

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
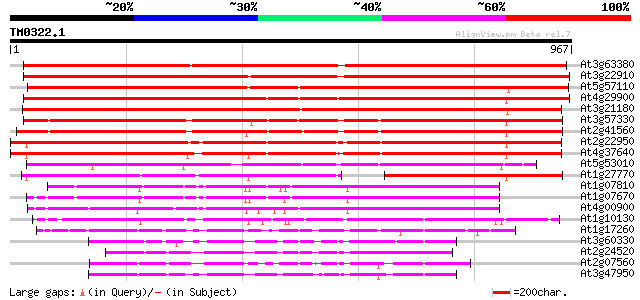
Query= TM0322.1
(967 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g63380 Ca2+-transporting ATPase -like protein 1332 0.0
At3g22910 calmodulin-stimulated calcium-ATPase, putative 1286 0.0
At5g57110 Ca2+-transporting ATPase-like protein (emb|CAB79748.1) 1018 0.0
At4g29900 Ca2+-transporting ATPase - like protein 1014 0.0
At3g21180 putative Ca2+-transporting ATPase 986 0.0
At3g57330 Ca2+-transporting ATPase-like protein 842 0.0
At2g41560 putative Ca2+-ATPase 820 0.0
At2g22950 pseudogene 784 0.0
At4g37640 plasma membrane-type calcium ATPase (ACA2) 782 0.0
At5g53010 Ca2+-transporting ATPase-like protein 518 e-147
At1g27770 envelope Ca2+-ATPase 387 e-107
At1g07810 ER-type Ca2+-pump protein 353 3e-97
At1g07670 endoplasmic reticulum-type calcium-transporting ATPase... 352 6e-97
At4g00900 Ca2+-transporting ATPase - like protein 346 4e-95
At1g10130 putative calcium ATPase 326 4e-89
At1g17260 H+-transporting ATPase AHA10 201 2e-51
At3g60330 plasma membrane H+-ATPase - like 182 1e-45
At2g24520 putative plasma membrane proton ATPase 181 2e-45
At2g07560 pseudogene 179 7e-45
At3g47950 H+-transporting ATPase - like protein 177 3e-44
>At3g63380 Ca2+-transporting ATPase -like protein
Length = 1033
Score = 1332 bits (3446), Expect = 0.0
Identities = 687/940 (73%), Positives = 787/940 (83%), Gaps = 14/940 (1%)
Query: 24 VDKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVR 83
+D+ +L ++K K+L GGVEGVA L T P KGI G++ + + RR+LFG+NTY +
Sbjct: 88 IDQEQLVEIMKGKDLPGIQALGGVEGVAASLRTNPTKGIHGNEQEVSRRRDLFGSNTYHK 147
Query: 84 PPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVS 143
PPPK L FV EA D TILILL CA SLGFGIKEHG EGWYEGGSIF+AVFLV+VVS
Sbjct: 148 PPPKGLLFFVYEAFKDLTILILLVCAIFSLGFGIKEHGIKEGWYEGGSIFVAVFLVIVVS 207
Query: 144 ALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFL 203
ALSNFRQ+RQFDKLSKISN+IKVEV+R+ R Q ISIFDV+VGDV++LKIGDQIPADGLFL
Sbjct: 208 ALSNFRQERQFDKLSKISNNIKVEVLRDSRRQHISIFDVVVGDVVFLKIGDQIPADGLFL 267
Query: 204 GGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSI 263
GHSLQVDESSMTGESDH+E++ PFL SG K+VDG+AQMLV +VG +T WGQ MSSI
Sbjct: 268 EGHSLQVDESSMTGESDHLEVDHKDNPFLFSGTKIVDGFAQMLVVSVGMSTTWGQTMSSI 327
Query: 264 SGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTD 323
+ D+SERTPLQ RLD LTS+IGKIGL VA LVL+VLL+RYFTGNTE E G +EY GSKT
Sbjct: 328 NQDSSERTPLQVRLDTLTSTIGKIGLTVAALVLVVLLVRYFTGNTEKE-GKREYNGSKTP 386
Query: 324 INDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGS 383
++ V N+VV IVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMM+DQAMVRKLSACETMGS
Sbjct: 387 VDTVVNSVVRIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMSDQAMVRKLSACETMGS 446
Query: 384 ATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYKP 443
ATVICTDKTGTLTLN+M+VTKFWLG E++ E+ + ++P VL+L +QG GLNTTGSV
Sbjct: 447 ATVICTDKTGTLTLNEMKVTKFWLGQESIHEDSTKMISPDVLDLLYQGTGLNTTGSVCVS 506
Query: 444 SAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETN 503
+ S PE SGSPTEKA+L W V +LGMDM+ +KQKH+VL VETF+S KKRSGV VR++++
Sbjct: 507 DSGSTPEFSGSPTEKALLSWTVLNLGMDMESVKQKHEVLRVETFSSAKKRSGVLVRRKSD 566
Query: 504 NTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEE-RSKIEKIIQGMAASSLRCIAFAYME 562
NTVHVHWKGAAEMVLAMCS+Y S G+ +D +S+I+ IIQGMAASSLRCIAFA+
Sbjct: 567 NTVHVHWKGAAEMVLAMCSHYYTSTGSVDLMDSTAKSRIQAIIQGMAASSLRCIAFAHKI 626
Query: 563 ISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNI 622
S VL EDGLTL+GIVGLKDPCRP V KAVETCKLAGV IKMITGDN+
Sbjct: 627 ASNDS----------VLEEDGLTLMGIVGLKDPCRPGVSKAVETCKLAGVTIKMITGDNV 676
Query: 623 FTAKAIATECGILDLNDAG--GVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMV 680
FTAKAIA ECGILD ND VVEGV+FRNYT+EERM+KVDKIRVMARSSP DKLLMV
Sbjct: 677 FTAKAIAFECGILDHNDKDEEDAVVEGVQFRNYTDEERMQKVDKIRVMARSSPSDKLLMV 736
Query: 681 QCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVL 740
+CL+ KGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNF SVATVL
Sbjct: 737 KCLRLKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFASVATVL 796
Query: 741 RWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALAL 800
+WGRCVYNNIQKFIQFQLTVNVAALVINFIAA+S+G+VPLT VQLLWVNLIMDTLGALAL
Sbjct: 797 KWGRCVYNNIQKFIQFQLTVNVAALVINFIAAISAGEVPLTAVQLLWVNLIMDTLGALAL 856
Query: 801 ATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSKEVKN 860
ATERPT EL+++KP+GRTE LIT +MWRNLL Q+LYQIAVLL+ QF G SIF+V KEVK+
Sbjct: 857 ATERPTNELLKRKPVGRTEALITNVMWRNLLVQSLYQIAVLLILQFKGMSIFSVRKEVKD 916
Query: 861 TLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADT 920
TLIFNTFVLCQVFNEFN+R MEK NVF+G+ +N LF+GI+ ITIVLQV+MVE L+KFADT
Sbjct: 917 TLIFNTFVLCQVFNEFNAREMEKKNVFKGLHRNRLFIGIIAITIVLQVIMVEFLKKFADT 976
Query: 921 ERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSKLFFTNAK 960
RLN QWG CI +A++SWPI + TK PV F + K
Sbjct: 977 VRLNGWQWGTCIALASLSWPIGFFTKFIPVSETPFLSYFK 1016
>At3g22910 calmodulin-stimulated calcium-ATPase, putative
Length = 1017
Score = 1286 bits (3327), Expect = 0.0
Identities = 654/947 (69%), Positives = 775/947 (81%), Gaps = 17/947 (1%)
Query: 24 VDKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVR 83
+D L+++VK+KN E GG G+ L + GI D+ RR FG+NTY R
Sbjct: 83 IDTETLNDLVKNKNQEKLESLGGPNGLVSALKSNTRLGINEEGDEIQRRRSTFGSNTYTR 142
Query: 84 PPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVS 143
P K HFV+EA D TILILLGCA LSLGFGIKEHG EGWY+GGSIF+AVFLVV VS
Sbjct: 143 QPSKGLFHFVVEAFKDLTILILLGCATLSLGFGIKEHGLKEGWYDGGSIFVAVFLVVAVS 202
Query: 144 ALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFL 203
A+SNFRQ+RQFDKLSK+S++IK++VVRNGR Q+ISIFD++VGD++ L IGDQ+PADG+F+
Sbjct: 203 AVSNFRQNRQFDKLSKVSSNIKIDVVRNGRRQEISIFDIVVGDIVCLNIGDQVPADGVFV 262
Query: 204 GGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSI 263
GH L VDESSMTGESDHVE+ FL SG K+ DG+ +M VT+VG NTAWGQMMS I
Sbjct: 263 EGHLLHVDESSMTGESDHVEVSLTGNTFLFSGTKIADGFGKMAVTSVGMNTAWGQMMSHI 322
Query: 264 SGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTD 323
S D +E+TPLQ+RLDKLTSSIGK+GL VAFLVLLVLLIRYFTG T+DE+GN+EY G T
Sbjct: 323 SRDTNEQTPLQSRLDKLTSSIGKVGLLVAFLVLLVLLIRYFTGTTKDESGNREYNGKTTK 382
Query: 324 INDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGS 383
+++ NAVV +VAAAVTI+VVAIPEGLPLAVTLTLAYSMKRMM D AMVRKLSACETMGS
Sbjct: 383 SDEIVNAVVKMVAAAVTIIVVAIPEGLPLAVTLTLAYSMKRMMKDNAMVRKLSACETMGS 442
Query: 384 ATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYKP 443
ATVICTDKTGTLTLNQM+VT FW GLE+ +++++ V+ELFHQGV +NTTGSV+K
Sbjct: 443 ATVICTDKTGTLTLNQMKVTDFWFGLES---GKASSVSQRVVELFHQGVAMNTTGSVFKA 499
Query: 444 SAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETN 503
A +E E SGSPTEKA+L WAV +L M M+++ ++H V+HVE FNSEKKRSGV ++K+
Sbjct: 500 KAGTEYEFSGSPTEKAILSWAVEELEMGMEKVIEEHDVVHVEGFNSEKKRSGVLMKKKGV 559
Query: 504 NTVH--VHWKGAAEMVLAMCSNYIDSNGTQKSL-DEERSKIEKIIQGMAASSLRCIAFAY 560
NT + VHWKGAAE +LAMCS + D +G + + ++++ + EKIIQ MAA SLRCIAFAY
Sbjct: 560 NTENNVVHWKGAAEKILAMCSTFCDGSGVVREMKEDDKIQFEKIIQSMAAKSLRCIAFAY 619
Query: 561 MEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGD 620
E +E + L+E+ L+LLGI+G+KDPCRP VKKAVE C+ AGV+IKMITGD
Sbjct: 620 SEDNE---------DNKKLKEEKLSLLGIIGIKDPCRPGVKKAVEDCQFAGVNIKMITGD 670
Query: 621 NIFTAKAIATECGILDLNDA--GGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLL 678
NIFTA+AIA ECGIL D V+EG +FRNYT+EER+EKV++I+VMARSSP DKLL
Sbjct: 671 NIFTARAIAVECGILTPEDEMNSEAVLEGEKFRNYTQEERLEKVERIKVMARSSPFDKLL 730
Query: 679 MVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVAT 738
MV+CLK+ GHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNF SVAT
Sbjct: 731 MVKCLKELGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFASVAT 790
Query: 739 VLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGAL 798
VL+WGRCVYNNIQKFIQFQLTVNVAALVINF+AAVS+GDVPLT VQLLWVNLIMDTLGAL
Sbjct: 791 VLKWGRCVYNNIQKFIQFQLTVNVAALVINFVAAVSAGDVPLTAVQLLWVNLIMDTLGAL 850
Query: 799 ALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSKEV 858
ALATE+PT +LM+KKPIGR PLIT IMWRNLLAQA YQI+VLLV QF G+SIFNV+++V
Sbjct: 851 ALATEKPTNDLMKKKPIGRVAPLITNIMWRNLLAQAFYQISVLLVLQFRGRSIFNVTEKV 910
Query: 859 KNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFA 918
KNTLIFNTFVLCQVFNEFN+RS+EK NVF+G+ KN LF+GI+ +T+VLQV+MVE L++FA
