

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0321.6
(497 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
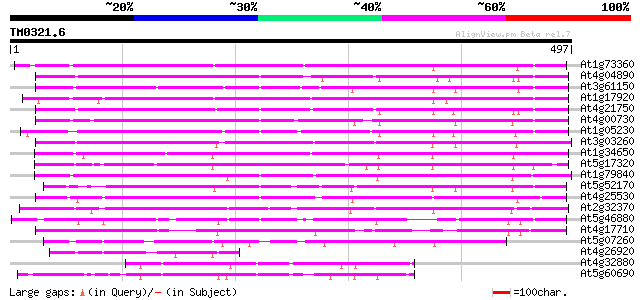
Score E
Sequences producing significant alignments: (bits) Value
At1g73360 putative homeobox protein 290 2e-78
At4g04890 putative homeotic protein 281 8e-76
At3g61150 homeobox protein 281 8e-76
At1g17920 unknown protein 280 2e-75
At4g21750 L1 specific homeobox gene ATML1/ovule-specific homeobo... 272 3e-73
At4g00730 homeodomain protein AHDP 269 2e-72
At1g05230 homeobox like protein 264 7e-71
At3g03260 unknown protein 261 5e-70
At1g34650 hypothetical protein 243 1e-64
At5g17320 homeobox protein 243 2e-64
At1g79840 homeobox protein (GLABRA2) 231 9e-61
At5g52170 homeodomain transcription factor-like 206 2e-53
At4g25530 homeodomain-containing transcription factor FWA 206 3e-53
At2g32370 putative homeodomain transcription factor 204 1e-52
At5g46880 homeobox protein 192 5e-49
At4g17710 GLABRA2 like protein 165 5e-41
At5g07260 unknown protein 113 3e-25
At4g26920 putative homeodomain protein 58 1e-08
At4g32880 HD-zip transcription factor (athb-8) 50 2e-06
At5g60690 REVOLUTA or interfascicular fiberless 1 48 1e-05
>At1g73360 putative homeobox protein
Length = 722
Score = 290 bits (741), Expect = 2e-78
Identities = 177/508 (34%), Positives = 287/508 (55%), Gaps = 28/508 (5%)
Query: 5 NDNVVNNPTRESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVR 64
N+N+ + P + ++K M +A AM+ELL LL T+EPLW ++ D +L
Sbjct: 215 NNNLQSQPNLAISD----MDKPIMTGIALTAMEELLRLLQTNEPLWTRT--DGCRDILNL 268
Query: 65 EIYAKKFPRITN-SENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKT 123
Y FPR +N +N E+S+ S +V + M LV +F+D KW ++ P+I++ +KT
Sbjct: 269 GSYENVFPRSSNRGKNQNFRVEASRSSGIVFMNAMALVDMFMDCVKWTELFPSIIAASKT 328
Query: 124 IQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFG 183
+ V G H GAL L++ E+ VLSPLV +RE+ LRYCQQ E G W++V+VS D
Sbjct: 329 LAVISSGMGGTHEGALHLLYEEMEVLSPLVATREFCELRYCQQTEQGSWIVVNVSYD-LP 387
Query: 184 NCASRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRW 243
S S + +FPSGC I+++PNG VTWVE +E + + L R+I+ IA+GA RW
Sbjct: 388 QFVSHSQSYRFPSGCLIQDMPNGYSKVTWVEHIETEEKELVHELYREIIHRGIAFGADRW 447
Query: 244 LLELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQN 303
+ LQR E+ A +S+ L G + +G++S++ L+ RM+ ++ ++
Sbjct: 448 VTTLQRMCERFASLSVP-ASSSRDLGGVILSPEGKRSMMRLAQRMISNYCLSVSRSNNTR 506
Query: 304 LLDPSNQLTTSIKLIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAG 363
S I++ AH++ +G V+ A ++ WLP +P+N+F FLKDE + +WD L+
Sbjct: 507 STVVSELNEVGIRVTAHKSPEPNGTVLCAATTFWLPNSPQNVFNFLKDERTRPQWDVLSN 566
Query: 364 GRPMRQKAHIS--INESNHVSILQTSSII-GEHMVIFQESSIDDLGGYVVYAPIDKLDAY 420
G +++ AHIS + N +S+L+ S+ +M+I QESS D G +VVY+P+D
Sbjct: 567 GNAVQEVAHISNGSHPGNCISVLRGSNATHSNNMLILQESSTDSSGAFVVYSPVDLAALN 626
Query: 421 MIVNGCDSSKKLILPSGFTISRDGREGN---------------EGSLLTMCFQVLVDEEG 465
+ ++G D S +L SGFTIS DG N GSL+T+ FQ++V
Sbjct: 627 IAMSGEDPSYIPLLSSGFTISPDGNGSNSEQGGASTSSGRASASGSLITVGFQIMVSNL- 685
Query: 466 SSSQVRKKTLESIDSSCMLMLQRIKDAL 493
++++ +++E++++ + +IK AL
Sbjct: 686 PTAKLNMESVETVNNLIGTTVHQIKTAL 713
>At4g04890 putative homeotic protein
Length = 743
Score = 281 bits (718), Expect = 8e-76
Identities = 174/503 (34%), Positives = 287/503 (56%), Gaps = 43/503 (8%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLW+ + N V +L E Y + FPR + L +
Sbjct: 247 DKPIIVELAVAAMEELVRMAQTGDPLWLSTDNSVE--ILNEEEYFRTFPRGIGPKPLGLR 304
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ S +V+ + + LV++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 305 SEASRQSAVVIMNHINLVEILMDVNQWSCVFSGIVSRALTLEVLSTGVAGNYNGALQVMT 364
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS ++ PSGC I+EL
Sbjct: 365 AEFQVPSPLVPTRENYFVRYCKQHSDGSWAVVDVSLDSLRPSTPILRTRRRPSGCLIQEL 424
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
PNG VTW+E +EV + + + +V++ +A+GAKRW+ L+R E+ A S S
Sbjct: 425 PNGYSKVTWIEHMEVDDR-SVHNMYKPLVQSGLAFGAKRWVATLERQCERLASSMAS--- 480
Query: 264 STEGLEGEVKLL---DGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAH 320
+ G++ ++ +GRKS++ L+ RMV SF + S + ++++
Sbjct: 481 ---NIPGDLSVITSPEGRKSMLKLAERMVMSFCSGVGASTAHAWTTMSTTGSDDVRVMTR 537
Query: 321 RNTILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
++ +D GIV+ A +S W+PV PK +F FL+DE + EWD L+ G +++ AHI+
Sbjct: 538 KS--MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRKEWDILSNGGMVQEMAHIA 595
Query: 375 INE--SNHVSILQTSS--IIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSK 430
N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D
Sbjct: 596 NGHEPGNCVSLLRVNSGNSSQSNMLILQESCTDASGSYVIYAPVDIVAMNVVLSGGDPDY 655
