+ +++ PT +K +R FLG AG+YRRF++ F +A P+ L + +A F +
Sbjct: 812 DPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDA-FEWTAVA 870
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
+AFE +K +L APV+ P + F + DA +GAVL Q+ + Y+ SR L
Sbjct: 871 QQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYI----SRQLK 926
Query: 1109 EAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLL 1168
Q + + EKELL V+FA K+R Y+L I+ TD +L++L ++ + P +W+
Sbjct: 927 GKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPK 986
Query: 1169 LQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFA 1228
L EFD EI R+GK+N VAD LSR+EG S+ +A++ E A
Sbjct: 987 LLEFDYEIQYRQGKENVVADALSRVEG------------SEVLHMAMTVVECDLLKDIQA 1034
Query: 1229 NFRVAGLIPHDLTWQQKKKFLHDAKSYL-WDDPFLFKICSDGVIRRCITEVDFEKILWHC 1287
+ + +T Q+ D+K Y W L + ++ + +LW
Sbjct: 1035 GYANDSQLQDIITALQRDP---DSKKYFSWSQNILRR--KSKIVVPANDNIKNTILLW-L 1088
Query: 1288 HGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQR-----TGNISRRNEMP 1342
HGS GGH SG + ++ FYW + ++ +A++ SC CQ+ + +P
Sbjct: 1089 HGSGVGGH-SGRDVTHQRVKGLFYWKGMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPLP 1147
Query: 1343 LKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAF-LK 1401
+ + + E+ +DF+ P S G I+V VD +SK ALS S + VA
Sbjct: 1148 IPDTIWSEV----SMDFIEGLPVSGGKTVIMVVVDRLSKAAHFIALSHPYSALTVAHAYL 1203
Query: 1402 KNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKR 1461
N+F G P +I+SD F + + GV K+++ YHPQ+ GQ E+ NR L+
Sbjct: 1204 DNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLET 1263
Query: 1462 ILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKA--CHLPVELEHKAYWA 1519
L + + + WS+ L A + Y T + + +PF +V+G+ HLP
Sbjct: 1264 YLRCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQVPPVHLPY--------- 1314
Query: 1520 IRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLV 1578
L + KVA R LQ E ++ L + + K++ D+ REF G V
Sbjct: 1315 ---LPGESKVAVVARSLQ--EREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDYV 1368
>At1g36590 hypothetical protein
Length = 1499
Score = 348 bits (894), Expect = 1e-95
Identities = 266/839 (31%), Positives = 406/839 (47%), Gaps = 70/839 (8%)
Query: 749 HKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGG 808
HKI L E P+ Q R + K+ + K + LL G + S S + SPV +V KK G
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQA-SSSPYASPVVLVKKKDG 649
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
WR+C+DYR LN +T KD FP+P I+ ++D L G + +D
Sbjct: 650 T------------------WRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKID 691
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
+GY+Q+ + P+D +KTAF G F Y MPFGL NAPATFQ M IF + +
Sbjct: 692 LRAGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVL 751
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
+F DD V+ + + +L V + + L KC F V LGH +S +GIE
Sbjct: 752 VFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIET 811
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
D AKI+ +++ PT +K +R FLG AG+YRRF++ F +A P+ L + +A F +
Sbjct: 812 DPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDA-FEWTAVA 870
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
+AFE +K +L APV+ P + F + DA +GAVL Q+ + Y+ SR L
Sbjct: 871 QQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYI----SRQLK 926
Query: 1109 EAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLL 1168
Q + + EKELL V+FA K+R Y+L I+ TD +L++L ++ + P +W+
Sbjct: 927 GKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPK 986
