

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0323a.10
(175 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
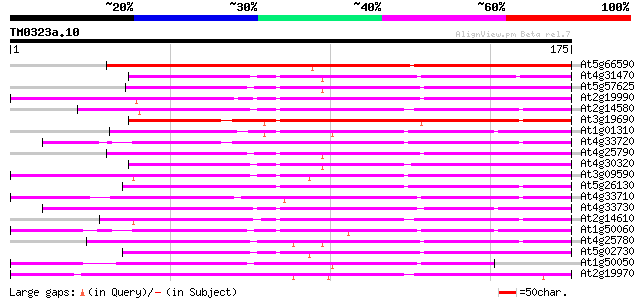
Score E
Sequences producing significant alignments: (bits) Value
At5g66590 Unknown protein (K1F13.27) 159 7e-40
At4g31470 pathogenesis-related protein homolog 110 3e-25
At5g57625 pathogenesis-related protein - like 110 4e-25
At2g19990 pathogenesis-related protein (PR-1) 109 7e-25
At2g14580 pathogenesis-related PR-1-like protein 108 2e-24
At3g19690 PR-1 protein, putative 106 6e-24
At1g01310 pathogenesis related protein, like 106 7e-24
At4g33720 pathogenesis-related protein 1 precursor, 19.3K 105 1e-23
At4g25790 putative pathogenesis-related protein 103 4e-23
At4g30320 PR-1-like protein 103 5e-23
At3g09590 putative pathogenesis-related protein 103 6e-23
At5g26130 pathogenesis-related protein - like 100 5e-22
At4g33710 pathogenesis-related protein 1 precursor, 18.9K 99 9e-22
At4g33730 pathogenesis-related protein - like 99 1e-21
At2g14610 pathogenesis-related PR-1-like protein 99 2e-21
At1g50060 branched-chain amino acid aminotransferase, putative 98 3e-21
At4g25780 putative pathogenesis-related protein 97 3e-21
At5g02730 pathogenesis related protein - like 96 1e-20
At1g50050 pathogenesis-related protein 1b precursor (pr-1b), put... 87 5e-18
At2g19970 putative pathogenesis-related protein 82 2e-16
>At5g66590 Unknown protein (K1F13.27)
Length = 185
Score = 159 bits (402), Expect = 7e-40
Identities = 77/147 (52%), Positives = 101/147 (68%), Gaps = 3/147 (2%)
Query: 31 PQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
P +SA AK F DAHN ARA VGV PL WS+ L A ARYQR++ C+ A+ + KYG
Sbjct: 40 PTISAAAKAFTDAHNKARAMVGVPPLVWSQTLEAAASRLARYQRNQKKCEFASLNPGKYG 99
Query: 91 SNQ--DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQ 148
+NQ G + TPS+AV+ WV EK FY+Y++++C A + C Y QVVWR SK++GCAQ
Sbjct: 100 ANQLWAKGLVAVTPSLAVETWVKEKPFYNYKSDTCAA-NHTCGVYKQVVWRNSKELGCAQ 158
Query: 149 LTCVVDKLILTICFYDPPGNINGESPY 175
TC + +LTICFY+PPGN+ G+ PY
Sbjct: 159 ATCTKESTVLTICFYNPPGNVIGQKPY 185
>At4g31470 pathogenesis-related protein homolog
Length = 185
Score = 110 bits (276), Expect = 3e-25
Identities = 62/139 (44%), Positives = 83/139 (59%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
++FL HN RA++ + PL+WS +LA A +AR +R CKL + S YG N G
Sbjct: 52 QQFLRPHNILRAKLRLPPLKWSNSLALYASRWARTRRG--DCKLIH-SGGPYGENLFWGS 108
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G GWTP AV W +E ++Y RT+ C A C HYTQ+VW+KS ++GCA C
Sbjct: 109 GKGWTPRDAVAAWASEMKYYDRRTSHCKANG-DCLHYTQLVWKKSSRIGCAISFCKTGDT 167
Query: 157 ILTICFYDPPGNINGESPY 175
+ IC YDPPGNI G+ P+
Sbjct: 168 FI-ICNYDPPGNIVGQPPF 185
