

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0316.5
(318 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
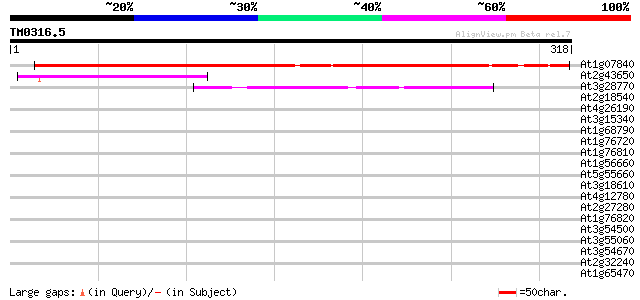
Sequences producing significant alignments: (bits) Value
At1g07840 unknown protein 295 2e-80
At2g43650 unknown protein 79 2e-15
At3g28770 hypothetical protein 42 6e-04
At2g18540 putative vicilin storage protein (globulin-like) 41 0.001
At4g26190 putative protein 39 0.003
At3g15340 hypothetical protein 39 0.003
At1g68790 putative nuclear matrix constituent protein 1 (NMCP1) 39 0.003
At1g76720 putative translation initiation factor IF-2 39 0.004
At1g76810 translation initiation factor IF-2 like protein 38 0.008
At1g56660 hypothetical protein 38 0.008
At5g55660 putative protein 37 0.011
At3g18610 unknown protein 37 0.014
At4g12780 auxilin-like protein 35 0.041
At2g27280 unknown protein 35 0.070
At1g76820 putative translation initiation factor IF-2 35 0.070
At3g54500 unknown protein 34 0.092
At3g55060 centromere protein - like 34 0.12
At3g54670 structural maintenance of chromosomes (SMC) - like pro... 34 0.12
At2g32240 putative myosin heavy chain 34 0.12
At1g65470 hypothetical protein, 3' partial 34 0.12
>At1g07840 unknown protein
Length = 312
Score = 295 bits (755), Expect = 2e-80
Identities = 162/304 (53%), Positives = 214/304 (70%), Gaps = 9/304 (2%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
+KEA LA++L+EMK LDVVR K+++LTA VK +PTA G SYLEAK+LLLL+YCQ L
Sbjct: 14 KKEAPQLASVLREMKNVLDVVRSKVEALTALVKANSFPTAGGISYLEAKHLLLLSYCQDL 73
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
VYY+LRKAKG SI+ HP+VRS+VEIR++LEKIRPIDKK QYQIQKL G
Sbjct: 74 VYYILRKAKGLSIDGHPLVRSLVEIRMFLEKIRPIDKKLQYQIQKLTTAGGPVTELAHSE 133
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNA 194
K + KSED+S Y+P PD+L +ED QE YRP KFAP SM +DKTSKQER+A
Sbjct: 134 GKGSCEAQKSEDLSNYKPKPDLLADKED--DQEDDGVYRPPKFAPMSM-EDKTSKQERDA 190
Query: 195 LRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEG-SSKEVERHIAKMEERAWQEEELF 253
R+EK +++ +++++ +++D+E++PEEIRDY G S E +R +A+ E + EEELF
Sbjct: 191 ARKEKHFFRQATENTYMKDVLDDLEDRPEEIRDYYGVESNEQKRFMAQYERQQKAEEELF 250
Query: 254 NRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNGRL 313
R P TK ++KREK LK S +GL LTE+F+D+IK F D+ GE+ + + R G
Sbjct: 251 TRAPRTKEDKKREKRLKSS-SGLHELTENFYDDIK---FLDKDGEKPRSFGR-NKRGGPF 305
Query: 314 KKRK 317
KKRK
Sbjct: 306 KKRK 309
>At2g43650 unknown protein
Length = 654
Score = 79.3 bits (194), Expect = 2e-15
Identities = 40/111 (36%), Positives = 66/111 (59%), Gaps = 3/111 (2%)
Query: 5 VNHLNNHDSKE---KEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLE 61
+N L+ + + A + LL E+ + ++ + KI + K+KEG+ YLE
Sbjct: 220 INSLSKEEQMDVVYSSAPEIVGLLSELNDAVEELESKINPVMNKLKEGEISLNGLARYLE 279
