

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0333b.7
(293 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
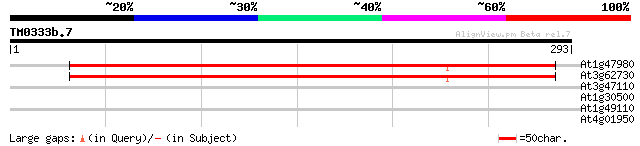
At1g47980 dessication-related protein 298 2e-81
At3g62730 unknown protein 270 5e-73
At3g47110 receptor protein kinase - like protein 33 0.18
At1g30500 transcription factor, putative 30 2.0
At1g49110 hypothetical protein 28 5.8
At4g01950 predicted protein of unknown function 28 7.6
>At1g47980 dessication-related protein
Length = 315
Score = 298 bits (763), Expect = 2e-81
Identities = 155/267 (58%), Positives = 191/267 (71%), Gaps = 13/267 (4%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
D L++F LNLE+LEAEF+ +G LG GLD P L GGP P+G Q A LDP+ RDI Q
Sbjct: 43 DRKLLEFPLNLEYLEAEFFLFGALGLGLDKVAPNLTMGGPSPIGAQKANLDPLTRDIILQ 102
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
FA+Q+ GHLRAI + VKGF RP L++SK+AFA+V++ AFG PPF+PYANS NYL+A+
Sbjct: 103 FAWQEVGHLRAIKKTVKGFARPQLDLSKKAFAKVMDKAFGVKFVPPFNPYANSYNYLIAS 162
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTV 211
Y++PYVGLTGYVG NPKLQ +R LV+GLLG +SGQDA+IR +LY R V PY TV
Sbjct: 163 YLVPYVGLTGYVGANPKLQCPASRKLVAGLLGVESGQDAVIRGMLYARAAHIVYPYGVTV 222
Query: 212 YEVTHRLSLLRNDLGK-------------KQAEGKVVGQVFGLNMYSLTYPRSPEEILRI 258
T ++S LRN LGK AEG+V+G V N SL++ R+PEEILRI
Sbjct: 223 AAFTDKISDLRNKLGKAGVKDEGLIVPKFMGAEGQVIGNVLVGNELSLSFDRTPEEILRI 282
Query: 259 VYGSGDEHVPGGFFPKGGNGNIAKSYL 285
VYGSG+E VPGGF+PKG +G IAKSYL
Sbjct: 283 VYGSGNESVPGGFYPKGADGEIAKSYL 309
>At3g62730 unknown protein
Length = 317
Score = 270 bits (691), Expect = 5e-73
Identities = 140/268 (52%), Positives = 182/268 (67%), Gaps = 14/268 (5%)
Query: 32 DVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQ 91
DVD V F +NLEF EAEF+ G G GLD + LA+GGPPP+G + A LDPI I +
Sbjct: 34 DVDRVHFAMNLEFTEAEFFLKGATGKGLDAYNATLAKGGPPPIGAKKANLDPITNRIIEE 93
Query: 92 FAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAA 151
F +Q+ GHLRAIT+ G PRPL+N+++E FA ++ A G+ NP FDPYANSLNYLLA+
Sbjct: 94 FGYQEIGHLRAITDMTGGIPRPLINLTRENFAVFMDRAVGRKSNPRFDPYANSLNYLLAS 153
Query: 152 YMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQW-T 210
Y +PYVGLTGYVGT P L + LV+GLLG +SGQDA+IRTLLYER+ +V+ Y T
Sbjct: 154 YYIPYVGLTGYVGTIPYLVYFNIKKLVAGLLGVESGQDAVIRTLLYERQNEKVEEYGGVT 213
Query: 211 VYEVTHRLSLLRNDLGK-------------KQAEGKVVGQVFGLNMYSLTYPRSPEEILR 257
V E+T+ +S LRN+LG AE + + + YSL+Y R+ +EILR
Sbjct: 214 VAELTNEISNLRNELGMCGIKDEGLCVPLWLGAENRTTSNILSADPYSLSYDRTAQEILR 273
Query: 258 IVYGSGDEHVPGGFFPKGGNGNIAKSYL 285
++YG+GDEH PGGF+P G NG IA+ +L
Sbjct: 274 VMYGTGDEHRPGGFWPCGANGRIARMFL 301
>At3g47110 receptor protein kinase - like protein
Length = 1025
Score = 33.1 bits (74), Expect = 0.18
Identities = 17/59 (28%), Positives = 27/59 (44%), Gaps = 7/59 (11%)
Query: 110 FPRPLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPK 168
FP P+ N+S F + ++F L P F +L L Y+G+ + GT P+
Sbjct: 241 FPPPIYNLSSLIFLSITGNSFSGTLRPDFGSLLPNLQIL-------YMGINSFTGTIPE 292
>At1g30500 transcription factor, putative
Length = 190
Score = 29.6 bits (65), Expect = 2.0
Identities = 21/72 (29%), Positives = 32/72 (44%), Gaps = 4/72 (5%)
Query: 52 YGTLGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAG-HLRAITEEVKGF 110
Y ++ L ++DP P GQY Y DP R IF+ G HL+ + + +G
Sbjct: 35 YESIVTSLVYSDPGTTNSMAP---GQYPYPDPYYRSIFAPPPQPYTGVHLQLMGVQQQGV 91
Query: 111 PRPLLNISKEAF 122
P P + + F
Sbjct: 92 PLPSDAVEEPVF 103
>At1g49110 hypothetical protein
Length = 212
Score = 28.1 bits (61), Expect = 5.8
Identities = 12/39 (30%), Positives = 24/39 (60%)
Query: 76 GQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPL 114
G Y Y++ ++ FA ++A +LRA+ +E++ F P+
Sbjct: 136 GLYKYVEGLIERKTVSFALREADNLRAVLKEMERFEVPV 174
>At4g01950 predicted protein of unknown function
Length = 520
Score = 27.7 bits (60), Expect = 7.6
Identities = 20/65 (30%), Positives = 26/65 (39%), Gaps = 9/65 (13%)
Query: 20 FIPIFCSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQYA 79
F P+F DV+ V H+ +FYGT GL DP P P
Sbjct: 411 FSPLFTEVSDVIVPVAVTVHVT--------FFYGTTASGLKALDPLFFLLDPYPT-YTIQ 461
Query: 80 YLDPI 84
+LDP+
Sbjct: 462 FLDPV 466
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,084,867
Number of Sequences: 26719
Number of extensions: 320077
Number of successful extensions: 647
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 640
Number of HSP's gapped (non-prelim): 7
length of query: 293
length of database: 11,318,596
effective HSP length: 99
effective length of query: 194
effective length of database: 8,673,415
effective search space: 1682642510
effective search space used: 1682642510
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0333b.7