>At5g57625 pathogenesis-related protein - like
Length = 207
Score = 110 bits (275), Expect = 4e-25
Identities = 60/140 (42%), Positives = 77/140 (54%), Gaps = 5/140 (3%)
Query: 37 AKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG 96
A+ FLD HN R+ +G+ PL W LA A +A +R C L + S YG N G
Sbjct: 72 ARLFLDPHNALRSGLGLPPLIWDGKLASYATWWANQRR--YDCSLTH-STGPYGENLFWG 128
Query: 97 -GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDK 155
G W P AVQ W+ E Y++ TNSC C HYTQ+VWR +K++GCA++ C
Sbjct: 129 SGSSWAPGFAVQSWIVEGRSYNHNTNSCDGSGM-CGHYTQMVWRDTKRLGCARVVCENGA 187
Query: 156 LILTICFYDPPGNINGESPY 175
+ C YDPPGN GE PY
Sbjct: 188 GVFITCNYDPPGNYVGEKPY 207
>At2g19990 pathogenesis-related protein (PR-1)
Length = 176
Score = 109 bits (273), Expect = 7e-25
Identities = 70/177 (39%), Positives = 92/177 (51%), Gaps = 6/177 (3%)
Query: 1 MSIFMQPFFSLALALFLLSAEAATVLDSVVPQLSAEAK--EFLDAHNWARAEVGVEPLEW 58
MS F F +AL L A T + + K E L HN ARA VGV P+ W
Sbjct: 4 MSFFGYSFIVVALFFDLTQAYRHTPAQPPKANANGDVKPQETLVVHNKARAMVGVGPMVW 63
Query: 59 SEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSY 118
+E LA A +A ++R R C + + S +G N G + VA + W+ EKE Y Y
Sbjct: 64 NETLATYAQSYA-HERAR-DCAMKH-SLGPFGENLAAGWGTMSGPVATEYWMTEKENYDY 120
Query: 119 RTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
+N+C C HYTQ+VWR S ++GCA + C D+ I IC YDPPGN G+ PY
Sbjct: 121 DSNTCGGDGV-CGHYTQIVWRDSVRLGCASVRCKNDEYIWVICSYDPPGNYIGQRPY 176
>At2g14580 pathogenesis-related PR-1-like protein
Length = 161
Score = 108 bits (269), Expect = 2e-24
Identities = 63/155 (40%), Positives = 91/155 (58%), Gaps = 8/155 (5%)
Query: 22 AATVLDSVVPQLSAEAKE-FLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCK 80
AA V VVP + ++++ +++AHN AR+++GV P++W E LA A +A + C+
Sbjct: 14 AALVGALVVPLKAQDSQQDYVNAHNQARSQIGVGPMQWDEGLAAYARNYANQLKG--DCR 71
Query: 81 LANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRK 140
L + S YG N G + AV WVNEK Y+Y TN+C + C HYTQVVWR
Sbjct: 72 LVH-SRGPYGENLAKSGGDLSGVAAVNLWVNEKANYNYDTNTC---NGVCGHYTQVVWRN 127
Query: 141 SKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
S ++GCA++ C I++ C YDPPGN + PY
Sbjct: 128 SVRLGCAKVRCNNGGTIIS-CNYDPPGNYANQKPY 161
>At3g19690 PR-1 protein, putative
Length = 161
Score = 106 bits (265), Expect = 6e-24
Identities = 56/140 (40%), Positives = 84/140 (60%), Gaps = 7/140 (5%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLG-CKLANFSHNKYGSNQDLG 96
++FL+AHN AR EVG++PL W + +A A A Y R+ C L + S+ +G N +
Sbjct: 27 QQFLEAHNEARNEVGLDPLVWDDEVAAYA---ASYANQRINDCALVH-SNGPFGENIAMS 82
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS-FPCWHYTQVVWRKSKQVGCAQLTCVVDK 155
+ A + W+NEK++Y Y +N+C P+ C HYTQVVW+ + ++GCA++ C
Sbjct: 83 SGEMSAEDAAEMWINEKQYYDYDSNTCNDPNGGTCLHYTQVVWKNTVRLGCAKVVCNSGG 142
