

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0328.16
(738 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
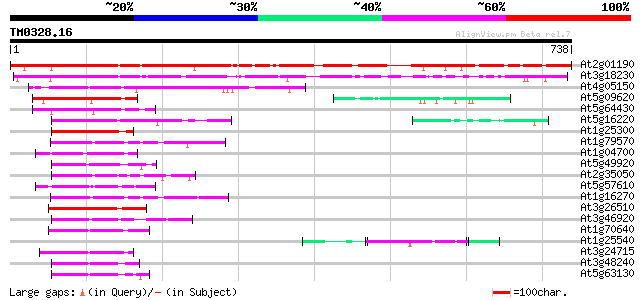
Score E
Sequences producing significant alignments: (bits) Value
At2g01190 unknown protein 603 e-173
At3g18230 unknown protein 338 6e-93
At4g05150 unknown protein 152 5e-37
At5g09620 putative protein 137 2e-32
At5g64430 unknown protein 129 6e-30
At5g16220 unknown protein 128 1e-29
At1g25300 113 4e-25
At1g79570 unknown protein 108 1e-23
At1g04700 putative Ser/Thr protein kinase 101 2e-21
At5g49920 unknown protein 100 4e-21
At2g35050 putative protein kinase 100 4e-21
At5g57610 putative protein 100 5e-21
At1g16270 putative Ser/Thr protein kinase 99 7e-21
At3g26510 unknown protein 99 1e-20
At3g46920 protein kinase - like protein 96 6e-20
At1g70640 unknown protein 93 5e-19
At1g25540 hypothetical protein 90 4e-18
At3g24715 hypothetical protein, 3' partial 89 7e-18
At3g48240 putative protein 86 6e-17
At5g63130 unknown protein 86 1e-16
>At2g01190 unknown protein
Length = 720
Score = 603 bits (1556), Expect = e-173
Identities = 383/788 (48%), Positives = 480/788 (60%), Gaps = 121/788 (15%)
Query: 1 MEAPPQPPTPSAAPPLS---SVAPPPQP-PPHL--NYADSVDSSPRSRNTDSWDD-PFPP 53
ME PP + +A S + AP P P PPH+ +Y +S+DSSPRSR TD WDD P P
Sbjct: 1 MEPPPSSLSSTAVASTSISAATAPVPPPLPPHVTSSYPESLDSSPRSRTTDGWDDLPAPS 60
Query: 54 -----------TSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLS 102
+SKLR MCSYGGHI+PRPHDKSLCY+GGDTRIVVV+R +SL + RLS
Sbjct: 61 GGGGGGGGSAVSSKLRFMCSYGGHILPRPHDKSLCYMGGDTRIVVVDRNSSLPSLIARLS 120
Query: 103 KTFLDGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLF 162
T LDGR FTLKYQLP+EDLDSLISVTTDEDL+NMI+EYDR T + ++S KPSR+RLFLF
Sbjct: 121 NTLLDGRSFTLKYQLPSEDLDSLISVTTDEDLDNMIEEYDR-TISASNSTKPSRLRLFLF 179
Query: 163 PTKPESAHSIPPQILDSSAKSDDWFLDALNGAGLLNRGFSDS-ASVNCLLGLDDVVAGNN 221
+KPE+ S+ QIL+SSAKSDDWFL+ALN AGLLNRGFSDS +VN LLGLDD +A +
Sbjct: 180 TSKPEATQSM-GQILESSAKSDDWFLNALNSAGLLNRGFSDSDTNVNRLLGLDDALALRS 238
Query: 222 NNIEQGGREGVEAGGGGASL----PGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSSP 277
N+ + R+G + A P + QDV+ +PDSPML+TSSSFGSTSSSP
Sbjct: 239 NSGDNNNRDGDDGSVKSAKQQQPPPPQQQEQQQGGQDVNCLPDSPMLDTSSSFGSTSSSP 298
Query: 278 ALANLPPIRTHAEDGGGGGVRVQQQDQKVLGIEEQFAQLGVAAGVGVGQKQDEGFVVITS 337
+LANLPPIR H E+ GGVR DQ+ LGIEEQFA+ V Q D+GF I+S
Sbjct: 299 SLANLPPIRVHVEE--PGGVRT-LPDQRNLGIEEQFARFNVG---NKHQLHDDGFAAISS 352
Query: 338 PPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYRKPPTP--TPQQQ 395
PPP PV T++ PA P+ +A VS ++Q RV+SDDERSDHGV GYRKPPTP PQ
Sbjct: 353 PPPMPV--TIALPA--APVTAATVSNEFQARVYSDDERSDHGVQAGYRKPPTPRSQPQNL 408
Query: 396 AQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQS 455
QQ H Q + S GG +LPSP+SVSSDSS++N M Q+P +YQE +
Sbjct: 409 PPQQAH----------QLKSNSGGGHELPSPNSVSSDSSMSNPMFHQRPSVYQEPI---- 454
Query: 456 GTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQ 515
++P+ S V +N SDP + + Q+ + Y+L QF+
Sbjct: 455 --AQIPSGSTVVTGMINPSDPSTLLSQHQNQDPAYILHPQFE------------------ 494
Query: 516 QHQQQQHQQQQHQQQQQQQQQQQYIHGT---QFIHHNPANA--IPAYYPVYPSQ--QQTH 568
QQ Q Q QQQ+IH Q+IHH+P++ +P Y VYPSQ QQ+
Sbjct: 495 ------------QQSAQSQPQQQFIHTAAPPQYIHHHPSSGLPVPTYIQVYPSQQPQQSF 542