Query: 62 AKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKK 112
K LLLL YCQS+ +YLL K++G I +HPV+ +VEI+ L+KI+ +D++
Sbjct: 280 VKQLLLLTYCQSITFYLLLKSEGQPIRDHPVLARLVEIKSLLDKIKELDEE 330
>At3g28770 hypothetical protein
Length = 2081
Score = 41.6 bits (96), Expect = 6e-04
Identities = 35/170 (20%), Positives = 77/170 (44%), Gaps = 14/170 (8%)
Query: 105 KIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLA 164
K + + K++++ K MK E+ KKE ++S+ K DM + +
Sbjct: 1077 KSKKKEDKKEHEDNKSMKKEED--------KKEKKKHEESKSRKKEEDKKDMEKLEDQNS 1128
Query: 165 PQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEE 224
++ D K + + ++ K+E+ +E E E+K ++ N++++K E
Sbjct: 1129 NKKKEDKNEKKKSQHVKLVKKESDKKEK----KENEEKSETKEIESSKSQKNEVDKK--E 1182
Query: 225 IRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRN 274
+ + K+ E+ + + EE+ ++ E + + E K++K KK +N
Sbjct: 1183 KKSSKDQQKKKEKEMKESEEKKLKKNEEDRKKQTSVEENKKQKETKKEKN 1232
Score = 38.1 bits (87), Expect = 0.006
Identities = 46/214 (21%), Positives = 90/214 (41%), Gaps = 16/214 (7%)
Query: 112 KQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGH-- 169
K++ + K K E++A+ + K+ K+++ +K RE+ +E
Sbjct: 990 KEENKDNKEKKESEDSASKNREKKEYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSK 1049
Query: 170 ---DAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIR 226
+ R +K A ++ K K+ N ++KE KE + + M E+K E+ +
Sbjct: 1050 KEKEESRDLK-AKKKEEETKEKKESENHKSKKKEDKKEHEDNKS----MKKEEDKKEKKK 1104
Query: 227 DYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKRE---KYLKKSRNGLDGLTESF 283
E S++ E ME+ E++ N+ K+E+K+ K +KK + +
Sbjct: 1105 HEESKSRKKEEDKKDMEKL---EDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEE 1161
Query: 284 FDEIKTLPFEDETGEQVMGPSKGSNRNGRLKKRK 317
E K + +V K S+++ + KK K
Sbjct: 1162 KSETKEIESSKSQKNEVDKKEKKSSKDQQKKKEK 1195
Score = 35.0 bits (79), Expect = 0.054
Identities = 47/193 (24%), Positives = 80/193 (41%), Gaps = 38/193 (19%)
Query: 125 ENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQ 184
EN T + SK++ K + SK N +M ED ++ + ++
Sbjct: 928 ENKDTINTSSKQKGKDKKKKKKESK---NSNMKKKEEDKKEYVNNELKKQED------NK 978
Query: 185 DKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEE 244
+T+K E + L+ E + KE K S D K E ++YE K + AK E+
Sbjct: 979 KETTKSENSKLKEENKDNKEKKESE-------DSASKNREKKEYE-EKKSKTKEEAKKEK 1030
Query: 245 RAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPS 304
+ Q+++ R ERK +K ++SR ++K E+ET E
Sbjct: 1031 KKSQDKK---REEKDSEERKSKKEKEESR------------DLKAKKKEEETKE------ 1069
Query: 305 KGSNRNGRLKKRK 317
K + N + KK++
Sbjct: 1070 KKESENHKSKKKE 1082
Score = 28.5 bits (62), Expect = 5.0
Identities = 35/152 (23%), Positives = 69/152 (45%), Gaps = 13/152 (8%)
Query: 130 SDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKT-- 187