Query: 156 LILTICFYDPPGNINGESPY 175
+T C YDPPGN GE P+
Sbjct: 143 TFIT-CNYDPPGNYIGEKPF 161
>At1g01310 pathogenesis related protein, like
Length = 294
Score = 106 bits (264), Expect = 7e-24
Identities = 59/146 (40%), Positives = 83/146 (56%), Gaps = 8/146 (5%)
Query: 32 QLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLG-CKLANFSHNKYG 90
+++ ++EFL AHN RA VG P +W LA A +A R+G C+L + S+ YG
Sbjct: 133 RVNRASREFLIAHNLVRARVGEPPFQWDGRLAAYARTWAN---QRVGDCRLVH-SNGPYG 188
Query: 91 SNQDLGGMG-WTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQL 149
N G W+P V W +E +FY + N+C P C HYTQ+VWR S +VGCA +
Sbjct: 189 ENIFWAGKNNWSPRDIVNVWADEDKFYDVKGNTC-EPQHMCGHYTQIVWRDSTKVGCASV 247
Query: 150 TCVVDKLILTICFYDPPGNINGESPY 175
C + + IC Y+PPGN GE+P+
Sbjct: 248 DC-SNGGVYAICVYNPPGNYEGENPF 272
>At4g33720 pathogenesis-related protein 1 precursor, 19.3K
Length = 163
Score = 105 bits (262), Expect = 1e-23
Identities = 69/165 (41%), Positives = 84/165 (50%), Gaps = 13/165 (7%)
Query: 11 LALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFA 70
LA+ FL+ DS PQ +FL HN ARAEVGV PL W E +A A
Sbjct: 12 LAITFFLVLIVHLKAQDS--PQ------DFLAVHNRARAEVGVGPLRWDEKVAAYA---R 60
Query: 71 RYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
Y R G S YG N T AV WV+E+ Y Y +N+C A C
Sbjct: 61 NYANQRKGDCAMKHSSGSYGENIAWSSGSMTGVAAVDMWVDEQFDYDYDSNTC-AWDKQC 119
Query: 131 WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S+++GCA++ C + +T C YDPPGN GE PY
Sbjct: 120 GHYTQVVWRNSERLGCAKVRCNNGQTFIT-CNYDPPGNWVGEWPY 163
>At4g25790 putative pathogenesis-related protein
Length = 210
Score = 103 bits (258), Expect = 4e-23
Identities = 55/146 (37%), Positives = 80/146 (54%), Gaps = 5/146 (3%)
Query: 31 PQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
P + ++FLD HN R +G+ PL W +A A +A +R C L + S YG
Sbjct: 69 PPTGSFEQQFLDPHNTVRGGLGLPPLVWDVKIASYATWWANQRR--YDCSLTH-STGPYG 125
Query: 91 SNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQL 149
N G G +T + AV+ W E + Y++ TN+C C HYTQ+VWR+++++GCA++
Sbjct: 126 ENLFWGSGSDFTSTFAVESWTVEAKSYNHMTNTCEGDGM-CGHYTQIVWRETRRLGCARV 184
Query: 150 TCVVDKLILTICFYDPPGNINGESPY 175
C + C YDPPGN GE PY
Sbjct: 185 VCENGAGVFITCNYDPPGNYVGEKPY 210
>At4g30320 PR-1-like protein
Length = 161
Score = 103 bits (257), Expect = 5e-23
Identities = 55/139 (39%), Positives = 79/139 (56%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
++F+ N ARA + ++PL+W LA+ A +A +R C L + S+ YG N G
Sbjct: 28 EQFMGPQNAARAHLRLKPLKWDAKLARYAQWWANQRRG--DCALTH-SNGPYGENLFWGS 84
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G W PS A W++E Y+YR+NSC S C HYTQ+VW+ ++++GCA + C
Sbjct: 85 GNRWGPSQAAYGWLSEARSYNYRSNSC--NSEMCGHYTQIVWKNTQKIGCAHVICNGGGG 142