Query: 1169 LQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFA 1228
L EFD EI R+GK+N VAD LSR+EG S+ +A++ E A
Sbjct: 987 LLEFDYEIQYRQGKENVVADALSRVEG------------SEVLHMAMTVVECDLLKDIQA 1034
Query: 1229 NFRVAGLIPHDLTWQQKKKFLHDAKSYL-WDDPFLFKICSDGVIRRCITEVDFEKILWHC 1287
+ + +T Q+ D+K Y W L + ++ + +LW
Sbjct: 1035 GYANDSQLQDIITALQRDP---DSKKYFSWSQNILRR--KSKIVVPANDNIKNTILLW-L 1088
Query: 1288 HGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQR-----TGNISRRNEMP 1342
HGS GGH SG + ++ FY + ++ +A++ SC CQ+ + +P
Sbjct: 1089 HGSGVGGH-SGRDVTHQRVKGLFYSKGMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPLP 1147
Query: 1343 LKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVA-FLK 1401
+ + + E+ +DF+ P S G I+V VD +SK ALS S + VA
Sbjct: 1148 IPDTIWSEV----SMDFIEGLPVSGGKTVIMVVVDRLSKAAHFIALSHPYSALTVAQAYL 1203
Query: 1402 KNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKR 1461
N+F G P +I+SD F + + GV K+++ YHPQ+ GQ E+ NR L+
Sbjct: 1204 DNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLET 1263
Query: 1462 ILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKA--CHLPVELEHKAYWA 1519
L + + + WS+ L A + Y T + + +PF +V+G+ HLP
Sbjct: 1264 YLRCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQVPPVHLPY--------- 1314
Query: 1520 IRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLV 1578
L + KVA R LQ E ++ L + + K++ D+ REF G V
Sbjct: 1315 ---LPGESKVAVVARSLQ--EREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDYV 1368
>At1g35370 hypothetical protein
Length = 1447
Score = 336 bits (861), Expect = 8e-92
Identities = 246/772 (31%), Positives = 387/772 (49%), Gaps = 68/772 (8%)
Query: 749 HKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGG 808
HKI L E P+ Q R KD + K + ++ +G I +S S + SPV +V KK G
Sbjct: 570 HKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQ-VSSSPFASPVVLVKKKDG 628
Query: 809 ITVVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
WR+C+DY LN +T KD F +P I+ ++D L G + +D
Sbjct: 629 T------------------WRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVVFSKID 670
Query: 869 GYSGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIE 928
+GY+Q+ + P+D +KTAF G F Y M FGL NAPATFQ M ++F D + +
Sbjct: 671 LRAGYHQVRMDPDDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVL 730
Query: 929 IFMDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEV 988
+F DD ++ + + +L LV + + L K H LGH +S + IE
Sbjct: 731 VFFDDILIYSSSIEEHKEHLRLVFEVMRLHKLFAKGSKEH--------LGHFISAREIET 782
Query: 989 DRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENC 1048
D AKI+ +++ PT +K +R FLG AG+YRRF+++F +A P+ + L K F +
Sbjct: 783 DPAKIQAVKEWPTPTTVKQVRGFLGFAGYYRRFVRNFGVIAGPL-HALTKTDGFCWSLEA 841
Query: 1049 LRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLN 1108
AF+++K L APV+ P + F + DA + AVL QK + Y+ SR L
Sbjct: 842 QSAFDTLKAVLCNAPVLALPVFDKQFMVETDACGQGIRAVLMQKGHPLAYI----SRQLK 897
Query: 1109 EAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLL 1168
Q + + EKELL +FA K+R Y+L I+ TD +L++L ++ + P +W+
Sbjct: 898 GKQLHLSIYEKELLAFIFAVRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPK 957
Query: 1169 LQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIP---IQEEFSDEKLLAVSTKEPLPWYV 1225
L EFD EI R+GK+N VAD LSR+EG + ++ +F E +A
Sbjct: 958 LLEFDYEIQYRQGKENLVADALSRVEGSEVLHMALSIVECDFLKEIQVA----------- 1006
Query: 1226 HFANFRVAGLIPHDLTWQQKKKFLHDAKS-YLWDDPFLFKICSDGVIRRCITEVDFEKIL 1284
+ G++ ++ Q+ DAK Y W L + ++ E+ + +
Sbjct: 1007 ----YESDGVLKDIISALQQHP---DAKKHYSWSQDILRR--KSKIVVPNDVEITNKLLQ 1057