Query: 569 PQHA--------QVYYVPAR-QAQAYNLSMQQA---NMGESGKPQTPPNPTTLVPQTAAY 616
QHA VYYV A + Y++ + Q+ + P PN T + P
Sbjct: 543 HQHAGRLDQQPYPVYYVTAPVPPRPYSMPVPQSPSVSDAAGSIPSNHPNSTMMPPPP--- 599
Query: 617 NNPMRNAPLPKTEMTAAG-YRTA--TGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIA 673
NN MR+ K EM AG Y TA GG+ + Q+P N QQQ++ YSQI HP QS
Sbjct: 600 NNHMRSVSSGKPEMGQAGVYTTAPGVGGAQMVHQIPTN----QQQFMGYSQIRHPPQS-- 653
Query: 674 PNSAPPANYAYDYVDPAHAQIYYSQPLAPTMPSQYQTMTA--AAMVLPEGS-SAQLPSDG 730
SA NY Y+YVD AH QIYY+QP+ +QYQTMT AMV+P+GS +A+LP++
Sbjct: 654 -GSAGNPNYGYEYVDNAHTQIYYTQPMG---HAQYQTMTGPPPAMVMPDGSAAAKLPAEN 709
Query: 731 MKQQTRPS 738
M QQ R S
Sbjct: 710 MTQQIRSS 717
>At3g18230 unknown protein
Length = 666
Score = 338 bits (867), Expect = 6e-93
Identities = 283/765 (36%), Positives = 382/765 (48%), Gaps = 146/765 (19%)
Query: 5 PQPP----TPSAAPPLSSVAPPPQPPPHLNYADSVDSSPRSRNTDSWDDPFP----PTSK 56
P PP TP+ P + + P P + +D SPR+ TD+ P P P +K
Sbjct: 4 PLPPLSQLTPNPTPAAAVSSAIPAPVKSVAVQQPIDGSPRAVATDTGGIPEPLAAVPGAK 63
Query: 57 LRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKYQ 116
LRLMCS+GGHI+PRPHDKSL Y GG+TRIVVV+R SL+ + +RLS L+GR FTLKYQ
Sbjct: 64 LRLMCSFGGHIMPRPHDKSLTYSGGETRIVVVDRRASLSSLRSRLSSMLLNGRSFTLKYQ 123
Query: 117 LPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQI 176
LP+EDLDSL+++TTDEDLENMI+EYDR ++A++ R+RLFLF K E+A ++ +
Sbjct: 124 LPSEDLDSLVTITTDEDLENMIEEYDR-AASSATATATQRLRLFLFANKLETAATM-GSL 181
Query: 177 LDSSAKSDDWFLDALNGAGLLNRGFSDSASV-NCLLGLDDVVAGNN--NNIEQGGREGVE 233
LD + KSD WF+DALN +GLL RG SDSA+V N L+ LD+ G N+E
Sbjct: 182 LDGT-KSDTWFVDALNQSGLLPRGLSDSAAVNNTLVNLDEASGGETEIQNLE------TN 234
Query: 234 AGGGGASLPGSFANCKNSKQD--VSSVPDSPMLETS-SSFGSTSSSPALANLPPIRTHAE 290
GG AN S Q+ +SS+PDSPM+E + SS GS+SSSP+ +NLPPIR
Sbjct: 235 VGGENNKRGDLVANGVISHQEMHMSSMPDSPMMEAAGSSIGSSSSSPSTSNLPPIR---- 290
Query: 291 DGGGGGVRVQQQDQKVLGIEEQFAQLGVAAGVGVGQKQDEGFVVITSPPPPPVPTTLSAP 350
VRV +DQ+ IEEQ AQ+ + + ++ D+G ++ + P P ++
Sbjct: 291 ------VRV-SEDQR---IEEQLAQM-TFSNMQTQRQFDDGVSLMANRPMMIPPGAMNDA 339
Query: 351 AMGVPINSAVVSGD---YQNRVFSDDERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQ 407
AM N+A G V +D+RS+ GV GYRKPP P
Sbjct: 340 AMA--YNNAPGDGTAPASNGHVSPEDDRSELGVSTGYRKPPLP----------------- 380
Query: 408 SHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPV 467
P R+ GG L SPDSV+SD+S+++A S KP+ YQ+Q R P PV
Sbjct: 381 MQPVTIPPRTVGGYGLTSPDSVASDTSISSATSFSKPMYYQDQ---PPALPRAP----PV 433
Query: 468 DPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQH 527
S S++ Q + H+L Q QQ
Sbjct: 434 AQPETTSAQSSQVLPQTENTSA-------QTTSHVLSQPGTYTTMDQQ------------ 474
Query: 528 QQQQQQQQQQQYIH-GTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYN 586
QQQ QQ ++H G Q+I H P+ Y P+Y QQQ +P VY + Q+Q Y
Sbjct: 475 --QQQPLAQQPFLHQGVQYIPH-PSQ----YIPIYSHQQQNYP----VYVMSVPQSQQYI 523
Query: 587 LSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLV 646
+G P PN + P N+ + E YR A PQ++
Sbjct: 524 ---------PAGTPPLYPN-----------SKPGTNS---RPEAAQNVYRAA---PPQVI 557
Query: 647 QVPANQLQHQQQYVTYSQIHHPSQSIAPN---SAPP------ANYA--YDYVDPAHAQIY 695
QLQ Q QY+ Y+ S + N AP ANY ++Y + + +Y
Sbjct: 558 -----QLQQQHQYMGYAGAPQHSTNANANYGTGAPQHSTNANANYGGPFEYTNSPNETVY 612
Query: 696 Y--SQPLAPT---MPSQYQTMT-AAAMVLPEGSSAQLPSDGMKQQ 734