S++ KK +S+K E+ + N D + ++ L +E + K ++ +
Sbjct: 681 SEVEVKKNDGSSEKGEEGKEN--NKDSMEDKK-LENKESQTDSKDDKSVDDKQEEAQIYG 737
Query: 188 --SKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEER 245
SK +++ + K+ KESK + +T N + K E ++ + S++VE+ K E +
Sbjct: 738 GESKDDKSVEAKGKK--KESKENKKTKTNENRVRNKEENVQGNKKESEKVEKG-EKKESK 794
Query: 246 AWQEEELFNRVPLTKHERKREKYLKKSRNGLD 277
+ E + L+ E + E K R+G D
Sbjct: 795 DAKSVETKDNKKLSSTENRDE---AKERSGED 823
>At2g18540 putative vicilin storage protein (globulin-like)
Length = 699
Score = 40.8 bits (94), Expect = 0.001
Identities = 31/140 (22%), Positives = 61/140 (43%), Gaps = 12/140 (8%)
Query: 134 SKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERN 193
++K +K E+++K R RE++ + + R + +++ ++E
Sbjct: 508 ARKREEEREKEEEMAKKREEERQRKEREEVERKRREEQERKRREEEARKREEERKREEEM 567
Query: 194 ALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELF 253
A RRE+E ++ + EE +IR+ + +E E + +ER +E E
Sbjct: 568 AKRREQERQRKER------------EEVERKIREEQERKREEEMAKRREQERQKKEREEM 615
Query: 254 NRVPLTKHERKREKYLKKSR 273
R + RKRE+ + K R
Sbjct: 616 ERKKREEEARKREEEMAKIR 635
Score = 38.5 bits (88), Expect = 0.005
Identities = 33/133 (24%), Positives = 60/133 (44%), Gaps = 8/133 (6%)
Query: 143 KSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEIL 202
+ E++ + R + RE+ +E +A R + + ++ ++E A +RE+E
Sbjct: 431 EEEEIERRRKEEEEARKREEAKRREEEEAKRREE-----EETERKKREEEEARKREEERK 485
Query: 203 KESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKM--EERAWQEEELFNRVPLTK 260
+E + + EE+ E+ R E +E E +AK EER +E E R +
Sbjct: 486 REEEEAKRREEERKKREEEAEQARKRE-EEREKEEEMAKKREEERQRKEREEVERKRREE 544
Query: 261 HERKREKYLKKSR 273
ERKR + + R
Sbjct: 545 QERKRREEEARKR 557
Score = 37.4 bits (85), Expect = 0.011
Identities = 41/188 (21%), Positives = 77/188 (40%), Gaps = 12/188 (6%)
Query: 98 EIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDML 157
E+ + +I ++++ +I++ K E A + ++E + + E+ R +
Sbjct: 416 ELSKLMREIEERKRREEEEIERRRKEEEEARKREEAKRREEEEAKRREEEETERKKREEE 475
Query: 158 VSR----------EDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKE-ILKESK 206
+R E+ +E R + +++ K+E A +RE+E KE +
Sbjct: 476 EARKREEERKREEEEAKRREEERKKREEEAEQARKREEEREKEEEMAKKREEERQRKERE 535
Query: 207 HSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKME-ERAWQEEELFNRVPLTKHERKR 265
R + + + EE R E K E + E ER +E E R + ERKR
Sbjct: 536 EVERKRREEQERKRREEEARKREEERKREEEMAKRREQERQRKEREEVERKIREEQERKR 595
Query: 266 EKYLKKSR 273
E+ + K R
Sbjct: 596 EEEMAKRR 603
Score = 35.0 bits (79), Expect = 0.054
Identities = 34/166 (20%), Positives = 73/166 (43%), Gaps = 11/166 (6%)
Query: 111 KKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHD 170
+K++ ++++ + + + ++K + E+++K R RE++ + +