Sbjct: 911 KNTLIFNTFVLCQVFNEFNARSLEKKNVFKGLHKNRLFIGIIVVTVVLQVVMVEFLKRFA 970
Query: 919 DTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSKLFFTNAKWVKSS 965
DTERLN QWG+CI IAA SWPI WL K PVP + FF+ KW K S
Sbjct: 971 DTERLNLGQWGVCIAIAAASWPIGWLVKSVPVPERHFFSYLKWKKRS 1017
>At5g57110 Ca2+-transporting ATPase-like protein (emb|CAB79748.1)
Length = 1074
Score = 1018 bits (2632), Expect = 0.0
Identities = 534/942 (56%), Positives = 676/942 (71%), Gaps = 15/942 (1%)
Query: 32 MVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLH 91
M KD N A ++GG +G+A++L T P KGI G DDD R+ ++G+NTY R K FL
Sbjct: 124 MSKDHNSGALEQYGGTQGLANLLKTNPEKGISGDDDDLLKRKTIYGSNTYPRKKGKGFLR 183
Query: 92 FVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQD 151
F+ +A +D T++IL+ A SL GIK G EGWY+GGSI AV LV+VV+A+S+++Q
Sbjct: 184 FLWDACHDLTLIILMVAAVASLALGIKTEGIKEGWYDGGSIAFAVILVIVVTAVSDYKQS 243
Query: 152 RQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVD 211
QF L+ +I +EV+R GR +ISI+D++VGDVI L IG+Q+PADG+ + GHSL +D
Sbjct: 244 LQFQNLNDEKRNIHLEVLRGGRRVEISIYDIVVGDVIPLNIGNQVPADGVLISGHSLALD 303
Query: 212 ESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERT 271
ESSMTGES V + K PFL+SG KV DG MLVT VG NT WG +M+SIS DN E T
Sbjct: 304 ESSMTGESKIVNKDANKDPFLMSGCKVADGNGSMLVTGVGVNTEWGLLMASISEDNGEET 363
Query: 272 PLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTDINDVCNAV 331
PLQ RL+ + + IG IGLAVA VL++LL RYFTG+T+D NG ++ KT + V + V
Sbjct: 364 PLQVRLNGVATFIGSIGLAVAAAVLVILLTRYFTGHTKDNNGGPQFVKGKTKVGHVIDDV 423
Query: 332 VSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDK 391
V ++ AVTIVVVA+PEGLPLAVTLTLAYSM++MMAD+A+VR+LSACETMGSAT IC+DK
Sbjct: 424 VKVLTVAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALVRRLSACETMGSATTICSDK 483
Query: 392 TGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEI 451
TGTLTLNQM V + + G + + + + T+ L +G+ NTTGS++ P + E
Sbjct: 484 TGTLTLNQMTVVESYAGGK---KTDTEQLPATITSLVVEGISQNTTGSIFVPEGGGDLEY 540
Query: 452 SGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWK 511
SGSPTEKA+L W V LGM+ + + + +LH FNSEKKR GVAV K + VHVHWK
Sbjct: 541 SGSPTEKAILGWGVK-LGMNFETARSQSSILHAFPFNSEKKRGGVAV-KTADGEVHVHWK 598
Query: 512 GAAEMVLAMCSNYIDSNGTQKSL-DEERSKIEKIIQGMAASSLRCIAFAYMEISEGGDYI 570
GA+E+VLA C +YID +G + D++ S + I MA +LRC+A A+
Sbjct: 599 GASEIVLASCRSYIDEDGNVAPMTDDKASFFKNGINDMAGRTLRCVALAFRTYEAEKVPT 658
Query: 571 EKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIAT 630
+ + VL ED L LL IVG+KDPCRP VK +V C+ AGV ++M+TGDN+ TA+AIA
Sbjct: 659 GEELSKWVLPEDDLILLAIVGIKDPCRPGVKDSVVLCQNAGVKVRMVTGDNVQTARAIAL 718
Query: 631 ECGIL--DLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGH 688
ECGIL D + + ++EG FR T+ ER + DKI VM RSSP DKLL+VQ L+++GH
Sbjct: 719 ECGILSSDADLSEPTLIEGKSFREMTDAERDKISDKISVMGRSSPNDKLLLVQSLRRQGH 778
Query: 689 VVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYN 748
VVAVTGDGTNDAPAL EADIGL+MGI GTEVAKESSDI+ILDDNF SV V+RWGR VY
Sbjct: 779 VVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKESSDIIILDDNFASVVKVVRWGRSVYA 838
Query: 749 NIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKE 808
NIQKFIQFQLTVNVAALVIN +AA+SSGDVPLT VQLLWVNLIMDTLGALALATE PT
Sbjct: 839 NIQKFIQFQLTVNVAALVINVVAAISSGDVPLTAVQLLWVNLIMDTLGALALATEPPTDH 898
Query: 809 LMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSKE-------VKNT 861
LM + P+GR EPLIT IMWRNLL QA+YQ++VLL F G SI + E VKNT
Sbjct: 899 LMGRPPVGRKEPLITNIMWRNLLIQAIYQVSVLLTLNFRGISILGLEHEVHEHATRVKNT 958
Query: 862 LIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTE 921
+IFN FVLCQ FNEFN+R ++ N+F+G++KN LF+GI+ IT+VLQV++VE L KFA T
Sbjct: 959 IIFNAFVLCQAFNEFNARKPDEKNIFKGVIKNRLFMGIIVITLVLQVIIVEFLGKFASTT 1018
Query: 922 RLNWEQWGICIGIAAVSWPIAWLTKLTPVPSKLFFTNAKWVK 963
+LNW+QW IC+GI +SWP+A + K PVP+ K +K
Sbjct: 1019 KLNWKQWLICVGIGVISWPLALVGKFIPVPAAPISNKLKVLK 1060
>At4g29900 Ca2+-transporting ATPase - like protein
Length = 1069
Score = 1014 bits (2623), Expect = 0.0
Identities = 534/957 (55%), Positives = 693/957 (71%), Gaps = 20/957 (2%)
Query: 24 VDKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVR 83
+ + ++ ++ +D+N+ A E GGV G++D+L T KGI G DDD R+ FG+NTY +
Sbjct: 116 IGQEQIVSISRDQNIGALQELGGVRGLSDLLKTNLEKGIHGDDDDILKRKSAFGSNTYPQ 175
Query: 84 PPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVS 143
+ F FV EA D T++IL+ A SL GIK G +GWY+G SI AV LV+VV+
Sbjct: 176 KKGRSFWRFVWEASQDLTLIILIVAAVASLALGIKTEGIEKGWYDGISIAFAVLLVIVVT 235
Query: 144 ALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFL 203
A S++RQ QF L++ +I++EV R+GR +ISI+D++VGDVI L IGDQ+PADG+ +
Sbjct: 236 ATSDYRQSLQFQNLNEEKRNIRLEVTRDGRRVEISIYDIVVGDVIPLNIGDQVPADGVLV 295
Query: 204 GGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSI 263
GHSL VDESSMTGES V+ K PFL+SG KV DG MLVT VG NT WG +M+S+
Sbjct: 296 AGHSLAVDESSMTGESKIVQKNSTKHPFLMSGCKVADGNGTMLVTGVGVNTEWGLLMASV 355
Query: 264 SGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTD 323
S DN TPLQ RL+ + + IG +GL VA +VL VL++RYFTG+T++E G ++ G KT
Sbjct: 356 SEDNGGETPLQVRLNGVATFIGIVGLTVAGVVLFVLVVRYFTGHTKNEQGGPQFIGGKTK 415
Query: 324 INDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGS 383
V + +V I AVTIVVVA+PEGLPLAVTLTLAYSM++MMAD+A+VR+LSACETMGS
Sbjct: 416 FEHVLDDLVEIFTVAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALVRRLSACETMGS 475
Query: 384 ATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVL-ELFHQGVGLNTTGSVYK 442
AT IC+DKTGTLTLN+M V + + GL+ + S++ P+ + +G+ NTTGSV++
Sbjct: 476 ATTICSDKTGTLTLNEMTVVECYAGLQKMDSPDSSSKLPSAFTSILVEGIAHNTTGSVFR 535
Query: 443 PSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKET 502
S E ++SGSPTE+A+L WA+ LGMD D LK + + FNSEKKR GVAV K
Sbjct: 536 -SESGEIQVSGSPTERAILNWAIK-LGMDFDALKSESSAVQFFPFNSEKKRGGVAV-KSP 592
Query: 503 NNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEER-SKIEKIIQGMAASSLRCIAFAYM 561
+++VH+HWKGAAE+VL C++Y+D + + + E++ ++ I MAA SLRC+A A+
Sbjct: 593 DSSVHIHWKGAAEIVLGSCTHYMDESESFVDMSEDKMGGLKDAIDDMAARSLRCVAIAFR 652
Query: 562 EISEGGDYI---EKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMIT 618
D I E+ R L ED L LL IVG+KDPCRP VK +V C+ AGV ++M+T
Sbjct: 653 TFE--ADKIPTDEEQLSRWELPEDDLILLAIVGIKDPCRPGVKNSVLLCQQAGVKVRMVT 710
Query: 619 GDNIFTAKAIATECGIL--DLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDK 676
GDNI TAKAIA ECGIL D + + ++EG FR+Y+EEER ++I VM RSSP DK
Sbjct: 711 GDNIQTAKAIALECGILASDSDASEPNLIEGKVFRSYSEEERDRICEEISVMGRSSPNDK 770
Query: 677 LLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSV 736
LL+VQ LK++GHVVAVTGDGTNDAPAL EADIGL+MGIQGTEVAKE SDI+ILDDNF SV
Sbjct: 771 LLLVQSLKRRGHVVAVTGDGTNDAPALHEADIGLAMGIQGTEVAKEKSDIIILDDNFESV 830
Query: 737 ATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLG 796
V+RWGR VY NIQKFIQFQLTVNVAALVIN +AA+S+G+VPLT VQLLWVNLIMDTLG
Sbjct: 831 VKVVRWGRSVYANIQKFIQFQLTVNVAALVINVVAAISAGEVPLTAVQLLWVNLIMDTLG 890
Query: 797 ALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV-- 854
ALALATE PT LM + P+GR EPLIT IMWRNL QA+YQ+ VLL+ F G SI ++
Sbjct: 891 ALALATEPPTDHLMDRAPVGRREPLITNIMWRNLFIQAMYQVTVLLILNFRGISILHLKS 950
Query: 855 ---SKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMV 911
++ VKNT+IFN FV+CQVFNEFN+R +++N+F G+L+NHLF+GI+ ITIVLQV++V
Sbjct: 951 KPNAERVKNTVIFNAFVICQVFNEFNARKPDEINIFRGVLRNHLFVGIISITIVLQVVIV 1010
Query: 912 ELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPS---KLFFTNAKWVKSS 965
E L FA T +L+WE W +CIGI ++SWP+A + KL PVP +F +W ++S
Sbjct: 1011 EFLGTFASTTKLDWEMWLVCIGIGSISWPLAVIGKLIPVPETPVSQYFRINRWRRNS 1067
>At3g21180 putative Ca2+-transporting ATPase
Length = 1090
Score = 986 bits (2548), Expect = 0.0
Identities = 526/945 (55%), Positives = 674/945 (70%), Gaps = 20/945 (2%)
Query: 23 DVDKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYV 82
D+D +L +M +++N+ ++GGV+GVA+ L + +GI + + R+ FG+NTY
Sbjct: 129 DIDLEKLVSMTRNQNMSNLQQYGGVKGVAEKLKSNMEQGINEDEKEVIDRKNAFGSNTYP 188
Query: 83 RPPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVV 142
+ K F F+ EA D T++IL+ A SL GIK G EGW +GGSI AV LV+VV
Sbjct: 189 KKKGKNFFMFLWEAWQDLTLIILIIAAVTSLALGIKTEGLKEGWLDGGSIAFAVLLVIVV 248
Query: 143 SALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLF 202
+A+S++RQ QF L+ +I++EV+R GR +ISI+DV+VGDVI L+IGDQ+PADG+
Sbjct: 249 TAVSDYRQSLQFQNLNDEKRNIQLEVMRGGRTVKISIYDVVVGDVIPLRIGDQVPADGVL 308
Query: 203 LGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSS 262