Query: 431 KLILPSGFTISRDGR----EGNE--------------GSLLTMCFQVLVDEEGSSSQVRK 472
+LPSGF I DG +GN+ GSLLT+ FQ+LVD ++++
Sbjct: 656 VALLPSGFAILPDGSVGGGDGNQHQEMVSTTSSGSCGGSLLTVAFQILVDSV-PTAKLSL 714
Query: 473 KTLESIDSSCMLMLQRIKDALHC 495
++ +++S ++RIK A+ C
Sbjct: 715 GSVATVNSLIKCTVERIKAAVSC 737
>At3g61150 homeobox protein
Length = 808
Score = 281 bits (718), Expect = 8e-76
Identities = 177/501 (35%), Positives = 286/501 (56%), Gaps = 37/501 (7%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGF-VLVREIYAKKFPRITNSENLYV 82
++S+ L +A AAM EL+ + T EPLWV+SS+ GF VL +E Y F R +
Sbjct: 313 QRSRYLDLALAAMDELVKMAQTREPLWVRSSDS--GFEVLNQEEYDTSFSRCVGPKQDGF 370
Query: 83 CEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLM 142
E+SK++ V+ + + LV+ +DS++WA++ P++VS+ T ++ G + +GAL LM
Sbjct: 371 VSEASKEAGTVIINSLALVETLMDSERWAEMFPSMVSRTSTTEIISSG-MGGRNGALHLM 429
Query: 143 WNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIRE 202
E+ +LSPLVP R+ FLR+C+Q G W +VDVS+DS +S S ++ PSGC +++
Sbjct: 430 HAELQLLSPLVPVRQVSFLRFCKQHAEGVWAVVDVSIDSIREGSSSS-CRRLPSGCLVQD 488
Query: 203 LPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYT 262
+ NG VTW+E E ++ + L R ++R +A+GA RW+ LQR E S T
Sbjct: 489 MANGYSKVTWIEHTEYDEN-HIHRLYRPLLRCGLAFGAHRWMAALQRQCECLTIL-MSST 546
Query: 263 ASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQ-----NLLDPSNQLTTSIKL 317
ST + +GRKS++ L+ RM +F + +Q N+ + + +
Sbjct: 547 VSTSTNPSPINC-NGRKSMLKLAKRMTDNFCGGVCASSLQKWSKLNVGNVDEDVRIMTRK 605
Query: 318 IAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+ GI++ A +S+W+PV+P+ LF FL +E ++EWD L+ G PM++ AHI+
Sbjct: 606 SVNNPGEPPGIILNAATSVWMPVSPRRLFDFLGNERLRSEWDILSNGGPMKEMAHIAKGH 665
Query: 376 NESNHVSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ SN VS+L+ S+I M+I QE+SID G VVYAP+D ++NG DS+ +
Sbjct: 666 DRSNSVSLLRASAINANQSSMLILQETSIDAAGAVVVYAPVDIPAMQAVMNGGDSAYVAL 725
Query: 434 LPSGFTISRDGREGNE-------------------GSLLTMCFQVLVDEEGSSSQVRKKT 474
LPSGF I +G+ G + GSLLT+ FQ+LV+ ++++ ++
Sbjct: 726 LPSGFAILPNGQAGTQRCAAEERNSIGNGGCMEEGGSLLTVAFQILVNSL-PTAKLTVES 784
Query: 475 LESIDSSCMLMLQRIKDALHC 495
+E++++ +Q+IK ALHC
Sbjct: 785 VETVNNLISCTVQKIKAALHC 805
>At1g17920 unknown protein
Length = 687
Score = 280 bits (715), Expect = 2e-75
Identities = 182/496 (36%), Positives = 287/496 (57%), Gaps = 19/496 (3%)
Query: 12 PTRESQESEVTVE--KSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAK 69
P+ SQ + V E KS M +A AM+ELL LL T+EPLW+K+ D VL E Y
Sbjct: 195 PSLPSQPNLVLSEMDKSLMTNIAVTAMEELLRLLQTNEPLWIKT--DGCRDVLNLENYEN 252
Query: 70 KFPRITNS----ENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQ 125
F R + S NL + E+S+ S +V + + LV + ++S K ++ P+IV+ +KT+
Sbjct: 253 MFTRSSTSGGKKNNLGM--EASRSSGVVFTNAITLVDMLMNSVKLTELFPSIVASSKTLA 310
Query: 126 VFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNC 185
V G NH AL LM E+ VLSPLV +RE+ LRYCQQ+E G W IV+VS + F
Sbjct: 311 VISSGLRGNHGDALHLMIEELQVLSPLVTTREFCVLRYCQQIEHGTWAIVNVSYE-FPQF 369
Query: 186 ASRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLL 245
S+S + +FPSGC I+++ NG VTWVE E + + + +DIV +A+GA+RW+
Sbjct: 370 ISQSRSYRFPSGCLIQDMSNGYSKVTWVEHGEFEEQEPIHEMFKDIVHKGLAFGAERWIA 429
Query: 246 ELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLL 305
LQR E+ T+S + L G + +G++S++ L++RMV +F + T
Sbjct: 430 TLQRMCERFTNLLEPATSSLD-LGGVIPSPEGKRSIMRLAHRMVSNFCLSVGTSNNTRST 488
Query: 306 DPSNQLTTSIKLIAHRNT-ILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGG 364
S I++ +H++ +G+V+ A +S WLP++P+N+F FLKDE + +WD L+ G
Sbjct: 489 VVSGLDEFGIRVTSHKSRHEPNGMVLCAATSFWLPISPQNVFNFLKDERTRPQWDVLSNG 548
Query: 365 RPMRQKAHIS--INESNHVSILQ--TSSIIGEHMVIFQESSID-DLGGYVVYAPIDKLDA 419
+++ AHI+ N N +S+L+ +S +M+I QES ID V+Y P+D
Sbjct: 549 NSVQEVAHITNGSNPGNCISVLRGFNASSSQNNMLILQESCIDSSSAALVIYTPVDLPAL 608
Query: 420 YMIVNGCDSSKKLILPSGFTISRDGREGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESID 479
+ ++G D+S ILPSGF IS DG GSL+T+ FQ++V +++ +++E+++
Sbjct: 609 NIAMSGQDTSYIPILPSGFAISPDGSSKGGGSLITVGFQIMVSGL-QPAKLNMESMETVN 667
Query: 480 SSCMLMLQRIKDALHC 495
+ + +IK L+C
Sbjct: 668 NLINTTVHQIKTTLNC 683
>At4g21750 L1 specific homeobox gene ATML1/ovule-specific homeobox
protein A20
Length = 762
Score = 272 bits (696), Expect = 3e-73
Identities = 175/513 (34%), Positives = 285/513 (55%), Gaps = 50/513 (9%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+K ++ +A AAM+EL+ + T +PLWV S N V +L E Y + FPR + + +
Sbjct: 256 DKPMIVELAVAAMEELVRMAQTGDPLWVSSDNSVE--ILNEEEYFRTFPRGIGPKPIGLR 313
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S++S +V+ + + L+++ +D ++W+ + IVS+A T++V G N++GAL++M
