

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.5
(210 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
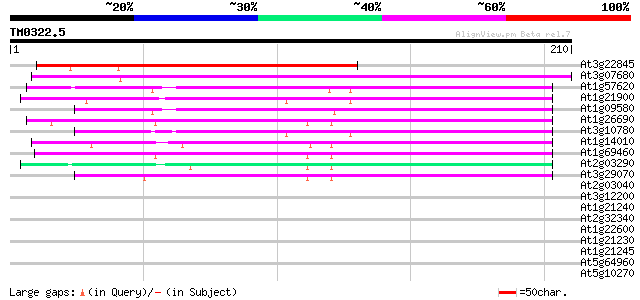
Score E
Sequences producing significant alignments: (bits) Value
At3g22845 unknown protein 187 3e-48
At3g07680 putative coated vesicle membrane protein 116 1e-26
At1g57620 unknown protein 82 3e-16
At1g21900 transmembrane protein, putative 75 2e-14
At1g09580 unknown protein 74 4e-14
At1g26690 unknown protein 63 1e-10
At3g10780 putative membrane protein 59 1e-09
At1g14010 transmembrane like protein 59 1e-09
At1g69460 unknown protein (At1g69460) 58 4e-09
At2g03290 putative Golgi-associated membrane trafficking protein 50 9e-07
At3g29070 transmembrane trafficking protein, putative 49 2e-06
At2g03040 unknown protein 40 0.001
At3g12200 putative protein 28 2.7
At1g21240 hypothetical protein 28 2.7
At2g32340 hypothetical protein 28 3.5
At1g22600 Similar to seed maturation protein PM27 28 4.6
At1g21230 hypothetical protein 28 4.6
At1g21245 wall-associated kinase 3 like protein 28 4.6
At5g64960 cdc2-like protein kinase 27 6.0
At5g10270 cdc2-like protein kinase 27 6.0
>At3g22845 unknown protein
Length = 135
Score = 187 bits (476), Expect = 3e-48
Identities = 86/125 (68%), Positives = 104/125 (82%), Gaps = 5/125 (4%)
Query: 11 VIFFLGTLIYN----VFSLSISLNDVECVSEHA-QSGDSVTGNFVVMDHDIFWSSDHPGI 65
V +G ++ N + SLS+++ND ECV E+ GD+V+GNFVV+DHDIFW SDHPG+
Sbjct: 10 VFVLIGLILLNSINQISSLSVTVNDEECVQEYVLYEGDTVSGNFVVVDHDIFWGSDHPGL 69
Query: 66 DFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGHIPSEH 125
DFTVT+P GN V +LKGTSGDKFEFKAP++G+YKFCFHNP STPETVSFYIHVGHIP+EH
Sbjct: 70 DFTVTSPAGNIVQTLKGTSGDKFEFKAPKSGMYKFCFHNPYSTPETVSFYIHVGHIPNEH 129
Query: 126 DLAKD 130
DLAKD
Sbjct: 130 DLAKD 134
>At3g07680 putative coated vesicle membrane protein
Length = 208
Score = 116 bits (290), Expect = 1e-26
Identities = 64/203 (31%), Positives = 101/203 (49%), Gaps = 1/203 (0%)
Query: 9 TLVIFFLGTLIYNVFSLSISLNDVECVSEHAQ-SGDSVTGNFVVMDHDIFWSSDHPGIDF 67
T+V+ L + ++ EC S A+ GD++ +FVV+ D W + G+D
Sbjct: 6 TIVLLGLLWSFQATLGIRFVIDREECFSHKAEYEGDTLHVSFVVIKSDSQWHFNEDGVDL 65
Query: 68 TVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGHIPSEHDL 127
+ P G +H + K +F + G+Y+FCF N ET+ F + +GH
Sbjct: 66 VIHGPTGEQIHDFREQISAKHDFVVQKKGVYRFCFTNKSPYHETIDFDVQLGHFAYYDQH 125
Query: 128 AKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLA 187
AKDEH P+ +I++L EAL +I EQ +L+A+ +R NE+ R + + E L
Sbjct: 126 AKDEHFTPLMEQISKLEEALYNIQFEQHWLEAQTDRQAIVNENMSKRAVHKALFESFALI 185
Query: 188 VTSILQVVYIRRLFSKSFAYNRV 210
S LQV +RRLF + +RV
Sbjct: 186 GASFLQVYLLRRLFERKLGMSRV 208
>At1g57620 unknown protein
Length = 212
Score = 81.6 bits (200), Expect = 3e-16
Identities = 55/204 (26%), Positives = 103/204 (49%), Gaps = 13/204 (6%)
Query: 7 ICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVM--DHDIFWSSDHPG 64
+ + +IF L + V+ L++ +CVSE QS V +++V+ +H IF P