+ GHSL +DESSMTGES V + K+PFL+SG KV DG MLVT VG NT WG +M+S
Sbjct: 309 ISGHSLAIDESSMTGESKIVHKDQ-KSPFLMSGCKVADGVGNMLVTGVGINTEWGLLMAS 367
Query: 263 ISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKT 322
IS D E TPLQ RL+ L + IG +GL+VA +VL+ LL+RYFTG T+D NG ++ T
Sbjct: 368 ISEDTGEETPLQVRLNGLATFIGIVGLSVALVVLVALLVRYFTGTTQDTNGATQFIKGTT 427
Query: 323 DINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMG 382
I+D+ + V I AVTIVVVA+PEGLPLAVTLTLAYSM++MMAD+A+VR+LSACETMG
Sbjct: 428 SISDIVDDCVKIFTIAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALVRRLSACETMG 487
Query: 383 SATVICTDKTGTLTLNQMRVTKFWLGLENV-VENFSNAMAPTVLELFHQGVGLNTTGSVY 441
SAT IC+DKTGTLTLNQM V + + G + V + + + P ++ L +GV NTTG+++
Sbjct: 488 SATTICSDKTGTLTLNQMTVVETYAGGSKMDVADNPSGLHPKLVALISEGVAQNTTGNIF 547
Query: 442 KPSAES-EPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRK 500
P + E EISGSPTEKA+L WA LGM D ++ + ++H FNSEKKR GVAV +
Sbjct: 548 HPKVDGGEVEISGSPTEKAILSWAYK-LGMKFDTIRSESAIIHAFPFNSEKKRGGVAVLR 606
Query: 501 ETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAASSLRCIAFAY 560
++ V +HWKGAAE+VLA C+ Y+DSNGT +S++ ++ I MA +SLRC+A A
Sbjct: 607 G-DSEVFIHWKGAAEIVLACCTQYMDSNGTLQSIESQKEFFRVAIDSMAKNSLRCVAIAC 665
Query: 561 --MEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMIT 618
E+++ ++ + L ED L LL IVG+KDPCRP V++AV C AGV ++M+T
Sbjct: 666 RTQELNQVPKE-QEDLDKWALPEDELILLAIVGIKDPCRPGVREAVRICTSAGVKVRMVT 724
Query: 619 GDNIFTAKAIATECGIL--DLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDK 676
GDN+ TAKAIA ECGIL D ++EG FR +E+ER + KI VM RSSP DK
Sbjct: 725 GDNLQTAKAIALECGILSSDTEAVEPTIIEGKVFRELSEKEREQVAKKITVMGRSSPNDK 784
Query: 677 LLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSV 736
LL+VQ L+K G VVAVTGDGTNDAPAL EADIGLSMGI GTEVAKESSDI+ILDDNF SV
Sbjct: 785 LLLVQALRKNGDVVAVTGDGTNDAPALHEADIGLSMGISGTEVAKESSDIIILDDNFASV 844
Query: 737 ATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLG 796
V+RWGR VY NIQKFIQFQLTVNVAAL+IN +AA+SSGDVPL VQLLWVNLIMDTLG
Sbjct: 845 VKVVRWGRSVYANIQKFIQFQLTVNVAALIINVVAAMSSGDVPLKAVQLLWVNLIMDTLG 904
Query: 797 ALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSK 856
ALALATE PT LM + P+GR EPLIT IMWRNLL Q+ YQ+AVLLV F G SI ++
Sbjct: 905 ALALATEPPTDHLMHRTPVGRREPLITNIMWRNLLVQSFYQVAVLLVLNFAGLSILGLNH 964
Query: 857 -------EVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVL 909
EVKNT+IFN FV+CQ+FNEFN+R +++NVF G+ KN LF+ IVG+T +LQ++
Sbjct: 965 ENHAHAVEVKNTMIFNAFVMCQIFNEFNARKPDEMNVFRGVNKNPLFVAIVGVTFILQII 1024
Query: 910 MVELLRKFADTERLNWEQW--GICIG-IAAVSWPIAWLTKLTPVP 951
+V L KFA T RL W+ W I IG + V WP+A + KL PVP
Sbjct: 1025 IVTFLGKFAHTVRLGWQLWLASIIIGLVRLVHWPLAIVGKLIPVP 1069
>At3g57330 Ca2+-transporting ATPase-like protein
Length = 1025
Score = 842 bits (2174), Expect = 0.0
Identities = 452/939 (48%), Positives = 635/939 (67%), Gaps = 36/939 (3%)
Query: 24 VDKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVR 83
V+ L++MV++ + ++ ++ GG EG+A + A+G+ S+ R +++G N Y
Sbjct: 95 VEADELASMVRNHDTKSLTKIGGPEGIAQKVSVSLAEGVRSSE--LHIREKIYGENRYTE 152
Query: 84 PPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVS 143
P + FL FV EAL D T++IL+ CA +S+G G+ G +G Y+G I L++ LVV+V+
Sbjct: 153 KPARSFLTFVWEALQDITLIILMVCAVVSIGVGVATEGFPKGMYDGTGILLSIILVVMVT 212
Query: 144 ALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFL 203
A+S+++Q QF L + I ++V R+G Q++SI D++VGDV++L IGDQ+PADG+F+
Sbjct: 213 AISDYKQSLQFRDLDREKKKIIIQVTRDGSRQEVSIHDLVVGDVVHLSIGDQVPADGIFI 272
Query: 204 GGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSI 263
G++L++DESS++GES+ + K PFLLSG KV +G A+MLVT VG T WG++M ++
Sbjct: 273 SGYNLEIDESSLSGESEPSHVNKEK-PFLLSGTKVQNGSAKMLVTTVGMRTEWGKLMDTL 331
Query: 264 SGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTD 323
S + TPLQ +L+ + + IGKIGL A L +VL IR+ K GS T+
Sbjct: 332 SEGGEDETPLQVKLNGVATIIGKIGLGFAVLTFVVLCIRFVV--------EKATAGSITE 383
Query: 324 -INDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMG 382
++ ++ A AVTI+VVA+PEGLPLAVTL+LA++MK++M+D+A+VR L+ACETMG
Sbjct: 384 WSSEDALTLLDYFAIAVTIIVVAVPEGLPLAVTLSLAFAMKQLMSDRALVRHLAACETMG 443
Query: 383 SATVICTDKTGTLTLNQMRVTKFWLGLENVVE----NFSNAMAPTVLELFHQGVGLNTTG 438
S+T ICTDKTGTLT N M V K W+ EN+ E NF ++ V + Q + NT
Sbjct: 444 SSTCICTDKTGTLTTNHMVVNKVWI-CENIKERQEENFQLNLSEQVKNILIQAIFQNTGS 502
Query: 439 SVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAV 498
V K E + +I GSPTE+A+L + + LG D+D +++HK+L +E FNS+KK+ V +
Sbjct: 503 EVVKDK-EGKTQILGSPTERAILEFGLL-LGGDVDTQRREHKILKIEPFNSDKKKMSV-L 559
Query: 499 RKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEER-SKIEKIIQGMAASSLRCIA 557
+ V KGA+E+VL MC +DSNG L EE+ + I +I+G A+ +LR +
Sbjct: 560 TSHSGGKVRAFCKGASEIVLKMCEKVVDSNGESVPLSEEKIASISDVIEGFASEALRTLC 619
Query: 558 FAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMI 617
Y ++ E PR L G TL+ +VG+KDP RP V++AV+TC+ AG+ ++M+
Sbjct: 620 LVYTDLDEA--------PRGDLPNGGYTLVAVVGIKDPVRPGVREAVQTCQAAGITVRMV 671
Query: 618 TGDNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKL 677
TGDNI TAKAIA ECGIL AGGV +EG +FRN E + KI+VMARS P+DK
Sbjct: 672 TGDNISTAKAIAKECGILT---AGGVAIEGSDFRNLPPHEMRAILPKIQVMARSLPLDKH 728
Query: 678 LMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVA 737
+V L+K G VVAVTGDGTNDAPAL EADIGL+MGI GTEVAKE++D++I+DDNF ++
Sbjct: 729 TLVNNLRKMGEVVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKENADVIIMDDNFATIV 788
Query: 738 TVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGA 797
V +WGR VY NIQKF+QFQLTVNV AL+INF++A +G PLT VQLLWVN+IMDTLGA
Sbjct: 789 NVAKWGRAVYINIQKFVQFQLTVNVVALIINFVSACITGSAPLTAVQLLWVNMIMDTLGA 848
Query: 798 LALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV--- 854
LALATE P + LM+++PIGRT IT+ MWRN++ Q++YQ+ VL + F GK I N+
Sbjct: 849 LALATEPPNEGLMKRQPIGRTASFITRAMWRNIIGQSIYQLIVLGILNFAGKQILNLNGP 908
Query: 855 -SKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVEL 913
S V NT+IFN+FV CQVFNE NSR +EK+NVFEG+ K+ +F+ ++ T+ QV++VE
Sbjct: 909 DSTIVLNTIIFNSFVFCQVFNEVNSREIEKINVFEGMFKSWVFVAVMTATVGFQVIIVEF 968
Query: 914 LRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPS 952
L FA T L+W+ W +CI I +VS +A K PV S
Sbjct: 969 LGAFASTVPLSWQHWLLCILIGSVSMILAVGLKCIPVES 1007
>At2g41560 putative Ca2+-ATPase
Length = 1030
Score = 820 bits (2119), Expect = 0.0
Identities = 444/954 (46%), Positives = 630/954 (65%), Gaps = 37/954 (3%)
Query: 12 KGTNHCSLVPHDVDKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAA 71
K T+ ++ L++MV+ + ++ ++ GGVE +A + ++GI S+
Sbjct: 83 KLTDEVKKAGFSIEADELASMVRKNDTKSLAQKGGVEELAKKVSVSLSEGIRSSE--VPI 140
Query: 72 RRELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGS 131
R ++FG N Y P + FL FV EAL+D T++IL+ CA +S+G G+ G G Y+G
Sbjct: 141 REKIFGENRYTEKPARSFLMFVWEALHDITLIILMVCAVVSIGVGVATEGFPRGMYDGTG 200
Query: 132 IFLAVFLVVVVSALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLK 191
I L++ LVV+V+A+S+++Q QF L + I V+V R+G Q+ISI D++VGDV++L
Sbjct: 201 ILLSILLVVMVTAISDYKQSLQFRDLDREKKKIIVQVTRDGSRQEISIHDLVVGDVVHLS 260
Query: 192 IGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVG 251
IGDQ+PADG+F+ G++L++DESS++GES+ + K PFLLSG KV +G A+MLVT VG
Sbjct: 261 IGDQVPADGIFISGYNLEIDESSLSGESEPSHVNKEK-PFLLSGTKVQNGSAKMLVTTVG 319
Query: 252 ANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDE 311
T WG++M ++ + TPLQ +L+ + + IGKIGL+ A L +VL IR+
Sbjct: 320 MRTEWGKLMETLVDGGEDETPLQVKLNGVATIIGKIGLSFAVLTFVVLCIRFVL------ 373
Query: 312 NGNKEYKGSKTD-INDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQA 370
+K GS T+ ++ ++ A +VTI+VVA+PEGLPLAVTL+LA++MK++M+D+A
Sbjct: 374 --DKATSGSFTNWSSEDALTLLDYFAISVTIIVVAVPEGLPLAVTLSLAFAMKKLMSDRA 431
Query: 371 MVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWL------GLENVVENFSNAMAPTV 424
+VR L+ACETMGS+T ICTDKTGTLT N M V K W+ E E+F ++ V
Sbjct: 432 LVRHLAACETMGSSTCICTDKTGTLTTNHMVVNKVWICDKVQERQEGSKESFELELSEEV 491
Query: 425 LELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHV 484
QG+ NT V K + +I GSPTE+A+L + + LG D + +++HK+L +
Sbjct: 492 QSTLLQGIFQNTGSEVVKDK-DGNTQILGSPTERAILEFGLL-LGGDFNTQRKEHKILKI 549
Query: 485 ETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEER-SKIEK 543
E FNS+KK+ V + KGA+E+VL MC N +DSNG L EER + I
Sbjct: 550 EPFNSDKKKMSVLIALPGGGA-RAFCKGASEIVLKMCENVVDSNGESVPLTEERITSISD 608
