

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.2
(856 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
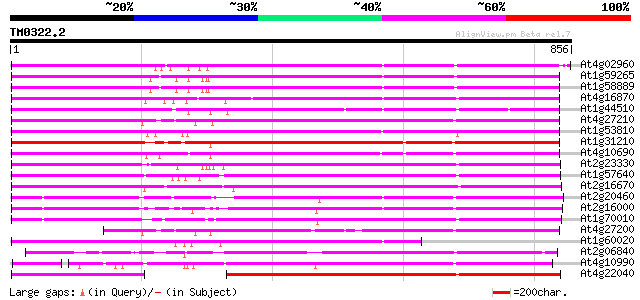
Score E
Sequences producing significant alignments: (bits) Value
At4g02960 putative polyprotein of LTR transposon 752 0.0
At1g59265 polyprotein, putative 746 0.0
At1g58889 polyprotein, putative 745 0.0
At4g16870 retrotransposon like protein 715 0.0
At1g44510 polyprotein, putative 714 0.0
At4g27210 putative protein 681 0.0
At1g53810 679 0.0
At1g31210 putative reverse transcriptase 675 0.0
At4g10690 retrotransposon like protein 656 0.0
At2g23330 putative retroelement pol polyprotein 590 e-168
At1g57640 578 e-165
At2g16670 putative retroelement pol polyprotein 572 e-163
At2g20460 putative retroelement pol polyprotein 571 e-163
At2g16000 putative retroelement pol polyprotein 567 e-162
At1g70010 hypothetical protein 545 e-155
At4g27200 putative protein 506 e-143
At1g60020 hypothetical protein 488 e-138
At2g06840 putative retroelement pol polyprotein 466 e-131
At4g10990 putative retrotransposon polyprotein 434 e-121
At4g22040 LTR retrotransposon like protein 431 e-121
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 752 bits (1941), Expect = 0.0
Identities = 430/935 (45%), Positives = 565/935 (59%), Gaps = 94/935 (10%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ + +D GGEF L Y + GI H
Sbjct: 531 FTRYTWLYPLKQKSQVKDTFIIFKSLVENRFQTRIGTLYSDNGGEFVVLRDYLSQHGISH 590
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVEMGL LL+HA + YW +AF VYLINRLPT +L
Sbjct: 591 FTSPPHTPEHNGLSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQL 650
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
+SPF KLF + + L++FGCACYP+LRPYN HKL SK+C F+GYS T +Y CL
Sbjct: 651 QSPFQKLFGQPPNYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHI 710
Query: 183 PTGRLYISKDVLFNEFRFPYKEL-FGCASSPQLSSVGSV----HSSIPLFSF-------- 229
PTGRLY S+ V F+E FP+ FG ++S + S + H+++P
Sbjct: 711 PTGRLYTSRHVQFDERCFPFSTTNFGVSTSQEQRSDSAPNWPSHTTLPTTPLVLPAPPCL 770
Query: 230 -PQLTTLSSPAPHSPN----TTLPESSSPASQSSTPILPEPTSPV---PQ---------- 271
P L T P P SP+ T + S+ P+S S+P EPT+P PQ
Sbjct: 771 GPHLDTSPRP-PSSPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQN 829
Query: 272 -------------SSPAPISPALSSPAPQS------------------SSTDSSSSVSPI 300
+SP+P SP +SP PQS S + SS+S P+
Sbjct: 830 SNSNSPILNNPNPNSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTPPL 889
Query: 301 -----------VN---PHNTHSMVTRAKDGIVKPRL---VPTLLLSQAEPTSAKLALKDP 343
VN P NTHSM TRAKDGI KP T L + +EP +A A+KD
Sbjct: 890 PPVLPAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSLAANSEPRTAIQAMKDD 949
Query: 344 KWYSAMQDEFNALHSNQTWTLVPLPSHRKAI-GCKWVFRIKENADGSVNRYKARLVAKGY 402
+W AM E NA N TW LVP P I GC+W+F K N+DGS+NRYKARLVAKGY
Sbjct: 950 RWRQAMGSEINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGY 1009
Query: 403 SQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPG 462
+Q G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L +EVYM QPPG
Sbjct: 1010 NQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDEVYMSQPPG 1069
Query: 463 FEAKDR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVY 521
F KDR VCRL KA+YGLKQAPRAW+ L+ LL GF S + SLF L +Y
Sbjct: 1070 FVDKDRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSISDTSLFVLQRGRSIIY 1129
Query: 522 VLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKY 581
+LVYVDDI+ITGN +L+H + L+ FS+K+ L YFLG++ + + + L L+Q +Y
Sbjct: 1130 MLVYVDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEAKRV-PQGLHLSQRRY 1188
Query: 582 IKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYS 641
DLL R +M AK ++TPM ++ KLT H + DP+ YR IVG+LQY+ TRP++SY+
Sbjct: 1189 TLDLLARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGSLQYLAFTRPDLSYA 1248
Query: 642 VNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPD 701
VN++ Q++ P ++HW A+KR+LRYL GT D G+ LK LSL A+ DADWA D D
Sbjct: 1249 VNRLSQYMHMPTDDHWNALKRVLRYLAGTPDHGIFLKK---GNTLSLHAYSDADWAGDTD 1305
Query: 702 DHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP- 760
D+ ST+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +EL W+ SLL EL I
Sbjct: 1306 DYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWICSLLTELGIQL 1365
Query: 761 HKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDIL 820
P+I CDN+ L NPV H+R KH+ LD F+R +V + +L + HV + DQ D L
Sbjct: 1366 SHPPVIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVHVSTHDQLADTL 1425
Query: 821 TKALSPLRFEDLRTKLNVTDKFTLAQPP*VCGGIL 855
TK LS + F++ K+ V + PP CGG+L
Sbjct: 1426 TKPLSRVAFQNFSRKIGV-----IKVPP-SCGGVL 1454
>At1g59265 polyprotein, putative
Length = 1466
Score = 746 bits (1927), Expect = 0.0
Identities = 430/915 (46%), Positives = 555/915 (59%), Gaps = 84/915 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ +D GGEF L YF+ GI H
Sbjct: 552 FTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISH 611
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL+HA I YW +AF VYLINRLPT +L
Sbjct: 612 LTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQL 671
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESPF KLF S + LR+FGCACYP+LRPYN HKL S++CVFLGYS T +Y CL
Sbjct: 672 ESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHL 731
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQ-----LSSVGSVHSSIPLFSFPQLTTLSS 237
T RLYIS+ V F+E FP+ S Q S V S H+++P + P L S
Sbjct: 732 QTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRT-PVLPAPSC 790
Query: 238 PAPHSPNTTLPESSSP-------------ASQSSTPILPEPTSP---VPQ---------- 271
PH T S+P + SS P PEPT+P PQ
Sbjct: 791 SDPHHAATPPSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQT 850
Query: 272 ---------------SSPAPISPALSSPAPQSSSTD------SSSSVSP----------- 299
SP+ ++ +LS+PA SSS+ SSSS SP
Sbjct: 851 QTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPP 910
Query: 300 ----IVN-----PHNTHSMVTRAKDGIVKPRLVPTL---LLSQAEPTSAKLALKDPKWYS 347
IVN P NTHSM TRAK GI+KP +L L +++EP +A ALKD +W +
Sbjct: 911 PLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRN 970
Query: 348 AMQDEFNALHSNQTWTLV-PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVE 406
AM E NA N TW LV P PSH +GC+W+F K N+DGS+NRYKARLVAKGY+Q
Sbjct: 971 AMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRP 1030
Query: 407 GFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK 466
G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L ++VYM QPPGF K
Sbjct: 1031 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDK 1090
Query: 467 DR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVY 525
DR + VC+L KALYGLKQAPRAW+ L+ LL GF S + SLF L VY+LVY
Sbjct: 1091 DRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVY 1150
Query: 526 VDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDL 585
VDDI+ITGN +L + + L+ FS+K L YFLG++ + + L L+Q +YI DL
Sbjct: 1151 VDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRV-PTGLHLSQRRYILDL 1209
Query: 586 LERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKV 645
L R +M AK ++TPM + KL+ + + DP+ YR IVG+LQY+ TRP+ISY+VN++
Sbjct: 1210 LARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRL 1269
Query: 646 CQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRS 705
QF+ P EEH A+KRILRYL GT + G+ LK LSL A+ DADWA D DD+ S
Sbjct: 1270 SQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKK---GNTLSLHAYSDADWAGDKDDYVS 1326
Query: 706 TSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTP 764
T+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +E+ W+ SLL EL I + P
Sbjct: 1327 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPP 1386
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
+I CDN+ L NPV H+R KH+ +D F+R +V + +L + HV + DQ D LTK L
Sbjct: 1387 VIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1446
Query: 825 SPLRFEDLRTKLNVT 839
S F++ +K+ VT
Sbjct: 1447 SRTAFQNFASKIGVT 1461
>At1g58889 polyprotein, putative
Length = 1466
Score = 745 bits (1923), Expect = 0.0
Identities = 429/915 (46%), Positives = 554/915 (59%), Gaps = 84/915 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PLK+KS TF+ F ++E + ++ +D GGEF L YF+ GI H
Sbjct: 552 FTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISH 611