Y P A T + S YQ+MT AAA S Q+ DG KQQ
Sbjct: 613 YHTQAPAANTAIPLASPYQSMTPAAAAAALADMSKQMALDGGKQQ 657
>At4g05150 unknown protein
Length = 477
Score = 152 bits (385), Expect = 5e-37
Identities = 129/392 (32%), Positives = 187/392 (46%), Gaps = 47/392 (11%)
Query: 25 PPPHLNYADSVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTR 84
PPP L+ DS+ SSPRS + ++R MC++GG I+PRP D LCYVGGD R
Sbjct: 36 PPPELDN-DSLASSPRSE--------YDSQPRVRFMCTFGGRILPRPPDNQLCYVGGDNR 86
Query: 85 IVVVERATSLADISTRLSKTFLDGRP-FTLKYQLPNEDLDSLISVTTDEDLENMIDEYDR 143
+V V R T+ A + ++L+K L G+ ++KYQLPNEDLD+LISV+TDED+ENM+DEYDR
Sbjct: 87 MVAVHRHTTFASLLSKLAK--LSGKSNISVKYQLPNEDLDALISVSTDEDVENMMDEYDR 144
Query: 144 TTTTTASSIKPSRIRLFLFPTK------PESAHSIPPQILDSSAKSDDWFLDALN-GAGL 196
+ + SR+RLFLF +S S +LDSS + WFLDALN G+
Sbjct: 145 VAQN--QNPRASRLRLFLFTKNVAGEEDNDSRASSISSLLDSSVNREQWFLDALNLGSSA 202
Query: 197 LNRGFSDSASVNCLLGLDDVVAGNNNNIEQ--GGREGVEAGGGGASLPGSFANCKNSKQ- 253
S+ S + V+ + + G + + L K ++
Sbjct: 203 AATAVSNGGSGRVFERVRSEVSSIVSEVPDYLFGLDNFDETAPPHELRDRDPRAKIQREV 262
Query: 254 DVSSVPDSPMLETSSSFGSTSSSPAL----ANLPP---IRTHAED------GGGGGVRVQ 300
S P SP + S +GSTSS+P + LPP I+ + + +V
Sbjct: 263 STLSDPGSPRRDVPSPYGSTSSAPVMRISTPELPPPVFIKPESPEPVSTPKSNPQPEQVM 322
Query: 301 QQDQKVLGIEEQFAQLGVAAGVGVGQKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAV 360
QQ + + Q+A G GQ+ I P VP ++ M N V
Sbjct: 323 QQSNLPVNSQWQYAP-------GPGQQVHYQGHTIHQSPVYYVPGSVPGNHMVQQGNHMV 375
Query: 361 VSGDYQ---NRVFSDDERSDHGVPVGYRKPPT 389
G++ ++ + H VP+GY +P T
Sbjct: 376 QPGNHMVQPVQMPGQYLQQYHHVPMGYHQPQT 407
>At5g09620 putative protein
Length = 531
Score = 137 bits (345), Expect = 2e-32
Identities = 75/147 (51%), Positives = 93/147 (63%), Gaps = 11/147 (7%)
Query: 30 NYADSVDSSPRSR-----NTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTR 84
+Y DS +SSPRSR N W+D K++LMCSYGG I PRPHD L YV GDT+
Sbjct: 8 SYPDSAESSPRSRDVEFENPSPWEDQQQQNYKVKLMCSYGGKIQPRPHDNQLTYVNGDTK 67
Query: 85 IVVVERATSLADISTRLSKTFL---DGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDEY 141
I+ V+R + ++LS DG + KYQLP EDLD+LISVT DEDLE+M+ EY
Sbjct: 68 IMSVDRGIRFPALVSKLSAVCSGGGDGGEISFKYQLPGEDLDALISVTNDEDLEHMMHEY 127
Query: 142 DRTTTTTASSIKPSRIRLFLFPTKPES 168
DR S KP+R+RLFLFP+ P S
Sbjct: 128 DRLLRL---STKPARMRLFLFPSSPIS 151
Score = 52.8 bits (125), Expect = 7e-07
Identities = 81/309 (26%), Positives = 108/309 (34%), Gaps = 87/309 (28%)
Query: 426 PDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHS-NPVDPKLNVSDPQSRIQVQQ 484
P V + + + P ++ +Q ++Q P H NP + + + + Q IQ++
Sbjct: 201 PSPVKVPQPVPEPVVLEPPQMFVDQRMLQ------PEHGVNPAEIQRQIQEFQM-IQIRD 253
Query: 485 HHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQ--HQQQQHQQQQ-HQQQQQQQQQQQ--- 538
EQ L Q Q QQQ + Q H QQ+ HQ Q QQ+ HQ Q QQQQQQ
Sbjct: 254 Q-EQQMLHQNQLHQQQQQQEAIHQNQLHHQQEVIHQNQLLQQEAIHQNQMLQQQQQQEAM 312
Query: 539 YIHGTQ---------FIHHNPANAIPAYYPV-----------------------YPSQQQ 566
Y T+ NPA PV P QQQ
Sbjct: 313 YRRKTEDEAGRYFPPTYTQNPAPVTNQQPPVGYWQGNTNNSNIQGNIYTTTSQNLPEQQQ 372
Query: 567 THPQHAQVYYVPARQAQA----YNLSMQQANMGESGKPQTP------------------- 603
Q QVY +PA Q+QA Y M+ G G +P
Sbjct: 373 ---QQQQVYMIPA-QSQAPGTLYQSVMRPTVQGNQGYYPSPVQRLHHPDAYMEQQNQPGY 428
Query: 604 ----PNPT---------TLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPA 650