Sbjct: 531 RKEREEVERKRREEQERKRREEEARKREEERKREEEMAKRREQERQRKEREEVERKIREE 590
Query: 171 AYRPVKFAPTSMDQDKTSKQERNAL---RREKEILKESKHSSFIRTLMNDIEEKPEEIRD 227
R + + + K+ER + +RE+E K + + IR + E + +E D
Sbjct: 591 QERKREEEMAKRREQERQKKEREEMERKKREEEARKREEEMAKIR----EEERQRKERED 646
Query: 228 YEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSR 273
E +E E + EE +EEE R + RK+E+ +K R
Sbjct: 647 VERKRREEE--AMRREEERKREEEAAKRA--EEERRKKEEEEEKRR 688
>At4g26190 putative protein
Length = 1067
Score = 39.3 bits (90), Expect = 0.003
Identities = 44/178 (24%), Positives = 83/178 (45%), Gaps = 14/178 (7%)
Query: 125 ENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQ 184
E+ AT D S + + E V Y N D ++ +DLA A + S+
Sbjct: 632 ESVATMDSESVQSLLYQSNGEGVKNYEGNADGEIASKDLASNIEDSATK-----GESVQD 686
Query: 185 DKTSKQERNALRREKEILKESKHSSFIRT--LMNDIEEKPEEIRDYEGSSKEVERHIAKM 242
DK +K+ + R+ E+ +++K + + T + +K + + D++ + E E I K
Sbjct: 687 DKNTKKRKK--DRKGEVDQDAKGAEGVSTVEVTTKKSKKKKNLLDHKTDNME-EDSIKKN 743
Query: 243 EERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESF--FDEIKTLPFEDETGE 298
E++ ++ ++K E K +K K+ +N LD T+ D++ +LP +DE E
Sbjct: 744 EKKEEVDQNDLGAEGVSKVEVKTKK-SKRKKNSLDHKTDDMEGKDDV-SLPRKDEEPE 799
>At3g15340 hypothetical protein
Length = 478
Score = 39.3 bits (90), Expect = 0.003
Identities = 59/283 (20%), Positives = 114/283 (39%), Gaps = 31/283 (10%)
Query: 8 LNNHDSKEKEAS-----PLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEA 62
L+ EKEAS +A L E+K+ LD + KI L+ KV + Q + L+
Sbjct: 194 LSEDSLAEKEASINRVKSMAVELNEVKKELDAITWKINHLSDKVGKSQ----NNLRVLDV 249
Query: 63 KNLLLL-----NYCQSLVYYLLR-KAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQ 116
K +L +Y + + + R K K + PV+R E+ +R ++ +
Sbjct: 250 KKAHILEERDRSYERIKMLRIQRDKGKAAFYQSLPVMRKARELAAS-GNVRDLEVFASSE 308
Query: 117 IQKLMKV--GENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRP 174
+ M + A D + P ++ + +P++ V E P +G +
Sbjct: 309 ADRFMTQWNDDKAFRDDYVKRISPSLCERQLNQDGRIKDPEVQVVWEKKVPVKGGEKVHE 368
Query: 175 VKFAPTSMDQDK-----TSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYE 229
+S + + T K+++ ++ + + S S + L + EKP++
Sbjct: 369 TNREDSSSNSSQYGSVITDKRKKETRKKAMDFNRSSAEESDVTDLEFSVYEKPKK----- 423
Query: 230 GSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKS 272
+EV+ K ER Q E+ R+ + + + +EK K+
Sbjct: 424 -EEEEVDEETLKEREREEQLEKA--RLAMERKRKLQEKAAAKA 463
>At1g68790 putative nuclear matrix constituent protein 1 (NMCP1)
Length = 1085
Score = 39.3 bits (90), Expect = 0.003
Identities = 72/308 (23%), Positives = 126/308 (40%), Gaps = 51/308 (16%)
Query: 12 DSKEKEASPLAALLKEM-----------KEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 60
++K +EA+ L +KE +E V+ K L K+KE + T +
Sbjct: 163 EAKLEEANALVIGMKEKALEVDRERAIAEEKFSVMNRKSSELERKLKEVE--TREKVHQR 220