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL+HA I YW +AF VYLINRLPT +L
Sbjct: 612 LTSPPHTPEHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQL 671
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
ESPF KLF S + LR+FGCACYP+LRPYN HKL S++CVFLGYS T +Y CL
Sbjct: 672 ESPFQKLFGTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHL 731
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQ-----LSSVGSVHSSIPLFSFPQLTTLSS 237
T RLYIS+ V F+E FP+ S Q S V S H+++P + P L S
Sbjct: 732 QTSRLYISRHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRT-PVLPAPSC 790
Query: 238 PAPHSPNTTLPESSSP-------------ASQSSTPILPEPTSP---VPQ---------- 271
PH T S+P + SS P PEPT+P PQ
Sbjct: 791 SDPHHAATPPSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQT 850
Query: 272 ---------------SSPAPISPALSSPAPQSSSTD------SSSSVSP----------- 299
SP+ ++ +LS+PA SSS+ SSSS SP
Sbjct: 851 QTHSSQNTSQNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPP 910
Query: 300 ----IVN-----PHNTHSMVTRAKDGIVKPRLVPTL---LLSQAEPTSAKLALKDPKWYS 347
IVN P NTHSM TRAK GI+KP +L L +++EP +A ALKD +W +
Sbjct: 911 PLAQIVNNNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRN 970
Query: 348 AMQDEFNALHSNQTWTLV-PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVE 406
AM E NA N TW LV P PSH +GC+W+F K N+DGS+NRYKAR VAKGY+Q
Sbjct: 971 AMGSEINAQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRP 1030
Query: 407 GFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK 466
G DY ETFSPV+K ++RI+L +AV + W I QLDVNNAFL G L ++VYM QPPGF K
Sbjct: 1031 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDK 1090
Query: 467 DR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVY 525
DR + VC+L KALYGLKQAPRAW+ L+ LL GF S + SLF L VY+LVY
Sbjct: 1091 DRPNYVCKLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVY 1150
Query: 526 VDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDL 585
VDDI+ITGN +L + + L+ FS+K L YFLG++ + + L L+Q +YI DL
Sbjct: 1151 VDDILITGNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRV-PTGLHLSQRRYILDL 1209
Query: 586 LERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKV 645
L R +M AK ++TPM + KL+ + + DP+ YR IVG+LQY+ TRP+ISY+VN++
Sbjct: 1210 LARTNMITAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRL 1269
Query: 646 CQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRS 705
QF+ P EEH A+KRILRYL GT + G+ LK LSL A+ DADWA D DD+ S
Sbjct: 1270 SQFMHMPTEEHLQALKRILRYLAGTPNHGIFLKK---GNTLSLHAYSDADWAGDKDDYVS 1326
Query: 706 TSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTP 764
T+G +V+LG + +SW SKKQ V RSSTEAEYR++ANT +E+ W+ SLL EL I + P
Sbjct: 1327 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPP 1386
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
+I CDN+ L NPV H+R KH+ +D F+R +V + +L + HV + DQ D LTK L
Sbjct: 1387 VIYCDNVGATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1446
Query: 825 SPLRFEDLRTKLNVT 839
S F++ +K+ VT
Sbjct: 1447 SRTAFQNFASKIGVT 1461
>At4g16870 retrotransposon like protein
Length = 1474
Score = 715 bits (1846), Expect = 0.0
Identities = 408/922 (44%), Positives = 536/922 (57%), Gaps = 93/922 (10%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
+TRYTW++PL++KS +TF+ F ++E + K++ + +D GGEF L + + GI H
Sbjct: 553 HTRYTWLYPLQQKSQVKSTFIAFKALVENRFQAKIRTLYSDNGGEFIALREFLVSNGISH 612
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL A + EYW +AF VYLINR+PT VL
Sbjct: 613 LTSPPHTPEHNGLSERKHRHIVETGLTLLTQASVPREYWPYAFAAAVYLINRMPTPVLSM 672
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLDP 183
ESPF KLF + + LR+FGC C+P+LRPY +KL S+ CVFLGYS T +Y C D
Sbjct: 673 ESPFQKLFGSKPNYERLRVFGCLCFPWLRPYTHNKLEERSRRCVFLGYSLTQTAYLCFDV 732
Query: 184 TG-RLYISKDVLFNEFRFPYKEL----------FGCASSPQLSSVGSVHSSIPLFSFPQL 232
RLY S+ V+F+E FP+ L F +SSP ++ + S S +P
Sbjct: 733 EHKRLYTSRHVVFDEASFPFSNLTSQNSLPTVTFEQSSSPLVTPILSSSSVLPSCLSSPC 792
Query: 233 TTL-------------------SSPAPHSPN--TTLP------ESSSPASQSSTPILPEP 265
T L +SPAP SP+ TT+ SSSP SS+ + EP
Sbjct: 793 TVLHQQQPPVTTPNSPHSSQPTTSPAPLSPHRSTTMDFQVPQVRSSSPLLSSSSSLNSEP 852
Query: 266 TSP------------------------------------------VPQSSPAPISPALSS 283
T+P P P P+ P ++
Sbjct: 853 TAPNENGPEPEAQSPPIGPLSNPTHEAFIGPLPNPNRNPTNEIEPTPAPHPKPVKPTTTT 912
Query: 284 PAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLL-----SQAEPTSAKL 338
P + +T S +S P N H+M TRAK+ I KP +L S +EPT+
Sbjct: 913 TTP-NRTTVSDASHQPTAPQQNQHNMKTRAKNNIKKPNTKFSLTATLPNRSPSEPTNVTQ 971
Query: 339 ALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLV 398
ALKD KW AM DEF+A N TW LVP S + +GCKWVF++K +G++++YKARLV
Sbjct: 972 ALKDKKWRFAMSDEFDAQQRNHTWDLVPHES-QLLVGCKWVFKLKYLPNGAIDKYKARLV 1030
Query: 399 AKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMV 458
AKG++Q G DY ETFSPV+K T+R++L +AV K W I QLDVNNAFL G L EEVYM
Sbjct: 1031 AKGFNQQYGVDYAETFSPVIKSTTIRLVLDVAVKKDWEIKQLDVNNAFLQGTLTEEVYMA 1090
Query: 459 QPPGFEAKDR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSG 517
QPPGF KDR + VCRL KA+YGLKQAPRAW+ LK L GF S + SLF
Sbjct: 1091 QPPGFIDKDRPTHVCRLRKAIYGLKQAPRAWYMELKQHLFNIGFVNSLSDASLFIYCHGT 1150
Query: 518 CSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLT 577
VYVLVYVDDII+TG+ + ++ L FS+K L YFLG++ + L L
Sbjct: 1151 TFVYVLVYVDDIIVTGSDKSSIDAVLTSLAERFSIKDPTDLHYFLGIEATRT-KQGLHLM 1209
Query: 578 QSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPE 637
Q KYIKDLL + +MA AK + TP+ ++ KLT HG + D S YRS+VG+LQY+ TRP+
Sbjct: 1210 QRKYIKDLLAKHNMADAKPVLTPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLAFTRPD 1269
Query: 638 ISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWA 697
I+Y+VN++ Q + QP E+HW A KR+LRYL GT G+ L S PL+L AF DADWA
Sbjct: 1270 IAYAVNRLSQLMPQPTEDHWQAAKRVLRYLAGTSTHGIFLDTTS---PLNLHAFSDADWA 1326
Query: 698 ADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQEL 757
D DD+ ST+ +++LG N +SW SKKQ VARSSTE+EYR +AN +E+ W+ SLL +L
Sbjct: 1327 GDSDDYVSTNAYVIYLGKNPISWSSKKQRGVARSSTESEYRAVANAASEVKWLCSLLSKL 1386
Query: 758 HIPHK-TPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQR 816
HI P I CDN+ L NPV H+R KH+ +D FVR + + +L + HV + DQ
Sbjct: 1387 HIRLPIRPSIFCDNIGATYLCANPVFHSRMKHIAIDYHFVRNMIQSGALRVSHVSTRDQL 1446
Query: 817 DDILTKALSPLRFEDLRTKLNV 838
D LTK LS F+ R K+ V
Sbjct: 1447 ADALTKPLSRAHFQSARFKIGV 1468
>At1g44510 polyprotein, putative
Length = 1459
Score = 714 bits (1843), Expect = 0.0
Identities = 401/906 (44%), Positives = 538/906 (59%), Gaps = 81/906 (8%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
YTRYTW++PL++KS TF+ F ++E + K++ + +D GGEF L + + GI H
Sbjct: 558 YTRYTWLYPLQQKSQVKATFIAFKALVENRFQAKIRTLYSDNGGEFIALRDFLVSNGISH 617
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ PHT NG+ ERKHRHIVE GL LL A + EYW +AF T VYLINR+PT VL
Sbjct: 618 LTSPPHTPEHNGLSERKHRHIVETGLTLLTQASVPREYWTYAFATAVYLINRMPTPVLCL 677
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
+SPF KLF S + + LR+FGC C+P+LRPY +KL SK CVFLGYS T +Y CLD
Sbjct: 678 QSPFQKLFGSSPNYQRLRVFGCLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDV 737
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQL-SSVGSVHSSIPLFS-FP-QLTTLSSPA 239
RLY S+ V+F+E +P+ S L + S SS P S FP + L SP
Sbjct: 738 DNNRLYTSRHVMFDESTYPFAASIREQSQSSLVTPPESSSSSSPANSGFPCSVLRLQSPP 797
Query: 240 PHSPNTTLPESSSPASQSSTPILPEPT-SPVPQ----------------SSPAPISPALS 282
SP T P P Q+ +P+ P T SP P S+ P +P +
Sbjct: 798 ASSPETPSP----PQQQNDSPVSPRQTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHEN 853
Query: 283 SPAPQSSSTDSSSSVSPIVNPH-------------------------------------- 304
P P++ S +S + P+ NP+
Sbjct: 854 GPEPEAQSNPNSPFIGPLPNPNPETNPSSSIEQRPVDKSTTTALPPNQTTIAATSNSRSQ 913
Query: 305 ---NTHSMVTRAKDGIVKPRLVPTLLLSQ-----AEPTSAKLALKDPKWYSAMQDEFNAL 356
N H M TR+K+ I KP+ +L ++ +EP + ALKD KW AM DEF+A
Sbjct: 914 PPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSEPNTVTQALKDKKWRFAMSDEFDAQ 973
Query: 357 HSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSP 416
N TW LVP + +GC+WVF++K +G +++YKARLVAKG++Q G DY ETFSP
Sbjct: 974 QRNHTWDLVPPNPTQHLVGCRWVFKLKYLPNGLIDKYKARLVAKGFNQQYGVDYAETFSP 1033
Query: 417 VVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLN 475