Sbjct: 11 LLSALIFSLSPICEAVW-LTVPHTGSKCVSEEIQSNVIVLADYLVISEEHSIF-----PT 64
Query: 65 IDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHV----GH 120
+ VT P G +H + T+ +F F ++G Y CF + F I++ G
Sbjct: 65 VSVKVTAPYGTVLHHRENTTNGQFAFTTQESGTYLACFEADAKSHGNKDFSINIDWKTGI 124
Query: 121 IPSEHD-LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYT 179
+ D +A+ E ++ + ++ +L A+E+I YL+ R+ R +E T +RV +Y+
Sbjct: 125 AAKDWDSIARKEKIEGVELEFKKLEGAVEAIHENLIYLRNREAEMRIVSEKTNSRVAWYS 184
Query: 180 VLEYILLAVTSILQVVYIRRLFSK 203
++ + V S LQ++Y+++ F K
Sbjct: 185 IMSLGICIVVSGLQILYLKQYFEK 208
>At1g21900 transmembrane protein, putative
Length = 216
Score = 75.5 bits (184), Expect = 2e-14
Identities = 54/205 (26%), Positives = 99/205 (47%), Gaps = 8/205 (3%)
Query: 5 SKICTLVIFFLGTLIYNVFSLSI-SLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHP 63
S T+V+FFL L+I + +CVSE QS V ++ V+D + P
Sbjct: 10 SLFLTVVLFFLTVNYGEAIWLTIPTTGGTKCVSEEIQSNVVVLADYYVVDEHN--PENTP 67
Query: 64 GIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCF----HNPMSTPETVSFYIHVG 119
+ VT+P GN +H + + +F F + G Y CF + ++ P T+ +G
Sbjct: 68 AVSSKVTSPYGNNLHHQENVTHGQFAFTTQEAGNYLACFWIDSSHHLANPITLGVDWKMG 127
Query: 120 HIPSEHD-LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFY 178
+ D +AK E ++ + +++ L + SI Y+K R+ R +E+T +RV ++
Sbjct: 128 IAAKDWDSVAKKEKIEGVELQLRRLEGLVLSIRENLNYIKDREAEMREVSETTNSRVAWF 187
Query: 179 TVLEYILLAVTSILQVVYIRRLFSK 203
+++ + V Q++Y++R F K
Sbjct: 188 SIMSLGVCVVVVGSQILYLKRYFHK 212
>At1g09580 unknown protein
Length = 217
Score = 74.3 bits (181), Expect = 4e-14
Identities = 48/186 (25%), Positives = 92/186 (48%), Gaps = 12/186 (6%)
Query: 25 LSISLNDVECVSEHAQSGDSVTGNFVVM--DHDIFWSSDHPGIDFTVTNPGGNAVHSLKG 82
L + +CVSE QS V +++++ DH++ P I VT+P GN +H+++
Sbjct: 33 LDVPPTGTKCVSEEIQSNVVVLADYLIISEDHEVM-----PTISVKVTSPYGNNLHNMEN 87
Query: 83 TSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGH-----IPSEHDLAKDEHLDPIN 137
+ +F F ++G Y CF + + I++ +AK E ++ +
Sbjct: 88 VTHGQFAFTTQESGNYLACFWADEKSHGNKNVSINIDWRTGIAAKDWASIAKKEKIEGVE 147
Query: 138 VKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYI 197
++I +L A+E+I YL+ R+ R +E T +RV +Y+++ + S QV+Y+
Sbjct: 148 LEIRKLEGAVEAIHENILYLRNREADMRTMSEKTNSRVAWYSIMSLGVCIAVSGFQVLYL 207
Query: 198 RRLFSK 203
++ F K
Sbjct: 208 KQYFEK 213
>At1g26690 unknown protein
Length = 214
Score = 63.2 bits (152), Expect = 1e-10
Identities = 44/203 (21%), Positives = 89/203 (43%), Gaps = 6/203 (2%)
Query: 7 ICTLVIFF-LGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMD-HDIFWSSDHPG 64
+CT+++F + + + + +C+SE +S G + V++ ++ S
Sbjct: 8 LCTILLFLAISSQVSQSLHFELQSGRTKCISEDIKSNSMTVGKYTVVNPNEAHPSPQSHK 67
Query: 65 IDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE---TVSFYIHVG-H 120
I VT+ GN H + +F F A ++G Y C+ PE ++ F G
Sbjct: 68 ISIRVTSSYGNTYHHAEDVESGQFAFTAVESGDYMACYTAVDHKPEVTLSIDFDWRTGVQ 127