P PT ++ PQ P P ++M Y T G + PA
Sbjct: 429 NVVQPQPTFSGGPQVMTSVGPQVMTSVGPPMGLQEPYSQMGKPVYYTVAGEGMMVQPPPA 488
Query: 651 NQLQHQQQY 659
Q QQQY
Sbjct: 489 QPQQQQQQY 497
>At5g64430 unknown protein
Length = 513
Score = 129 bits (324), Expect = 6e-30
Identities = 81/179 (45%), Positives = 105/179 (58%), Gaps = 27/179 (15%)
Query: 30 NYADSVDSSPRSRNTDSWDDPFPP-----------TSKLRLMCSYGGHIVPRPHDKSLCY 78
+Y DS DSSPRSR + +D+P PP + K++ MCSYGG I PRPHD L Y
Sbjct: 8 SYPDSTDSSPRSREIE-FDNPPPPWDDQNQNQQQHSYKVKFMCSYGGKIQPRPHDNQLTY 66
Query: 79 VGGDTRIVVVERATSLADISTRLSKTFLDGR----PFTLKYQLPNEDLDSLISVTTDEDL 134
V G+T+I+ V+R ++++LS G T KYQLP EDLD+LISVT D+DL
Sbjct: 67 VNGETKILSVDRGIRFPVLASKLSTVCGGGDGGGGEVTFKYQLPGEDLDALISVTNDDDL 126
Query: 135 ENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKSD-DWFLDALN 192
E+M+ EYDR S KP+R+RLFLFP SS +SD D F++ALN
Sbjct: 127 EHMMHEYDRLLRL---SSKPARMRLFLFPASSGFGS-------QSSTQSDRDRFVEALN 175
Score = 33.5 bits (75), Expect = 0.44
Identities = 57/237 (24%), Positives = 83/237 (34%), Gaps = 39/237 (16%)
Query: 506 QFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQ-----FIHHNPANAIPAYY-- 558
Q D + + +Q Q Q+ H + Q+QQQQQ+ ++ + I P Y
Sbjct: 234 QPDHVVNPMEIQRQMQEFQRMHIRDQEQQQQQEAVYRRKSNEDGLIETGGGYFSPPYIQN 293
Query: 559 PVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQT------PPNPTTLVPQ 612
P P QQT PQ QA Q + P T P P ++P
Sbjct: 294 PAPPPPQQTIPQ-------TNPQAPPMTTFWQGNHNPGVVFPTTTHGLGLPEQPVYMIPS 346
Query: 613 ---TAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPS 669
+ Y+ P P G G PQ+ +V ++ TY + H
Sbjct: 347 PSPSPVYHAPPPPQPQGVIRPILTGQVNQGGYYPQVQRVASD---------TYREPHQSP 397
Query: 670 QSIA-PNSAPPANYAYDYVDPAHAQIYYSQPLAPTMPSQYQTMTAAAMV--LPEGSS 723
++A P AP A P Q + S P P QY + V LP+ S+
Sbjct: 398 YNVAQPLQAPIGTKAAQQQQPPPPQPFTSGP----PPPQYTAVPPPRQVVGLPDTSA 450
>At5g16220 unknown protein
Length = 476
Score = 128 bits (321), Expect = 1e-29
Identities = 92/250 (36%), Positives = 130/250 (51%), Gaps = 37/250 (14%)
Query: 56 KLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERA--TSLADISTRLSKTFLDGRPFTL 113
KLR+MC YGG IV P KS YVGGDTRIV + + TS A + + L+ T PF +
Sbjct: 30 KLRVMCRYGGSIVSPPQTKSPRYVGGDTRIVAIPSSAETSFASLVSHLTVTLKISYPFKV 89
Query: 114 KYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIP 173
KYQLP+++LDSLISV DED++ M++E+ ++ SSI SRIRLFLFP K + ++
Sbjct: 90 KYQLPDQELDSLISVEADEDVQIMMEEHG--YLSSESSIPQSRIRLFLFPLKSQQSNEAG 147
Query: 174 PQILDSSAKSDDWFLDALN------------GAGLLNRGFSDSASVNCLLGLDDVVAGNN 221
DS + +D L +L ++ V+ L ++ +
Sbjct: 148 ASQGDSDQCKVETDIDWLGIEESNKPIREELTQPVLQHPKTEMWFVDALKSVEMMQTRRT 207
Query: 222 NNIEQGGREGVEAGGGGASLPGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSSPALAN 281
N+ G +G G + G +S MLET+SSFGSTSSS + +N
Sbjct: 208 NSGTSG------SGDGNGGICGQ---------------ESMMLETNSSFGSTSSSVSSSN 246
Query: 282 LPPIRTHAED 291
LPPI++ ED
Sbjct: 247 LPPIKSSGED 256
Score = 44.3 bits (103), Expect = 2e-04
Identities = 52/184 (28%), Positives = 73/184 (39%), Gaps = 36/184 (19%)
Query: 530 QQQQQQQQQYIHGTQFIHHNPANAIPA--YYPVYPSQQQTHPQHAQVYYVPARQAQAYNL 587
Q QQQ Q G +I N +PA Y+ Q PQ +YY+P Q Y+
Sbjct: 324 QAQQQHIQVIYTGQPYITGNSPMTLPATAYHHTNHVYYQRPPQPYPIYYIPVEQ---YSS 380
Query: 588 SMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQ 647