V+K T+R++L +AV K W + QLDVNNAFL G L EEVYM QPPGF KDR S VCRL
Sbjct: 1034 VIKATTIRVVLDVAVKKNWPLKQLDVNNAFLQGTLTEEVYMAQPPGFVDKDRPSHVCRLR 1093
Query: 476 KALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCS-VYVLVYVDDIIITGN 534
KA+YGLKQAPRAW+ LK LL GF S + SLF +YS G + +Y+LVYVDDII+TG+
Sbjct: 1094 KAIYGLKQAPRAWYMELKQHLLNIGFVNSLADTSLF-IYSHGTTLLYLLVYVDDIIVTGS 1152
Query: 535 SLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYA 594
+ +++ L FS+K L YFLG++ + L L Q KY+ DLL + +M A
Sbjct: 1153 DHKSVSAVLSSLAERFSIKDPTDLHYFLGIEATRT-NTGLHLMQRKYMTDLLAKHNMLDA 1211
Query: 595 KGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLE 654
K ++TP+ ++ KLT HG + D S YRS+VG+LQY+ TRP+I+++VN++ QF+ QP
Sbjct: 1212 KPVATPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLAFTRPDIAFAVNRLSQFMHQPTS 1271
Query: 655 EHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLG 714
+HW A KR+LRYL GT G+ L S P+ L AF DADWA D D+ ST+ +++LG
Sbjct: 1272 DHWQAAKRVLRYLAGTTTHGIFLNSSS---PIHLHAFSDADWAGDSADYVSTNAYVIYLG 1328
Query: 715 PNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHI--PHKTPLILCDNMS 772
N +SW SKKQ V+RSSTE+EYR +AN +E+ W+ SLL ELHI PH P I CDN+
Sbjct: 1329 RNPISWSSKKQRGVSRSSTESEYRAVANAASEIRWLCSLLTELHIRLPH-GPTIFCDNIG 1387
Query: 773 TVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDL 832
+ NPV H+R KH+ LD FVR + +++L + HV + DQ D LTK+LS F
Sbjct: 1388 ATYICANPVFHSRMKHIALDYHFVRGMIQSRALRVSHVSTNDQLADALTKSLSRPHFLSA 1447
Query: 833 RTKLNV 838
R+K+ V
Sbjct: 1448 RSKIGV 1453
>At4g27210 putative protein
Length = 1318
Score = 681 bits (1758), Expect = 0.0
Identities = 386/874 (44%), Positives = 522/874 (59%), Gaps = 49/874 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
Y+R++W++PLK KSD Y F+ FHK++E QL++K+ V Q DGGGEF + + GI
Sbjct: 356 YSRFSWIYPLKLKSDFYNIFLAFHKLVENQLSQKISVFQCDGGGEFVSHKFLQHLQSHGI 415
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
+ +CPHT QNG+ ERKHRH+VE+GL++L +H+ ++W AF T +LIN LPT L
Sbjct: 416 QQQLSCPHTPQQNGLAERKHRHLVELGLSMLFQSHVPHKFWVEAFFTANFLINLLPTSAL 475
Query: 122 -DNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKC 180
++ SP+ KL+ D LR FG AC+P LR Y +K + S +CVFLGY+ +K Y+C
Sbjct: 476 KESISPYEKLYDKKPDYTSLRSFGSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRC 535
Query: 181 L-DPTGRLYISKDVLFNEFRFP----YKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTL 235
L PTGRLYIS+ V+F+E +P YK L +P L++ S P +T
Sbjct: 536 LYPPTGRLYISRHVIFDESVYPFSHTYKHLHPQPRTPLLAAWLRSSDS------PAPSTS 589
Query: 236 SSPAPHSPNTTLPESSSPASQSSTPILPE--PTSPVPQSSPAPI--SPAL---------- 281
+SP+ SP T + P Q TP+LP P S V +S SP
Sbjct: 590 TSPSSRSPLFTSADFP-PLPQRKTPLLPTLVPISSVSHASNITTQQSPDFDSERTTDFDS 648
Query: 282 -----SSPAPQSSSTDSSSSVSPIVNPHNTHS------MVTRAKDGIVKPRLVPTLL--- 327
SS + Q+ S + VN H TH+ MVTRAK GI KP L
Sbjct: 649 ASIGDSSHSSQAGSDSEETIQQASVNVHQTHASTNVHPMVTRAKVGISKPNPRYVFLSHK 708
Query: 328 LSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENAD 387
+S EP + ALK P W AM +E QTW+LVP S +G KWVFR K +AD
Sbjct: 709 VSYPEPKTVTAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHAD 768
Query: 388 GSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFL 447
G++N+ KAR+VAKG+ Q EG DY ET+SPVV+ TVR++L LA + W I Q+DV NAFL
Sbjct: 769 GTLNKLKARIVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFL 828
Query: 448 NGKLEEEVYMVQPPGF-EAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCC 506
+G L+E VYM QP GF + VC L+K++YGLKQ+PRAWF++ LL+FGF S
Sbjct: 829 HGDLKETVYMTQPAGFVDPSKPDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCSKS 888
Query: 507 NPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQV 566
+PSLF + + +L+YVDD++ITGNS L L+A LN EF + +G L YFLG+QV
Sbjct: 889 DPSLFIYAHNNNLILLLLYVDDMVITGNSSQTLTSLLAALNKEFRMTDMGQLHYFLGIQV 948
Query: 567 QHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVG 626
Q L ++Q KY +DLL ASM + + TP+ H +DP+ +RSI G
Sbjct: 949 QR-QQNGLFMSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAG 1007
Query: 627 ALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPL 686
LQY+T+TRP+I ++VN VCQ + QP + +KRILRY+KGT+ G+ S P
Sbjct: 1008 KLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISY---SRDSPT 1064
Query: 687 SLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L A+ D+DW RS G F+G NLVSW SKK V+RSSTEAEY++L++ +E
Sbjct: 1065 LLQAYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASE 1124
Query: 747 LLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSL 805
+LW+ +LL+EL IP TP + CDN+S V LT NP H RTKH ++D FVRE+V K+L
Sbjct: 1125 ILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERVALKAL 1184
Query: 806 IIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVT 839
+++H+P +Q DI TK+L F LR KL VT
Sbjct: 1185 VVKHIPGSEQIADIFTKSLPYEAFIHLRGKLGVT 1218
>At1g53810
Length = 1522
Score = 679 bits (1753), Expect = 0.0
Identities = 384/885 (43%), Positives = 532/885 (59%), Gaps = 51/885 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
Y+R+TW +PLK KSD ++TFV F K++E QL K+K+ Q DGGGEF + + GI
Sbjct: 545 YSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDHGI 604
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
+CP+T QNG+ ERKHRHIVE+GL+++ + + L+YW +F T ++IN LPT L
Sbjct: 605 QQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTSSL 664
Query: 122 DN-ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKC 180
DN ESP+ KL+ + + LR+FGCACYP LR Y + K S +CVFLGY+ +K Y+C
Sbjct: 665 DNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYNEKYKGYRC 724
Query: 181 L-DPTGRLYISKDVLFNEFRFPYKELFGCA----SSPQLSS-VGSVH---------SSIP 225
L PTGR+YIS+ V+F+E P++ ++ +P L + S H S P
Sbjct: 725 LYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVTPTQPDQSRYP 784
Query: 226 LFSFPQLTTL---SSPAPHSPNTTLPESSSPASQSSTPIL-----PEPTSPV-------- 269
+ S PQ T ++PA + T P +S SQ + I PE T+ +
Sbjct: 785 VSSIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVSGSPERTTGLDSASIGDS 844
Query: 270 ---PQSSPAPISPALSSPA--PQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVP 324
P + + SPA SSPA PQ S + + N H+MVTR K+GI KP
Sbjct: 845 YHSPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNKRY 904
Query: 325 TLL---LSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFR 381
LL +S EP + ALK P W +AMQ+E +TWTLVP + +G WVFR
Sbjct: 905 VLLTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVLGSMWVFR 964
Query: 382 IKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLD 441
K +ADGS+++ KARLVAKG+ Q EG DY ET+SPVV+ TVR+IL +A W + Q+D
Sbjct: 965 TKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLKWELKQMD 1024
Query: 442 VNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGLKQAPRAWFERLKAALLKFG 500
V NAFL+G L E VYM QP GF K + VC L+K+LYGLKQ+PRAWF+R LL+FG
Sbjct: 1025 VKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFSNFLLEFG 1084
Query: 501 FTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDY 560
F S +PSLF S+ + +L+YVDD++ITGN+ L HL+A LN EF +K +G + Y
Sbjct: 1085 FICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMKDMGQVHY 1144
Query: 561 FLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSL 620
FLG+Q+Q +D L ++Q KY +DLL ASMA + TP+ + +DP+
Sbjct: 1145 FLGIQIQ-TYDGGLFMSQQKYAEDLLITASMANCSPMPTPLPLQLDRVSNQDEVFSDPTY 1203
Query: 621 YRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPC 680
+RS+ G LQY+T+TRP+I ++VN VCQ + QP + +KRILRY+KGTV G+
Sbjct: 1204 FRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVSMGIQYNSN 1263
Query: 681 S------LHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTE 734
S L A+ D+D+A + RS G F+G N++SW SKKQ V+RSSTE
Sbjct: 1264 SSSVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQPTVSRSSTE 1323
Query: 735 AEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRTKHMELDI 793
AEYR+L+ T +E+ W+ S+L+E+ + TP + CDN+S V LT NP H RTKH ++D
Sbjct: 1324 AEYRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHARTKHFDVDH 1383
Query: 794 FFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
++RE+V K+L+++H+P Q DI TK+L F LR KL V
Sbjct: 1384 HYIRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKLGV 1428
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 675 bits (1741), Expect = 0.0
Identities = 358/852 (42%), Positives = 520/852 (61%), Gaps = 46/852 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