Sbjct: 314 SEASRESTVVIMNHINLIEILMDVNQWSSVFCGIVSRALTLEVLSTGVAGNYNGALQVMT 373
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRIREL 203
E V SPLVP+RE +F+RYC+Q G W +VDVSLDS + + +++ PSGC I+EL
Sbjct: 374 AEFQVPSPLVPTRENYFVRYCKQHSDGIWAVVDVSLDSL-RPSPITRSRRRPSGCLIQEL 432
Query: 204 PNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTA 263
NG VTWVE +EV + + + +V +A+GAKRW+ L R E+ A S S
Sbjct: 433 QNGYSKVTWVEHIEVDDR-SVHNMYKPLVNTGLAFGAKRWVATLDRQCERLASSMASNIP 491
Query: 264 STEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT 323
+ + + +GRKS++ L+ RMV SF + S + ++++ ++
Sbjct: 492 ACD--LSVITSPEGRKSMLKLAERMVMSFCTGVGASTAHAWTTLSTTGSDDVRVMTRKS- 548
Query: 324 ILD------GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS--I 375
+D GIV+ A +S W+PV PK +F FL+DE ++EWD L+ G +++ AHI+
Sbjct: 549 -MDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRSEWDILSNGGLVQEMAHIANGR 607
Query: 376 NESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ N VS+L+ +S +M+I QES D G YV+YAP+D + ++++G D +
Sbjct: 608 DPGNSVSLLRVNSGNSGQSNMLILQESCTDASGSYVIYAPVDIIAMNVVLSGGDPDYVAL 667
Query: 434 LPSGFTISRDGR--------------------EGNE-----------GSLLTMCFQVLVD 462
LPSGF I DG EGN GSLLT+ FQ+LVD
Sbjct: 668 LPSGFAILPDGSARGGGGSANASAGAGVEGGGEGNNLEVVTTTGSCGGSLLTVAFQILVD 727
Query: 463 EEGSSSQVRKKTLESIDSSCMLMLQRIKDALHC 495
++++ ++ +++S ++RIK AL C
Sbjct: 728 SV-PTAKLSLGSVATVNSLIKCTVERIKAALAC 759
>At4g00730 homeodomain protein AHDP
Length = 590
Score = 269 bits (688), Expect = 2e-72
Identities = 170/499 (34%), Positives = 280/499 (56%), Gaps = 43/499 (8%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+KS +L +A AM EL+ L + EPLWVKS D L ++ Y + F ++++ +
Sbjct: 106 QKSVLLELALTAMDELVKLAQSEEPLWVKSL-DGERDELNQDEYMRTF---SSTKPTGLA 161
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ S +V+ + + LV+ +DS++W ++ P V++A T V G +GAL+LM
Sbjct: 162 TEASRTSGMVIINSLALVETLMDSNRWTEMFPCNVARATTTDVISGGMAGTINGALQLMN 221
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSF-GNCASRSVAKKFPSGCRIRE 202
E+ VLSPLVP R FLR+C+Q G W +VDVS+D N V ++ PSGC +++
Sbjct: 222 AELQVLSPLVPVRNVNFLRFCKQHAEGVWAVVDVSIDPVRENSGGAPVIRRLPSGCVVQD 281
Query: 203 LPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYT 262
+ NG VTWVE E ++ Q+ L R ++R+ + +G++RWL LQR E C +
Sbjct: 282 VSNGYSKVTWVEHAEYDEN-QIHQLYRPLLRSGLGFGSQRWLATLQRQCE---CLAILIS 337
Query: 263 ASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNL-------LDPSNQLTTSI 315
+S + GRKS++ L+ RM +F ++ + N +DP ++ T
Sbjct: 338 SSVTSHDNTSITPGGRKSMLKLAQRMTFNFCSGISAPSVHNWSKLTVGNVDPDVRVMT-- 395
Query: 316 KLIAHRNTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQK 370
R ++ D GIV+ A +S+WLP P+ L+ FL++E + EWD L+ G PM++
Sbjct: 396 -----RKSVDDPGEPPGIVLSAATSVWLPAAPQRLYDFLRNERMRCEWDILSNGGPMQEM 450
Query: 371 AHISINESNHVSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDS 428
AHI+ + VS+L+++++ M+I QE+ ID G VVYAP+D ++++NG DS
Sbjct: 451 AHITKGQDQGVSLLRSNAMNANQSSMLILQETCIDASGALVVYAPVDIPAMHVVMNGGDS 510
Query: 429 SKKLILPSGFTISRDG------------REGNEGSLLTMCFQVLVDEEGSSSQVRKKTLE 476
S +LPSGF + DG R GSLLT+ FQ+LV+ ++++ +++E
Sbjct: 511 SYVALLPSGFAVLPDGGIDGGGSGDGDQRPVGGGSLLTVAFQILVNNL-PTAKLTVESVE 569
Query: 477 SIDSSCMLMLQRIKDALHC 495
++++ +Q+I+ AL C
Sbjct: 570 TVNNLISCTVQKIRAALQC 588
>At1g05230 homeobox like protein
Length = 721
Score = 264 bits (675), Expect = 7e-71
Identities = 174/504 (34%), Positives = 279/504 (54%), Gaps = 32/504 (6%)
Query: 10 NNPTR--ESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIY 67
NNP +S + +K ++ ++ AAM+EL+ ++ EPLW VL E Y
Sbjct: 229 NNPNDLLKSITAPTESDKPVIIDLSVAAMEELMRMVQVDEPLWKS-------LVLDEEEY 281
Query: 68 AKKFPRITNSENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVF 127
A+ FPR E+S++S +V+ + + +V++ +D ++W+ I +VS+A T+ V
Sbjct: 282 ARTFPRGIGPRPAGYRSEASRESAVVIMNHVNIVEILMDVNQWSTIFAGMVSRAMTLAVL 341
Query: 128 DVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSF-GNCA 186
G N++GAL++M E V SPLVP+RE +F RYC+Q G W +VD+SLDS N
Sbjct: 342 STGVAGNYNGALQVMSAEFQVPSPLVPTRETYFARYCKQQGDGSWAVVDISLDSLQPNPP 401
Query: 187 SRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLE 246
+R ++ SGC I+ELPNG VTWVE VEV + L + +V A+GAKRW+
Sbjct: 402 AR--CRRRASGCLIQELPNGYSKVTWVEHVEVDDR-GVHNLYKHMVSTGHAFGAKRWVAI 458
Query: 247 LQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLD 306
L R E+ A T + G G + +GR+S++ L+ RMV SF ++
Sbjct: 459 LDRQCERLA--SVMATNISSGEVGVITNQEGRRSMLKLAERMVISFCAGVSASTAHTWTT 516
Query: 307 PSNQLTTSIKLIAHRNTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFL 361
S ++++ R ++ D GIV+ A +S W+PV PK +F FL+DE + EWD L
Sbjct: 517 LSGTGAEDVRVMT-RKSVDDPGRPPGIVLSAATSFWIPVPPKRVFDFLRDENSRNEWDIL 575