Query: 157 ILTICFYDPPGNINGESPY 175
+ C YDPPGN G PY
Sbjct: 143 VFLTCNYDPPGNFLGRKPY 161
>At3g09590 putative pathogenesis-related protein
Length = 186
Score = 103 bits (256), Expect = 6e-23
Identities = 67/190 (35%), Positives = 93/190 (48%), Gaps = 19/190 (10%)
Query: 1 MSIFMQPFFSLALALFLLSAEAATVLDSVVPQLSAEA-------------KEFLDAHNWA 47
MS+ + S L LFLL + +VL + +P A +EFL AHN A
Sbjct: 1 MSVTLSHPSSCLLLLFLLFSGHPSVLGTSIPDAVNTAARLVNRARRAKLSREFLQAHNDA 60
Query: 48 RAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSN--QDLGGMGWTPSVA 105
R GV L W LA+ A +A+ ++ C + + S YG N W+P
Sbjct: 61 RVSSGVPTLGWDRDLARFADKWAKQRKS--DCSMIH-SGGPYGENIFWHRRKKTWSPEKV 117
Query: 106 VQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDP 165
V W E+ Y +TN+C AP C HYTQ+VWR++ VGCA++ C + L +C YDP
Sbjct: 118 VTRWFEERFNYDVKTNTC-APGKMCGHYTQMVWRETTAVGCARVKCHNGRGYLVVCEYDP 176
Query: 166 PGNINGESPY 175
GN GE P+
Sbjct: 177 RGNYEGERPF 186
>At5g26130 pathogenesis-related protein - like
Length = 164
Score = 100 bits (248), Expect = 5e-22
Identities = 55/140 (39%), Positives = 79/140 (56%), Gaps = 3/140 (2%)
Query: 36 EAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDL 95
+ +++LD HN AR +VGV P++W + A +A+ ++ +N S+ YG N
Sbjct: 28 QPQDYLDEHNRARTQVGVPPMKWHAGAEQYAWNYAQQRKGDCSLTHSN-SNGLYGENLAW 86
Query: 96 GGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDK 155
G + + AV+ WVNEK Y Y +N+C + C HYTQVVWR S+ VGCA++ C
Sbjct: 87 SGGALSGAEAVKLWVNEKSDYIYASNTC-SDGKQCGHYTQVVWRTSEWVGCAKVKCDNGG 145
Query: 156 LILTICFYDPPGNINGESPY 175
+T C Y PPGN G PY
Sbjct: 146 TFVT-CNYYPPGNYRGRWPY 164
>At4g33710 pathogenesis-related protein 1 precursor, 18.9K
Length = 166
Score = 99.4 bits (246), Expect = 9e-22
Identities = 62/176 (35%), Positives = 93/176 (52%), Gaps = 11/176 (6%)
Query: 1 MSIFMQPFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSE 60
M +F+ P + LAL L+ A A + + +++LD HN AR +V V ++W
Sbjct: 1 MKMFISPQTLVLLALALVLAFAVPL------KAQDRRQDYLDVHNHARDDVSVPHIKWHA 54
Query: 61 ALAKDAGLFARYQRDRLGCKLANF-SHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYR 119
A+ A +A QR + C+L + S +YG N + + AV+ WV EK Y ++
Sbjct: 55 GAARYAWNYA--QRRKRDCRLIHSNSRGRYGENLAWSSGDMSGAAAVRLWVREKSDYFHK 112
Query: 120 TNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
+N+C A C HYTQVVW+ S+ VGCA++ C +T C Y PGN+ G PY
Sbjct: 113 SNTCRAGK-QCGHYTQVVWKNSEWVGCAKVKCDNGGTFVT-CNYSHPGNVRGRRPY 166
>At4g33730 pathogenesis-related protein - like
Length = 172
Score = 99.0 bits (245), Expect = 1e-21
Identities = 60/165 (36%), Positives = 83/165 (49%), Gaps = 5/165 (3%)
Query: 11 LALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFA 70
L L +L + ++ S PQ +L HN ARA V V+PL W +A A +A