Y+RY+W +PL KS+ + F++F K++E QLN K+KV Q+DGGGEF L + + GI
Sbjct: 541 YSRYSWFYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGI 600
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
HR +CP+T QNG+ ERKHRH+VE+GL++L H+H ++W +F T Y+INRLP+ VL
Sbjct: 601 HHRISCPYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVL 660
Query: 122 DNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCL 181
N SP+ LF D LR+FG ACYP LRP +K S +CVFLGY++ +K Y+C
Sbjct: 661 KNLSPYEALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCF 720
Query: 182 -DPTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAP 240
PTG++YIS++V+FNE P+KE + S +P +S P L
Sbjct: 721 YPPTGKVYISRNVIFNESELPFKEKY--------------QSLVPQYSTPLLQAWQ---- 762
Query: 241 HSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPI 300
H+ + + ++P S PI + + + ++ L+ P P S++ S V+P+
Sbjct: 763 HNKISEISVPAAPVQLFSKPI------DLNTYAGSQVTEQLTDPEPTSNNEGSDEEVNPV 816
Query: 301 VNPH--------NTHSMVTRAKDGIVKPRLVPTLLLSQ---AEPTSAKLALKDPKWYSAM 349
N+H+M TR+K GI KP L+ S+ AEP + A+K P W A+
Sbjct: 817 AEEIAANQEQVINSHAMTTRSKAGIQKPNTRYALITSRMNTAEPKTLASAMKHPGWNEAV 876
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFD 409
+E N +H TW+LVP + KWVF+ K + DGS+++ KARLVAKG+ Q EG D
Sbjct: 877 HEEINRVHMLHTWSLVPPTDDMNILSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVD 936
Query: 410 YKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF-EAKDR 468
Y ETFSPVV+ T+R++L ++ SKGW I QLDV+NAFL+G+L+E V+M QP GF + +
Sbjct: 937 YLETFSPVVRTATIRLVLDVSTSKGWPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKP 996
Query: 469 SLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDD 528
+ VCRL KA+YGLKQAPRAWF+ LL +GF S +PSLF + G +Y+L+YVDD
Sbjct: 997 THVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDD 1056
Query: 529 IIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLER 588
I++TG+ +L+ L+ L FS+K LG YFLG+Q++ + L L Q+ Y D+L++
Sbjct: 1057 ILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLGIQIED-YANGLFLHQTAYATDILQQ 1115
Query: 589 ASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQF 648
A M+ + TP+ Q+L + A+P+ +RS+ G LQY+TITRP+I ++VN +CQ
Sbjct: 1116 AGMSDCNPMPTPL--PQQLDNLNSELFAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQR 1173
Query: 649 LSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSG 708
+ P + +KRILRY+KGT+ GL P + L+L A+ D+D A + RST+G
Sbjct: 1174 MHSPTTSDFGLLKRILRYIKGTIGMGL---PIKRNSTLTLSAYSDSDHAGCKNTRRSTTG 1230
Query: 709 SLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPL-IL 767
+ LG NL+SW +K+Q V+ SSTEAEYR L E+ W+ LL++L IP P +
Sbjct: 1231 FCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAREITWISFLLRDLGIPQYLPTQVY 1290
Query: 768 CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPL 827
CDN+S V L+ NP LH R+KH + D ++RE+V + QH+ + Q D+ TK+L
Sbjct: 1291 CDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGLIETQHISATFQLADVFTKSLPRR 1350
Query: 828 RFEDLRTKLNVT 839
F DLR+KL V+
Sbjct: 1351 AFVDLRSKLGVS 1362
>At4g10690 retrotransposon like protein
Length = 1515
Score = 656 bits (1692), Expect = 0.0
Identities = 366/887 (41%), Positives = 517/887 (58%), Gaps = 59/887 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKP--LTPYFTNLGI 61
Y+R+TW +PLK KSD ++ FV F +++E Q K+ + Q DGGGEF + + GI
Sbjct: 542 YSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGGEFVSYKFVAHLASCGI 601
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
+CPHT QNGI ER+HR++ E+GL+L+ H+ + + W AF T +L N LP+ L
Sbjct: 602 KQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEAFFTSNFLSNLLPSSTL 661
Query: 122 -DNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKC 180
DN+SP+ L LR+FG ACYPYLRPY +K S CVFLGY+ +K Y+C
Sbjct: 662 SDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSLLCVFLGYNNKYKGYRC 721
Query: 181 LDP-TGRLYISKDVLFNEFRFPYKELFG----CASSPQLSSVGSVHSSIPL--------- 226
L P TG++YI + VLF+E +FPY +++ + SP ++ SS L
Sbjct: 722 LHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQKGFSSTALSRETPSTNV 781
Query: 227 --FSFPQLTTLSS-PAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSS 283
FP T SS P +PN ++ ++ + P SP+ +S P P S+
Sbjct: 782 EDIIFPSATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVVPPSPITSTS-LPTQPEEST 840
Query: 284 PAPQSSSTDSSSSVSPIVNPHN---------------------------THSMVTRAKDG 316
STDS +++S + P + +H M+TRAK G
Sbjct: 841 SDQNHYSTDSETAISSAMTPQSINVSLFEDSDFPPLQSVISSTTAAPETSHPMITRAKSG 900
Query: 317 IVKPRLVPTLLLSQA---EPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKA 373
I KP L ++ EP S K ALKD W +AM +E +H TW LVP +
Sbjct: 901 ITKPNPKYALFSVKSNYPEPKSVKEALKDEGWTNAMGEEMGTMHETDTWDLVPPEMVDRL 960
Query: 374 IGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSK 433
+GCKWVF+ K N+DGS++R KARLVA+GY Q EG DY ET+SPVV+ TVR IL +A
Sbjct: 961 LGCKWVFKTKLNSDGSLDRLKARLVARGYEQEEGVDYVETYSPVVRSATVRSILHVATIN 1020
Query: 434 GWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGLKQAPRAWFERL 492
W + QLDV NAFL+ +L+E V+M QPPGFE R VC+L KA+Y LKQAPRAWF++
Sbjct: 1021 KWSLKQLDVKNAFLHDELKETVFMTQPPGFEDPSRPDYVCKLKKAIYDLKQAPRAWFDKF 1080
Query: 493 KAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSL 552
+ LLK+GF S +PSLF +++L+YVDD+I+TGN+ +LQ L+ L+ EF +
Sbjct: 1081 SSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLYVDDMILTGNNDVLLQQLLNILSTEFRM 1140
Query: 553 KQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGA 612
K +G L YFLG+Q H H+ L L+Q KY DLL A M+ + TP+ L +
Sbjct: 1141 KDMGALHYFLGIQA-HYHNDGLFLSQEKYTSDLLVNAGMSDCSSMPTPL--QLDLLQGNN 1197
Query: 613 NYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVD 672
+P+ +R + G LQY+T+TRP+I ++VN VCQ + P + +KRIL YLKGT+
Sbjct: 1198 KPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFVCQKMHAPTMSDFHLLKRILHYLKGTMT 1257
Query: 673 FGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSS 732
G++L S + L + D+DWA D RST G FLG N++SW +K+ V++SS
Sbjct: 1258 MGINL---SSNTDSVLRCYSDSDWAGCKDTRRSTGGFCTFLGYNIISWSAKRHPTVSKSS 1314
Query: 733 TEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILCDNMSTVALTHNPVLHTRTKHMEL 791
TEAEYR L+ +E+ W+ LLQE+ +P + P + CDN+S V L+ NP LH+R+KH ++
Sbjct: 1315 TEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIPEMYCDNLSAVYLSANPALHSRSKHFQV 1374
Query: 792 DIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNV 838
D ++VRE+V +L ++H+P+ Q DI TK+L F DLR KL V
Sbjct: 1375 DYYYVRERVALGALTVKHIPASQQLADIFTKSLPQAPFCDLRFKLGV 1421
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 590 bits (1520), Expect = e-168
Identities = 352/918 (38%), Positives = 505/918 (54%), Gaps = 93/918 (10%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+R W + + K++ NF M E Q K+VK +TD G EF LTPYF GI+H
Sbjct: 586 YSRSVWTYLMSSKTEVSQLIKNFCAMSERQFGKQVKAFRTDNGTEFMCLTPYFQTHGILH 645
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+ +C T QNG VERKHRHI+ + A L ++ +++W + LT +LINR P+ VL
Sbjct: 646 QTSCVDTPQQNGRVERKHRHILNVARACLFQGNLPVKFWGESILTATHLINRTPSAVLKG 705
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
++P+ LF LR FGC CY ++RP N K + S++CVF+GY K+++ D
Sbjct: 706 KTPYELLFGERPSYDMLRSFGCLCYAHIRPRNKDKFTSRSRKCVFIGYPHGKKAWRVYDL 765
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHS 242
TG+++ S+DV F+E +PY A++ Q S+V + P+ + +S+ S
Sbjct: 766 ETGKIFASRDVRFHEDIYPY------ATATQ-SNVPLPPPTPPMVNDDWFLPISTQVD-S 817
Query: 243 PNTTLPESSSPA----------------SQSSTPILPEPTSPVPQSSPAPISPALSSPAP 286
N SSSPA S S+ P+ E S VP SSP+ +S
Sbjct: 818 TNVDSSSSSSPAQSGSIDQPPRSIDQSPSTSTNPVPEEIGSIVPSSSPSRSIDRSTSDLS 877
Query: 287 QSSSTD------SSSSVSP----------------------IVNP--------HNTHSM- 309
S +T+ SS+ SP + N HN HS
Sbjct: 878 ASDTTELLSTGESSTPSSPGLPELLGKGCREKKKSVLLKDFVTNTTSKKKTASHNIHSPS 937
Query: 310 --------VTRAKDGIVKPRLVP------------------TLLLSQAEPTSAKLALKDP 343
+ + D + L P +L EP K A+
Sbjct: 938 QVLPSGLPTSLSADSVSGKTLYPLSDFLTNSGYSANHIAFMAAILDSNEPKHFKDAILIK 997
Query: 344 KWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYS 403
+W AM E +AL +N TW + LP +KAI KWV+++K N+DG++ R+KARLV G
Sbjct: 998 EWCEAMSKEIDALEANHTWDITDLPHGKKAISSKWVYKLKYNSDGTLERHKARLVVMGNH 1057
Query: 404 QVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF 463