QA P P+T++ N ++P+ +T A P+
Sbjct: 381 RHVQA---------LPVKPSTVL------NYHQVDSPVVRTSSPLA---------PEF-- 414
Query: 648 VPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANYAYDYVDP---AHAQIYYSQPLAPTM 704
++Q+ + V S ++ S Y D D AHAQIY SQP APT+
Sbjct: 415 --SSQVYPLSKPVDSSVQTSSEATLNTTSRDAFIYNTDVDDDNDIAHAQIYKSQPPAPTL 472
Query: 705 PSQY 708
PSQY
Sbjct: 473 PSQY 476
>At1g25300
Length = 272
Score = 113 bits (282), Expect = 4e-25
Identities = 59/108 (54%), Positives = 74/108 (67%), Gaps = 8/108 (7%)
Query: 56 KLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKY 115
KL L+CSYGG I+P P +KSL Y+GG+TR+V+V R S D LS L GR F+LKY
Sbjct: 10 KLHLLCSYGGRIMPLPPEKSLHYIGGETRLVIVPRGISFLDFFKLLSDKLLSGRSFSLKY 69
Query: 116 QLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFP 163
+LP+ D DSLI+V+ +EDL+NMI EYD T + RIRLFLFP
Sbjct: 70 KLPSCDFDSLITVSDNEDLQNMIAEYDST--------RLRRIRLFLFP 109
>At1g79570 unknown protein
Length = 1248
Score = 108 bits (269), Expect = 1e-23
Identities = 72/236 (30%), Positives = 121/236 (50%), Gaps = 19/236 (8%)
Query: 54 TSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTL 113
T+K++++CS+GG I+PRP D L YVGG+T I+ + + S ++ ++ + + R +
Sbjct: 174 TAKVKILCSFGGKILPRPGDSKLRYVGGETHIISIRKDISWQELRQKILEIYYQTR--VV 231
Query: 114 KYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIP 173
KYQLP EDLD+L+SV+++EDL+NM++EY+ S ++R+FLF +
Sbjct: 232 KYQLPGEDLDALVSVSSEEDLQNMLEEYNEMENRGGS----QKLRMFLFSISDMDDALL- 286
Query: 174 PQILDSSAKSDDWFLDALNGAGLLNRGFSDSASVNCLLGLDDVVAGNNNNIEQGGREGV- 232
+ + S+ ++ A+NG + S + LLGLD A N ++ EG+
Sbjct: 287 -GVNKNDGDSEFQYVVAVNGMDI------GSGKNSTLLGLDSSSANNLAELDVRNTEGIN 339
Query: 233 ----EAGGGGASLPGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSSPALANLPP 284
+ G GAS + S Q S+P S L S S ++ ++PP
Sbjct: 340 TIAGDVVGVGASQLMVNGFQQTSAQQSESIPPSSSLHYSQSIPLNAAYQLQQSVPP 395
>At1g04700 putative Ser/Thr protein kinase
Length = 1042
Score = 101 bits (251), Expect = 2e-21
Identities = 59/136 (43%), Positives = 82/136 (59%), Gaps = 9/136 (6%)
Query: 34 SVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATS 93
+V SPR + D + P L+L+CS+GG I+ RP D L Y+GG+TRI+ + +
Sbjct: 100 NVADSPRKMFQTAISDVYLP-EVLKLLCSFGGRILQRPGDGKLRYIGGETRIISIRKHVG 158
Query: 94 LADISTRLSKTF-LDGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSI 152
L ++ + KT+ L P T+KYQLP EDLD+LISV +DEDL +MI+EY T S
Sbjct: 159 LNEL---MHKTYALCNHPHTIKYQLPGEDLDALISVCSDEDLLHMIEEYQEAETKAGS-- 213
Query: 153 KPSRIRLFLFPTKPES 168
RIR+FL P+ S
Sbjct: 214 --QRIRVFLVPSTESS 227
>At5g49920 unknown protein
Length = 288
Score = 100 bits (248), Expect = 4e-21
Identities = 57/143 (39%), Positives = 85/143 (58%), Gaps = 20/143 (13%)
Query: 55 SKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLK 114
SK++ MCS+GG I+PRP D L YVGG+TR+V V S +++ +L T + LK
Sbjct: 8 SKVKFMCSFGGRILPRPSDSVLKYVGGETRVVAVSPDISFSELMKKL--TAITENDIVLK 65
Query: 115 YQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHS--- 171
YQ+ EDLD+L+SV +DED+++MI+EY+R T ++R FLFP P S
Sbjct: 66 YQIIPEDLDALVSVKSDEDVKHMIEEYNRHET--------PKLRTFLFPANPVVLESQLG 117
Query: 172 -IPPQILDSSAKSDDWFLDALNG 193
I PQ ++ +++A+NG
Sbjct: 118 PIEPQTIEQR------YIEAING 134
>At2g35050 putative protein kinase
Length = 1257
Score = 100 bits (248), Expect = 4e-21