Sbjct: 13 LLLINYLTQIDVSSAQYSQYPQSHEYPDSYLRPHNAARAAVKVKPLRWDFGIATVAQDYA 72
Query: 71 RYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
+ C L + S YG N G + + AV WV+EK +Y + +NSC P+ C
Sbjct: 73 NHLASG-PCSLEH-SSGPYGENLAFGSGDMSAAQAVAMWVHEKSYYDFYSNSCHGPA--C 128
Query: 131 WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S ++GC + C + + +C YDP GN G PY
Sbjct: 129 GHYTQVVWRGSARLGCGKAKC-NNGASIVVCNYDPAGNYIGARPY 172
>At2g14610 pathogenesis-related PR-1-like protein
Length = 161
Score = 98.6 bits (244), Expect = 2e-21
Identities = 59/148 (39%), Positives = 83/148 (55%), Gaps = 8/148 (5%)
Query: 29 VVPQLSAEA-KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHN 87
V+P + ++ +++L HN AR VGV P++W E +A A +A R C+L + S
Sbjct: 21 VLPSKAQDSPQDYLRVHNQARGAVGVGPMQWDERVAAYARSYAEQLRGN--CRLIH-SGG 77
Query: 88 KYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCA 147
YG N G + AV WV+EK Y+Y N+C + C HYTQVVWRKS ++GCA
Sbjct: 78 PYGENLAWGSGDLSGVSAVNMWVSEKANYNYAANTC---NGVCGHYTQVVWRKSVRLGCA 134
Query: 148 QLTCVVDKLILTICFYDPPGNINGESPY 175
++ C I++ C YDP GN E PY
Sbjct: 135 KVRCNNGGTIIS-CNYDPRGNYVNEKPY 161
>At1g50060 branched-chain amino acid aminotransferase, putative
Length = 161
Score = 97.8 bits (242), Expect = 3e-21
Identities = 64/176 (36%), Positives = 92/176 (51%), Gaps = 16/176 (9%)
Query: 1 MSIFMQPFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSE 60
M+ F PF + FL+ A A PQ ++L++HN ARA+VGV + W
Sbjct: 1 MNTFKTPFLVIVAISFLVVATNA----QNTPQ------DYLNSHNTARAQVGVPNVVWDT 50
Query: 61 ALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSV-AVQDWVNEKEFYSYR 119
LA A ++ +++ C L + S+ YG N G ++ AV+ WV+EK +YSY
Sbjct: 51 TLAAYALNYSNFRK--ADCNLVH-SNGPYGENLAKGSSSSFSAISAVKLWVDEKPYYSYA 107
Query: 120 TNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
N+C C HYTQVVWR S ++GCA++ C + C Y+ PGN GE PY
Sbjct: 108 YNNCTGGK-QCLHYTQVVWRDSVKIGCARVQC-TNTWWFVSCNYNSPGNWVGEYPY 161
>At4g25780 putative pathogenesis-related protein
Length = 190
Score = 97.4 bits (241), Expect = 3e-21
Identities = 57/155 (36%), Positives = 81/155 (51%), Gaps = 8/155 (5%)
Query: 25 VLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANF 84
+ ++ P ++ +FL HN RA PL W L A +A +R C L +
Sbjct: 40 ICKNLCPGCDHDSLQFLFRHNLVRAARFEPPLIWDRRLQNYAQGWANQRRG--DCALRHS 97
Query: 85 SHN---KYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRK 140
N G N G G W+P+ AV W +EK FY Y +N+C A C HYTQ+VW+
Sbjct: 98 VSNGEFNLGENIYWGYGANWSPADAVVAWASEKRFYHYGSNTCDAGQM-CGHYTQIVWKS 156
Query: 141 SKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
+++VGCA++ C + +T C YDPPGN G+ PY
Sbjct: 157 TRRVGCARVVCDNGGIFMT-CNYDPPGNYIGQKPY 190
>At5g02730 pathogenesis related protein - like
Length = 205
Score = 95.5 bits (236), Expect = 1e-20