Query: 1285 W-HCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQ--RTGNISRRNEM 1341
W HC G G SG + + ++S FYW + ++ +AF+ SC CQ ++ N + +
Sbjct: 1058 WLHCSGM---GGRSGRDASHQRVKSLFYWKGMVKDIQAFIRSCGTCQQCKSDNAAYPGLL 1114
Query: 1342 PLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVV--AF 1399
I + DV +DF+ P S G I+V VD +SK AL+ S + V AF
Sbjct: 1115 QPLPIPDKIWCDV-SMDFIEGLPNSGGKSVIMVVVDRLSKAAHFVALAHPYSALTVAQAF 1173
Query: 1400 LKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNREL 1459
L N++ G P +I+SD F + ++ + GV+ ++S+ YHPQ+ GQ E+ NR L
Sbjct: 1174 L-DNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKLQGVELRMSSAYHPQSDGQTEVVNRCL 1232
Query: 1460 KRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKA--CHLP 1509
+ L + ++ W++ L A + Y T + + +PF LV+G+A HLP
Sbjct: 1233 ENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSSQMTPFELVYGQAPPIHLP 1284
>At2g11680 putative retroelement pol polyprotein
Length = 301
Score = 327 bits (838), Expect = 4e-89
Identities = 164/303 (54%), Positives = 219/303 (72%), Gaps = 19/303 (6%)
Query: 970 MVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLA 1029
MVR+GIVLGHK+S+KG++V++AK++V+ +L PP K IRSFLGHAGFYRRFIKDFSKLA
Sbjct: 1 MVREGIVLGHKISKKGVQVNKAKVDVMMQLQPPKTGKDIRSFLGHAGFYRRFIKDFSKLA 60
Query: 1030 KPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVL 1089
+P+T LL KEA F F+E CL AF+S KE+LV+AP++ AP+ PFEIMCD SD A+GAVL
Sbjct: 61 RPLTRLLCKEAEFIFEEECLTAFKSTKEALVSAPIVQAPNRDYPFEIMCDDSDYAVGAVL 120
Query: 1090 CQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAAL 1149
QK +R L+VIYYAS +++ Q Y TTEKELL VVFA EKFR Y++G KV V+TDHAAL
Sbjct: 121 GQKIDRKLHVIYYASGTMDDTQVRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDHAAL 180
Query: 1150 RHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVADHLSRLEGGACSPIPIQEEFSD 1209
RH++AK+D+KPRL+RW+LLLQEFD+EI+D++G +N VADHLSR+ +PI + +
Sbjct: 181 RHIYAKKDTKPRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMR--IEDEVPIDDSMPE 238
Query: 1210 EKLLAV-----------------STKEPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDA 1252
E+L+A+ + ++ L WY + G P +L+ +KKKF D
Sbjct: 239 EQLMAIQQLNESAQINKSLDQVCAVEDKLMWYADHVKYLFGGEEPPNLSSYEKKKFFKDI 298
Query: 1253 KSY 1255
+
Sbjct: 299 NHF 301
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 309 bits (791), Expect = 1e-83
Identities = 188/474 (39%), Positives = 265/474 (55%), Gaps = 27/474 (5%)
Query: 751 ILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGIT 810
I LE P+ + R+ P+ ++K++ LL G I P S S W +PV V KK G
Sbjct: 483 IELEPGTAPLSKAPYRMAPAEMAELKKQLKDLLGKGFIRP-STSPWGAPVLFVKKKDG-- 539
Query: 811 VVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGY 870
+R+CIDYR LN VT K+ +PLP ID++LD+L G + +D
Sbjct: 540 ----------------SFRLCIDYRELNRVTVKNRYPLPRIDELLDQLRGATCFSKIDLT 583
Query: 871 SGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIF 930
SGY+QI +A D KTAF YG F + MPFGL NAPA F R M ++F + ++ + IF
Sbjct: 584 SGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIF 643
Query: 931 MDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDR 990
+DD V+ + + +L V+++ +E L KC F R+ LGH VS +G+ VD
Sbjct: 644 IDDILVYSKSPEEQEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDP 703
Query: 991 AKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLR 1050
KIE I PTN IRSFLG AG+YRRF+K F+ +A+PMT L K+ PF + + C
Sbjct: 704 EKIEAIRDWPRPTNATEIRSFLGWAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSQECEE 763
Query: 1051 AFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEA 1110