Identities = 68/200 (34%), Positives = 111/200 (55%), Gaps = 25/200 (12%)
Query: 55 SKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLK 114
++ + +CS+GG ++PRP D+ L YVGG+TRI+ + + S ++ ++ + F + R T+K
Sbjct: 175 NRAKFLCSFGGKVIPRPRDQKLRYVGGETRIIRISKTISFQELMHKMKEIFPEAR--TIK 232
Query: 115 YQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKP-ESAHSIP 173
YQLP EDLD+L+SV++DEDL+NM++E S KP R+FLF + E A +
Sbjct: 233 YQLPGEDLDALVSVSSDEDLQNMMEEC--IVFGNGGSEKP---RMFLFSSSDIEEAQFV- 286
Query: 174 PQILDSSAKSDDWFLDALNGAGLLNR----GFSDSASVNCLLGLDDVVAGN-NNNIEQGG 228
+ + S+ ++ A+NG L +R G S + LD+++ GN + I++
Sbjct: 287 --MEHAEGDSEVQYVVAVNGMDLSSRRSSLGLSPPGN-----NLDELLHGNFDRKIDRAA 339
Query: 229 REGVEAG----GGGASLPGS 244
E A G SLP S
Sbjct: 340 TEPAVASLTPLAGNESLPAS 359
>At5g57610 putative protein
Length = 1054
Score = 99.8 bits (247), Expect = 5e-21
Identities = 63/159 (39%), Positives = 96/159 (59%), Gaps = 11/159 (6%)
Query: 34 SVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATS 93
SV+SS S + D+P +++ +CS+ G I+PRP D L YVGG+TRIV V R
Sbjct: 5 SVNSSVTSLVSSLNDEPH----RVKFLCSFLGSILPRPQDGKLRYVGGETRIVSVNRDIR 60
Query: 94 LADISTRLSKTFLDGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIK 153
++ +++ + + DG LKYQ P+EDLD+L+SV D+D+ NM++EYD+ S
Sbjct: 61 YEELMSKMRELY-DGAA-VLKYQQPDEDLDALVSVVNDDDVTNMMEEYDK----LGSGDG 114
Query: 154 PSRIRLFLFPTKPESAHSIPPQILDSSAKSDDWFLDALN 192
+R+R+FLF T PE S+ D +S+ ++DALN
Sbjct: 115 FTRLRIFLFST-PEQDGSLHYVERDDQRESERRYVDALN 152
>At1g16270 putative Ser/Thr protein kinase
Length = 1147
Score = 99.4 bits (246), Expect = 7e-21
Identities = 72/234 (30%), Positives = 116/234 (48%), Gaps = 16/234 (6%)
Query: 54 TSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTL 113
T+K++++CS+GG I+PRP D L YVGG+T I+ + + S ++ ++ + + R +
Sbjct: 162 TAKVKVLCSFGGKILPRPGDSKLRYVGGETHIISIRKDISWQELRQKVLEIYY--RTHVV 219
Query: 114 KYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIP 173
KYQLP EDLD+L+SV+ DEDL NM++EY+ S ++R+FLF +
Sbjct: 220 KYQLPGEDLDALVSVSCDEDLLNMMEEYNEMENRGGS----QKLRMFLFSVSDLDGALL- 274
Query: 174 PQILDSSAKSDDWFLDALNGAGLLNRGFSDSASVNCLLGLDDVVAGNNNNIEQGGREGVE 233
+ S S+ ++ A+N L S S + L GLD A N ++ EG+
Sbjct: 275 -GVNKSDVDSEFQYVVAVNDMDL------GSRSNSTLNGLDSSSANNLAELDVRNTEGIN 327
Query: 234 AGGGGASLPGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSSPALANLPPIRT 287
G G + L G ++S Q S P + + S + +PP T
Sbjct: 328 -GVGPSQLTGIDFQ-QSSMQYSESAPPTSFAQYPQSIPHNGAFQFQQAVPPNAT 379
>At3g26510 unknown protein
Length = 196
Score = 98.6 bits (244), Expect = 1e-20
Identities = 57/134 (42%), Positives = 84/134 (62%), Gaps = 10/134 (7%)
Query: 52 PPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTF--LDGR 109
P S ++ +CSYGG I+PR D L Y GG TR++ V R+ S +++++++++ + G
Sbjct: 5 PTNSTIKFLCSYGGKILPRYPDGKLRYNGGHTRVLAVPRSVSFSELASKMAEMCGGVGGS 64
Query: 110 -PFTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPE- 167
T++ QLP EDLD+L+S+T+DEDL N+I+EYD SS P +IR+FL P K
Sbjct: 65 ITVTIRCQLPTEDLDALVSITSDEDLVNLIEEYD-----LVSSSSPMKIRVFLNPPKSAA 119
Query: 168 -SAHSIPPQILDSS 180
S S PP L SS
Sbjct: 120 GSKKSPPPLALPSS 133
>At3g46920 protein kinase - like protein
Length = 1171
Score = 96.3 bits (238), Expect = 6e-20
Identities = 61/187 (32%), Positives = 107/187 (56%), Gaps = 21/187 (11%)