Query: 362 AGGRPMRQKAHIS--INESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYVVYAPIDKL 417
+ G +++ AHI+ + N VS+L+ +S +M+I QES D +V+YAP+D +
Sbjct: 576 SNGGVVQEMAHIANGRDTGNCVSLLRVNSANSSQSNMLILQESCTDPTASFVIYAPVDIV 635
Query: 418 DAYMIVNGCDSSKKLILPSGFTISRDGRE------GNEGSLLTMCFQVLVDEEGSSSQVR 471
+++NG D +LPSGF I DG G+ GSLLT+ FQ+LVD ++++
Sbjct: 636 AMNIVLNGGDPDYVALLPSGFAILPDGNANSGAPGGDGGSLLTVAFQILVDSV-PTAKLS 694
Query: 472 KKTLESIDSSCMLMLQRIKDALHC 495
++ ++++ ++RIK ++ C
Sbjct: 695 LGSVATVNNLIACTVERIKASMSC 718
>At3g03260 unknown protein
Length = 699
Score = 261 bits (668), Expect = 5e-70
Identities = 173/499 (34%), Positives = 270/499 (53%), Gaps = 32/499 (6%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
+ S + +A +A++EL L E WVKS D +V+ E Y + + + +
Sbjct: 207 DMSLLSEIAASAVEELKRLFLAEEQFWVKSCIDET-YVIDTESYERFSHAVKHFSSTTAH 265
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVG-SIENHSGALRLM 142
ESSK +V + L+++FLD +KW ++ PTIV+KA TI V G I + L++M
Sbjct: 266 VESSKAVTVVHVEAINLIQMFLDPEKWKELFPTIVNKANTIHVLGSGLPIRGNCNVLQVM 325
Query: 143 WNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDS---FGNCASRSVAKKFPSGCR 199
W ++H+LSPLVP+RE+ +R CQ++E G W+I DVS + FGN A K PSGC
Sbjct: 326 WEQLHILSPLVPAREFMVVRCCQEIEKGIWIIADVSHRANFDFGNAA----CYKRPSGCL 381
Query: 200 IRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEF 259
I+ LP+ V W+E VEV + RD++ YGAKRW++ L+R E+ A S
Sbjct: 382 IQALPDAHSKVMWIEHVEVDHKLDTHKIYRDLLSGGSGYGAKRWIVTLERMCERMALSSI 441
Query: 260 SYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQG-IQNLLDPSNQLTTSIKLI 318
++ E + + R+SV+ L RMV +F ++L G I N + SI++
Sbjct: 442 QTLPPSDRSE-VITTGEARRSVMKLGERMVKNFNEMLTMSGKIDFPQQSKNGVRVSIRMN 500
Query: 319 AHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHI--SIN 376
GIVV A SS+ +P+TP +F FL++ + +WD L+ G + + A I +
Sbjct: 501 IEAGQ-PPGIVVSASSSLAIPLTPLQVFAFLQNLDTRQQWDILSYGTVVNEIARIVTGSS 559
Query: 377 ESNHVSILQTSSIIGEH-------------MVIFQESSIDDLGGYVVYAPIDKLDAYMIV 423
E+N V+IL+ E+ M++ Q+ +D LGG +VYAP+D + V
Sbjct: 560 ETNCVTILRVHPTHEENNDKMVVQDSCKDDMLMLQDCYMDALGGMIVYAPMDMATMHFAV 619
Query: 424 NG-CDSSKKLILPSGFTISRDGREG---NEGSLLTMCFQVLVDEEGS-SSQVRKKTLESI 478
+G D S ILPSGF IS DGR + G+LLT+ FQ+LV + + S +V +K+++++
Sbjct: 620 SGEVDPSHIPILPSGFVISSDGRRSTVEDGGTLLTVAFQILVSGKANRSREVNEKSVDTV 679
Query: 479 DSSCMLMLQRIKDALHCSD 497
+ +QRIK L+C +
Sbjct: 680 SALISSTIQRIKGLLNCPE 698
>At1g34650 hypothetical protein
Length = 708
Score = 243 bits (621), Expect = 1e-64
Identities = 162/492 (32%), Positives = 264/492 (52%), Gaps = 23/492 (4%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVR----EIYAKKFPRITNSE 78
+EK++M +A A+ E++ L+ +W+KS+ D G ++ + Y K + +
Sbjct: 220 LEKNRMFEIAKNAVAEVMSLIQMEHSMWIKSTID--GRAIIDPGNYKRYFTKNSHLKSRS 277
Query: 79 NLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGA 138
L ESS + ++V LV +FL+++KWA + PTIV++AKTI V D + +
Sbjct: 278 ALQSHHESSMEVVVVQMDARNLVDMFLNTEKWARLFPTIVTEAKTIHVLDSMDHPRQTFS 337
Query: 139 LRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVS--LDSFGNCASRSVAKKFPS 196
R+++ ++H+LSPLV RE+ LR CQQ++ WLI DVS L + ++ + K PS
Sbjct: 338 -RVVYEQLHILSPLVLPREFIILRTCQQMKEDLWLIADVSCYLQNVEFESTAPICTKRPS 396
Query: 197 GCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKAC 256
G I+ LP+G VTW+E VEV L RD++ YGA+RW LQR E+ +
Sbjct: 397 GVLIQALPHGRSKVTWIEHVEVTDKVWPHQLYRDLLYGGFGYGARRWTATLQRMCERLSL 456
Query: 257 SEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIK 316
+ T+ G VK ++GR+SV++L RM+ +F ++ +L S + ++
Sbjct: 457 YSMTDFPPTD-YPGVVKTIEGRRSVMSLGERMLKNFAWIMKMSDKLDLPQQSGANNSGVR 515
Query: 317 LIAHRNTIL---DGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHI 373
+ NT G++V A SS+ LP+ P ++ FL++ + +WD G P+ + A
Sbjct: 516 ISVRTNTEAGQPPGLIVCAGSSLSLPLPPLQVYDFLRNLEVRHQWDVHCQGNPVTEAARF 575
Query: 374 --SINESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNG-CDSSK 430
++ N+V+ LQ SS+ ++I Q+ ID LGG VVYAP++ AY ++G D S
Sbjct: 576 VTGPDQKNNVTFLQPSSVGEYKLMILQDGFIDALGGMVVYAPMNLNTAYSAISGQVDPST 635
Query: 431 KLILPSGFTISRDGR------EGNEGSLLTMCFQVLVDEEGSSSQVR-KKTLESIDSSCM 483
ILPSGF ISRD +G +LLT+ FQ+ V + + K + ++++
Sbjct: 636 IPILPSGFIISRDSHPSSSEVDGGSMTLLTLAFQIFVTGPSYYTDLNLKDSATTVNTLVS 695
Query: 484 LMLQRIKDALHC 495
+QRIK L+C
Sbjct: 696 SAVQRIKAMLNC 707
>At5g17320 homeobox protein
Length = 718
Score = 243 bits (619), Expect = 2e-64
Identities = 181/497 (36%), Positives = 265/497 (52%), Gaps = 37/497 (7%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYV 82
+EK ML A A+ E+L L+ + +W KSS D V+ +Y K F + TN+
Sbjct: 234 LEKIAMLEAAEKAVSEVLSLIQMDDTMWKKSSIDD-RLVIDPGLYEKYFTK-TNTNGR-- 289