F S+KE L + PV+ P+ P+ + DAS + LG VL Q + VI YASR L +
Sbjct: 764 GFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQHGK----VIAYASRQLMKH 819
Query: 1111 QRNYTTTEKELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQ 1170
+ NY T + E+ V+FA + +R Y+ G KV V TDH +L+++F + + R RW+ L+
Sbjct: 820 EGNYPTHDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVA 879
Query: 1171 EFDLEIIDRRGKDNSVADHLSRLEGGAC---SPIPIQEEFSDEKLLAVSTKEPL 1221
++DLEI GK N V D LSR GA S + E +L AV+ +EPL
Sbjct: 880 DYDLEIAYHPGKANVVVDALSRKRVGAALGQSVEVLVSEIGALRLCAVA-REPL 932
>At2g07410 putative retroelement pol polyprotein
Length = 411
Score = 290 bits (743), Expect = 4e-78
Identities = 159/283 (56%), Positives = 189/283 (66%), Gaps = 37/283 (13%)
Query: 1360 MGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGG 1419
MG FP S+G YILVAVDYVSKWVEA TND+KVV+ K IFTRFGV +ISDG
Sbjct: 1 MGSFPSSYGNKYILVAVDYVSKWVEAIVSPTNDAKVVLKLFKTIIFTRFGVHMVVISDGR 60
Query: 1420 THFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLD 1479
HF N+AFE+LL+K+GVKHKV+T YHPQTSGQVEISNRE+K ILEK V +RKD S KLD
Sbjct: 61 KHFINKAFENLLKKHGVKHKVATSYHPQTSGQVEISNREIKAILEKTVGVTRKDLSAKLD 120
Query: 1480 DALWAY----RTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRL 1535
DALWAY +TAFKTPIGT+PF+L++ K+ HLP+ELE KA WA++ +NFD K A E+RL
Sbjct: 121 DALWAYKTAFKTAFKTPIGTTPFNLLYAKSGHLPIELEFKAMWAVKLMNFDIKTAEEERL 180
Query: 1536 LQLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQLVLLFNSRLRLFPGKLKSR 1595
+QLN+LDE RL AYES+ YK K KLKSR
Sbjct: 181 IQLNDLDEIRLEAYESSKNYKGKR-------------------------------KLKSR 209
Query: 1596 WSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVYQGGEVL 1638
GPF V + P+GAV + FTVNGQRLK Y ++L
Sbjct: 210 LFGPFCVTEIRPYGAVTL--AVKSGDFTVNGQRLKKYLEDQIL 250
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 290 bits (742), Expect = 5e-78
Identities = 186/521 (35%), Positives = 278/521 (52%), Gaps = 40/521 (7%)
Query: 705 KPVILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAIC-MHKILLEENYKPIVQP 763
+P + V++ ++ EK+++ + ++G+ P+ I LE P+ +
Sbjct: 423 RPTSGSLVISAVQAEKMIEKGCEAYLEFEDVFQSLQGLPPSRSDPFTIELEPGTAPLSKA 482
Query: 764 QRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTR 823
R+ P+ ++K++ LL A PV V KK G
Sbjct: 483 PYRMAPAEMAELKKQLEDLLGA-------------PVLFVKKKDG--------------- 514
Query: 824 QVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQ 883
+R+CIDYR LN VT K+ +PLP ID++LD+L G + +D SGY+ I +A D
Sbjct: 515 ---SFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADV 571
Query: 884 EKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDA 943
KTAF YG F + MPFGL NAPA F R M ++F ++++ + IF+DD V+ + +
Sbjct: 572 RKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEE 631
Query: 944 CLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPT 1003
+L V+++ +E L KC F R+ LGH VS +G+ VD KIE I PT
Sbjct: 632 HEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTPT 691
Query: 1004 NIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAP 1063
N IRSFLG AG+YRRF+K F+ +A+PMT L K+ PF + C F S+KE L + P
Sbjct: 692 NATEIRSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTP 751
Query: 1064 VIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLG 1123
V+ P+ P+ + DAS + LG VL Q+ + VI YASR L + + NY T + E+
Sbjct: 752 VLALPEHGEPYMVYTDASGVGLGCVLMQRGK----VIAYASRQLRKHEGNYPTHDLEMAA 807
Query: 1124 VVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKD 1183
V+FA + +R Y+ G V V TDH +L+++F + + R +W+ L+ ++DLEI GK
Sbjct: 808 VIFALKIWRSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLEIAYHPGKA 867