Query: 544 IIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKA 603
II+G A+ +LR + Y ++ E P L + G T++ +VG+KDP RP V++A
Sbjct: 609 IIEGFASEALRTLCLVYKDLDEA--------PSGELPDGGYTMVAVVGIKDPVRPGVREA 660
Query: 604 VETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVD 663
V+TC+ AG+ ++M+TGDNI TAKAIA ECGI GG+ +EG EFR+ + E +
Sbjct: 661 VQTCQAAGITVRMVTGDNISTAKAIAKECGIYT---EGGLAIEGSEFRDLSPHEMRAIIP 717
Query: 664 KIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKES 723
KI+VMARS P+DK +V L+K G VVAVTGDGTNDAPAL EADIGL+MGI GTEVAKE+
Sbjct: 718 KIQVMARSLPLDKHTLVSNLRKIGEVVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKEN 777
Query: 724 SDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTV 783
+D++I+DDNF ++ V RWGR VY NIQKF+QFQLTVNV AL+INF++A +G PLT V
Sbjct: 778 ADVIIMDDNFKTIVNVARWGRAVYINIQKFVQFQLTVNVVALIINFVSACITGSAPLTAV 837
Query: 784 QLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLV 843
QLLWVN+IMDTLGALALATE P + LM++ PI RT ITK MWRN+ Q++YQ+ VL +
Sbjct: 838 QLLWVNMIMDTLGALALATEPPNEGLMKRAPIARTASFITKTMWRNIAGQSVYQLIVLGI 897
Query: 844 FQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGI 899
F GKS+ + S V NT+IFN+FV CQVFNE NSR +EK+NVF+G+ + +F +
Sbjct: 898 LNFAGKSLLKLDGPDSTAVLNTVIFNSFVFCQVFNEINSREIEKINVFKGMFNSWVFTWV 957
Query: 900 VGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSK 953
+ +T+V QV++VE L FA T L+W+ W + I I +++ +A + K PV S+
Sbjct: 958 MTVTVVFQVIIVEFLGAFASTVPLSWQHWLLSILIGSLNMIVAVILKCVPVESR 1011
>At2g22950 pseudogene
Length = 1015
Score = 784 bits (2024), Expect = 0.0
Identities = 456/969 (47%), Positives = 620/969 (63%), Gaps = 38/969 (3%)
Query: 1 IISKRNSHYVSKGTNHCSLVPHDVDKA-------RLSNMVKDKNLEAYSEFGGVEGVADV 53
++SK ++S + VP +V A L ++V+ +++ GGV+G++
Sbjct: 66 LVSKAAFQFISGVSPSDYKVPEEVKAAGFDICADELGSIVEGHDVKKLKFHGGVDGLSGK 125
Query: 54 LGTIPAKGI-LGSDDDTAARRELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLS 112
L P G+ G + + R+ELFG N + + F FV EAL D T++IL CA +S
Sbjct: 126 LKACPNAGLSTGEPEQLSKRQELFGINKFAESELRSFWVFVWEALQDMTLMILGVCAFVS 185
Query: 113 LGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQDRQFDKLSKISNDIKVEVVRNG 172
L GI G +G ++G I ++ LVV V+A S++RQ QF L K I V+V RNG
Sbjct: 186 LIVGIATEGWPQGSHDGLGIVASILLVVFVTATSDYRQSLQFRDLDKEKKKITVQVTRNG 245
Query: 173 RPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFL 232
Q++SI+D+L GDV++L IGDQ+PADGLFL G S+ +DESS+TGES+ V + + PFL
Sbjct: 246 FRQKMSIYDLLPGDVVHLAIGDQVPADGLFLSGFSVVIDESSLTGESEPVMVTA-QNPFL 304
Query: 233 LSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVA 292
LSG KV DG +MLVT VG T WG++M+++S + TPLQ +L+ + + IGKIGL+ A
Sbjct: 305 LSGTKVQDGSCKMLVTTVGMRTQWGKLMATLSEGGDDETPLQVKLNGVATIIGKIGLSFA 364
Query: 293 FLVLLVLLIRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPL 352
+ VL+ F + + S D ++ + A AVTIVVVA+PEGLPL
Sbjct: 365 IVTFAVLVQGMFMRKL---SLGPHWWWSGDDALEL----LEYFAIAVTIVVVAVPEGLPL 417
Query: 353 AVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTK--FWLGLE 410
AVTL+LA++MK+MM D+A+VR L+ACETMGSAT IC+DKTGTLT N M V K + ++
Sbjct: 418 AVTLSLAFAMKKMMNDKALVRHLAACETMGSATTICSDKTGTLTTNHMTVVKSCICMNVQ 477
Query: 411 NVVENFSNAMAP---TVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSD 467
+V S+ + L+L Q + NT G V + + EI G+PTE A+L +S
Sbjct: 478 DVASKSSSLQSDIPEAALKLLLQLIFNNTGGEVVV-NERGKTEILGTPTETAILELGLS- 535
Query: 468 LGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDS 527
LG E +Q +KV+ VE FNS KKR GV + + H KGA+E+VLA C I+S
Sbjct: 536 LGGKFQEERQSNKVIKVEPFNSTKKRMGVVIELPEGGRIRAHTKGASEIVLAACDKVINS 595
Query: 528 NGTQKSLDEERSKIEKI-IQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTL 586
+G LD+E K + I A +LR + AYM+I E G ++G P E G T
Sbjct: 596 SGEVVPLDDESIKFLNVTIDEFANEALRTLCLAYMDI-ESGFSADEGIP-----EKGFTC 649
Query: 587 LGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVE 646
+GIVG+KDP RP V+++VE C+ AG+ ++M+TGDNI TAKAIA ECGIL + G+ +E
Sbjct: 650 IGIVGIKDPVRPGVRESVELCRRAGIMVRMVTGDNINTAKAIARECGILTDD---GIAIE 706
Query: 647 GVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKK-GHVVAVTGDGTNDAPALKE 705
G FR +EE +E + KI+VMARSSPMDK +V+ L+ VVAVTGDGTNDAPAL E
Sbjct: 707 GPVFREKNQEEMLELIPKIQVMARSSPMDKHTLVKQLRTTFDEVVAVTGDGTNDAPALHE 766
Query: 706 ADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAAL 765
ADIGL+MGI GTEVAKE +D++ILDDNF+++ TV +WGR VY NIQKF+QFQLTVNV AL
Sbjct: 767 ADIGLAMGIAGTEVAKEIADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVAL 826
Query: 766 VINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKI 825
++NF +A +G PLT VQLLWVN+IMDTLGALALATE P ELM++ P+GR IT
Sbjct: 827 IVNFSSACLTGSAPLTAVQLLWVNMIMDTLGALALATEPPNNELMKRMPVGRRGNFITNA 886
Query: 826 MWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFNEFNSRSM 881
MWRN+L QA+YQ ++ + Q GKS+F + S V NTLIFN FV CQVFNE +SR M
Sbjct: 887 MWRNILGQAVYQFIIIWILQAKGKSMFGLVGSDSTLVLNTLIFNCFVFCQVFNEVSSREM 946
Query: 882 EKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPI 941
E+++VF+GIL N++F+ ++G T+ Q++++E L FA T L QW I + + PI
Sbjct: 947 EEIDVFKGILDNYVFVVVIGATVFFQIIIIEFLGTFASTTPLTIVQWFFSIFVGFLGMPI 1006
Query: 942 AWLTKLTPV 950
A K PV
Sbjct: 1007 AAGLKKIPV 1015
>At4g37640 plasma membrane-type calcium ATPase (ACA2)
Length = 1014
Score = 782 bits (2019), Expect = 0.0
Identities = 451/973 (46%), Positives = 617/973 (63%), Gaps = 47/973 (4%)
Query: 1 IISKRNSHYVSKGTNHCSLVPHDVDKA-------RLSNMVKDKNLEAYSEFGGVEGVADV 53
++SK ++S + VP DV A L ++V+ +++ GGV+G+A
Sbjct: 66 LVSKAAFQFISGVSPSDYTVPEDVKAAGFEICADELGSIVESHDVKKLKFHGGVDGLAGK 125
Query: 54 LGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLSL 113
L P G+ + R+ELFG N + + F FV EAL D T++IL CA +SL
Sbjct: 126 LKASPTDGLSTEAAQLSQRQELFGINKFAESEMRGFWVFVWEALQDMTLMILGVCAFVSL 185
Query: 114 GFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQDRQFDKLSKISNDIKVEVVRNGR 173
GI G +G ++G I ++ LVV V+A S++RQ QF L K I V+V RNG
Sbjct: 186 IVGIATEGWPKGSHDGLGIAASILLVVFVTATSDYRQSLQFRDLDKEKKKITVQVTRNGF 245
Query: 174 PQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLL 233
Q++SI+D+L GD+++L IGDQ+PADGLFL G S+ +DESS+TGES+ V + + PFL+
Sbjct: 246 RQKLSIYDLLPGDIVHLAIGDQVPADGLFLSGFSVVIDESSLTGESEPVMVNA-QNPFLM 304
Query: 234 SGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAF 293
SG KV DG +M++T VG T WG++M++++ + TPLQ +L+ + + IGKIGL A
Sbjct: 305 SGTKVQDGSCKMMITTVGMRTQWGKLMATLTEGGDDETPLQVKLNGVATIIGKIGLFFAV 364
Query: 294 LVLLVLLIRYF-----TGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPE 348
+ VL+ F TG +G++ + ++ A AVTIVVVA+PE
Sbjct: 365 VTFAVLVQGMFMRKLSTGTHWVWSGDEALE------------LLEYFAIAVTIVVVAVPE 412
Query: 349 GLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLG 408
GLPLAVTL+LA++MK+MM D+A+VR L+ACETMGSAT IC+DKTGTLT N M V K +
Sbjct: 413 GLPLAVTLSLAFAMKKMMNDKALVRHLAACETMGSATTICSDKTGTLTTNHMTVVKSCIC 472
Query: 409 LE-----NVVENFSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLW 463
+ N + + + + ++L Q + NT G V + + E+ G+PTE A+L
Sbjct: 473 MNVQDVANKGSSLQSEIPESAVKLLIQSIFNNTGGEVVV-NKHGKTELLGTPTETAILEL 531
Query: 464 AVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSN 523
+S LG E ++ +KV+ VE FNS KKR GV + + H KGA+E+VLA C
Sbjct: 532 GLS-LGGKFQEERKSYKVIKVEPFNSTKKRMGVVIELPEGGRMRAHTKGASEIVLAACDK 590
Query: 524 YIDSNGTQKSLDEERSKIEKI-IQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLRED 582
++S+G LDEE K + I A +LR + AYM+I EGG P +
Sbjct: 591 VVNSSGEVVPLDEESIKYLNVTINEFANEALRTLCLAYMDI-EGGF-----SPDDAIPAS 644
Query: 583 GLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGG 642
G T +GIVG+KDP RP VK++VE C+ AG+ ++M+TGDNI TAKAIA ECGIL + G
Sbjct: 645 GFTCVGIVGIKDPVRPGVKESVELCRRAGITVRMVTGDNINTAKAIARECGILTDD---G 701
Query: 643 VVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKK-GHVVAVTGDGTNDAP 701
+ +EG FR +EE +E + KI+VMARSSPMDK +V+ L+ VVAVTGDGTNDAP
Sbjct: 702 IAIEGPVFREKNQEELLELIPKIQVMARSSPMDKHTLVKQLRTTFDEVVAVTGDGTNDAP 761
Query: 702 ALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVN 761
AL EADIGL+MGI GTEVAKES+D++ILDDNF+++ TV +WGR VY NIQKF+QFQLTVN
Sbjct: 762 ALHEADIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVN 821
Query: 762 VAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPL 821
V ALV+NF +A +G PLT VQLLWVN+IMDTLGALALATE P ELM++ P+GR
Sbjct: 822 VVALVVNFSSACLTGSAPLTAVQLLWVNMIMDTLGALALATEPPNDELMKRLPVGRRGNF 881
Query: 822 ITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFNEFN 877