Query: 83 CEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGAL-RL 141
ESSKD ++V L+ +FL ++KWA + PTIV++AKTI V D S+++ R+
Sbjct: 290 -PESSKDVVVVQMDAGNLIDIFLTAEKWARLFPTIVNEAKTIHVLD--SVDHRGKTFSRV 346
Query: 142 MWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVS--LDSFGNCASRSVAKKFPSGCR 199
++ ++H+LSPLVP RE+ LR CQQ+E W+I DVS L + S + K PSG
Sbjct: 347 IYEQLHILSPLVPPREFMILRTCQQIEDNVWMIADVSCHLPNIEFDLSFPICTKRPSGVL 406
Query: 200 IRELPNGCCMVTWVEQVEVGQS-FQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSE 258
I+ LP+G VTW+E V V + + L RD++ YGA+RW + L+RT E+ S
Sbjct: 407 IQALPHGFSKVTWIEHVVVNDNRVRPHKLYRDLLYGGFGYGARRWTVTLERTCERLIFST 466
Query: 259 FSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTS---I 315
G V+ + GR SV++L RM+ +F ++ + N LD S Q T+ I
Sbjct: 467 SVPALPNNDNPGVVQTIRGRNSVMHLGERMLRNFAWMMK---MVNKLDFSPQSETNNSGI 523
Query: 316 KLIAHRNTIL---DGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAH 372
++ N G++V A SS+ LP+ P ++ FLK+ + +WD L G P + A
Sbjct: 524 RIGVRINNEAGQPPGLIVCAGSSLSLPLPPVQVYDFLKNLEVRHQWDVLCHGNPATEAAR 583
Query: 373 I--SINESNHVSILQTS-SIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNG-CDS 428
N N VS L+ S I ++I Q+S D LGG V YAP+D A ++G D
Sbjct: 584 FVTGSNPRNTVSFLEPSIRDINTKLMILQDSFKDALGGMVAYAPMDLNTACAAISGDIDP 643
Query: 429 SKKLILPSGFTISRDGR------EGNEGSLLTMCFQVLVD----EEGSSSQVRKKTLESI 478
+ ILPSGF ISRDGR EG +LLT+ FQ+LV ++ +V T+ ++
Sbjct: 644 TTIPILPSGFMISRDGRPSEGEAEGGSYTLLTVAFQILVSGPSYSPDTNLEVSATTVNTL 703
Query: 479 DSSCMLMLQRIKDALHC 495
SS +QRIK L C
Sbjct: 704 ISS---TVQRIKAMLKC 717
>At1g79840 homeobox protein (GLABRA2)
Length = 747
Score = 231 bits (588), Expect = 9e-61
Identities = 152/500 (30%), Positives = 263/500 (52%), Gaps = 30/500 (6%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRI-TNSENLY 81
+EKS++ ++N A EL + + EP+W++S + +L + Y K+FP+ +S
Sbjct: 252 LEKSRIAEISNRATLELQKMATSGEPMWLRSV-ETGREILNYDEYLKEFPQAQASSFPGR 310
Query: 82 VCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENH-SGALR 140
E+S+D+ +V +L + F+D +W + ++SKA T+ V G + GA++
Sbjct: 311 KTIEASRDAGIVFMDAHKLAQSFMDVGQWKETFACLISKAATVDVIRQGEGPSRIDGAIQ 370
Query: 141 LMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVA----KKFPS 196
LM+ E+ +L+P+VP+RE +F+R C+Q+ +W IVDVS+ + + + +K PS
Sbjct: 371 LMFGEMQLLTPVVPTREVYFVRSCRQLSPEKWAIVDVSVSVEDSNTEKEASLLKCRKLPS 430
Query: 197 GCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKAC 256
GC I + NG VTWVE ++V S + L R +V +A+GA+ W+ LQ E+
Sbjct: 431 GCIIEDTSNGHSKVTWVEHLDVSAS-TVQPLFRSLVNTGLAFGARHWVATLQLHCERLVF 489
Query: 257 SEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIK 316
+ + + L V L GRKSV+ ++ RM SF++ + + + ++
Sbjct: 490 FMATNVPTKDSLG--VTTLAGRKSVLKMAQRMTQSFYRAIAASSYHQWTKITTKTGQDMR 547
Query: 317 LIAHRNTI----LDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAH 372
+ + +N G++V A SS+WLPV+P LF F +DE R+ EWD L+ G ++ A+
Sbjct: 548 VSSRKNLHDPGEPTGVIVCASSSLWLPVSPALLFDFFRDEARRHEWDALSNGAHVQSIAN 607
Query: 373 ISINESNHVSI-LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKK 431
+S + S+ +QT + + + Q+SS + VVYAP+D +++ G D S
Sbjct: 608 LSKGQDRGNSVAIQTVKSREKSIWVLQDSSTNSYESVVVYAPVDINTTQLVLAGHDPSNI 667
Query: 432 LILPSGFTISRDG--------------REGNEGSLLTMCFQVLVDEEGSSSQVRKKTLES 477
ILPSGF+I DG R GSLLT+ Q L++ ++++ +++ES
Sbjct: 668 QILPSGFSIIPDGVESRPLVITSTQDDRNSQGGSLLTLALQTLIN-PSPAAKLNMESVES 726
Query: 478 IDSSCMLMLQRIKDALHCSD 497
+ + + L IK +L D
Sbjct: 727 VTNLVSVTLHNIKRSLQIED 746
>At5g52170 homeodomain transcription factor-like
Length = 682
Score = 206 bits (525), Expect = 2e-53
Identities = 153/488 (31%), Positives = 249/488 (50%), Gaps = 58/488 (11%)
Query: 31 VANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVCEESSKDS 90
+A AM ELL L LW SS G + FP S+++
Sbjct: 224 LAMEAMDELLKLAELETSLW--SSKSEKGSM-------NHFP-------------GSRET 261
Query: 91 LLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLS 150
LV+ + + LV+ +D++KWA++ IV+ A T++V GS + +G++ LM E V+S
Sbjct: 262 GLVLINSLALVETLMDTNKWAEMFECIVAVASTLEVISNGSDGSRNGSILLMQAEFQVMS 321
Query: 151 PLVPSREYFFLRYCQQVEAGEWLIVDVSLD---SFGNCASRSVAKKFPSGCRIRELPNGC 207
PLVP ++ FLRYC+Q G W +VDVS D N S +K FPSGC I+++ NGC
Sbjct: 322 PLVPIKQKKFLRYCKQHGDGLWAVVDVSYDINRGNENLKSYGGSKMFPSGCIIQDIGNGC 381
Query: 208 CMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTASTEG 267
VTW+E E +S L + ++ +++ GA +WL LQR C F+ S+E
Sbjct: 382 SKVTWIEHSEYEES-HTHSLYQPLLSSSVGLGATKWLATLQR-----QCESFTMLLSSED 435
Query: 268 LEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGI---QNLLDPSNQLTTSIKLIAHRNTI 324
G G KS++ L+ RM +F+ + I + LL + + +++ ++