Query: 61 EAKNLLLLNYCQSLVYYLLRK-----AKGFSIEEH---PVVRSIV--EIRLYLEKIRPID 110
E +L+ V+Y R+ K ++EE V RSI E R+ +E R I+
Sbjct: 221 EHLSLVTEREAHEAVFYKQREDLQEWEKKLTLEEDRLSEVKRSINHREERV-MENERTIE 279
Query: 111 KKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHD 170
KK++ K+ + A S+L K+E + K D+S + + + ++ D+ +E H+
Sbjct: 280 KKEKILENLQQKI--SVAKSELTEKEESIKI-KLNDISLKEKDFEAMKAKVDIKEKELHE 336
Query: 171 AYRPVKFAPTSMDQDKTSKQERNAL---RREKEILKESKHSSFIRTLMNDIEEKPEEIRD 227
+ M+ K ++ L RRE E+ E R+L ++E K EI
Sbjct: 337 -FEENLIEREQMEIGKLLDDQKAVLDSRRREFEMELEQMR----RSLDEELEGKKAEIEQ 391
Query: 228 YEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEI 287
+ E +AK E K+E+ +KK LD ++ ++
Sbjct: 392 LQVEISHKEEKLAKREAAL----------------EKKEEGVKKKEKDLDARLKTVKEKE 435
Query: 288 KTLPFEDE 295
K L E++
Sbjct: 436 KALKAEEK 443
Score = 31.6 bits (70), Expect = 0.59
Identities = 45/221 (20%), Positives = 94/221 (42%), Gaps = 29/221 (13%)
Query: 78 LLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKE 137
L +K +G +E + + ++ + ++ +KK + ++L++ + L + E
Sbjct: 410 LEKKEEGVKKKEKDLDARLKTVKEKEKALKAEEKKLHMENERLLE--DKECLRKLKDEIE 467
Query: 138 PVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRR 197
+ ++ ++ S+ R + L + +E + R +D+ KQE L +
Sbjct: 468 EIGTETTKQESRIREEHESL----RITKEERVEFLRLQSELKQQIDK---VKQEEELLLK 520
Query: 198 EKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVP 257
E+E LK+ K E+ E +++E K+ + E A + E+L N
Sbjct: 521 EREELKQDK-------------ERFE--KEWEALDKKRANITREQNEVAEENEKLRNLQI 565
Query: 258 LTKHERKREKY-----LKKSRNGLDGLTESFFDEIKTLPFE 293
KH KRE+ LK+ +G+ ESF +++ L +
Sbjct: 566 SEKHRLKREEMTSRDNLKRELDGVKMQKESFEADMEDLEMQ 606
>At1g76720 putative translation initiation factor IF-2
Length = 1224
Score = 38.9 bits (89), Expect = 0.004
Identities = 39/174 (22%), Positives = 68/174 (38%), Gaps = 24/174 (13%)
Query: 127 AATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGH----------------- 169
A + P+ + P +S + +V K + P + E+ +EG
Sbjct: 282 AELGETPAAERPASS--TPEVEKVQAQPGPVAPVENAGEKEGEKETVETAAAKKKKKKKE 339
Query: 170 -DAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEI--- 225
D + A TS + K KQE + + K++K + + + + E E +
Sbjct: 340 KDKEKKAAAAATSSVEAKEEKQEESVTEPLQPKKKDAKGKAAEKKIPKHVREMQEALARR 399
Query: 226 -RDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDG 278
E KE E + K EE ++EEL + K +RK ++ K R L+G
Sbjct: 400 QEAEERKKKEEEEKLRKEEEERRRQEELEAQAEEAKRKRKEKEKEKLLRKKLEG 453
>At1g76810 translation initiation factor IF-2 like protein
Length = 1280
Score = 37.7 bits (86), Expect = 0.008
Identities = 39/173 (22%), Positives = 68/173 (38%), Gaps = 24/173 (13%)
Query: 127 AATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVK---------- 176
AA + P+ + P +S E+ + P+ + E+ +EG + K