IT MWRN+L QA+YQ V+ + Q GK++F + S + NTLIFN FV CQVFNE +
Sbjct: 882 ITNAMWRNILGQAVYQFIVIWILQAKGKAMFGLDGPDSTLMLNTLIFNCFVFCQVFNEIS 941
Query: 878 SRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAV 937
SR ME+++VF+GIL N++F+ ++G T+ Q++++E L FA T L QW I I +
Sbjct: 942 SREMEEIDVFKGILDNYVFVVVIGATVFFQIIIIEFLGTFASTTPLTITQWIFSIFIGFL 1001
Query: 938 SWPIAWLTKLTPV 950
PIA K PV
Sbjct: 1002 GMPIAAGLKTIPV 1014
>At5g53010 Ca2+-transporting ATPase-like protein
Length = 1095
Score = 518 bits (1333), Expect = e-147
Identities = 323/924 (34%), Positives = 501/924 (53%), Gaps = 80/924 (8%)
Query: 29 LSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKI 88
L +VK+++LEA + + GV G++++L T GI DD+ RR +G+NTY K
Sbjct: 147 LVQLVKERSLEALNRYNGVHGLSNLLKTDLKVGIDRRDDEILLRRNAYGSNTYPCKKGKT 206
Query: 89 FLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWY-EGGSIFLAVFLVV------- 140
F +F+ A + +L+++ A IK G +GWY E + + VF ++
Sbjct: 207 FWYFLWRASQFSHLLVIMFAAVFFSLLRIKTKGILDGWYIEACIVLVTVFHIIAIEEIIW 266
Query: 141 ----------VVSALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYL 190
+++A++ ++Q +F KL++ + +EV+R GR ++SI+D++VGD++ L
Sbjct: 267 KQSCLFYLSPILAAVAEYKQSCRFIKLTEEKRTVYLEVIRGGRRVRVSIYDIVVGDIVPL 326
Query: 191 KIGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAV 250
K G Q+PADG+ +SL+V E +T + V+ + PFLLSG+K+++G MLVT+V
Sbjct: 327 KNGCQVPADGVLFVANSLKVAEQEVTASDEIVQKDLQTNPFLLSGSKLIEGIGTMLVTSV 386
Query: 251 GANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLV------------ 298
G NT WG M +S E P Q L L S + A + +
Sbjct: 387 GMNTEWGLKME-VSQKTDEEKPFQGYLKWLAISASWFVVLFASVACSIQVGGSSAPSWQG 445
Query: 299 ----LLIRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAV 354
+ RYF+G T+ +G + T ++ V++ ++ + +VVA+P GL +AV
Sbjct: 446 PNNRFISRYFSGVTKKSDGTPMFIYGITTADEAIEFVITSLSFGIATIVVAVPVGLSIAV 505
Query: 355 TLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVE 414
L +Y ++ +K+ + + M V W G + +
Sbjct: 506 RLNSSYHFPYFISFAKTTKKMRKDKVL------------------MSVVDVWAGGIRMQD 547
Query: 415 NFSNAMAPTVL-ELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMD 473
+ PT L EL +G+ NT GSV + +EPE+ GSPTE+A+L + + LGM D
Sbjct: 548 MDDVSQLPTFLKELIIEGIAQNTNGSVVFETGVTEPEVYGSPTEQAILNFG-NKLGMKFD 606
Query: 474 ELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKS 533
+ + V H FN +KK GVA++ T+ HVHWKG+A+ +L+ C Y+D ++
Sbjct: 607 DARSASLVRHTIPFNPKKKYGGVALQLGTH--AHVHWKGSAKTILSSCEGYMDGANNSRA 664
Query: 534 LDEERSK-IEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGL 592
++E++ K E I+ M+ LRC A AY E G + L LL IVG+
Sbjct: 665 INEQKRKSFEGTIENMSKEGLRCAALAYQPC-------ELGSLPTITEPRNLVLLAIVGI 717
Query: 593 KDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVE-GVEFR 651
KDPCRP + A++ C V + M+T ++ TA+AIA ECGIL DA G + G +FR
Sbjct: 718 KDPCRPGTRDAIQLCNSGSVKVCMVTDNDGLTAQAIAIECGIL--TDASGRNIRTGAQFR 775
Query: 652 NYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLS 711
++ ER + I V A+SSP D LL+VQ LKK+GH+VA TG G +D L+EAD+ L+
Sbjct: 776 ELSDLEREQIAGDILVFAQSSPNDNLLLVQALKKRGHIVAATGMGIHDPKTLREADVSLA 835
Query: 712 MGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIA 771
MG+ GT AKE+SD +ILDDNF ++ + W R +YNN+QK I F+LTV+V+AL + +
Sbjct: 836 MGVGGTAAAKENSDFIILDDNFATIVKCIIWSRSLYNNVQKSILFRLTVSVSALAVCVVE 895
Query: 772 AVSSGDVPLTTVQLLWVNLIMDTLGALALA-TERPTKELMQKKPIGRTEPLITKIMWRNL 830
V PL VQ L VNLI+D LGALALA R LM K P+G +PLITK MW +
Sbjct: 896 VVVYDAFPLNAVQFLLVNLIIDILGALALAYRPRSDHHLMGKPPVGIRDPLITKTMWSKM 955
Query: 831 LAQALYQIAVLLVFQF-------YGKSIFNVSKEVKNTLIFNTFVLCQVFNEFNSRSMEK 883
+ Q Y + L++ +G++ ++++ NTLIFN+FV VFNEF +S+++
Sbjct: 956 IIQVFYLVLSLVLINSEKLLKLKHGQT--GNAEKMMNTLIFNSFVFYLVFNEFEIQSVDQ 1013
Query: 884 LNVFEGILKNHLFLGIVGITIVLQ 907
F+ +L+ ++FL + TI+ Q
Sbjct: 1014 --TFKEVLRENMFLVTITSTIISQ 1035
>At1g27770 envelope Ca2+-ATPase
Length = 946
Score = 387 bits (995), Expect = e-107
Identities = 237/568 (41%), Positives = 344/568 (59%), Gaps = 28/568 (4%)
Query: 20 VPHDVDKA-------RLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAAR 72
+P +V KA L ++V+ +L+ GG EG+ + L T A GI S+D + R
Sbjct: 87 LPEEVRKAGFEICPDELGSIVEGHDLKKLKIHGGTEGLTEKLSTSIASGISTSEDLLSVR 146
Query: 73 RELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSI 132
+E++G N + P + F FV EAL DTT++IL CA +SL GI G G ++G I
Sbjct: 147 KEIYGINQFTESPSRGFWLFVWEALQDTTLMILAACAFVSLIVGILMEGWPIGAHDGLGI 206
Query: 133 FLAVFLVVVVSALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKI 192
++ LVV V+A S++RQ QF L I V+V R+ Q+ISI+D+L GDV++L I
Sbjct: 207 VASILLVVFVTATSDYRQSLQFKDLDAEKKKIVVQVTRDKLRQKISIYDLLPGDVVHLGI 266
Query: 193 GDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGA 252
GDQIPADGLF+ G S+ ++ESS+TGES+ V + ++ PFLLSG KV DG +MLVT VG
Sbjct: 267 GDQIPADGLFISGFSVLINESSLTGESEPVSVS-VEHPFLLSGTKVQDGSCKMLVTTVGM 325
Query: 253 NTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDEN 312
T WG++M+++S + TPLQ +L+ + + IGKIGL A ++ +L++ +N
Sbjct: 326 RTQWGKLMATLSEGGDDETPLQVKLNGVATIIGKIGLFFA-VITFAVLVQGLANQKRLDN 384
Query: 313 GNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMV 372
+ + D A++ A AVTIVVVA+PEGLPLAVTL+LA++MK+MM D+A+V
Sbjct: 385 SHWIWTA------DELMAMLEYFAVAVTIVVVAVPEGLPLAVTLSLAFAMKKMMNDKALV 438
Query: 373 RKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLE-------NVVENFSNAMAPTVL 425
R L+ACETMGSAT IC+DKTGTLT N M V K + + + F++ + + +
Sbjct: 439 RNLAACETMGSATTICSDKTGTLTTNHMTVVKACICEQAKEVNGPDAAMKFASGIPESAV 498
Query: 426 ELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVE 485
+L Q + NT G + ++ EI G+PTE A+L + +S LG D E++Q V+ VE
Sbjct: 499 KLLLQSIFTNTGGEIVVGKG-NKTEILGTPTETALLEFGLS-LGGDFQEVRQASNVVKVE 556
Query: 486 TFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEE-RSKIEKI 544
FNS KKR GV + + H KGA+E+VL C YI+ +G LDE+ S ++ I
Sbjct: 557 PFNSTKKRMGVVIELPERH-FRAHCKGASEIVLDSCDKYINKDGEVVPLDEKSTSHLKNI 615
Query: 545 IQGMAASSLRCIAFAYMEISEGGDYIEK 572
I+ A+ +LR + AY EI G ++ EK
Sbjct: 616 IEEFASEALRTLCLAYFEI--GPEFREK 641
Score = 353 bits (906), Expect = 3e-97
Identities = 177/311 (56%), Positives = 235/311 (74%), Gaps = 5/311 (1%)
Query: 647 GVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKK-GHVVAVTGDGTNDAPALKE 705
G EFR ++EE ++ + K++VMARSSPMDK +V+ L+ VVAVTGDGTNDAPAL E
Sbjct: 635 GPEFREKSDEELLKLIPKLQVMARSSPMDKHTLVRLLRTMFQEVVAVTGDGTNDAPALHE 694
Query: 706 ADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAAL 765
ADIGL+MGI GTEVAKES+D++ILDDNF+++ TV +WGR VY NIQKF+QFQLTVNV AL
Sbjct: 695 ADIGLAMGISGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVAL 754
Query: 766 VINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKI 825
++NF++A +G+ PLT VQLLWVN+IMDTLGALALATE P +LM++ P+GR I+ +
Sbjct: 755 IVNFLSACLTGNAPLTAVQLLWVNMIMDTLGALALATEPPQDDLMKRSPVGRKGNFISNV 814
Query: 826 MWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFNEFNSRSM 881
MWRN+L Q+LYQ+ ++ Q GK++F + S NTLIFN FV CQVFNE +SR M
Sbjct: 815 MWRNILGQSLYQLVIIWCLQTKGKTMFGLDGPDSDLTLNTLIFNIFVFCQVFNEISSREM 874
Query: 882 EKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPI 941
EK++VF+GILKN++F+ ++ T+V QV+++ELL FADT LN QW + I + + P+
Sbjct: 875 EKIDVFKGILKNYVFVAVLTCTVVFQVIIIELLGTFADTTPLNLGQWLVSIILGFLGMPV 934
Query: 942 AWLTKLTPVPS 952
A K+ PV S
Sbjct: 935 AAALKMIPVGS 945
>At1g07810 ER-type Ca2+-pump protein
Length = 1061
Score = 353 bits (906), Expect = 3e-97
Identities = 269/852 (31%), Positives = 444/852 (51%), Gaps = 89/852 (10%)
Query: 65 SDDDTAARRELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGE 124
S D+ R +++G N +P +LE NDT + ILL A +S F + E
Sbjct: 47 SSDEVLKRHQIYGLNELEKPEGTSIFKLILEQFNDTLVRILLAAAVIS--FVLAFFDGDE 104
Query: 125 GWYEGGSIF---LAVFLVVVVSALSNFRQDRQFDKLSKISNDIKVE---VVRNG-RPQQI 177
G G + F L +FL+++V+A+ Q+ +K + +I+ + V+R+G + +
Sbjct: 105 GGEMGITAFVEPLVIFLILIVNAIVGIWQETNAEKALEALKEIQSQQATVMRDGTKVSSL 164
Query: 178 SIFDVLVGDVIYLKIGDQIPADG--LFLGGHSLQVDESSMTGESDHV-----------EI 224
+++ GD++ L++GD++PAD + L +L+V++ S+TGES+ V +I
Sbjct: 165 PAKELVPGDIVELRVGDKVPADMRVVALISSTLRVEQGSLTGESEAVSKTTKHVDENADI 224
Query: 225 EPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSI--SGDNSERTPLQARLDKLTS 282
+ K + +G VV+G LVT G NT G++ S I + + E TPL+ +L++
Sbjct: 225 QGKKC-MVFAGTTVVNGNCICLVTDTGMNTEIGRVHSQIQEAAQHEEDTPLKKKLNEFGE 283
Query: 283 SIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIV 342