Sbjct: 436 HTGLSHA--GTKSILKLAQRMKLNFYSGITASCIHKWEKLL--AENVGQDTRILTRKSLE 491
Query: 325 LDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHI--SINESNHVS 382
GIV+ A +S+WLPVT + LF FL D + +WD L+ G M + E + VS
Sbjct: 492 PSGIVLSAATSLWLPVTQQRLFEFLCDGKCRNQWDILSNGASMENTLLVPKGQQEGSCVS 551
Query: 383 ILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTI 440
+L+ + M+I QE+ D G VVYAP+D +++G DS+ +LPSGF+I
Sbjct: 552 LLRAAGNDQNESSMLILQETWNDVSGALVVYAPVDIPSMNTVMSGGDSAYVALLPSGFSI 611
Query: 441 SRDG---------------REGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLM 485
DG + ++G LLT+ FQ+LV+ ++++ +++E++++
Sbjct: 612 LPDGSSSSSDQFDTDGGLVNQESKGCLLTVGFQILVNSL-PTAKLNVESVETVNNLIACT 670
Query: 486 LQRIKDAL 493
+ +I+ AL
Sbjct: 671 IHKIRAAL 678
>At4g25530 homeodomain-containing transcription factor FWA
Length = 686
Score = 206 bits (523), Expect = 3e-53
Identities = 146/486 (30%), Positives = 236/486 (48%), Gaps = 28/486 (5%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVV---GFVLVREIYAKKFPRITNSENL 80
E S L +A A++EL+ L P W+ + +V G + E Y F +T
Sbjct: 209 ETSIFLNLAITALRELITLGEVDCPFWM--IDPIVRSKGVSKIYEKYRSSFNNVTKPPGQ 266
Query: 81 YVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALR 140
V E+S+ LV + + LVK +D+ KW ++ IV A T +V GS SG+L+
Sbjct: 267 IV--EASRAKGLVPMTCVTLVKTLMDTGKWVNVFAPIVPVASTHKVLSTGSGGTKSGSLQ 324
Query: 141 LMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRI 200
+ E V+SPLVP R+ F+RYC+++ G W++VDV+ +K+ PSG I
Sbjct: 325 QIQAEFQVISPLVPKRKVTFIRYCKEIRQGLWVVVDVTPTQNPTLLPYGCSKRLPSGLII 384
Query: 201 RELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEK-KACSEF 259
+L NG VTW+EQ E +S + L + ++ I GAKRWL LQR E S
Sbjct: 385 DDLSNGYSQVTWIEQAEYNES-HIHQLYQPLIGYGIGLGAKRWLATLQRHCESLSTLSST 443
Query: 260 SYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQ-----NLLDPSNQLTTS 314
+ T + GL + G ++ L+ RM ++++ + + + + + + ++
Sbjct: 444 NLTEISPGLSAK-----GATEIVKLAQRMTLNYYRGITSPSVDKWQKIQVENVAQNMSFM 498
Query: 315 IKLIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
I+ + L GIV+ A +S+WLPV LF F+ + EWD L M + I
Sbjct: 499 IRKNVNEPGELTGIVLSASTSVWLPVNQHTLFAFISHLSFRHEWDILTNDTTMEETIRIQ 558
Query: 375 INESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLIL 434
H +I+ I+ M++ QE D G VVYAP++ ++ G +S L
Sbjct: 559 -KAKRHGNIISLLKIVNNGMLVLQEIWNDASGAMVVYAPVETNSIELVKRGENSDSVKFL 617
Query: 435 PSGFTISRDGREGN-------EGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQ 487
PSGF+I DG G+ G LLT Q+LV +++ + + T++S+++ +
Sbjct: 618 PSGFSIVPDGVNGSYHRGNTGGGCLLTFGLQILVGINPTAALI-QGTVKSVETLMAHTIV 676
Query: 488 RIKDAL 493
+IK AL
Sbjct: 677 KIKSAL 682
>At2g32370 putative homeodomain transcription factor
Length = 721
Score = 204 bits (518), Expect = 1e-52
Identities = 152/505 (30%), Positives = 255/505 (50%), Gaps = 35/505 (6%)
Query: 9 VNNPTRESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYA 68
V N +RE+ K ++ +A AM+ELL + +EPLW+ N L + Y
Sbjct: 231 VGNHSRETTGPADANTKPIIMELAFGAMEELLVMAQVAEPLWMGGFNGT-SLALNLDEYE 289
Query: 69 KKF-----PRITNSENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKT 123
K F PR+ E+S+++ LV +V++ + + W+ + IV +A+T
Sbjct: 290 KTFRTGLGPRLGGFRT-----EASRETALVAMCPTGIVEMLMQENLWSTMFAGIVGRART 344
Query: 124 IQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFG 183
+ + N +G L++M E VLSPLV +RE +F+RYC+Q G W +VD+S+D
Sbjct: 345 HEQIMADAAGNFNGNLQIMSAEYQVLSPLVTTRESYFVRYCKQQGEGLWAVVDISIDHLL 404
Query: 184 NCASRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRW 243
+ ++ PSGC I+E+ +G VTWVE VEV + + + ++ A+ A RW
Sbjct: 405 PNINLKCRRR-PSGCLIQEMHSGYSKVTWVEHVEVDDAGSYSIFEK-LICTGQAFAANRW 462
Query: 244 LLELQRTFEKKACSEFSYTASTEGLEGEVKLLD-GRKSVINLSNRMVGSFFK-VLNTQG- 300
+ L R C S ST+ + L + G+ S++ ++ R+ +FF + N G
Sbjct: 463 VGTLVR-----QCERISSILSTDFQSVDSALTNHGKMSMLKIAERIARTFFAGMTNATGS 517
Query: 301 -IQNLLDPSNQLTTSIKLIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWD 359
I + ++ + ++K + G+++ A +S WLP P +F FL++ + WD
Sbjct: 518 TIFSGVEGEDIRVMTMKSVNDPGK-PPGVIICAATSFWLPAPPNTVFDFLREATHRHNWD 576
Query: 360 FLAGGRPMRQKAHIS--INESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKL 417
L G M + A I+ I++ N S+L+ M+I QE+S D +V+YAP+D
Sbjct: 577 VLCNGEMMHKIAEITNGIDKRNCASLLRHGHTSKSKMMIVQETSTDPTASFVLYAPVDMT 636
Query: 418 DAYMIVN-GCDSSKKLILPSGFTISRD------GREGNEGSLLTMCFQVLVDEEGSSSQV 470
+ ++ G D +ILPSGF I D G+EG GSLLT+ FQ+LV E G +++
Sbjct: 637 SMDITLHGGGDPDFVVILPSGFAIFPDGTGKPGGKEG--GSLLTISFQMLV-ESGPEARL 693
Query: 471 RKKTLESIDSSCMLMLQRIKDALHC 495
++ + ++ ++RIKD C
Sbjct: 694 SVSSVATTENLIRTTVRRIKDLFPC 718
>At5g46880 homeobox protein
Length = 783
Score = 192 bits (487), Expect = 5e-49