Query: 1184 NSVADHLSRLEGGAC---SPIPIQEEFSDEKLLAVSTKEPL 1221
N VAD LS GA S + E +L AV+ +EPL
Sbjct: 868 NVVADALSHKRVGAAPGQSVEALVSEIGALRLCAVA-REPL 907
Score = 89.4 bits (220), Expect = 2e-17
Identities = 89/334 (26%), Positives = 151/334 (44%), Gaps = 35/334 (10%)
Query: 1322 FVESCDRCQRTGNISRRNEMPLKNILEIEL----FDVWGIDFMGPFPPSFGC*YILVAVD 1377
+V CD CQ + +++P + + + +D IDF+ P S I V VD
Sbjct: 946 WVAECDVCQL---VKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLPVSRTKDAIWVIVD 1002
Query: 1378 YVSKWVEASALSTNDSKVVVAFLKK---NIFTRFGVPRAIISDGGTHFCNRAFESLLEKY 1434
++K A+ D V+A KK I GVP +I+SD + F + + + +
Sbjct: 1003 RLTKSAHFLAIRKTDGAAVLA--KKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEM 1060
Query: 1435 GVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIG 1494
G K ++ST YHPQT GQ E + + L+ +L V W+ L +AY ++ IG
Sbjct: 1061 GTKVQMSTAYHPQTYGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPASIG 1120
Query: 1495 TSPFHLVFGKACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASI 1554
+PF ++ + C P+ L +I ++ + R+L+LN +
Sbjct: 1121 MAPFEALYERPCRTPLCLTQVGERSIYGADYVQETTERIRVLKLNMKEA----------- 1169
Query: 1555 YKEKTKKWHDRKILNREFVSGQLVLLFNSRLR-----LFPGKLKSRWSGPF-VVKRVFPH 1608
+++ + + D++ EF G V L + LR + KL R+ GPF +V+RV P
Sbjct: 1170 -QDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSISETKLSPRYMGPFRIVERVGP- 1227
Query: 1609 GAVEVENPET----KNIFTVNGQRLKVYQGGEVL 1638
A +E P+ +F V+ R +++ EVL
Sbjct: 1228 VAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEVL 1261
>At1g20390 hypothetical protein
Length = 1791
Score = 273 bits (698), Expect = 6e-73
Identities = 162/495 (32%), Positives = 253/495 (50%), Gaps = 20/495 (4%)
Query: 700 EEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISPAICMHKILLEENYKP 759
E + V + + ++P +L+ +L+ + W+I+D+KGI PAI H++ ++ +KP
Sbjct: 718 ESDPTRCVGVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNVDPTFKP 777
Query: 760 IVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNEL 819
+ Q +R+L P V +E+ KLL AG I + EW++ VV KK G
Sbjct: 778 VKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNG----------- 826
Query: 820 IPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVA 879
KWRVC+DY LN KD +PLP ID++++ +G+ F+D +SGYNQI +
Sbjct: 827 -------KWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMH 879
Query: 880 PEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGP 939
+DQEKT+F G + YK M FGL NA AT+QR + + +D I +E+++DD V
Sbjct: 880 KDDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSL 939
Query: 940 NFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKL 999
+ + +L+ + LN KC F V G LG+ V+++GIE + +I I +L
Sbjct: 940 KPEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILEL 999
Query: 1000 TPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESL 1059
P N + ++ G RFI + P NLL++ A F +D++ AFE +K+ L
Sbjct: 1000 PSPRNAREVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRAQFDWDKDSEEAFEKLKDYL 1059
Query: 1060 VTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEK 1119
T P++V P+ + SD A+ +VL ++ I+Y S+ L EA+ Y EK
Sbjct: 1060 STPPILVKPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEK 1119
Query: 1120 ELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDR 1179
L VV + K RPY + V TD LR R+ +W + L E+D++ R
Sbjct: 1120 AALAVVTSARKLRPYFQSHTIAVLTDQ-PLRVALHSPSQSGRMTKWAVELSEYDIDFRPR 1178
Query: 1180 RG-KDNSVADHLSRL 1193
K +AD L L
Sbjct: 1179 PAMKSQVLADFLIEL 1193
Score = 189 bits (481), Expect = 9e-48