+ I + LV L+ ++YF + + +K S + C I AV +
Sbjct: 284 VLTMIIGLICALVWLIN-VKYFLSWEYVDGWPRNFKFSF----EKCTYYFEI---AVALA 335
Query: 343 VVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRV 402
V AIPEGLP +T LA ++M A+VRKL + ET+G TVIC+DKTGTLT NQM V
Sbjct: 336 VAAIPEGLPAVITTCLALGTRKMAQKNALVRKLPSVETLGCTTVICSDKTGTLTTNQMAV 395
Query: 403 TKF--------WLGLENV-----------VENFSNAMAPTVLELFHQGVGLNTTGSVYKP 443
+K L NV +E++ L++ + + +V +
Sbjct: 396 SKLVAMGSRIGTLRSFNVEGTSFDPRDGKIEDWPMGRMDANLQMIAKIAAICNDANVEQ- 454
Query: 444 SAESEPEISGSPTEKAMLLWA--------VSDLGMDMD---------ELKQKHKVLHVET 486
++ + G PTE A+ + +++ D D EL+Q+ L
Sbjct: 455 -SDQQFVSRGMPTEAALKVLVEKMGFPEGLNEASSDGDVLRCCRLWSELEQRIATLE--- 510
Query: 487 FNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDE-ERSKIEKII 545
F+ ++K GV V + N + + KGA E VL ++ +G+++ LD+ R I + +
Sbjct: 511 FDRDRKSMGVMVDSSSGNKLLLV-KGAVENVLERSTHIQLLDGSKRELDQYSRDLILQSL 569
Query: 546 QGMAASSLRCIAFAYMEI-SEGGDYI---EKGKPRQVLR-------EDGLTLLGIVGLKD 594
+ M+ S+LRC+ FAY ++ S+ Y + +Q+L E L +G VGL+D
Sbjct: 570 RDMSLSALRCLGFAYSDVPSDFATYDGSEDHPAHQQLLNPSNYSSIESNLIFVGFVGLRD 629
Query: 595 PCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLN-DAGGVVVEGVEFRNY 653
P R V++A+ C+ AG+ + +ITGDN TA+AI E G+ + + D + G+EF +
Sbjct: 630 PPRKEVRQAIADCRTAGIRVMVITGDNKSTAEAICREIGVFEADEDISSRSLTGIEFMDV 689
Query: 654 TEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMG 713
+++ + + +R+ P K +V+ LK+ G VVA+TGDG NDAPALK ADIG++MG
Sbjct: 690 QDQKNHLRQTGGLLFSRAEPKHKQEIVRLLKEDGEVVAMTGDGVNDAPALKLADIGVAMG 749
Query: 714 IQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAV 773
I GTEVAKE+SD+V+ DDNF+++ + GR +YNN++ FI++ ++ N+ + F+ A
Sbjct: 750 ISGTEVAKEASDMVLADDNFSTIVAAVGEGRSIYNNMKAFIRYMISSNIGEVASIFLTAA 809
Query: 774 SSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITK-IMWRNLLA 832
+ VQLLWVNL+ D A AL P K++M+K P + LIT I++R ++
Sbjct: 810 LGIPEGMIPVQLLWVNLVTDGPPATALGFNPPDKDIMKKPPRRSDDSLITAWILFRYMVI 869
Query: 833 QALYQIAVLLVF 844
+A + VF
Sbjct: 870 GLYVGVATVGVF 881
>At1g07670 endoplasmic reticulum-type calcium-transporting ATPase 4
(ECA4)
Length = 1061
Score = 352 bits (903), Expect = 6e-97
Identities = 277/888 (31%), Positives = 451/888 (50%), Gaps = 95/888 (10%)
Query: 30 SNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIF 89
S +VK A+ + V + G KG+ S D+ R +++G N +P
Sbjct: 16 SELVKSDTFPAWGK--DVSECEEKFGVSREKGL--STDEVLKRHQIYGLNELEKPEGTSI 71
Query: 90 LHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIF---LAVFLVVVVSALS 146
+LE NDT + ILL A +S F + EG G + F L +FL+++V+A+
Sbjct: 72 FKLILEQFNDTLVRILLAAAVIS--FVLAFFDGDEGGEMGITAFVEPLVIFLILIVNAIV 129
Query: 147 NFRQDRQFDKLSKISNDIKVE---VVRNG-RPQQISIFDVLVGDVIYLKIGDQIPADG-- 200
Q+ +K + +I+ + V+R+G + + +++ GD++ L++GD++PAD
Sbjct: 130 GIWQETNAEKALEALKEIQSQQATVMRDGTKVSSLPAKELVPGDIVELRVGDKVPADMRV 189
Query: 201 LFLGGHSLQVDESSMTGESDHV-----------EIEPLKAPFLLSGAKVVDGYAQMLVTA 249
+ L +L+V++ S+TGES+ V +I+ K + +G VV+G LVT
Sbjct: 190 VALISSTLRVEQGSLTGESEAVSKTTKHVDENADIQGKKC-MVFAGTTVVNGNCICLVTD 248
Query: 250 VGANTAWGQMMSSI--SGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGN 307
G NT G++ S I + + E TPL+ +L++ + I + LV L+ ++YF
Sbjct: 249 TGMNTEIGRVHSQIQEAAQHEEDTPLKKKLNEFGEVLTMIIGLICALVWLIN-VKYFLSW 307
Query: 308 TEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMA 367
+ + +K S + C I AV + V AIPEGLP +T LA ++M
Sbjct: 308 EYVDGWPRNFKFSF----EKCTYYFEI---AVALAVAAIPEGLPAVITTCLALGTRKMAQ 360
Query: 368 DQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKF--------WLGLENV------- 412
A+VRKL + ET+G TVIC+DKTGTLT NQM V+K L NV
Sbjct: 361 KNALVRKLPSVETLGCTTVICSDKTGTLTTNQMAVSKLVAMGSRIGTLRSFNVEGTSFDP 420
Query: 413 ----VENFSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDL 468
+E++ L++ + + +V K ++ + G PTE A+ + V +
Sbjct: 421 RDGKIEDWPTGRMDANLQMIAKIAAICNDANVEK--SDQQFVSRGMPTEAALKV-LVEKM 477
Query: 469 GMD------------------MDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHW 510
G EL+Q+ L F+ ++K GV V + + +
Sbjct: 478 GFPEGLNEASSDGNVLRCCRLWSELEQRIATLE---FDRDRKSMGVMVDSSSGKKLLLV- 533
Query: 511 KGAAEMVLAMCSNYIDSNGTQKSLDE-ERSKIEKIIQGMAASSLRCIAFAYMEI-SEGGD 568
KGA E VL ++ +G+ + LD+ R I + + M+ S+LRC+ FAY ++ S+
Sbjct: 534 KGAVENVLERSTHIQLLDGSTRELDQYSRDLILQSLHDMSLSALRCLGFAYSDVPSDFAT 593
Query: 569 YI---EKGKPRQVLR-------EDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMIT 618
Y + +Q+L E L +G VGL+DP R V++A+ C+ AG+ + +IT
Sbjct: 594 YDGSEDHPAHQQLLNPSNYSSIESNLVFVGFVGLRDPPRKEVRQAIADCRTAGIRVMVIT 653
Query: 619 GDNIFTAKAIATECGILDLN-DAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKL 677
GDN TA+AI E G+ + + D + G EF + +++ + + +R+ P K
Sbjct: 654 GDNKSTAEAICREIGVFEADEDISSRSLTGKEFMDVKDQKNHLRQTGGLLFSRAEPKHKQ 713
Query: 678 LMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVA 737
+V+ LK+ G VVA+TGDG NDAPALK ADIG++MGI GTEVAKE+SD+V+ DDNF+++
Sbjct: 714 EIVRLLKEDGEVVAMTGDGVNDAPALKLADIGVAMGISGTEVAKEASDLVLADDNFSTIV 773
Query: 738 TVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGA 797
+ GR +YNN++ FI++ ++ N+ + F+ A + VQLLWVNL+ D A
Sbjct: 774 AAVGEGRSIYNNMKAFIRYMISSNIGEVASIFLTAALGIPEGMIPVQLLWVNLVTDGPPA 833
Query: 798 LALATERPTKELMQKKPIGRTEPLITK-IMWRNLLAQALYQIAVLLVF 844
AL P K++M+K P + LIT I++R ++ +A + VF
Sbjct: 834 TALGFNPPDKDIMKKPPRRSDDSLITAWILFRYMVIGLYVGVATVGVF 881
>At4g00900 Ca2+-transporting ATPase - like protein
Length = 1054
Score = 346 bits (887), Expect = 4e-95
Identities = 284/891 (31%), Positives = 447/891 (49%), Gaps = 92/891 (10%)
Query: 32 MVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLH 91
M ++K+ A+S VE T KG+ + +D RR+ +G N + K H
Sbjct: 1 MEEEKSFSAWS--WSVEQCLKEYKTRLDKGL--TSEDVQIRRQKYGFNELAKEKGKPLWH 56
Query: 92 FVLEALNDTTILILLGCAGLS--LGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFR 149
VLE +DT + ILLG A +S L F +EHG G G+ F+ V L+++++A+
Sbjct: 57 LVLEQFDDTLVKILLGAAFISFVLAFLGEEHGSGSGFEAFVEPFVIV-LILILNAVVGVW 115
Query: 150 QDRQFDKLSKISNDIKVE---VVRNGRP-QQISIFDVLVGDVIYLKIGDQIPADGLFLG- 204
Q+ +K + +++ E V+R+G + +++ GD++ L +GD++PAD G
Sbjct: 116 QESNAEKALEALKEMQCESAKVLRDGNVLPNLPARELVPGDIVELNVGDKVPADMRVSGL 175
Query: 205 -GHSLQVDESSMTGES------------DHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVG 251
+L+V++SS+TGE+ D E++ K + +G VV+G +VT++G
Sbjct: 176 KTSTLRVEQSSLTGEAMPVLKGANLVVMDDCELQG-KENMVFAGTTVVNGSCVCIVTSIG 234
Query: 252 ANTAWGQMMSSISGDNSER--TPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTE 309
+T G++ I + E TPL+ +LD+ S ++ A+ + +LV +I Y +
Sbjct: 235 MDTEIGKIQRQIHEASLEESETPLKKKLDEFGS---RLTTAICIVCVLVWMINYKNFVSW 291
Query: 310 DENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQ 369
D + K + C I AV + V AIPEGLP +T LA ++M
Sbjct: 292 DVVDGYKPVNIKFSF-EKCTYYFKI---AVALAVAAIPEGLPAVITTCLALGTRKMAQKN 347
Query: 370 AMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWL---------------------- 407
A+VRKL + ET+G TVIC+DKTGTLT NQM T+F+
Sbjct: 348 AIVRKLPSVETLGCTTVICSDKTGTLTTNQMSATEFFTLGGKTTTTRVFSVSGTTYDPKD 407
Query: 408 -GLENVVENFSNAMAPTVLEL---------FHQGVGLNTTGSVYKPSAESEPEISGSP-- 455
G+ + N +A V E+ F++G TG + + + E G P
Sbjct: 408 GGIVDWGCNNMDANLQAVAEICSICNDAGVFYEGKLFRATGLPTEAALKVLVEKMGIPEK 467
Query: 456 --TEKAMLLWAVSDLGMDM-----DELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHV 508
+E + SD G + D ++ K + F+ +K V V E N +
Sbjct: 468 KNSENIEEVTNFSDNGSSVKLACCDWWNKRSKKVATLEFDRVRKSMSVIV-SEPNGQNRL 526
Query: 509 HWKGAAEMVLAMCSNYIDSNGTQKSLDEE-RSKIEKIIQGMAASSLRCIAFAYM-EISEG 566
KGAAE +L S ++G+ +LDE R I K M + LRC+ AY E+ E
Sbjct: 527 LVKGAAESILERSSFAQLADGSLVALDESSREVILKKHSEMTSKGLRCLGLAYKDELGEF 586
Query: 567 GDYIEKGKP--RQVLR-------EDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMI 617
DY + P +++L E L +G+VGL+DP R V +A+E C+ AG+ + +I
Sbjct: 587 SDYSSEEHPSHKKLLDPSSYSNIETNLIFVGVVGLRDPPREEVGRAIEDCRDAGIRVMVI 646
Query: 618 TGDNIFTAKAIATECGILDLN-DAGGVVVEGVEFRNYTEEERMEKVDKI--RVMARSSPM 674
TGDN TA+AI E + N D G EF + R E + K +V +R+ P
Sbjct: 647 TGDNKSTAEAICCEIRLFSENEDLSQSSFTGKEFMSLPASRRSEILSKSGGKVFSRAEPR 706
Query: 675 DKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFN 734
K +V+ LK+ G +VA+TGDG NDAPALK ADIG++MGI GTEVAKE+SD+V+ DDNF+
Sbjct: 707 HKQEIVRMLKEMGEIVAMTGDGVNDAPALKLADIGIAMGITGTEVAKEASDMVLADDNFS 766
Query: 735 SVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDT 794