Query: 121 IPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTV 180
S +AK ++ + + L E + SI E YL+ R+ + N +T +++ + +
Sbjct: 128 SKSWSSVAKKSQVEVMEFDVKRLIETVNSIHEEMFYLREREEEMQNLNRATNSKMAWLSF 187
Query: 181 LEYILLAVTSILQVVYIRRLFSK 203
L + + +Q V+++ F K
Sbjct: 188 LSLFVCLGVAGMQFVHLKTFFEK 210
>At3g10780 putative membrane protein
Length = 217
Score = 59.3 bits (142), Expect = 1e-09
Identities = 48/187 (25%), Positives = 87/187 (45%), Gaps = 10/187 (5%)
Query: 25 LSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTS 84
L++ + CV E Q+ V +++ +D D F P +D VT+P G ++ + +
Sbjct: 29 LTVPESGERCVYEEIQANVVVVLDYICID-DAFTQLG-PTLDVRVTSPYGKELYKIANVT 86
Query: 85 GDKFEFKAPQNGIYKFCF-------HNPMSTPETVSFYIHVGHIPSEHD-LAKDEHLDPI 136
+ F ++G + C H+ +++ VS +G + D +AK E ++ +
Sbjct: 87 HGQAAFTTSESGTFLACLAMHHDQSHHSVNSSVIVSLDWKMGIRAKDWDSVAKKEKIEAM 146
Query: 137 NVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVY 196
++I E +I YL+ R+ R NE T TRV ++ + V SI QV+Y
Sbjct: 147 ELEIRRSTEYASAIRANILYLRIREAYMREINEKTNTRVNQLGLMSLGVAIVVSISQVLY 206
Query: 197 IRRLFSK 203
++R F K
Sbjct: 207 LKRYFLK 213
>At1g14010 transmembrane like protein
Length = 212
Score = 59.3 bits (142), Expect = 1e-09
Identities = 47/206 (22%), Positives = 86/206 (40%), Gaps = 15/206 (7%)
Query: 9 TLVIFFLGTLIYNVFSLSISL--NDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHP--- 63
++V+ L L S+ L +C+SE + G + +++ DHP
Sbjct: 7 SIVLLILSILSPVTLSIRYELLSGHTKCISEEIHANAMTIGKYSIINPH----EDHPLPS 62
Query: 64 --GIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPET---VSFYIHV 118
+ VT+P G A H G +F F A + G Y CF PET + F
Sbjct: 63 SHKVTVRVTSPQGTAYHESDGVESGQFSFVAVETGDYISCFSAVDHKPETTLIIDFDWRT 122
Query: 119 G-HIPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIF 177
G H ++AK ++ + ++ +L E + I E YL+ R+ N +T +++ +
Sbjct: 123 GIHTKDWSNVAKKSQVETMEFEVKKLFETVNGIHDEMFYLRDREEEMHNLNIATNSKMAW 182
Query: 178 YTVLEYILLAVTSILQVVYIRRLFSK 203
+ + + + LQ +++ F K
Sbjct: 183 LSFVSLAVCLSVAGLQFWHLKTFFQK 208
>At1g69460 unknown protein (At1g69460)
Length = 214
Score = 57.8 bits (138), Expect = 4e-09
Identities = 44/199 (22%), Positives = 87/199 (43%), Gaps = 5/199 (2%)
Query: 10 LVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMD-HDIFWSSDHPGIDFT 68
L+I + + I + + +C++E +S G + + + H+ I
Sbjct: 12 LLILAIWSPISHSLHFDLHSGRTKCIAEDIKSNSMTVGKYNIDNPHEGQALPQTHKISVK 71
Query: 69 VTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE---TVSFYIHVG-HIPSE 124
VT+ GN H + +F F A + G Y CF PE ++ F G S
Sbjct: 72 VTSNSGNNYHHAEQVDSGQFAFSAVEAGDYMACFTAVDHKPEVSLSIDFEWKTGVQSKSW 131
Query: 125 HDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYI 184
++AK ++ + ++ L + + SI E YL+ R+ + N ST T++ + +VL +
Sbjct: 132 ANVAKKSQVEVMEFEVKSLLDTVNSIHEEMYYLRDREEEMQDLNRSTNTKMAWLSVLSFF 191
Query: 185 LLAVTSILQVVYIRRLFSK 203
+ + +Q ++++ F K
Sbjct: 192 VCIGVAGMQFLHLKTFFEK 210
>At2g03290 putative Golgi-associated membrane trafficking protein
Length = 213
Score = 50.1 bits (118), Expect = 9e-07