Identities = 116/399 (29%), Positives = 198/399 (49%), Gaps = 9/399 (2%)
Query: 1219 EPLPWYVHFANFRVAGLIPHDLTWQQKKKFLHDAKSYLWDDPFLFKICSDGVIRRCITEV 1278
+P W V ++ G +P D W ++ + AK L + L K+ + G + C+
Sbjct: 1376 DPTDWRVEIRDYLSDGTLPSD-KWTARRLRIKAAKYTLMKE-HLLKVSAFGAMLNCLHGT 1433
Query: 1279 DFEKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRR 1338
+ +I+ H + G H G A K+ + GFYWPT+ + + F C++CQR +
Sbjct: 1434 EINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQ 1493
Query: 1339 NEMPLKNILEIELFDVWGIDFMGPFPPSFGC*YILVAVDYVSKWVEASALSTNDSKVVVA 1398
L+ + F W +D +GP P S +ILV DY +KWVEA + +T + V
Sbjct: 1494 PTELLRAGVAPYPFMRWAMDIVGPMPASRQKRFILVMTDYFTKWVEAESYATIRANDVQN 1553
Query: 1399 FLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRE 1458
F+ K I R G+P II+D G+ F + +FE+ + ++ STP +PQ +GQ E +N+
Sbjct: 1554 FVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEATNKT 1613
Query: 1459 LKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYW 1518
+ L+K ++ + W+ +LD LW+YRT ++ +PF +G P E+ Y
Sbjct: 1614 ILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEV---GYS 1670
Query: 1519 AIRKLNFDWKVASEKRLL--QLNELDEFRLRAYESASIYKEKTKKWHDRKILNREFVSGQ 1576
++R+ R++ +L++L+E R A Y+ K +++K+ NR F G
Sbjct: 1671 SLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGD 1730
Query: 1577 LVL--LFNSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEV 1613
LVL +F + + GKL + W G + V ++ G E+
Sbjct: 1731 LVLRKVFENTAEINAGKLGANWEGSYQVSKIVRPGDYEL 1769
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 263 bits (673), Expect = 5e-70
Identities = 152/386 (39%), Positives = 220/386 (56%), Gaps = 23/386 (5%)
Query: 751 ILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGIT 810
I LE PI + R+ P+ ++K++ +LL G I P S S W +PV V KK G
Sbjct: 146 IELEPGTTPISKAPYRMAPAEMAELKKQLEELLAKGFIRP-SSSPWGAPVLFVKKKDG-- 202
Query: 811 VVANENNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGY 870
+R+CIDYR LN VT K+ +PLP ID+++D+L G Q++ +D
Sbjct: 203 ----------------SFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLA 246
Query: 871 SGYNQICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIF 930
SGY+QI + P D KTAF YG F + MPFGL NAPA F + M +F D ++ + IF
Sbjct: 247 SGYHQIPIEPTDVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIF 306
Query: 931 MDDFSVFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDR 990
+DD V +++A +L VL+R +E L K F R LGH +S++G+ VD
Sbjct: 307 IDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFSFWQRSVGFLGHVISDQGVSVDP 366
Query: 991 AKIEVIEKLTPPTNIKGIRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLR 1050
KI I++ P N IRSFLG AG+YRRF+ F+ +A+P+T L K+ F + + C +
Sbjct: 367 EKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEK 426
Query: 1051 AFESIKESLVTAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEA 1110
+F +K L+ APV+V P+ P+ + DAS + LG VL QK VI YASR L +
Sbjct: 427 SFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGS----VIAYASRQLRKH 482
Query: 1111 QRNYTTTEKELLGVVFACEKFRPYIL 1136
++NY T + E+ VVFA + +R Y++
Sbjct: 483 EKNYPTHDLEMAAVVFALKIWRSYLI 508
Score = 99.0 bits (245), Expect = 2e-20
Identities = 97/365 (26%), Positives = 164/365 (44%), Gaps = 50/365 (13%)
Query: 1281 EKILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQRTGNISRRNE 1340
E+IL H S + H G + L+ ++W + ++ +V C CQ + ++
Sbjct: 553 EEILREAHQSKFSIH-PGSNKMYRDLKRYYHWVGMRKDVARWVAKCPTCQL---VKAEHQ 608
Query: 1341 MP---LKNILEIEL-FDVWGIDFMGPFPPSFGC*Y--ILVAVDYVSKWVEASALSTNDSK 1394
+P L+N+ E +D +DF+ P + + V VD ++K A+S D