Identities = 158/515 (30%), Positives = 256/515 (49%), Gaps = 75/515 (14%)
Query: 2 ETLNDNVVNNPTRESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVG-- 59
+T N+N NN +E + +E A + +QEL + +T EPLW+K +D +G
Sbjct: 301 QTANNNNNNNMLLADEEKVIAME------FAVSCVQELTKMCDTEEPLWIKKKSDKIGGE 354
Query: 60 -FVLVREIYAKKFPRITNSENLY--VCEESSKDSLLVMASGMELVKVFLDSDKWADILPT 116
L E Y + FP ++N E+SK + +V+ + + LV FL++DKW+++ +
Sbjct: 355 ILCLNEEEYMRLFPWPMENQNNKGDFLREASKANAVVIMNSITLVDAFLNADKWSEMFCS 414
Query: 117 IVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQ-VEAGEWLIV 175
IV++AKT+Q+ G + SG+L L VLSPLVP+RE +FLRY +Q E G W IV
Sbjct: 415 IVARAKTVQIISSG-VSGASGSLLL------VLSPLVPTREAYFLRYVEQNAETGNWAIV 467
Query: 176 DVSLDSFG------NCASRSVAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVR 229
D +DSF N + +K PSGC I+++PNG V WVE VEV + +
Sbjct: 468 DFPIDSFHDQMQPMNTITHEYKRK-PSGCIIQDMPNGYSQVKWVEHVEVDEK-HVHETFA 525
Query: 230 DIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMV 289
+ V++ +A+GA RWL LQR E+ A S A L + R++++ LS R+V
Sbjct: 526 EYVKSGMAFGANRWLDVLQRQCERIA----SLMA-----RNITDLGEARRNIMRLSQRLV 576
Query: 290 GSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNT---ILDGIVVIAVSSIWLPVTPKNLF 346
+F ++T Q+ S ++++ + G+V+ AVS+ WLP + +F
Sbjct: 577 KTFCVNISTAYGQSWTALSETTKDTVRITTRKMCEPGQPTGVVLCAVSTTWLPFSHHQVF 636
Query: 347 LFLKDEFRKAEWDFLAGGRPMRQKAHISINESNHVSILQTSSIIGEHMVIFQESSIDDLG 406
++D+ +H S++ ++S +++ QES ID+ G
Sbjct: 637 DLIRDQ--------------------------HHQSLVASNSWHNVELML-QESCIDNSG 669
Query: 407 GYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTI----SRDGREGNEGS----LLTMCFQ 458
+VY+ +D +NG DSS ILP GF+I +G N S LLT+ Q
Sbjct: 670 SLIVYSTVDVDSIQQAMNGEDSSNIPILPLGFSIVPVNPPEGISVNSHSPPSCLLTVGIQ 729
Query: 459 VLVDEEGSSSQVRKKTLESIDSSCMLMLQRIKDAL 493
VL +++ T+ +I++ + +I AL
Sbjct: 730 VLASNV-PTAKPNLSTVTTINNHLCATVNQITSAL 763
>At4g17710 GLABRA2 like protein
Length = 661
Score = 165 bits (418), Expect = 5e-41
Identities = 135/485 (27%), Positives = 230/485 (46%), Gaps = 76/485 (15%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVC 83
EK+ + +A + +EL + + +EPLW K D L E Y K F +++
Sbjct: 232 EKAIDMELAVSCARELAKMCDINEPLWNKKRLDNESVCLNEEEYKKMFLWPLMNDDDRFR 291
Query: 84 EESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMW 143
E+S+ + ++M + + LVK FLD+DKW+++ IVS AKT Q+ G+ SG L L
Sbjct: 292 REASRANAVIMLNCITLVKAFLDADKWSEMFFPIVSSAKTAQIISSGA-SGPSGTLLL-- 348
Query: 144 NEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSF--GNCASRSVAKKFPSGCRIR 201
Q E G+W++VD +D + + ++ PSGC I+
Sbjct: 349 ---------------------QNAEEGKWMVVDFPIDRIKPASATTTDQYRRKPSGCIIQ 387
Query: 202 ELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSY 261
+ NG VTWVE VEV + D +VR+ V + +A+GA+RWL L+R E+ A +
Sbjct: 388 AMRNGYSQVTWVEHVEVEEKHVQDEVVREFVESGVAFGAERWLSVLKRQCERMASLMATN 447
Query: 262 TASTEGLE----GEVKLLDGRKSVINLSNRMVGSF-FKVLNTQGIQNLLDPSNQLTTSIK 316
G+ + ++ RK+++ LS RMV +F ++N+ G D ++K
Sbjct: 448 ITDLGGIYITCFTMIPSVEARKNLMKLSQRMVKTFCLNIINSHGQAPTKD-------TVK 500
Query: 317 LIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISIN 376
+++ + + G+V AVS LP + + +F L+D R ++ + N
Sbjct: 501 IVSRK--VCGGLVPCAVSVTLLPYSHQQVFDLLRDNQRLSQ---------------VESN 543
Query: 377 ESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPS 436
S++V ++ QE+ D+ G +VY+ +D + + +NG D S+ +LP
Sbjct: 544 SSHNVELM------------LQETCTDNSGSLLVYSTVDPVAVQLAMNGEDPSEIPLLPV 591
Query: 437 GFTI----SRDGREGNEGS----LLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQR 488
GF++ DG EG+ S LLT+ QVL ++ ++ T+ I+ + R
Sbjct: 592 GFSVVPVNPSDGVEGSSVSSPSCLLTVAIQVL-GSNVTTERLDLSTVSVINHRICATVNR 650
Query: 489 IKDAL 493
I AL
Sbjct: 651 ITSAL 655
>At5g07260 unknown protein
Length = 541
Score = 113 bits (282), Expect = 3e-25
Identities = 105/425 (24%), Positives = 185/425 (42%), Gaps = 45/425 (10%)
Query: 31 VANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVCEESSKDS 90
+ + +++E++FL P+W + G + + E Y+K FP + +V E S+ S
Sbjct: 95 MTSLSLKEVVFLARQRTPMWTSN-----GRLNLDEYYSKLFPWYARNAPGFV-HEVSRAS 148
Query: 91 LLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLS 150
V LV ++ W I P+I++ DV G ++ N + +S
Sbjct: 149 AFVPCDASSLVANLMNHVSWQKIFPSIIA--------DVSVESQQRGLQKINVNFMPQIS 200
Query: 151 PLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCAS--RSVAKKFPSGCRIRELPNGCC 208
PL+ +R LR + +E W I ++S+ F + A R +FPSG I+ + NG
Sbjct: 201 PLIQTRNVKLLRRSRHIEDDTWAIAEISM-YFSSYAQHLRPEYMRFPSGYLIQHIANGIS 259
Query: 209 MVT----WVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTAS 264
VT WV + E G + +N +GA+RWL LQ+ + S
Sbjct: 260 KVTILDHWVYKEEEGMN---------TFNSNSEFGAQRWLTALQKHYYNTC------PVS 304