++ + + GR +YNN++ FI++ ++ NV ++ F+ A + VQLLWVNL+ D
Sbjct: 767 TIVSAVAEGRSIYNNMKAFIRYMISSNVGEVISIFLTAALGIPECMIPVQLLWVNLVTDG 826
Query: 795 LGALALATERPTKELMQKKPIGRTEPLITK-IMWRNLLAQALYQIAVLLVF 844
A AL ++M+K P + LI ++ R L+ + +A + +F
Sbjct: 827 PPATALGFNPADIDIMKKPPRKSDDCLIDSWVLIRYLVIGSYVGVATVGIF 877
>At1g10130 putative calcium ATPase
Length = 992
Score = 326 bits (835), Expect = 4e-89
Identities = 284/1003 (28%), Positives = 461/1003 (45%), Gaps = 136/1003 (13%)
Query: 39 EAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLHFVLEALN 98
+AY+ V V D G P KG+ SD L+G N F VL+ +
Sbjct: 3 DAYAR--SVSEVLDFFGVDPTKGL--SDSQVVHHSRLYGRNGTP------FWKLVLKQFD 52
Query: 99 DTTILILLGCAGLSLGFGIKEHGPG-EGWYEGGSIFLAVFLVVVVSALSNFRQDRQFDKL 157
D + IL+ A +S + G + E I L + V ++ ++ ++L
Sbjct: 53 DLLVKILIVAAIVSFVLALANGETGLTAFLEPFVILLILAANAAVGVITETNAEKALEEL 112
Query: 158 SKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPAD--GLFLGGHSLQVDESSM 215
+I V+RNG + +++ GD++ + +G +IPAD + + ++ +VD++ +
Sbjct: 113 RAYQANIAT-VLRNGCFSILPATELVPGDIVEVTVGCKIPADLRMIEMSSNTFRVDQAIL 171
Query: 216 TGESDHVE-----------IEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSIS 264
TGES VE + K L SG VV G + +V VG+NTA G + S+
Sbjct: 172 TGESCSVEKDVDCTLTTNAVYQDKKNILFSGTDVVAGRGRAVVIGVGSNTAMGSIHDSML 231
Query: 265 GDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTDI 324
+ E TPL+ +LD+ S + K+ + LV +V + G+ D + +KG+
Sbjct: 232 QTDDEATPLKKKLDEFGSFLAKVIAGICVLVWVVNI-----GHFSDPSHGGFFKGA---- 282
Query: 325 NDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSA 384
+ AV + V AIPEGLP VT LA K+M A+VR L + ET+G
Sbjct: 283 -------IHYFKIAVALAVAAIPEGLPAVVTTCLALGTKKMARLNAIVRSLPSVETLGCT 335
Query: 385 TVICTDKTGTLTLNQMRVTKFWLGL---------ENVVENFSNAMAPTVLELFHQGVGL- 434
TVIC+DKTGTLT N M V+K + E V + A TV + + L
Sbjct: 336 TVICSDKTGTLTTNMMSVSKICVVQSAEHGPMINEFTVSGTTYAPEGTVFDSNGMQLDLP 395
Query: 435 ----------------NTTGSVYKPSAESEPEISGSPTEKAMLLWA--VSDLGMDMD--- 473
N + Y P +S +I G TE A+ + A V G D
Sbjct: 396 AQSPCLHHLAMCSSLCNDSILQYNPDKDSYEKI-GESTEVALRVLAEKVGLPGFDSMPSA 454
Query: 474 -ELKQKH--------------KVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVL 518
+ KH K ++V F ++K V + + + KGA E ++
Sbjct: 455 LNMLSKHERASYCNHYWENQFKKVYVLEFTRDRKMMSVLCSHKQMDVMFS--KGAPESII 512
Query: 519 AMCSNYI-DSNGTQKSLDEE-RSKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPR 576
A C+ + + +G+ L R+++E +LRC+A A+ + G I
Sbjct: 513 ARCNKILCNGDGSVVPLTAAGRAELESRFYSFGDETLRCLALAFKTVPHGQQTISYDN-- 570
Query: 577 QVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILD 636
E+ LT +G+VG+ DP R V+ A+ C AG+ + ++TGDN TA+++ + G D
Sbjct: 571 ----ENDLTFIGLVGMLDPPREEVRDAMLACMTAGIRVIVVTGDNKSTAESLCRKIGAFD 626
Query: 637 -LNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGD 695
L D G+ EF ++ + ++ + +R P K ++V+ L+K+ VVA+TGD
Sbjct: 627 NLVDFSGMSYTASEFERLPAVQQTLALRRMTLFSRVEPSHKRMLVEALQKQNEVVAMTGD 686
Query: 696 GTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQ 755
G NDAPALK+ADIG++MG GT VAK +SD+V+ DDNF S+ + GR +YNN ++FI+
Sbjct: 687 GVNDAPALKKADIGIAMG-SGTAVAKSASDMVLADDNFASIVAAVAEGRAIYNNTKQFIR 745
Query: 756 FQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPI 815
+ ++ N+ +V F+AAV L VQLLWVNL+ D L A A+ + ++M+ KP
Sbjct: 746 YMISSNIGEVVCIFVAAVLGIPDTLAPVQLLWVNLVTDGLPATAIGFNKQDSDVMKAKPR 805
Query: 816 GRTEPLITKIMWRNLLAQALY----------------------QIAVLLVFQ-------F 846
E ++T ++ L +Y + L+ F+
Sbjct: 806 KVGEAVVTGWLFFRYLVIGVYVGLATVAGFIWWFVYSDGGPKLTYSELMNFETCALRETT 865
Query: 847 YGKSIFNVSKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVL 906
Y SIF +T+ V+ ++FN N+ S + + N +G + +T++L
Sbjct: 866 YPCSIF--EDRHPSTVAMTVLVVVEMFNALNNLSENQSLLVITPRSNLWLVGSIILTMLL 923
Query: 907 QVLM--VELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKL 947
VL+ V L L+W +W + +S+P+ + +L
Sbjct: 924 HVLILYVHPLAVLFSVTPLSWAEW---TAVLYLSFPVIIIDEL 963
>At1g17260 H+-transporting ATPase AHA10
Length = 946
Score = 201 bits (510), Expect = 2e-51
Identities = 205/840 (24%), Positives = 376/840 (44%), Gaps = 109/840 (12%)
Query: 47 VEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLHFVLEALNDTTILILL 106
+E V + L T P +G+L D + R ++FG N F+ F+ N + ++
Sbjct: 27 LEEVFEYLRTSP-QGLLSGDAEE--RLKIFGPNRLEEKQENRFVKFLGFMWNPLS-WVME 82
Query: 107 GCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQDRQFDKLSKISNDI-- 164
A +++ G W + I V L+++ + +S F ++ + + + +
Sbjct: 83 AAALMAIALA-NSQSLGPDWEDFTGI---VCLLLINATISFFEENNAGNAAAALMARLAL 138
Query: 165 KVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEI 224
K V+R+G+ Q+ ++ GD+I +K+GD IPAD L G L++D+S +TGES + +
Sbjct: 139 KTRVLRDGQWQEQDASILVPGDIISIKLGDIIPADARLLEGDPLKIDQSVLTGES--LPV 196
Query: 225 EPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSI 284
K + SG+ G + +V A G+ T +G+ + + T + ++ +SI
Sbjct: 197 TKKKGEQVFSGSTCKQGEIEAVVIATGSTTFFGKTARLV-----DSTDVTGHFQQVLTSI 251
Query: 285 GKIGL-AVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVV 343
G + ++A ++L ++I + +++ + IN++ + +++
Sbjct: 252 GNFCICSIAVGMVLEIIIMFPV----------QHRSYRIGINNL-----------LVLLI 290
Query: 344 VAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVT 403
IP +P +++TLA R+ A+ ++++A E M V+C DKTGTLTLN + V
Sbjct: 291 GGIPIAMPTVLSVTLAIGSHRLSQQGAITKRMTAIEEMAGMDVLCCDKTGTLTLNSLTVD 350
Query: 404 KFWLGLENVVENFSNAM-APTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLL 462
K N++E F + M T+L L + L E++ I
Sbjct: 351 K------NLIEVFVDYMDKDTILLLAGRASRL-----------ENQDAIDA--------- 384
Query: 463 WAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCS 522
A+ + D E + + +H FN KR+ + +++ + KGA E VL +C
Sbjct: 385 -AIVSMLADPREARANIREIHFLPFNPVDKRTAITY-IDSDGKWYRATKGAPEQVLNLC- 441
Query: 523 NYIDSNGTQKSLDEERSKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLRED 582
+ +E ++ II A LR +A AY EI E + G R
Sbjct: 442 ---------QQKNEIAQRVYAIIDRFAEKGLRSLAVAYQEIPEKSNNSPGGPWR------ 486
Query: 583 GLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGG 642
G++ L DP R + + + GV +KMITGD + AK G+
Sbjct: 487 ---FCGLLPLFDPPRHDSGETILRALSLGVCVKMITGDQLAIAKETGRRLGM-----GTN 538
Query: 643 VVVEGVEFRNYTEEERMEKVDKIRVMARS----SPMDKLLMVQCLKKKGHVVAVTGDGTN 698
+ + +E VD++ MA P K +V+ L++ HVV +TGDG N
Sbjct: 539 MYPSSSLLGHNNDEHEAIPVDELIEMADGFAGVFPEHKYEIVKILQEMKHVVGMTGDGVN 598
Query: 699 DAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQL 758
DAPALK+ADIG+++ T+ A+ S+DIV+ D + + + + R ++ ++ + + +
Sbjct: 599 DAPALKKADIGIAVA-DATDAARSSADIVLTDPGLSVIISAVLTSRAIFQRMRNYTVYAV 657
Query: 759 TVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATER----PTKELMQKKP 814
++ + L +A + D P V ++ I++ + ++ +R PT E +
Sbjct: 658 SITIRILGFTLLALIWEYDFPPFMVLII---AILNDGTIMTISKDRVRPSPTPESWKLNQ 714
Query: 815 IGRTEPLITKIMWRNLLAQALYQIAVLLVF---QFYGKSIFNVSKEVKNTLIFNTFVLCQ 871
I T +I + L+ Y I V F F+ KSI N S++V + + ++ Q
Sbjct: 715 IFATGIVIGTYL--ALVTVLFYWIIVSTTFFEKHFHVKSIANNSEQVSSAMYLQVSIISQ 772
>At3g60330 plasma membrane H+-ATPase - like
Length = 961
Score = 182 bits (461), Expect = 1e-45
Identities = 153/644 (23%), Positives = 295/644 (45%), Gaps = 94/644 (14%)
Query: 136 VFLVVVVSALSNFRQDRQFDKLSKISNDI--KVEVVRNGRPQQISIFDVLVGDVIYLKIG 193
V L+++ S +S ++ + + + + K + VR+G+ +I +++ GD++ +K+G
Sbjct: 103 VVLLLINSTISFVEENNAGNAAAALMAQLAPKAKAVRDGKWNEIDAAELVPGDIVSIKLG 162
Query: 194 DQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGAN 253
D IPAD L G L++D++++TGES V P + + SG+ G + +V A G +
Sbjct: 163 DIIPADARLLEGDPLKIDQATLTGESLPVTKNPGASVY--SGSTCKQGEIEAVVIATGVH 220
Query: 254 TAWGQMMSSISGDNSERTPLQARLDKLTSSIGK-------IGLAVAFLVLLVLLIRYFTG 306
T +G+ + + T K+ ++IG +G+A+ +V+ L
Sbjct: 221 TFFGKAAHLV-----DSTTHVGHFQKVLTAIGNFCICSIAVGMAIEIVVIYGL------- 268
Query: 307 NTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMM 366
+ +G + I+++ + +++ IP +P +++T+A R+
Sbjct: 269 ---------QKRGYRVGIDNL-----------LVLLIGGIPIAMPTVLSVTMAIGAHRLA 308
Query: 367 ADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLE 426
A+ ++++A E M V+C+DKTGTLTLN++ V K ++E
Sbjct: 309 QQGAITKRMTAIEEMAGMDVLCSDKTGTLTLNKLSVDK------------------NLIE 350
Query: 427 LFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVET 486
+F +G+ + + +A E + + +ML D E + K LH
Sbjct: 351 VFKRGIDRDMAVLMAARAARLENQDAIDTAIVSML--------SDPKEARAGIKELHFLP 402
Query: 487 FNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQ 546
F+ +R+ + + +H KGA E +L M N + E + K+ I