Q EG D+KETF+PV K TVR IL++A +K W +HQ+DV+NAFL+G LEEEVYM PPGF
Sbjct: 1058 QKEGVDFKETFAPVAKLTTVRTILAVAAAKDWEVHQMDVHNAFLHGDLEEEVYMRLPPGF 1117
Query: 464 EAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVL 523
+ D S VCRL K+LYGLKQAPR WF +L AL GFT S + SLF+L + ++VL
Sbjct: 1118 KCSDPSKVCRLRKSLYGLKQAPRCWFSKLSTALRNIGFTQSYEDYSLFSLKNGDTIIHVL 1177
Query: 524 VYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIK 583
VYVDD+I+ GN+L + ++L+ F +K LG L YFLG++V D L+Q KY
Sbjct: 1178 VYVDDLIVAGNNLDAIDRFKSQLHKCFHMKDLGKLKYFLGLEVSRGPD-GFCLSQRKYAL 1236
Query: 584 DLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVN 643
D+++ + K + P+ N KL +P YR +VG Y+TITRP++SY+V+
Sbjct: 1237 DIVKETGLLGCKPSAVPIALNHKLASITGPVFTNPEQYRRLVGRFIYLTITRPDLSYAVH 1296
Query: 644 KVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDH 703
+ QF+ PL HW A R++RYLKG+ G+ L+ S L + A+CD+D+ A P
Sbjct: 1297 ILSQFMQAPLVAHWEAALRLVRYLKGSPAQGIFLRSDS---SLIINAYCDSDYNACPLTR 1353
Query: 704 RSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKT 763
RS S +V+LG + +SW +KKQ V+ SS EAEYR +A T+ EL W+++LL++L + H +
Sbjct: 1354 RSLSAYVVYLGDSPISWKTKKQDTVSYSSAEAEYRAMAYTLKELKWLKALLKDLGVHHSS 1413
Query: 764 PLIL-CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTK 822
P+ L CD+ + + + NPV H RTKH+E D VR+ VL+K + +H+ + DQ D+LTK
Sbjct: 1414 PMKLHCDSEAAIHIAANPVFHERTKHIESDCHKVRDAVLDKLITTEHIYTEDQVADLLTK 1473
Query: 823 ALSPLRFEDLRTKLNVTD 840
+L FE L + L VTD
Sbjct: 1474 SLPRPTFERLLSTLGVTD 1491
>At1g57640
Length = 1444
Score = 578 bits (1490), Expect = e-165
Identities = 331/876 (37%), Positives = 490/876 (55%), Gaps = 51/876 (5%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+R WV+ + KS+T +F ++E Q + ++K+V++D G EF + YF + GI H
Sbjct: 576 YSRGVWVYLMTDKSETQKHLKDFIALVERQFDTEIKIVRSDNGTEFLCMREYFLHKGIAH 635
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+C T HQNG VERKHRHI+ + AL +++ +++W L+ YLINR P+++L
Sbjct: 636 ETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQFWGECILSAAYLINRTPSMLLQG 695
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
+SP+ L++ + LR+FG CY + + + K + S+ CVF+GY K ++ D
Sbjct: 696 KSPYEMLYKTAPKYSHLRVFGSLCYAHNQNHKGDKFAARSRRCVFVGYPHGQKGWRLFDL 755
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLS---SPA 239
+ ++S+DV+F E FPY ++ C + V V + T + A
Sbjct: 756 EEQKFFVSRDVIFQETEFPYSKM-SCNEEDERVLVDCVGPPFIEEAIGPRTIIGRNIGEA 814
Query: 240 PHSPNTTL--------PESSSPAS----QSSTPILPEPTS-----PVPQSSPAPISPALS 282
PN ESSSP+ S P L T P+ ++PAPI S
Sbjct: 815 TVGPNVATGPIIPEINQESSSPSEFVSLSSLDPFLASSTVQTADLPLSSTTPAPIQLRRS 874
Query: 283 SPAPQS-----------------SSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPT 325
S Q S SSSS+ PI + H + K +
Sbjct: 875 SRQTQKPMKLKNFVTNTVSVESISPEASSSSLYPIEKYVDCHRFTSSHKAFLAA------ 928
Query: 326 LLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKEN 385
+ + EPT+ A+ D W AM E +L NQT+++V LP ++A+G KWV++IK
Sbjct: 929 -VTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFSIVNLPPGKRALGNKWVYKIKYR 987
Query: 386 ADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNA 445
+DG++ RYKARLV G Q EG DY ETF+PV K TVR+ L +A ++ WH+HQ+DV+NA
Sbjct: 988 SDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTVRLFLGVAAARDWHVHQMDVHNA 1047
Query: 446 FLNGKLEEEVYMVQPPGFEAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASC 505
FL+G L+EEVYM P GF+ D S VCRL+K+LYGLKQAPR WF +L +AL ++GFT S
Sbjct: 1048 FLHGDLKEEVYMKLPQGFQCDDPSKVCRLHKSLYGLKQAPRCWFSKLSSALKQYGFTQSL 1107
Query: 506 CNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQ 565
+ SLF+ + G V+VLVYVDD+II+G+ + + L + F +K LG L YFLG++
Sbjct: 1108 SDYSLFSYNNDGIFVHVLVYVDDLIISGSCPDAVAQFKSYLESCFHMKDLGLLKYFLGIE 1167
Query: 566 VQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIV 625
V + + L+Q KY+ D++ + A+ + P+ N KL+ + ++D S YR +V
Sbjct: 1168 VSR-NAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQNHKLSLSTSPLLSDSSRYRRLV 1226
Query: 626 GALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQP 685
G L Y+ +TRPE+SYSV+ + QF+ P ++HW A R++RYLK G+ L S
Sbjct: 1227 GRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRVVRYLKSNPGQGILLSSTS---T 1283
Query: 686 LSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVA 745
L + +CD+D+AA P RS +G V LG +SW +KKQ V+RSS EAEYR +A
Sbjct: 1284 LQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQPTVSRSSAEAEYRAMAFLTQ 1343
Query: 746 ELLWVQSLLQELHIPHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKS 804
EL+W++ +L +L + H + I D+ S +AL+ NPV H RTKH+E+D F+R+ +L+
Sbjct: 1344 ELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHERTKHVEVDCHFIRDAILDGI 1403
Query: 805 LIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVTD 840
+ VPS Q DILTKAL KL + D
Sbjct: 1404 IATSFVPSHKQLADILTKALGEKEVRYFLRKLGILD 1439
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 572 bits (1473), Expect = e-163
Identities = 327/863 (37%), Positives = 477/863 (54%), Gaps = 32/863 (3%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+R W++ L KS+ NF M + Q N K+K V++D G EF LT +F G++H
Sbjct: 476 YSRGVWLYLLNDKSEAPCHLKNFFAMTDRQFNVKIKTVRSDNGTEFLCLTKFFQEQGVIH 535
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+C T +N VERKHRH++ + AL A++ +++W LT YLINR P+ VL++
Sbjct: 536 ERSCVATPERNDRVERKHRHLLNVARALRFQANLPIQFWGECVLTAAYLINRTPSSVLND 595
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
+P+ +L + LR+FG CY + R K + S+ CVF+GY K ++ D
Sbjct: 596 STPYERLHKKQPRFDHLRVFGSLCYAHNRNRGGDKFAERSRRCVFVGYPHGQKGWRLFDL 655
Query: 183 PTGRLYISKDVLFNEFRFPYK---------ELFGCASSPQLSSV--GSVHSSIPLFSFPQ 231
++S+DV+F+E FP++ E +P + + VH P
Sbjct: 656 EQNEFFVSRDVVFSELEFPFRISHEQNVIEEEEEALWAPIVDGLIEEEVHLGQNAGPTPP 715
Query: 232 LTTLSSPAPHSPNTTLPESS-SPASQSSTPILPEP-TSPVPQSSPAPISPALSSPAPQSS 289
+ S P SP+ T S S +S T ++P P TS SSP+ + P ++
Sbjct: 716 ICVSS---PISPSATSSRSEHSTSSPLDTEVVPTPATSTTSASSPSSPTNLQFLPLSRAK 772
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLAL------KDP 343
T + + P V P S +A +K +V T + Q P+ L D
Sbjct: 773 PTTAQAVAPPAVPPPRRQSTRNKAPPVTLKDFVVNT-TVCQESPSKLNSILYQLQKRDDT 831
Query: 344 KWYSAMQDEF---NALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAK 400
+ +SA + +A N TWT+ LP ++AIG +WV+++K N+DGSV RYKARLVA
Sbjct: 832 RRFSASHTTYVAIDAQEENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARLVAL 891
Query: 401 GYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQP 460
G Q EG DY ETF+PV K TVR+ L +AV + W IHQ+DV+NAFL+G L EEVYM P
Sbjct: 892 GNKQKEGEDYGETFAPVAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYMKLP 951
Query: 461 PGFEAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSV 520
PGFEA + VCRL KALYGLKQAPR WFE+L AL ++GF S + SLFTL +
Sbjct: 952 PGFEASHPNKVCRLRKALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGSVRI 1011
Query: 521 YVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSK 580
+L+YVDD+IITGNS Q L + F +K LG L YFLG++V + + Q K
Sbjct: 1012 KILIYVDDLIITGNSQRATQQFKEYLASCFHMKDLGPLKYFLGIEVAR-STTGIYICQRK 1070
Query: 581 YIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISY 640
Y D++ + K + P+ N KL + + DP YR +VG L Y+ +TR ++++
Sbjct: 1071 YALDIISETGLLGVKPANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTRLDLAF 1130
Query: 641 SVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADP 700
SV+ + +F+ +P E+HWAA R++RYLK G+ L+ Q + +CD+DWA DP
Sbjct: 1131 SVHILARFMQEPREDHWAAALRVVRYLKADPGQGVFLRRSGDFQ---ITGWCDSDWAGDP 1187
Query: 701 DDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP 760
RS +G V G + +SW +KKQ V++SS EAEYR ++ +ELLW++ LL L +
Sbjct: 1188 MSRRSVTGYFVQFGDSPISWKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLGVS 1247
Query: 761 HKTPLIL-CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDI 819
H P+I+ CD+ S + + NPV H RTKH+E+D FVR++ + + +HV + Q DI
Sbjct: 1248 HVQPMIMCCDSKSAIYIATNPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLADI 1307
Query: 820 LTKALSPLRFEDLRTKLNVTDKF 842
TK L F R KL + + +
Sbjct: 1308 FTKPLGRDCFSAFRIKLGIRNLY 1330
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 571 bits (1472), Expect = e-163
Identities = 339/850 (39%), Positives = 477/850 (55%), Gaps = 51/850 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
++R TW++ LK KSD T F F ++E Q + +VK V++D E T ++ GIV