Identities = 53/142 (37%), Positives = 76/142 (53%), Gaps = 6/142 (4%)
Query: 36 EAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSN--Q 93
++ EFL AHN AR G L W + LA+ A +A+ ++ CK+ + S YG N +
Sbjct: 56 QSAEFLLAHNAARVASGASNLRWDQGLARFASKWAKQRKS--DCKMTH-SGGPYGENIFR 112
Query: 94 DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVV 153
W+P V W++E Y N+C + + C HYTQ+VWR + VGCA+ C
Sbjct: 113 YQRSENWSPRRVVDKWMDESLNYDRVANTCKSGAM-CGHYTQIVWRTTTAVGCARSKCDN 171
Query: 154 DKLILTICFYDPPGNINGESPY 175
++ L IC Y P GN GESP+
Sbjct: 172 NRGFLVICEYSPSGNYEGESPF 193
>At1g50050 pathogenesis-related protein 1b precursor (pr-1b),
putative
Length = 226
Score = 87.0 bits (214), Expect = 5e-18
Identities = 52/152 (34%), Positives = 76/152 (49%), Gaps = 14/152 (9%)
Query: 1 MSIFMQPFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSE 60
M+ F PF + FL+ A A +++L+ HN ARA+VGV + W
Sbjct: 1 MNTFKTPFLIIVAISFLVLATNA----------QNAQQDYLNTHNTARAQVGVANVVWDT 50
Query: 61 ALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGGMG-WTPSVAVQDWVNEKEFYSYR 119
+A A +A ++ + C L + YG N G +T AV WVNEK +Y+Y
Sbjct: 51 VVAAYATNYANARK--VDCSLTPSTGGSYGENLANGNNALFTGVAAVNLWVNEKPYYNYT 108
Query: 120 TNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTC 151
N+C+ C HYTQVVW S ++GCA++ C
Sbjct: 109 ANACIGAQ-QCKHYTQVVWSNSVKIGCARVLC 139
>At2g19970 putative pathogenesis-related protein
Length = 177
Score = 81.6 bits (200), Expect = 2e-16
Identities = 60/182 (32%), Positives = 83/182 (44%), Gaps = 12/182 (6%)
Query: 1 MSIFMQPFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSE 60
MS F F L L LL+ A L V P + ++ L HN RA VGV PL+W++
Sbjct: 1 MSSFRYTFVLLTLLSVLLTR--AYGLPRVRPIKDVQPRKTLKVHNQIRAAVGVAPLKWNK 58
Query: 61 ALAKDAGLFARYQRDRLGCKLANFSHN--KYGSNQDLGGM----GWTPSVAVQDWVNEKE 114
+A A FA Q C ++ H+ YG N G + + +A + W+ EK
Sbjct: 59 TVAAYAQKFANRQAKAGVCDYSSMRHSDGPYGENIAAGWVQPKDQMSGPIAAKYWLTEKP 118
Query: 115 FYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDP-PGNINGES 173
Y + TN C C HYTQ+V +S +GC C ++LI +C Y P P
Sbjct: 119 NYDHATNKC---KDVCGHYTQMVANQSLSLGCGSFRCHENELIYIVCNYYPMPVGDENTR 175
Query: 174 PY 175
PY
Sbjct: 176 PY 177
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,033,756
Number of Sequences: 26719
Number of extensions: 155138
Number of successful extensions: 348
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 271
Number of HSP's gapped (non-prelim): 26
length of query: 175
length of database: 11,318,596
effective HSP length: 93
effective length of query: 82
effective length of database: 8,833,729
effective search space: 724365778
effective search space used: 724365778
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0323a.10