Sbjct: 609 VPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGA 668
Query: 1395 VVVAFLKKNIFTRF-GVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVE 1453
++A + R G+P +I+SD T F ++ + + + G + +ST YHPQT GQ E
Sbjct: 669 EIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLSTAYHPQTDGQSE 728
Query: 1454 ISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELE 1513
+ + L+ +L V +W + L +AY +F+ IG SP+ ++G+AC P+
Sbjct: 729 RTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALYGRACRTPL--- 785
Query: 1514 HKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKE---KTKKWHDRKILNR 1570
W E+RL +DE R KE + K + +++
Sbjct: 786 ------------CWTPVGERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKEL 833
Query: 1571 EFVSGQLVLL----------FNSRLRLFPGKLKSRWSGPF-VVKRVFPHGAV--EVENPE 1617
EF G LV L F SR +L P R+ GP+ V++RV GAV +++ P
Sbjct: 834 EFQVGDLVYLKAMTYKGAGRFTSRKKLSP-----RYVGPYKVIERV---GAVAYKLDLPP 885
Query: 1618 TKNIF 1622
N+F
Sbjct: 886 KLNVF 890
>At2g06890 putative retroelement integrase
Length = 1215
Score = 256 bits (654), Expect = 8e-68
Identities = 162/505 (32%), Positives = 253/505 (50%), Gaps = 31/505 (6%)
Query: 828 WRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTA 887
WR+C D R +N+VT K P+P +D MLD L G + +D SGY+QI + D+ KTA
Sbjct: 469 WRMCFDCRAINNVTVKYCHPIPRLDDMLDELHGSSIFSKIDLKSGYHQIRMNEGDEWKTA 528
Query: 888 FTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGPNFDACLGN 947
F +G++ + MPFGL +AP+TF R M + I + ++ DD V+ + + +
Sbjct: 529 FKTKHGLYEWLVMPFGLTHAPSTFMRLMNHVLRAFIGIFVIVYFDDILVYSESLREHIEH 588
Query: 948 LALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKLTPPTNIKG 1007
L VL ++ L N +KC F + + LG VS G++VD K++ I P +
Sbjct: 589 LDSVLNVLRKEELYANLKKCTFCTDNLVFLGFVVSADGVKVDEEKVKAIRDWPSPKTVGE 648
Query: 1008 IRSFLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVA 1067
+RSF G AGFYRRF KDFS + P+T +++K+ F +++ AF+S+K+ L APV++
Sbjct: 649 VRSFHGLAGFYRRFFKDFSTIVAPLTEVMKKDVGFKWEKAQEEAFQSLKDKLTNAPVLIL 708
Query: 1068 PDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFA 1127
++ FEI CDAS + +GAVL Q ++ +I + S L A NY T +KEL +V A
Sbjct: 709 SEFLKTFEIECDASGIGIGAVLMQDQK----LIAFFSEKLGGATLNYPTYDKELYALVRA 764
Query: 1128 CEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRRGKDNSVA 1187
++++ Y+ ++HTDH +L+HL +Q R RWV ++ F I ++GKDN VA
Sbjct: 765 LQRWQHYLWPKVFVIHTDHESLKHLKGQQKLNKRHARWVEFIETFAYVIKYKKGKDNVVA 824
Query: 1188 DHLSRLEGGACSPIPIQEEFSDEKLLAVSTKEPLPWYVHFANFRVAGLIPHDLTWQQKKK 1247
D LS ++ +ST + F + HD K
Sbjct: 825 DALS------------------QRYTLLSTLNVK--LMGFEQIKEVYETDHDFQEVYKAC 864
Query: 1248 FLHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKI-LWHCHGSSYGGHFSGERTAAKVL 1306
+ Y D FLF R C+ + + HG GHF +T +V+
Sbjct: 865 EKFASGRYFRQDKFLFY-----ENRLCVPNCSLRDLFVREAHGGGLMGHFGIAKT-LEVM 918
Query: 1307 QSGFYWPTLHRNSRAFVESCDRCQR 1331
F WP + + + C+ C++
Sbjct: 919 TEHFRWPHMKCDVKRICGRCNTCKQ 943
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,514,176
Number of Sequences: 26719
Number of extensions: 1674567
Number of successful extensions: 5529
Number of sequences better than 10.0: 213
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 63
Number of HSP's that attempted gapping in prelim test: 4996
Number of HSP's gapped (non-prelim): 448
length of query: 1648
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1535
effective length of database: 8,299,349
effective search space: 12739500715
effective search space used: 12739500715
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0335b.5