Sbjct: 654 HSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKELA-FTEFYKAKGIVS 712
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CP T QN +VERKH+HI+ + AL+ +++SL YW LT V+LINR P+ +L N
Sbjct: 713 FHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFLINRTPSALLSN 772
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
++PF L D L+ FGC CY HK S+ CVFLGY K YK LD
Sbjct: 773 KTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKGYKLLDL 832
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGS-VHSSIPLFSFPQLTTLSSPAPH 241
+ ++IS++V F+E ELF ASS Q ++ S V + + S T P+P
Sbjct: 833 ESNVVHISRNVEFHE------ELFPLASSQQSATTASDVFTPMDPLSSGNSITSHLPSPQ 886
Query: 242 -SPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPI 300
SP+T + S ++ + V + PIS +LS S +++S I
Sbjct: 887 ISPSTQI--SKRRITKFPAHLQDYHCYFVNKDDSHPISSSLSYSQISPSHMLYINNISKI 944
Query: 301 VNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQ 360
P + H AKD +W A+ E A+
Sbjct: 945 PIPQSYH----EAKD--------------------------SKEWCGAIDQEIGAMERTD 974
Query: 361 TWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKP 420
TW + LP +KA+GCKWVF +K +ADGS+ R+KAR+VAKGY+Q EG DY ETFSPV K
Sbjct: 975 TWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKM 1034
Query: 421 ITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGF-EAKDRSL----VCRLN 475
TV+++L ++ SK W+++QLD++NAFLNG LEE +YM P G+ + K SL VCRL
Sbjct: 1035 ATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLK 1094
Query: 476 KALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNS 535
K++YGLKQA R WF + +LL GF + +LF + +LVYVDDI+I +
Sbjct: 1095 KSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCIGSEFIVLLVYVDDIVIASTT 1154
Query: 536 LPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAK 595
Q L L A F L++LG L YFLG++V + + L+Q KY +LL A M K
Sbjct: 1155 EQAAQSLTEALKASFKLRELGPLKYFLGLEVART-SEGISLSQRKYALELLTSADMLDCK 1213
Query: 596 GISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEE 655
S PM N +L+K+ + D +YR +VG L Y+TITRP+I+++VNK+CQF S P
Sbjct: 1214 PSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTA 1273
Query: 656 HWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGP 715
H AAV ++L+Y+KGTV GL S L+L + DADW PD RST+G +F+G
Sbjct: 1274 HLAAVYKVLQYIKGTVGQGLFY---SAEDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGS 1330
Query: 716 NLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVA 775
+L+SW SKKQ V+RSS EAEYR LA E+ W+ +LL L + P++ D+ + V
Sbjct: 1331 SLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVHSGVPILYSDSTAAVY 1390
Query: 776 LTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
+ NPV H RTKH+E+D VREK+ N L + HV + DQ DILTK L P +F L +K
Sbjct: 1391 IATNPVFHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAHLLSK 1450
Query: 836 LNVTDKFTLA 845
+++ + F +
Sbjct: 1451 MSIQNIFVFS 1460
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 567 bits (1462), Expect = e-162
Identities = 338/853 (39%), Positives = 485/853 (56%), Gaps = 57/853 (6%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
++R TW++ LK KS+ T F F + +E Q KVK V++D E K T ++ GIV
Sbjct: 643 HSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELK-FTSFYAEKGIVS 701
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CP T QN +VERKH+HI+ + AL+ + + L W LT V+LINR P+ +L N
Sbjct: 702 FHSCPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMN 761
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
++P+ L + + LR FGC CY P HK S+ C+FLGY + +K YK +D
Sbjct: 762 KTPYEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDL 821
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFS--FPQLTTLSSPAP 240
+ ++IS++V F+E E+F A +P GS SS+ LF+ P + + S
Sbjct: 822 ESNTVFISRNVQFHE------EVFPLAKNP-----GS-ESSLKLFTPMVPVSSGIISDTT 869
Query: 241 HSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPI-----SPALSSPAPQSSSTDSSS 295
HSP SS P+ S P P+ +S + PA + + S SST S S
Sbjct: 870 HSP------SSLPSQISDLP--PQISSQRVRKPPAHLNDYHCNTMQSDHKYPISSTISYS 921
Query: 296 SVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNA 355
+SP +H + I K +P PT+ A +W A+ E A
Sbjct: 922 KISP------SHMCYI---NNITK---IPI-------PTNYAEAQDTKEWCEAVDAEIGA 962
Query: 356 LHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFS 415
+ TW + LP +KA+GCKWVF +K ADG++ RYKARLVAKGY+Q EG DY +TFS
Sbjct: 963 MEKTNTWEITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFS 1022
Query: 416 PVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAK-----DRSL 470
PV K T++++L ++ SK W + QLDV+NAFLNG+LEEE++M P G+ + ++
Sbjct: 1023 PVAKMTTIKLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNV 1082
Query: 471 VCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDII 530
V RL +++YGLKQA R WF++ ++LL GF + + +LF G V VLVYVDDI+
Sbjct: 1083 VLRLKRSIYGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDGEFVIVLVYVDDIV 1142
Query: 531 ITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERAS 590
I S L +L+ F L+ LG L YFLG++V + + Q KY +LL+
Sbjct: 1143 IASTSEAAAAQLTEELDQRFKLRDLGDLKYFLGLEVART-TAGISICQRKYALELLQSTG 1201
Query: 591 MAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLS 650
M K +S PMI N K+ K + + D YR IVG L Y+TITRP+I+++VNK+CQF S
Sbjct: 1202 MLACKPVSVPMIPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSS 1261
Query: 651 QPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSL 710
P H A R+L+Y+KGTV GL S L+L F D+DWA+ D RST+
Sbjct: 1262 APRTTHLTAAYRVLQYIKGTVGQGLFY---SASSDLTLKGFADSDWASCQDSRRSTTSFT 1318
Query: 711 VFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDN 770
+F+G +L+SW SKKQ V+RSS EAEYR LA E++W+ +LL L P++ D+
Sbjct: 1319 MFVGDSLISWRSKKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQASPPVPILYSDS 1378
Query: 771 MSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFE 830
+ + + NPV H RTKH++LD VRE++ N L + HV + DQ DILTK L P +FE
Sbjct: 1379 TAAIYIATNPVFHERTKHIKLDCHTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFE 1438
Query: 831 DLRTKLNVTDKFT 843
L++K+++ + F+
Sbjct: 1439 HLKSKMSILNIFS 1451
>At1g70010 hypothetical protein
Length = 1315
Score = 545 bits (1405), Expect = e-155
Identities = 322/849 (37%), Positives = 464/849 (53%), Gaps = 38/849 (4%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+R TWV+ L+ KSD T F M+E Q +K V++D E T ++ + GIV
Sbjct: 492 YSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELN-FTQFYHSKGIVP 550
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+CP T QN +VERKH+HI+ + +L +HI + YW LT VYLINRLP +L++
Sbjct: 551 YHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRLPAPILED 610
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
+ PF L + +++FGC CY P + HK S +K C F+GY + K YK LD
Sbjct: 611 KCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGFKGYKLLDL 670
Query: 183 PTGRLYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHS 242
T + +S+ V+F+E FP+ +GS S FP L +P P
Sbjct: 671 ETHSIIVSRHVVFHEELFPF--------------LGSDLSQEEQNFFPDL----NPTPPM 712
Query: 243 PNTTLPESSSPASQSSTPILPE--PTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPI 300
+ + S SS ILP PT+ VP+ S PA S S
Sbjct: 713 QRQSSDHVNPSDSSSSVEILPSANPTNNVPEPSVQTSHRKAKKPAYLQDYYCHSVVSS-- 770
Query: 301 VNPHNTHSMVTRAKDGIVKPRLVPTLLLSQA-EPTSAKLALKDPKWYSAMQDEFNALHSN 359
PH ++ D I P L L + EP++ A K W AM EF+ L
Sbjct: 771 -TPHEIRKFLSY--DRINDPYLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEGT 827
Query: 360 QTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVK 419
TW + LP+ ++ IGC+W+F+IK N+DGSV RYKARLVA+GY+Q EG DY ETFSPV K
Sbjct: 828 HTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVAK 887
Query: 420 PITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKD-----RSLVCRL 474
+V+++L +A + QLD++NAFLNG L+EE+YM P G+ ++ + VCRL
Sbjct: 888 LNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCRL 947
Query: 475 NKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGN 534
K+LYGLKQA R W+ + + LL GF S C+ + F S G + VLVY+DDIII N
Sbjct: 948 KKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFLCVLVYIDDIIIASN 1007
Query: 535 SLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYA 594
+ + L +++ + F L+ LG L YFLG+++ DK + ++Q KY DLL+