Sbjct: 308 AALGETPAAERPASSTPVEEKAA---QPEPVAPVENAGEKEGEEETAAAKKKKKKKEKEK 364
Query: 177 -------FAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEI---- 225
A TS + K KQE + + K++K + + + + E E +
Sbjct: 365 EKKAAAAAAATSSVEVKEEKQEESVTEPLQPKKKDAKGKAAEKKIPKHVREMQEALARRQ 424
Query: 226 RDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDG 278
E KE E + K EE ++EEL + K +RK ++ K R L+G
Sbjct: 425 EAEERKKKEEEEKLRKEEEERRRQEELEAQAEEAKRKRKEKEKEKLLRKKLEG 477
>At1g56660 hypothetical protein
Length = 522
Score = 37.7 bits (86), Expect = 0.008
Identities = 48/215 (22%), Positives = 87/215 (40%), Gaps = 9/215 (4%)
Query: 111 KKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHD 170
K ++ + ++L + E + K E +K++ K + + D+ +E+L ++G
Sbjct: 117 KGKEKKHEELEEEKEGKKKKNKKEKDESGPEEKNKKADKEKKHEDVSQEKEELEEEDGKK 176
Query: 171 AYRPVKFAPTSMDQDKTSKQER----NALRREKEILKESKHSSFIRTLMNDIEEKPEEIR 226
+ K + ++ K K+E+ + E + +K K L + EEK +E
Sbjct: 177 NKKKEKDESGTEEKKKKPKKEKKQKEESKSNEDKKVKGKKEKGEKGDLEKEDEEKKKEHD 236
Query: 227 DYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLK--KSRNGLDGLTE--S 282
+ + KE + K +E+ E + P + + K E K K G G E
Sbjct: 237 ETDQEMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPE 296
Query: 283 FFDEIKTLPFEDETGEQVMGPSKGSNRNGRLKKRK 317
DE K D T EQ M ++ G+ KK K
Sbjct: 297 KEDEGKKTKEHDAT-EQEMDDEAADHKEGKKKKNK 330
Score = 37.0 bits (84), Expect = 0.014
Identities = 32/138 (23%), Positives = 71/138 (51%), Gaps = 5/138 (3%)
Query: 184 QDKTSKQERNALRREKEILKESKHSSFIRT--LMNDIE-EKPEEIRDYEGSSKEVERHIA 240
Q K K+E+ + + EK++ ++ K + + T + DI+ E+PE + E ++E ++ +
Sbjct: 360 QKKNKKKEKKSEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAEKKEEDDTEEKKK--S 417
Query: 241 KMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQV 300
K+E +E + + K+++K K K + + + +S +I+ ++E ++
Sbjct: 418 KVEGGESEEGKKKKKKDKKKNKKKDTKEPKMTEDEEEKKDDSKDVKIEGSKAKEEKKDKD 477
Query: 301 MGPSKGSNRNGRLKKRKA 318
+ KG N G+LK + A
Sbjct: 478 VKKKKGGNDIGKLKTKLA 495
>At5g55660 putative protein
Length = 759
Score = 37.4 bits (85), Expect = 0.011
Identities = 44/212 (20%), Positives = 88/212 (40%), Gaps = 15/212 (7%)
Query: 84 GFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDK 143
G + EE V++ VE + P ++++ ++ ++ G+ + ++++ V DK
Sbjct: 114 GVATEEDAVMKESVESADNKDAENPEGEQEKESKEEKLEGGKANGNEEGDTEEKLVGGDK 173
Query: 144 SEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILK 203
+DV + ++ ++ A +E ++A + ++E N KE K
Sbjct: 174 GDDVDEAEKVENVDEDDKEEALKEKNEA-------------ELAEEEETNKGEEVKEANK 220
Query: 204 ESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHER 263
E + + ++E+K E +D E KE E+ K +E+ ++ ++ K
Sbjct: 221 EDDVEADTKVAEPEVEDKKTESKD-ENEDKEEEKEDEKEDEKEESNDDKEDKKEDIKKSN 279
Query: 264 KREKYLKKSRNGLDGLTESFFD-EIKTLPFED 294