Query: 265 TEGLEGEVKLLDG--RKSVINLSNRMVGSFFK-VLNTQGIQ-NLLDPSNQLTTSIKLIAH 320
+ +++ D RK+++NLS+ MV F V G + N L+ +I++
Sbjct: 305 IPSIGHNIQIFDQICRKNLLNLSSFMVNVFCSGVCGITGQRWNRLNTVGVSANNIRMFTQ 364
Query: 321 RNTILDGIVVIAVSSIWLP---VTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISI-- 375
+ + GI + VS+ L P+ +F + ++ W +L + M++ I
Sbjct: 365 ESRGMSGIPCVLVSATGLARMHTKPEVMFGLINGAEKQEIWSYLESAKDMKELIRIGRHP 424
Query: 376 NESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILP 435
N N VS+ + + QE+ D+ G +++ ++ +NG D S +LP
Sbjct: 425 NSWNEVSVFSIEWKGSKEWYLIQETYYDESGAMIIHTCVEAPYFAAAINGGDLSGVELLP 484
Query: 436 SGFTI 440
SGFTI
Sbjct: 485 SGFTI 489
>At4g26920 putative homeodomain protein
Length = 461
Score = 57.8 bits (138), Expect = 1e-08
Identities = 42/172 (24%), Positives = 86/172 (49%), Gaps = 17/172 (9%)
Query: 36 MQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVCEESSKDSLLVMA 95
+ E++ L PLW +S + + + + E Y++ FP + +V E+S+ S ++
Sbjct: 72 VNEIIALATLESPLWRRSQREEM--LTLNEYYSRFFPWYAKNVPRFV-HEASRASEVIHV 128
Query: 96 SGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMWNE--VHVLSPLV 153
L+ + +W I P++V SIE+ + +R++ + + +++P++
Sbjct: 129 DASWLLTKLKNPMRWVTIFPSLVGNV---------SIESSNDDVRMIIDMEFLTLITPVI 179
Query: 154 PSREYFFLRYCQQVEAGEWLIVDVS--LDSFGNCASRSVAKKFPSGCRIREL 203
P+R+ LRYC ++ W+I D+S L S+ + R +FPSG I+ +
Sbjct: 180 PTRKVKVLRYCHRIANDTWIIADISMYLSSYSD-DLRPEFLRFPSGFIIKHV 230
>At4g32880 HD-zip transcription factor (athb-8)
Length = 833
Score = 50.4 bits (119), Expect = 2e-06
Identities = 61/274 (22%), Positives = 121/274 (43%), Gaps = 23/274 (8%)
Query: 103 VFLDSDKWADIL---PTIVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYF 159
V LD + A+IL P + +++ + +V S N G L L++ +++ + L P+R+++
Sbjct: 212 VGLDPTRVAEILKDKPCWLRDCRSLDIVNVLSTAN-GGTLELIYMQLYAPTTLAPARDFW 270
Query: 160 FLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKF------PSGCRIRELPNGCCMVTWV 213
LRY +E G +I + SL++ N S + F PSG IR G ++ V
Sbjct: 271 MLRYTSVMEDGSLVICERSLNNTQNGPSMPPSPHFVRAEILPSGYLIRPCEGGGSILHIV 330
Query: 214 EQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTASTEGLEGEVK 273
+ ++ + + + ++R + ++ + + L+ + ++ E S T
Sbjct: 331 DHFDL-EPWSVPEVLRSLYESSTLLAQRTTMAALR--YLRQISQEISQPNVTGWGRRPAA 387
Query: 274 LLDGRKSVINLSNRMVGSF----FKVLNTQGIQNLL-----DPSNQLTTSIKLIAHRNTI 324
L + + N V F + +L + GI ++ P+ + TS A+ T
Sbjct: 388 LRALSQRLSKGFNEAVNGFSDEGWSILESDGIDDVTLLVNSSPTKMMMTSSLPFANGYTS 447
Query: 325 LDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEW 358
+ V+ A +S+ L P ++ L E R+ EW
Sbjct: 448 MPSAVLCAKASMLLQNVPPSILLRFLREHRQ-EW 480
>At5g60690 REVOLUTA or interfascicular fiberless 1
Length = 842
Score = 47.8 bits (112), Expect = 1e-05
Identities = 83/372 (22%), Positives = 157/372 (41%), Gaps = 50/372 (13%)
Query: 8 VVNNPTRESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIY 67
VVN+P+ ES V LR AN+ L T K++ V +V + +
Sbjct: 132 VVNDPSCES----VVTTPQHSLRDANSPAGLLSIAEETLAEFLSKATGTAVDWVQMPGM- 186
Query: 68 AKKFPRITNSENLYVCEESSKDSLLVMASGMELVKVFLDSDKWADIL---PTIVSKAKTI 124
K P +S ++ + + + A G+ V L+ K A+IL P+ +++
Sbjct: 187 -KPGP---DSVGIFAISQRC-NGVAARACGL----VSLEPMKIAEILKDRPSWFRDCRSL 237
Query: 125 QVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFG- 183
+VF + N G + L++ + + + L P+R+++ LRY ++ G +++ + SL G
Sbjct: 238 EVFTMFPAGN-GGTIELVYMQTYAPTTLAPARDFWTLRYTTSLDNGSFVVCERSLSGSGA 296
Query: 184 --NCASRSV---AKKFPSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAY 238
N AS S A+ SG IR G ++ V+ + + +++ + ++R + ++
Sbjct: 297 GPNAASASQFVRAEMLSSGYLIRPCDGGGSIIHIVDHLNL-EAWSVPDVLRPLYESSKVV 355
Query: 239 GAKRWLLELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVI--NLSNRMVGSFFKVL 296
K + L+ + ++ E + GEV GR+ + S R+ F +
Sbjct: 356 AQKMTISALR--YIRQLAQESN---------GEVVYGLGRQPAVLRTFSQRLSRGFNDAV 404
Query: 297 NTQGIQNL----LDPSNQLTTSIKLIAHRNTI------LDGIVVIAVSSIWLPVTPKNLF 346
N G D + + +I H N I L G++ S + V P L
Sbjct: 405 NGFGDDGWSTMHCDGAEDIIVAINSTKHLNNISNSLSFLGGVLCAKASMLLQNVPPAVLI 464
Query: 347 LFLKDEFRKAEW 358
FL++ ++EW
Sbjct: 465 RFLRE--HRSEW 474
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,539,539
Number of Sequences: 26719
Number of extensions: 432370
Number of successful extensions: 1176
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1039
Number of HSP's gapped (non-prelim): 36
length of query: 497
length of database: 11,318,596
effective HSP length: 103
effective length of query: 394
effective length of database: 8,566,539
effective search space: 3375216366
effective search space used: 3375216366
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0321.6