Sbjct: 1008 NDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVR-SDKGIHISQRKYALDLLDETGQLGC 1066
Query: 595 KGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLE 654
K S PM + + YR ++G L Y+ ITRP+I+++VNK+ QF P +
Sbjct: 1067 KPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRK 1126
Query: 655 EHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLG 714
H AV +IL+Y+KGT+ GL S L L + +AD+ + D RSTSG +FLG
Sbjct: 1127 AHLQAVYKILQYIKGTIGQGLFYSATS---ELQLKVYANADYNSCRDSRRSTSGYCMFLG 1183
Query: 715 PNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTP-LILCDNMST 773
+L+ W S+KQ +V++SS EAEYR+L+ EL+W+ + L+EL +P P L+ CDN +
Sbjct: 1184 DSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEAA 1243
Query: 774 VALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLR 833
+ + +N V H RTKH+E D VRE++L + H+ + Q D TK L P F L
Sbjct: 1244 IHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKPLYPSHFHRLI 1303
Query: 834 TKLNVTDKF 842
+K+ + + F
Sbjct: 1304 SKMGLLNIF 1312
>At4g27200 putative protein
Length = 819
Score = 506 bits (1304), Expect = e-143
Identities = 309/731 (42%), Positives = 415/731 (56%), Gaps = 61/731 (8%)
Query: 144 GCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCL-DPTGRLYISKDVLFNEFRFP- 201
G AC+P LR Y +K + S +CVFLGY+ +K Y+CL PTGRLYIS+ V+F+E +P
Sbjct: 15 GSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRCLYPPTGRLYISRHVIFDESVYPF 74
Query: 202 ---YKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSS 258
YK L +P L++ S P +T +SP+ SP T + P Q
Sbjct: 75 SHTYKHLHPQPRTPLLAAWLRSSDS------PAPSTSTSPSSRSPLFTSADFP-PLPQRK 127
Query: 259 TPILPE--PTSPVPQSSPAPI--SPAL---------------SSPAPQSSSTDSSSSVSP 299
TP+LP P S V +S SP SS + Q+ S +
Sbjct: 128 TPLLPTLVPISSVSHASNITTQQSPDFDSERTTDFDSASIGDSSHSSQAGSDSEETIQQA 187
Query: 300 IVNPH------NTHSMVTRAKDGIVKPRLVPTLL---LSQAEPTSAKLALKDPKWYSAMQ 350
VN H N H MVTRAK GI KP L +S EP + ALK P W AM
Sbjct: 188 SVNVHQTPASTNVHPMVTRAKVGISKPNPRYVFLSHKVSYPEPKTVTAALKHPGWTGAMT 247
Query: 351 DEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDY 410
+E QTW+LVP S +G KWVFR K +ADG++N+ KAR+VAKG+ Q EG DY
Sbjct: 248 EEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHADGTLNKLKARIVAKGFLQEEGIDY 307
Query: 411 KETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSL 470
ET+SPVV+ TVR++L LA + W I Q+DV NAFL+G L+E VYM QP A +R
Sbjct: 308 LETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFLHGDLKETVYMTQP----AANRDH 363
Query: 471 VCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDII 530
VC L+K++YGLKQ+PRAWF++ LL+FGF +PSLF +Y+ ++ +L+
Sbjct: 364 VCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCRKSDPSLF-IYAHNNNLILLLLSQ--- 419
Query: 531 ITGNSLPVLQHLIAKLNAEFSLKQLGTLDY-FLGVQVQHLHDKSLLLTQSKYIKDLLERA 589
L L+A LN EF + +G FLG+QVQ L ++Q KY +DLL A
Sbjct: 420 -------TLTSLLAALNKEFRMTDMGQHSLTFLGIQVQR-QQNGLFMSQQKYAEDLLIAA 471
Query: 590 SMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFL 649
SM + + TP+ H +DP+ +RSI G LQY+T+TRP+I ++VN VCQ +
Sbjct: 472 SMEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAGKLQYLTLTRPDIQFAVNFVCQKM 531
Query: 650 SQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGS 709
QP + +KRILRY+KGT+ G+ S P L A+ D+DW RS G
Sbjct: 532 HQPTISDFHLLKRILRYIKGTITMGISY---SRDSPTLLQAYSDSDWGNCKQTRRSVGGL 588
Query: 710 LVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIP-HKTPLILC 768
F+G NLVSW SKK V+RSSTEAEY++L++ +E+LW+ +LL+EL IP TP + C
Sbjct: 589 CTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASEILWLSTLLRELRIPLPDTPELFC 648
Query: 769 DNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLR 828
DN+S V LT NP H RTKH ++D FVRE+V K+L+++H+P +Q DI TK+L
Sbjct: 649 DNLSAVYLTANPAFHARTKHFDIDFHFVRERVALKALVVKHIPGSEQIADIFTKSLPYEA 708
Query: 829 FEDLRTKLNVT 839
F LR KL VT
Sbjct: 709 FIHLRGKLGVT 719
>At1g60020 hypothetical protein
Length = 1194
Score = 488 bits (1257), Expect = e-138
Identities = 285/687 (41%), Positives = 379/687 (54%), Gaps = 63/687 (9%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
YTRY+W +P+K+KS F+ F ++ + +K+ + +D GGEF L + ++ GI H
Sbjct: 509 YTRYSWFYPIKQKSHVKDVFMTFKALVANKFQRKIIHLYSDNGGEFIALRSFLSSNGITH 568
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
T PHT NGI ERKHRHIVE GL LL A + YW +AF +YLINR+ + V+
Sbjct: 569 LTTPPHTPEHNGISERKHRHIVETGLTLLGQASMPKSYWSYAFTIAIYLINRMSSDVIGG 628
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
SP+ +LF + + LR+FGC C+P+LRPY HKL CVFLGYS T +Y CL+
Sbjct: 629 ISPYKRLFGQAPNYLKLRVFGCLCFPWLRPYTTHKLDDRPAPCVFLGYSQTQSAYLCLNR 688
Query: 183 PTGRLYISKDVLFNEFRFPY-KELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPA-- 239
TGR+Y S+ V F E +P+ K ++ + S+ S+ +++P F QL ++ P
Sbjct: 689 TTGRVYTSRHVQFVENTYPFTKPTLDPFTNLEESNNHSITTTVPSPPFVQLPSVPPPTRD 748
Query: 240 PHSPNTTLPESS----SPASQSSTPILPEP------------------------------ 265
PH P + P S SP S SS + P
Sbjct: 749 PHQPPPSQPAPSPSPLSPPSMSSPVMTSSPQFSSNRDSTTLHGDYSHVDYGLSSPSNPPG 808
Query: 266 --TSPVPQSSPA----------------PISPALSSPAPQS-SSTDSSSSVSPIVNPHNT 306
TSP SP+ P SP+ SS +P S SS SP P N
Sbjct: 809 PITSPTTSKSPSEPTSSPSHSNQPNKTPPNSPSSSSSSPTPIPSPSPQSSNSPPPPPQNQ 868
Query: 307 HSMVTRAKDGIVKP----RLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTW 362
HSM TRAK+ I KP L T PT+ AL+DP W +AM +EFNA N T+
Sbjct: 869 HSMRTRAKNNITKPIKKLTLAATPKGKSKIPTTVAEALRDPNWRNAMSEEFNAGLRNSTY 928
Query: 363 TLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPIT 422
LVP H+ +G +W+F IK N DGS+NRYKAR +AKG+ Q G DY TFSPV+K T
Sbjct: 929 DLVPPKPHQNFVGTRWIFTIKYNPDGSINRYKARFLAKGFHQQHGLDYSNTFSPVIKSTT 988
Query: 423 VRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDR-SLVCRLNKALYGL 481
V+ +L +AVS+ W I QLD+NNAFL G+L E+VY+ QPPGF DR + VC L KAL+GL
Sbjct: 989 VQTVLDIAVSRSWDIRQLDINNAFLQGRLTEDVYVAQPPGFINPDRPNYVCHLKKALHGL 1048
Query: 482 KQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQH 541
KQAPRAW++ L+ LL GFT S N SLF + +Y+LVYVDD +ITG++ ++
Sbjct: 1049 KQAPRAWYQELRGFLLTCGFTNSVANTSLFIRQHNKDYIYILVYVDDFLITGSNSNLIAQ 1108
Query: 542 LIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPM 601
I L FSLK LG L YFL ++ L L Q +Y+ DLL + M AK +STPM
Sbjct: 1109 FITCLANRFSLKDLGQLSYFLEIEATRT-KAGLHLMQRRYVLDLLTKTKMLDAKTVSTPM 1167
Query: 602 ISNQKLTKHGANYMADPSLYRSIVGAL 628
KLT + +P YR I+G+L
Sbjct: 1168 SPTPKLTLTSGTPIDNPGGYRQILGSL 1194
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 466 bits (1200), Expect = e-131
Identities = 299/826 (36%), Positives = 432/826 (52%), Gaps = 103/826 (12%)
Query: 25 NFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVHRFTCPHTHHQNGIVERKHRHI 84
NF M + Q K V+ +++D G EF T YF GI+H +C T QN VERKHRHI
Sbjct: 357 NFCAMADRQFRKPVRSIRSDNGTEFMCHTSYFQEHGILHETSCVDTPQQNARVERKHRHI 416
Query: 85 VEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDNESPFTKLF--QVSFDVKFLRI 142
+ + L + P+ VL ++P+ LF Q S+D+ LR
Sbjct: 417 LNVARTCLFQGNFP-----------------TPSPVLKGKTPYEVLFGKQPSYDM--LRT 457
Query: 143 FGCACYPYLRPYNAHKLSLHSKECVFLGY---STTHKSYKCLDPTGRLYISKDVLFNEFR 199
FGC CY ++RP + K + S++C+F+GY + T ++ +DPT S D E
Sbjct: 458 FGCLCYAHIRPRDKDKFASRSRKCIFIGYPHETATPNTHDSIDPTS---TSSD----ENN 510
Query: 200 FPYKELFGCASSPQL-SSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSS 258
P + + A P SS+ S H H+ + + SS
Sbjct: 511 TPPEPVTPQAEQPHSPSSISSPH-----------------IVHNKGSVHSRHLNEDHDSS 553
Query: 259 TPILPE-------PTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVT 311
+P LPE P P P ++ + S SS S S+VSP V+ ++ M
Sbjct: 554 SPGLPELLGKGHRPKHP-PVYLKDYVAHKVHSSPHTSSPGLSDSNVSPTVSANHIAFMAA 612
Query: 312 RAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHR 371
+L E K + +W AMQ E AL +N TW + LP +
Sbjct: 613 ---------------ILDSNEQNHFKDDVLIKEWCDAMQKEIEALEANHTWDVTDLPHGK 657
Query: 372 KAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAV 431
KAI KWV+++K N+DG++ R+KARLV G Q EG D+KETF+PV K TVR++L++A
Sbjct: 658 KAISSKWVYKLKFNSDGTLERHKARLVVMGNHQKEGIDFKETFAPVAKMTTVRLLLAVAA 717
Query: 432 SKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSLVCRLNKALYGLKQAPRAWFER 491
+K W + Q+DV+NAFL+G LE+E +LYGLKQAPR WF +
Sbjct: 718 AKDWDVFQMDVHNAFLHGDLEQE----------------------SLYGLKQAPRCWFAK 755
Query: 492 LKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFS 551