Identities = 45/208 (21%), Positives = 84/208 (39%), Gaps = 13/208 (6%)
Query: 5 SKICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPG 64
S I L+I L ++ + + +C+ E G + +++ + +HP
Sbjct: 6 SSILLLIIALLSPRTLSM-RYELKSSKTKCIGEEIHENAMSIGKYFIVNPN---EDNHPL 61
Query: 65 ID-----FTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE---TVSFYI 116
D V P G +H +F F A +NG Y C PE T+ F
Sbjct: 62 PDSHKIIVKVMPPQGKNLHEADKVEAGQFSFTAYENGSYVACITAIDYKPETTLTIDFDW 121
Query: 117 HVG-HIPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRV 175
G H ++AK +D + ++ L + + SI E YL+ R+ + N ST +++
Sbjct: 122 KTGVHSKEWTNVAKKSQVDMMEYQVKTLMDTVISIHEEMYYLREREEEMQELNRSTNSKM 181
Query: 176 IFYTVLEYILLAVTSILQVVYIRRLFSK 203
+ + ++ + LQ +++ F K
Sbjct: 182 AWLSFGSLVVCLSVAGLQFWHLKTFFEK 209
>At3g29070 transmembrane trafficking protein, putative
Length = 204
Score = 48.9 bits (115), Expect = 2e-06
Identities = 39/184 (21%), Positives = 79/184 (42%), Gaps = 5/184 (2%)
Query: 25 LSISLNDVECVSEHAQSGDSVTGNF-VVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGT 83
L + + +C+S+ ++ G + +V ++ + TV++P G + H +
Sbjct: 17 LDMESGNTKCISDDIKTNYMTVGTYSIVNPNEGHHLPPSHKLFVTVSSPKGKSHHHAENV 76
Query: 84 SGDKFEFKAPQNGIYKFCFHNPMSTPE---TVSFYIHVG-HIPSEHDLAKDEHLDPINVK 139
KF F A + G Y CF P P V F G +AK + + V+
Sbjct: 77 ESGKFVFTAEETGDYMTCFVAPGYRPTAKFAVDFEWKSGVEAKDWTTIAKRGQITMLEVE 136
Query: 140 IAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRR 199
+ +L + E+I E L R+ + N ST +R+ ++L +++ + LQ+ +++
Sbjct: 137 VRKLLDVTETIHEEMFQLIEREREMQELNRSTNSRMAALSLLSFVVTMSVAGLQLRHLKS 196
Query: 200 LFSK 203
+
Sbjct: 197 FLER 200
>At2g03040 unknown protein
Length = 166
Score = 39.7 bits (91), Expect = 0.001
Identities = 38/162 (23%), Positives = 62/162 (37%), Gaps = 7/162 (4%)
Query: 5 SKICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMD--HDIFWSSDH 62
S I L+I L ++ + + +C+ E G + +++ D D
Sbjct: 6 SSILLLIIALLSPRTLSM-RYELKSSKTKCIGEEIHENAMSIGKYFIVNPNEDHHPLPDS 64
Query: 63 PGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE---TVSFYIHVG 119
I V P G +H +F F A +NG Y C PE T+ F G
Sbjct: 65 HKIIVKVMPPQGKNLHEADNVEAGQFSFTAYENGSYVACITAIDYKPETTLTIDFDWKTG 124
Query: 120 -HIPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKAR 160
H ++AK +D + ++ L + + SI E YL+ R
Sbjct: 125 VHSKEWTNVAKKSQVDMMEYQVKTLMDTVISIHEEMYYLRER 166
>At3g12200 putative protein
Length = 571
Score = 28.5 bits (62), Expect = 2.7
Identities = 25/101 (24%), Positives = 45/101 (43%), Gaps = 5/101 (4%)
Query: 44 SVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFH 103
S+ ++V D + +D+ FT GGN +++K G F P+ I+K+
Sbjct: 72 SLKNPYIVHYEDSWIDNDNNACIFTAYYEGGNMANAIKKARGKLF----PEERIFKWLAQ 127
Query: 104 NPMSTPETVS-FYIHVGHIPSEHDLAKDEHLDPINVKIAEL 143
++ S +H+ S L KD+H+ N +A+L
Sbjct: 128 LLLAVNYLHSNRVVHMDLTCSNIFLPKDDHVQLGNYGLAKL 168
>At1g21240 hypothetical protein
Length = 741
Score = 28.5 bits (62), Expect = 2.7