Query: 56 KLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKY 115
K++ +CSY G I+PRP D L YVGG TRIV V++ + ++ + + G P +KY
Sbjct: 74 KVKFLCSYNGKIIPRPSDGMLRYVGGQTRIVSVKKNVRFDEFEQKMIQVY--GHPVVVKY 131
Query: 116 QLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQ 175
QLP+EDLD+L+SV++ ED++NM++E+++ SS ++R+FLF
Sbjct: 132 QLPDEDLDALVSVSSSEDIDNMMEEFEK--LVERSSDGSGKLRVFLFDA----------- 178
Query: 176 ILDSSAKSDDWFLDALNGAGL-LNRGFSDSASVNCLLGLDDVVAGNNN-NIEQGGREGVE 233
SS++ DD F G G+ + + + ++ + ++ + V +G++N N + G + V+
Sbjct: 179 ---SSSEVDDSFGILEYGDGVDIGQRYVEAVN-GVVVSKESVASGSSNPNSDFSGVDVVD 234
Query: 234 AGGGGAS 240
+ G G S
Sbjct: 235 SLGVGQS 241
>At1g70640 unknown protein
Length = 174
Score = 93.2 bits (230), Expect = 5e-19
Identities = 55/135 (40%), Positives = 81/135 (59%), Gaps = 11/135 (8%)
Query: 52 PPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPF 111
P T+ ++ +CSYGG I PR D L Y GGDTR++ V RA S ++ +L + + G
Sbjct: 3 PNTTTVKFLCSYGGRITPRYPDGKLRYQGGDTRVLSVTRAISFTELKKKLGE--ICGIAV 60
Query: 112 T-LKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKP--ES 168
T L+ QLP +DLD+L++V +DEDL+N+++EYD T +I +FL P K +
Sbjct: 61 TSLRCQLPTDDLDALVTVRSDEDLKNLMEEYDLAITAQV------KIHVFLSPLKSTRTT 114
Query: 169 AHSIPPQILDSSAKS 183
A+S PP SS+ S
Sbjct: 115 ANSSPPPSTTSSSSS 129
>At1g25540 hypothetical protein
Length = 809
Score = 90.1 bits (222), Expect = 4e-18
Identities = 63/143 (44%), Positives = 72/143 (50%), Gaps = 11/143 (7%)
Query: 469 PKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQ---- 524
P++ Q + Q+ Q +Q +QQQ QQQHL QQQ Q Q QQQQHQQQQ QQ
Sbjct: 649 PQIPNQQQQQQQQLHQQQQQQQQIQQQQQQQQHLQQQQMPQLQQQQQQHQQQQQQQHQLS 708
Query: 525 --QQHQQQQQQQQQQQYIHG-TQFIHHNPANAIPAYYPVYPSQQQTHP-QHAQVYYVPAR 580
Q HQQQQQQQQQQQ H TQ HH+ + P+ QQQT P Q P
Sbjct: 709 QLQHHQQQQQQQQQQQQQHQLTQLQHHHQQQQQAS--PLNQMQQQTSPLNQMQQQTSPLN 766
Query: 581 QAQAYNLSMQQANMGESGKPQTP 603
Q Q QQ MG Q P
Sbjct: 767 QMQQQQ-QPQQMVMGGQAFAQAP 788
Score = 72.4 bits (176), Expect = 9e-13
Identities = 71/216 (32%), Positives = 83/216 (37%), Gaps = 61/216 (28%)
Query: 386 KPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPV 445
KP P QQQ QQQLH Q Q Q Q QQ Q+
Sbjct: 648 KPQIPNQQQQQQQQLHQQQQQQQQIQQQQQ--------------------------QQQH 681
Query: 446 IYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
+ Q+Q+ PQ + Q QQH QQQ QQ L Q
Sbjct: 682 LQQQQM------------------------PQLQQQQQQH-------QQQQQQQHQLSQL 710
Query: 506 QFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAY-YPVYPSQ 564
Q QQQ QQQQ QQQQHQ Q Q QQQQQ ++ Q +P N + P+ Q
Sbjct: 711 QHHQQQQQQQQQQQQQHQLTQLQHHHQQQQQASPLNQMQ-QQTSPLNQMQQQTSPLNQMQ 769
Query: 565 QQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKP 600
QQ PQ Q+ AQA S Q G+ P
Sbjct: 770 QQQQPQ--QMVMGGQAFAQAPGRSQQGGGGGQPNMP 803
Score = 68.6 bits (166), Expect = 1e-11
Identities = 60/184 (32%), Positives = 74/184 (39%), Gaps = 26/184 (14%)
Query: 471 LNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQ----- 525
L+VSD R+ + + Q QQ QQQ QQQ QQQQ QQQQ QQQ
Sbjct: 626 LSVSDKACRLIGMLFPGDMVVFKPQIPNQQQQQQQQLHQQQQQQQQIQQQQQQQQHLQQQ 685
Query: 526 -----QHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPAR 580
Q QQQQ QQQQQQ +Q HH QQQ Q Q
Sbjct: 686 QMPQLQQQQQQHQQQQQQQHQLSQLQHH---------------QQQQQQQQQQQQQHQLT 730