Sbjct: 403 FSPANRRTALTY-LDGEGKMHRVSKGAPEEILDMAHNKL----------EIKEKVHATID 451
Query: 547 GMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVET 606
A LR + AY E+ + GD +G P + ++ L DP R + + +E
Sbjct: 452 KFAERGLRSLGLAYQEVPD-GDVKGEGGP--------WDFVALLPLFDPPRHDSAQTIER 502
Query: 607 CKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVDKIR 666
GV +KMITGD + AK G+ ++ + +E +E D
Sbjct: 503 ALHLGVSVKMITGDQLAIAKETGRRLGMGTNMYPSSSLLSDNNTEGVSVDELIENADG-- 560
Query: 667 VMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDI 726
A P K +V+ L+ + H+ +TGDG NDAPALK+ADIG+++ T+ A+ +SDI
Sbjct: 561 -FAGVFPEHKYEIVKRLQSRKHICGMTGDGVNDAPALKKADIGIAVD-DATDAARGASDI 618
Query: 727 VILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFI 770
V+ + + + + + R ++ ++ + + +++ + +V+ F+
Sbjct: 619 VLTEPGLSVIISAVLTSRAIFQRMKNYTIYAVSITI-RIVMGFM 661
>At2g24520 putative plasma membrane proton ATPase
Length = 931
Score = 181 bits (458), Expect = 2e-45
Identities = 150/599 (25%), Positives = 271/599 (45%), Gaps = 76/599 (12%)
Query: 165 KVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEI 224
K +V+R+ + + ++ GDVI +K+GD IPAD L G L++D+SS+TGES V
Sbjct: 113 KTKVLRDNQWSEQEASILVPGDVISIKLGDIIPADARLLDGDPLKIDQSSLTGESIPVTK 172
Query: 225 EPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSI 284
P F SG+ G + +V A G +T +G+ + N K+ +SI
Sbjct: 173 NPSDEVF--SGSICKQGEIEAIVIATGVHTFFGKAAHLVDNTNQI-----GHFQKVLTSI 225
Query: 285 GKIGL-AVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVV 343
G + ++A +++ LL+ Y +G + + +++
Sbjct: 226 GNFCICSIALGIIVELLVMYPIQRRRYRDG---------------------IDNLLVLLI 264
Query: 344 VAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVT 403
IP +P +++T+A R+ A+ ++++A E M V+C DKTGTLTLN++ V
Sbjct: 265 GGIPIAMPSVLSVTMATGSHRLFQQGAITKRMTAIEEMAGMDVLCCDKTGTLTLNKLTVD 324
Query: 404 KFWLGLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLW 463
K ++E+F +GVG + ++ E + + ML
Sbjct: 325 K------------------NLVEVFAKGVGKEHVFLLAARASRIENQDAIDAAIVGML-- 364
Query: 464 AVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSN 523
D E + + +H FN KR+ + +++ H KGA E +L +C+
Sbjct: 365 ------ADPKEARAGVREVHFFPFNPVDKRTALTY-VDSDGNWHRASKGAPEQILNLCN- 416
Query: 524 YIDSNGTQKSLDEERSKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDG 583
++ R K+ +I A LR +A A E+ E G P Q
Sbjct: 417 ---------CKEDVRRKVHGVIDKFAERGLRSLAVARQEVLE-KKKDAPGGPWQ------ 460
Query: 584 LTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGV 643
L+G++ L DP R + + + GV++KMITGD + K G+
Sbjct: 461 --LVGLLPLFDPPRHDSAETIRRALNLGVNVKMITGDQLAIGKETGRRLGMGTNMYPSSA 518
Query: 644 VVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPAL 703
++ V+ + E ++K A P K +V L+++ H+ +TGDG NDAPAL
Sbjct: 519 LLGQVKDSSLGALPVDELIEKADGFAGVFPEHKYEIVHRLQQRNHICGMTGDGVNDAPAL 578
Query: 704 KEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNV 762
K+ADIG+++ + T+ A+ +SDIV+ + + + + + R ++ ++ + + +++ +
Sbjct: 579 KKADIGIAV-VDATDAARGASDIVLTEPGLSVIISAVLTSRAIFQRMKNYTIYAVSITI 636
>At2g07560 pseudogene
Length = 949
Score = 179 bits (454), Expect = 7e-45
Identities = 161/671 (23%), Positives = 306/671 (44%), Gaps = 104/671 (15%)
Query: 138 LVVVVSALSNFRQDRQFDKLSKISNDI--KVEVVRNGRPQQISIFDVLVGDVIYLKIGDQ 195
L+++ S +S ++ + + + ++ K +V+R+GR + ++ GD+I +K+GD
Sbjct: 105 LLIINSTISFIEENNAGNAAAALMANLAPKTKVLRDGRWGEQEAAILVPGDLISIKLGDI 164
Query: 196 IPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTA 255
+PAD L G L++D+S++TGES + + + SG+ G + +V A G +T
Sbjct: 165 VPADARLLEGDPLKIDQSALTGES--LPATKHQGDEVFSGSTCKQGEIEAVVIATGVHTF 222
Query: 256 WGQMMSSISGDNSERTPLQARLDKLTSSIGKIGL-AVAFLVLLVLLIRYFTGNTEDENGN 314
+G+ + N+ K+ ++IG + ++ +L+ ++I Y + + +G
Sbjct: 223 FGKAAHLVDSTNNV-----GHFQKVLTAIGNFCICSIGIGMLIEIIIMYPIQHRKYRDG- 276
Query: 315 KEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRK 374
+ + +++ IP +P +++T+A R+ A+ ++
Sbjct: 277 --------------------IDNLLVLLIGGIPIAMPTVLSVTMAIGSHRLSQQGAITKR 316
Query: 375 LSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQGVGL 434
++A E M V+C+DKTGTLTLN++ V K N++E FS + + L
Sbjct: 317 MTAIEEMAGMDVLCSDKTGTLTLNKLTVDK------NLIEVFSKDVDKDYVILL------ 364
Query: 435 NTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRS 494
S E++ I S V+ LG D E + +H FN +KR+
Sbjct: 365 ----SARASRVENQDAIDTS---------IVNMLG-DPKEARAGITEVHFLPFNPVEKRT 410
Query: 495 GVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAASSLR 554
+ +TN H KGA E ++ +C D G E + + +II A LR
Sbjct: 411 AITY-IDTNGEWHRCSKGAPEQIIELC----DLKG------ETKRRAHEIIDKFAERGLR 459
Query: 555 CIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDI 614
+ A + E D G P + +G++ L DP R + + + GV++
Sbjct: 460 SLGVARQRVPE-KDKESAGTPWE--------FVGLLPLFDPPRHDSAETIRRALDLGVNV 510
Query: 615 KMITGDNIFTAKAIATECG----------ILDLND--AGGVVVEGVEFRNYTEEERMEKV 662
KMITGD + K G +L+ D GGV V+ E +
Sbjct: 511 KMITGDQLAIGKETGRRLGMGTNMYPSSSLLENKDDTTGGVPVD-------------ELI 557
Query: 663 DKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKE 722
+K A P K +V+ L+++ H+V +TGDG NDAPALK+ADIG+++ T+ A+
Sbjct: 558 EKADGFAGVFPEHKYEIVRKLQERKHIVGMTGDGVNDAPALKKADIGIAVD-DATDAARS 616
Query: 723 SSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTT 782
+SDIV+ + + + + + R ++ ++ + + +++ + +V+ F+ + +
Sbjct: 617 ASDIVLTEPGLSVIVSAVLTSRAIFQRMKNYTIYAVSITI-RIVLGFMLVALIWEFDFSP 675
Query: 783 VQLLWVNLIMD 793
+L + ++ D
Sbjct: 676 FMVLIIAILND 686
>At3g47950 H+-transporting ATPase - like protein
Length = 960
Score = 177 bits (449), Expect = 3e-44
Identities = 156/650 (24%), Positives = 294/650 (45%), Gaps = 103/650 (15%)
Query: 136 VFLVVVVSALSNFRQDRQFDKLSKISNDI--KVEVVRNGRPQQISIFDVLVGDVIYLKIG 193
+ L+V+ S +S ++ + + + + K +V+R+GR + ++ GD+I +K+G
Sbjct: 108 ITLLVINSTISFIEENNAGNAAAALMARLAPKAKVLRDGRWGEQDAAILVPGDIISIKLG 167
Query: 194 DQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGAN 253
D +PAD L G L++D+S++TGES + + + SG+ G + +V A G +
Sbjct: 168 DIVPADARLLEGDPLKIDQSALTGES--LPVTKSSGDGVYSGSTCKQGEIEAVVIATGVH 225
Query: 254 TAWGQMMSSISGDNSERTPLQARLDKLTSSIGKI---GLAVAFLVLLVLLIRYFTGNTED 310
T +G+ + N ++ ++IG +AV L+ +V++
Sbjct: 226 TFFGKAAHLVDTTNQI-----GHFQQVLTAIGNFCICSIAVGMLIEIVVMYPI------- 273
Query: 311 ENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQA 370
+++ + I+++ + +++ IP +P +++T+A R+ A
Sbjct: 274 -----QHRAYRPGIDNL-----------LVLLIGGIPIAMPTVLSVTMAIGSHRLSQQGA 317
Query: 371 MVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQ 430
+ ++++A E M V+C+DKTGTLTLN++ V K ++E+F +
Sbjct: 318 ITKRMTAIEEMAGMDVLCSDKTGTLTLNKLTVDK------------------NLIEVFMK 359
Query: 431 GVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSE 490
GV +T + ++ E + + ML D + + + +H FN
Sbjct: 360 GVDADTVVLMAARASRLENQDAIDAAIVGML--------ADPKDARAGIQEVHFLPFNPT 411
Query: 491 KKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAA 550
KR+ + NT V KGA E +L + N KS E R + +I A
Sbjct: 412 DKRTALTYIDNEGNTHRVS-KGAPEQILNLAHN--------KSEIERR--VHAVIDKFAE 460
Query: 551 SSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLA 610
LR +A AY ++ EG G P Q +G++ L DP R + + +
Sbjct: 461 RGLRSLAVAYQDVPEGRK-DSAGGPWQ--------FVGLMPLFDPPRHDSAETIRRALNL 511
Query: 611 GVDIKMITGDNIFTAKAIATECG----------ILDLNDAGGVVVEGVEFRNYTEEERME 660
GV +KMITGD + K G +L N +V V+ E
Sbjct: 512 GVSVKMITGDQLAIGKETGRRLGMGTNMYPSSALLGQNKDESIVALPVD----------E 561
Query: 661 KVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVA 720
++K A P K +V+ L+ + H+ +TGDG NDAPALK+ADIG+++ T+ A
Sbjct: 562 LIEKADGFAGVFPEHKYEIVKRLQARKHICGMTGDGVNDAPALKKADIGIAVA-DATDAA 620
Query: 721 KESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFI 770
+ +SDIV+ + + + + + R ++ ++ + + +++ + +V+ F+
Sbjct: 621 RSASDIVLTEPGLSVIISAVLTSRAIFQRMKNYTIYAVSITI-RIVLGFM 669
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,691,895
Number of Sequences: 26719
Number of extensions: 881804
Number of successful extensions: 2479
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 50
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 2194
Number of HSP's gapped (non-prelim): 110
length of query: 967
length of database: 11,318,596
effective HSP length: 109
effective length of query: 858
effective length of database: 8,406,225
effective search space: 7212541050
effective search space used: 7212541050
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0322.1