Identities = 15/90 (16%), Positives = 39/90 (42%), Gaps = 15/90 (16%)
Query: 95 NGIYKFCFHNPMSTPETVSFYIHVGHIPSEHDLAKDEHLDPINVK--------------- 139
+G CF P ++ VS+++ H++ D+ L+ N+K
Sbjct: 611 SGQKALCFERPQASKHLVSYFVSATEENRLHEIIDDQVLNEDNLKEIQEAARIAAECTRL 670
Query: 140 IAELREALESIITEQKYLKARDNRHRYTNE 169
+ E R ++ + + + L+ +H+++++
Sbjct: 671 MGEERPRMKEVAAKLEALRVEKTKHKWSDQ 700
>At2g32340 hypothetical protein
Length = 302
Score = 28.1 bits (61), Expect = 3.5
Identities = 15/42 (35%), Positives = 22/42 (51%)
Query: 20 YNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSD 61
Y FSL+ SL +V+ + DS G+ ++ DH I S D
Sbjct: 9 YGDFSLTNSLVNVDKEDLRREEADSDAGSVIICDHSICDSGD 50
>At1g22600 Similar to seed maturation protein PM27
Length = 385
Score = 27.7 bits (60), Expect = 4.6
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 1/68 (1%)
Query: 129 KDEHLDPINVKIAELREALESIITEQKYL-KARDNRHRYTNESTKTRVIFYTVLEYILLA 187
KD ++ + E+ A E++ E+K KAR+ R + +TK VLE + +A
Sbjct: 116 KDRTASDLSDETPEMTVAREALEVEEKVSWKAREARGKVNERATKKAHRVQKVLEKVQIA 175
Query: 188 VTSILQVV 195
V I VV
Sbjct: 176 VRGIGTVV 183
>At1g21230 hypothetical protein
Length = 733
Score = 27.7 bits (60), Expect = 4.6
Identities = 17/93 (18%), Positives = 43/93 (45%), Gaps = 15/93 (16%)
Query: 95 NGIYKFCFHNPMSTPETVSFYIHV-----------GHIPSEHDLAKDEHLDPINVK---- 139
+G CF P S+ VS+++ G + +E++ + + I V+
Sbjct: 604 SGEKALCFERPQSSKHLVSYFVSAMKENRLHEIIDGQVMNEYNQREIQESARIAVECTRI 663
Query: 140 IAELREALESIITEQKYLKARDNRHRYTNESTK 172
+ E R +++ + E + L+ + +H+++++ K
Sbjct: 664 MGEERPSMKEVAAELEALRVKTTKHQWSDQYPK 696
>At1g21245 wall-associated kinase 3 like protein
Length = 166
Score = 27.7 bits (60), Expect = 4.6
Identities = 14/52 (26%), Positives = 27/52 (51%), Gaps = 2/52 (3%)
Query: 95 NGIYKFCFHNPMSTPETVSFYIHVGHIPSEHDLAKDEHLDPINVKIAELREA 146
+G CF P ++ VS+++ H++ D+ L+ N+K E++EA
Sbjct: 37 SGQKALCFERPQASKHLVSYFVSATEENRLHEIIDDQVLNEDNLK--EIQEA 86
>At5g64960 cdc2-like protein kinase
Length = 513
Score = 27.3 bits (59), Expect = 6.0
Identities = 11/21 (52%), Positives = 14/21 (66%)
Query: 49 FVVMDHDIFWSSDHPGIDFTV 69
F MDHD+ +D PG+ FTV
Sbjct: 118 FEYMDHDLTGLADRPGLRFTV 138
>At5g10270 cdc2-like protein kinase
Length = 505
Score = 27.3 bits (59), Expect = 6.0
Identities = 11/21 (52%), Positives = 14/21 (66%)
Query: 49 FVVMDHDIFWSSDHPGIDFTV 69
F MDHD+ +D PG+ FTV
Sbjct: 118 FEYMDHDLTGLADRPGLRFTV 138
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,763,701
Number of Sequences: 26719
Number of extensions: 199249
Number of successful extensions: 468
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 450
Number of HSP's gapped (non-prelim): 22
length of query: 210
length of database: 11,318,596
effective HSP length: 95
effective length of query: 115
effective length of database: 8,780,291
effective search space: 1009733465
effective search space used: 1009733465
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0322.5