Query: 581 QAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATG 640
Q Q ++ QQA+ + QT P + QT+ N + + M + A G
Sbjct: 731 QLQHHHQQQQQASPLNQMQQQTSP-LNQMQQQTSPLNQMQQQQQPQQMVMGGQAFAQAPG 789
Query: 641 GSPQ 644
S Q
Sbjct: 790 RSQQ 793
Score = 30.0 bits (66), Expect = 4.9
Identities = 25/95 (26%), Positives = 37/95 (38%), Gaps = 8/95 (8%)
Query: 643 PQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANYAYDYVDPAHAQIYYSQPLAP 702
PQL Q Q QQQ SQ+ H Q + + H Q + PL
Sbjct: 688 PQLQQQQQQHQQQQQQQHQLSQLQHHQQQQQQQQQQQQQHQLTQLQHHHQQQQQASPL-- 745
Query: 703 TMPSQYQTMTAAAMVLPEGSSAQLPSDGMKQQTRP 737
+Q Q T+ + + +S P + M+QQ +P
Sbjct: 746 ---NQMQQQTSPLNQMQQQTS---PLNQMQQQQQP 774
>At3g24715 hypothetical protein, 3' partial
Length = 800
Score = 89.4 bits (220), Expect = 7e-18
Identities = 49/124 (39%), Positives = 74/124 (59%), Gaps = 6/124 (4%)
Query: 40 RSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADIST 99
+S N F K++ +CS+GG I+PR D+ L YVGG+T I+ + + S ++
Sbjct: 159 QSNNFTGGGGDFDRFGKVKFLCSFGGRIMPRSTDEKLKYVGGETHIISIRKNLSWEELKK 218
Query: 100 RLSKTFLDGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRL 159
+ S + + ++KYQLP ++LDSLISV++DEDL+NMI+EY+ S R RL
Sbjct: 219 KTSA--ICQQLHSIKYQLPGDELDSLISVSSDEDLQNMIEEYNGLERLEGS----QRPRL 272
Query: 160 FLFP 163
FL P
Sbjct: 273 FLIP 276
>At3g48240 putative protein
Length = 180
Score = 86.3 bits (212), Expect = 6e-17
Identities = 49/117 (41%), Positives = 69/117 (58%), Gaps = 10/117 (8%)
Query: 55 SKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLK 114
+ L+ +CSYGG I+PR D L YVGG TR++ V + S +++ +L + G LK
Sbjct: 12 NSLKFLCSYGGRILPRSIDGKLRYVGGFTRVLSVHHSISFTELTMKLEE--FCGYSVELK 69
Query: 115 YQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHS 171
QLPN DL++LISV +DEDL N+++EYDR + +IR L P K S S
Sbjct: 70 CQLPNGDLETLISVKSDEDLVNIVEEYDR--------VYGGKIRAILSPPKQMSPRS 118
>At5g63130 unknown protein
Length = 192
Score = 85.5 bits (210), Expect = 1e-16
Identities = 54/132 (40%), Positives = 78/132 (58%), Gaps = 15/132 (11%)
Query: 55 SKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLK 114
S L+ +CSYGG I+PR D L YVGG TR++ V+R+ S +++ +L + G L+
Sbjct: 13 SSLKFLCSYGGRILPRSTDGKLRYVGGHTRVLSVDRSISFSELMKKLYE--FCGYSVDLR 70
Query: 115 YQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAH---S 171
QLPN DL++LISV ++E+L +++EYDR I ++IR L P P S+H S
Sbjct: 71 CQLPNGDLETLISVKSEEELAEIVEEYDR--------ICGAKIRAVLSP--PRSSHKTES 120
Query: 172 IPPQILDSSAKS 183
P D S KS
Sbjct: 121 SPSSSGDRSPKS 132
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.127 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,521,709
Number of Sequences: 26719
Number of extensions: 925521
Number of successful extensions: 15548
Number of sequences better than 10.0: 384
Number of HSP's better than 10.0 without gapping: 222
Number of HSP's successfully gapped in prelim test: 173
Number of HSP's that attempted gapping in prelim test: 6194
Number of HSP's gapped (non-prelim): 3571
length of query: 738
length of database: 11,318,596
effective HSP length: 107
effective length of query: 631
effective length of database: 8,459,663
effective search space: 5338047353
effective search space used: 5338047353
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0328.16