KR K + G E D E KT F D
Sbjct: 280 KRGKGKTEKTRGKTKSDEEKKDIEPKTPFFSD 311
>At3g18610 unknown protein
Length = 636
Score = 37.0 bits (84), Expect = 0.014
Identities = 39/185 (21%), Positives = 84/185 (45%), Gaps = 20/185 (10%)
Query: 109 IDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKS----EDVSKYRPN----------P 154
+ KKQ+ ++ +++ + A +P K E +SD S E+ +K P+
Sbjct: 37 VTKKQKKELIDVVQ--KEKAEKTVPKKVESSSSDASDSDEEEKTKETPSKLKDESSSEEE 94
Query: 155 DMLVSREDLAP-QEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRT 213
D S E++AP ++ + + K +S D D TS +E ++++ +L+++K S
Sbjct: 95 DDSSSDEEIAPAKKRPEPIKKAKVESSSSDDDSTSDEETAPVKKQPAVLEKAKVESSSSD 154
Query: 214 LMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSR 273
+ +E+ ++ ++ + + ++ + +EE VP+ K EK +S
Sbjct: 155 DDSSSDEETVPVKKQPAVLEKAKIESSSSDDDSSSDEE---TVPMKKQTAVLEKAKAESS 211
Query: 274 NGLDG 278
+ DG
Sbjct: 212 SSDDG 216
Score = 35.8 bits (81), Expect = 0.032
Identities = 30/98 (30%), Positives = 42/98 (42%), Gaps = 4/98 (4%)
Query: 108 PIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRP-NPDMLVSREDLAPQ 166
P+ KK+ + K K +++ + S EP + K V +P D S ED +
Sbjct: 248 PVVKKKPTTVVKDAKAESSSSEEESSSDDEPTPAKKPTVVKNAKPAAKDSSSSEEDSDEE 307
Query: 167 EGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKE 204
E D P K A S KTSKQE ++ E KE
Sbjct: 308 ESDDEKPPTKKAKVS---SKTSKQESSSDESSDESDKE 342
Score = 31.2 bits (69), Expect = 0.78
Identities = 32/126 (25%), Positives = 59/126 (46%), Gaps = 10/126 (7%)
Query: 104 EKIRPIDKKQQYQIQKLM---KVGENAATSD---LPSKKEPVASDKSEDVSKYRPNPDML 157
E+I P KK+ I+K ++ +TSD P KK+P +K++ S + D
Sbjct: 101 EEIAPA-KKRPEPIKKAKVESSSSDDDSTSDEETAPVKKQPAVLEKAKVESS--SSDDDS 157
Query: 158 VSREDLAPQEGHDAY-RPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMN 216
S E+ P + A K +S D D +S +E ++++ +L+++K S +
Sbjct: 158 SSDEETVPVKKQPAVLEKAKIESSSSDDDSSSDEETVPMKKQTAVLEKAKAESSSSDDGS 217
Query: 217 DIEEKP 222
+E+P
Sbjct: 218 SSDEEP 223
>At4g12780 auxilin-like protein
Length = 904
Score = 35.4 bits (80), Expect = 0.041
Identities = 47/177 (26%), Positives = 72/177 (40%), Gaps = 20/177 (11%)
Query: 107 RPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVAS--DKSEDVSKYRPNPDMLVSREDLA 164
RPI KK + +S S P AS D+ +D S R N D +
Sbjct: 384 RPIKKKVNEPSIPTSAYHSHVPSSGRASVNSPTASQMDELDDFSIGR-NQTAANGYPDPS 442
Query: 165 PQEGHDAYRPVKFAPTSMDQ--DKTSKQERNAL-RREKEILKESKHSSFIRTLMNDIEEK 221
E D + + +M DK + R+A RREKE LK S+ +
Sbjct: 443 SGEDSDVFSTAAASAAAMKDAMDKAEAKFRHAKERREKENLKASR------------SRE 490
Query: 222 PEEIRDYEGSSKEV-ERHIAKMEERAWQEEELFNRVPLTKHERKRE-KYLKKSRNGL 276
+ +Y+ +E+ E+ + ERA +E E+ K ER+RE K +++ R L
Sbjct: 491 GDHTENYDSRERELREKQVRLDRERAEREAEMEKAQEREKEEREREQKRIERERERL 547
>At2g27280 unknown protein
Length = 764