L AL K GFT S + SLF+L G ++ LVYVDD II GN+L + H L+ F
Sbjct: 756 LSTALRKLGFTQSYEDYSLFSLNRDGTVIHFLVYVDDFIIVGNNLKAIDHFKEHLHKCFH 815
Query: 552 LKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHG 611
+K LG L YFLG++V D L+Q KY D++ A + K + PM + KL
Sbjct: 816 MKDLGKLKYFLGLEVSRGAD-GFCLSQQKYALDIINEAGLLGYKPSAVPMELHHKLGSIS 874
Query: 612 ANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTV 671
+ +P+ YR +V Y+TITRP++SY+V+ + QF+ PLE HW A R++RYLKG+
Sbjct: 875 SPVFDNPAQYRRLVDRFIYLTITRPDLSYAVHILSQFMQTPLEAHWHATLRLVRYLKGSP 934
Query: 672 DFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARS 731
D G+ L+ + LSL A+CD+D+ P RS S +++LG +SW +KKQ V+ S
Sbjct: 935 DQGILLRS---DRALSLTAYCDSDYNPCPRTRRSLSAYVLYLGDTPISWKTKKQDTVSSS 991
Query: 732 STEAEYRNLANTVAELLWVQSLLQELHIPHKTPLIL-CDNMSTVALTHNPVLHTRTKHME 790
S EAEYR +A T+ E+ W+++L+ L + H P++L CD+ + + + NPV H RTKH+E
Sbjct: 992 SAEAEYRAMAYTLKEIKWLKALMTTLGVDHTQPILLFCDSQAAIHIAANPVFHERTKHIE 1051
Query: 791 LDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTKL 836
D VR+ V +K + H+ + D+LTKAL FE L + L
Sbjct: 1052 KDCHQVRDAVTDKVISTPHIST----TDLLTKALPRPTFERLLSTL 1093
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 434 bits (1115), Expect = e-121
Identities = 287/801 (35%), Positives = 429/801 (52%), Gaps = 75/801 (9%)
Query: 91 LLAHAHISLEYWDHA----FLTVVYLINRLPTV-VLDNESPFTKLFQVSFDVKFLRIFGC 145
LL H + + ++ H FLT+V R V ++ N+S + +F V + F +
Sbjct: 234 LLWHQRLGIRHYLHYRNLYFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKLIFTQYN-- 291
Query: 146 ACYPYLRPYNAHKLS----------LHSKECVFLG--------YSTTHKSYKCLD-PTGR 186
A +R N +L+ +H C + Y + +K YK LD +
Sbjct: 292 AKIKAIRSDNVKELAFTKFVKEQGMIHQFSCAYTPQQNSVVERYPSGYKGYKVLDLESHS 351
Query: 187 LYISKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAP---HSP 243
+ I+++V+F+E +FP+K S SV +SI P S P +
Sbjct: 352 ISITRNVVFHETKFPFK-----TSKFLKESVDMFPNSILPLPAPLHFVESMPLDDDLRAD 406
Query: 244 NTTLPESSSPASQSSTPILPEP------------TSPVP-------QSSPAPIS------ 278
+ S+S +S SS P LP T+ VP +PA +S
Sbjct: 407 DNNASTSNSASSASSIPPLPSTVNTQNTDALDIDTNSVPIARPKRNAKAPAYLSEYHCNS 466
Query: 279 -PALSSPAPQSSST--DSSSSVSP--IVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEP 333
P LSS +P +S++ SSS+ P I P+ + ++ K + + + + EP
Sbjct: 467 VPFLSSLSPTTSTSIETPSSSIPPKKITTPYPMSTAISYDKLTPLFHSYICAYNV-ETEP 525
Query: 334 TSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRY 393
+ A+K KW A +E +AL N+TW + L + +GCKWVF IK N DGS+ RY
Sbjct: 526 KAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNVVGCKWVFTIKYNPDGSIERY 585
Query: 394 KARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEE 453
KARLVA+G++Q EG DY ETFSPV K +V+++L LA + GW + Q+DV+NAFL+G+L+E
Sbjct: 586 KARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAATGWSLTQMDVSNAFLHGELDE 645
Query: 454 EVYMVQPPGFE-----AKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNP 508
E+YM P G+ + VCRL K+LYGLKQA R W++RL + L F S +
Sbjct: 646 EIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQASRQWYKRLSSVFLGANFIQSPADN 705
Query: 509 SLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQH 568
++F S + VLVYVDD++I N +++L L +EF +K LG +FLG+++
Sbjct: 706 TMFVKVSCTSIIVVLVYVDDLMIASNDSSAVENLKELLRSEFKIKDLGPARFFLGLEIAR 765
Query: 569 LHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGAL 628
+ + + Q KY ++LLE ++ K S PM N LTK + + + YR +VG L
Sbjct: 766 -SSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLTKEMGTLLPNATSYRELVGRL 824
Query: 629 QYVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSL 688
Y+ ITRP+I+++V+ + QFLS P + H A ++LRYLKG GL S L L
Sbjct: 825 LYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLKGNPGQGLMY---SASSELCL 881
Query: 689 IAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELL 748
F DADW D RS +G ++LG +L++W SKKQS+V+RSSTE+EYR+LA E++
Sbjct: 882 NGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLAQATCEII 941
Query: 749 WVQSLLQELHIPHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLII 807
W+Q LL++LH+ P + CDN S + L NPV H RTKH+E+D VR+++ L
Sbjct: 942 WLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQIKAGKLKT 1001
Query: 808 QHVPSLDQRDDILTKALSPLR 828
HVP+ +Q DILTK L P++
Sbjct: 1002 LHVPTGNQLADILTKPLHPVQ 1022
Score = 60.5 bits (145), Expect = 4e-09
Identities = 29/75 (38%), Positives = 44/75 (58%), Gaps = 1/75 (1%)
Query: 5 TRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVHR 64
TR TWV+ +K KS+ F F K+I Q N K+K +++D E T + G++H+
Sbjct: 261 TRTTWVYMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKEL-AFTKFVKEQGMIHQ 319
Query: 65 FTCPHTHHQNGIVER 79
F+C +T QN +VER
Sbjct: 320 FSCAYTPQQNSVVER 334
>At4g22040 LTR retrotransposon like protein
Length = 1109
Score = 431 bits (1108), Expect = e-121
Identities = 225/510 (44%), Positives = 323/510 (63%), Gaps = 5/510 (0%)
Query: 332 EPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVN 391
EPT+ A+ D W AM E +L NQT+++V LP ++A+G KWV++IK +DG++
Sbjct: 599 EPTTYNEAMVDKAWREAMSAEIESLRVNQTFSIVNLPPGKRALGNKWVYKIKYRSDGAIE 658
Query: 392 RYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKL 451
RYKARLV G Q EG DY ETF+PV K TVR+ L +A ++ WH+HQ+DV+NAFL+G L
Sbjct: 659 RYKARLVVLGNCQKEGVDYDETFAPVAKMSTVRLFLGVAAARDWHVHQMDVHNAFLHGDL 718
Query: 452 EEEVYMVQPPGFEAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLF 511
+EEVYM P GF+ D S VCRL+K+LYGLKQAPR WF +L +AL ++GFT S + SLF
Sbjct: 719 KEEVYMKLPQGFQCDDPSKVCRLHKSLYGLKQAPRCWFSKLSSALKQYGFTQSLSDYSLF 778
Query: 512 TLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHD 571
+ + G V+VLVYVDD+II+G+ + + L + F +K LG L YFLG++V +
Sbjct: 779 SYNNDGVFVHVLVYVDDLIISGSCPDAVAQFKSYLESCFHMKDLGLLKYFLGIEVSR-NA 837
Query: 572 KSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYV 631
+ L+Q KY+ D++ + A+ + P+ N KL+ + ++D S YR +VG L Y+
Sbjct: 838 QGFYLSQRKYVLDIISEMGLLGARPSAFPLEQNHKLSLSTSPLLSDSSRYRRLVGRLIYL 897
Query: 632 TITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAF 691
+TRPE+SYSV+ + QF+ P ++HW A R++RYLK G+ L S L + +
Sbjct: 898 AVTRPELSYSVHTLAQFMQNPRQDHWNAAIRVVRYLKSNPGQGILLSSTS---TLQINGW 954
Query: 692 CDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQ 751
CD+D+AA P RS +G V LG +SW +KKQ ++RSS EAEYR +A EL+W++
Sbjct: 955 CDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQPTISRSSAEAEYRAMAFLTQELMWLK 1014
Query: 752 SLLQELHIPHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHV 810
+L +L + H + I D+ S +AL+ NPV H RTKH+E+D F+R+ +L+ + V
Sbjct: 1015 RVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHERTKHVEVDCHFIRDAILDGIIATSFV 1074
Query: 811 PSLDQRDDILTKALSPLRFEDLRTKLNVTD 840
PS Q DILTKAL KL + D
Sbjct: 1075 PSHKQLADILTKALGEKEVRYFLRKLGILD 1104
Score = 158 bits (400), Expect = 1e-38
Identities = 72/203 (35%), Positives = 120/203 (58%), Gaps = 1/203 (0%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLTPYFTNLGIVH 63
Y+R WV+ + KS+T +F ++E Q + ++K V++D G EF + YF + GI H
Sbjct: 376 YSRGVWVYLMTDKSETQKHLKDFMALVERQFDTEIKTVRSDNGTEFLCMREYFLHKGIAH 435
Query: 64 RFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDN 123
+C T HQNG VERKHRHI+ + AL +++ +++W L+ YLINR P+++L
Sbjct: 436 ETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQFWGECILSAAYLINRTPSMLLQG 495
Query: 124 ESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD- 182
+SP+ L++ + LR+FG CY + + + K + S+ CVF+GY K ++ D
Sbjct: 496 KSPYEMLYKTAPKYSHLRVFGSLCYAHNQNHKGDKFAARSRRCVFVGYPHGQKGWRLFDL 555
Query: 183 PTGRLYISKDVLFNEFRFPYKEL 205
+ ++S+DV+F E FPY ++
Sbjct: 556 EEQKFFVSRDVIFQETEFPYSKM 578
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,775,734
Number of Sequences: 26719
Number of extensions: 902442
Number of successful extensions: 8081
Number of sequences better than 10.0: 415
Number of HSP's better than 10.0 without gapping: 216
Number of HSP's successfully gapped in prelim test: 210
Number of HSP's that attempted gapping in prelim test: 4170
Number of HSP's gapped (non-prelim): 1718
length of query: 856
length of database: 11,318,596
effective HSP length: 108
effective length of query: 748
effective length of database: 8,432,944
effective search space: 6307842112
effective search space used: 6307842112
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0322.2