Score = 34.7 bits (78), Expect = 0.070
Identities = 47/238 (19%), Positives = 89/238 (36%), Gaps = 24/238 (10%)
Query: 86 SIEEHPVVRSIVEI------RLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPV 139
++EE P S E+ + L +++ ++++ IQ LMK E + +
Sbjct: 61 ALEEDPSAFSYDEVYDDMKQKAVLPRMQDREERKPRYIQNLMKQAERREKEHEIVYERKL 120
Query: 140 ASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQER-NALRRE 198
A ++ +D + + +E Q K +ER LR E
Sbjct: 121 AKEREKDEHLFSDKEKFVTGAYKRKLEE----------------QKKWLAEERLRELREE 164
Query: 199 KE-ILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVP 257
++ + K+ S F + ++ E+ E E +R K+EE+ E+ R
Sbjct: 165 RDDVTKKKDLSDFYFNIGKNVAFGAREVEAKEAEKLEEQRKAEKLEEQRKAEKLEELRKE 224
Query: 258 LTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNGRLKK 315
+T+ E+KR+ K+ S ++ L E E+ MG R +K+
Sbjct: 225 VTRVEKKRKSPEKEVSPDSGEFGSSRSKSLEPLEAEQAVSEKEMGSDGTEERKSSIKE 282
>At1g76820 putative translation initiation factor IF-2
Length = 1146
Score = 34.7 bits (78), Expect = 0.070
Identities = 28/105 (26%), Positives = 46/105 (43%), Gaps = 4/105 (3%)
Query: 178 APTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEI----RDYEGSSK 233
A TS + K KQE + + K++K + + + + E E + E K
Sbjct: 259 AATSSVEAKEEKQEESVTEPLQPRKKDAKGKAAEKKIPKHVREIQEALARRQEAKERKKK 318
Query: 234 EVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDG 278
E E + K EE ++EEL + K +RK ++ K R L+G
Sbjct: 319 EEEEKLRKEEEERRRQEELDAQAEEAKRKRKEKEKEKLLRKKLEG 363
>At3g54500 unknown protein
Length = 648
Score = 34.3 bits (77), Expect = 0.092
Identities = 31/115 (26%), Positives = 53/115 (45%), Gaps = 11/115 (9%)
Query: 78 LLRKAKGFSIEEHPVVRSIVEIRLYLE-KIRPIDKKQQYQIQKLMKVGENAATS------ 130
L KA G S + P VR ++ Y E K +P + Q YQ K+MK + TS
Sbjct: 220 LTGKANGLSSQSVPSVRVTLKADQYREHKGQPSVEDQPYQQNKMMKFSKMPGTSEARPFQ 279
Query: 131 DLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEG----HDAYRPVKFAPTS 181
+L ++ P ++ + V++ P L++ L+ EG H ++ P ++ S
Sbjct: 280 ELYGQRIPFSNSAGKCVNQLAPPQSSLMAVNLLSESEGSGTSHYSHMPNQYMANS 334
>At3g55060 centromere protein - like
Length = 896
Score = 33.9 bits (76), Expect = 0.12
Identities = 47/204 (23%), Positives = 83/204 (40%), Gaps = 29/204 (14%)
Query: 59 YLEAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPID-KKQQYQI 117
YL+ + L +LN L Y LL+ KG + + P Y +K D +Q+ I
Sbjct: 627 YLQEQGLSMLNESSQLCYKLLKFIKG-KLTQLP--------ETYQDKNSVKDGLSEQFMI 677
Query: 118 QKLMKV-GENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVK 176
+ MKV G T +L + V S + + N + + + +E A +
Sbjct: 678 ESEMKVHGIRRGTENLKRSLQTVTSVVASNSESSSSNTGRPREQRNQSVEENLRAELSAE 737
Query: 177 FAPTSMDQDKTSKQERN---------ALRREKEILKESKHSSF---------IRTLMNDI 218
TS+ ++K +E+ A R EIL+ SS ++ L + +
Sbjct: 738 TLITSLVREKLYSKEKEIEQLQAELAAAVRGNEILRCEVQSSLDNLSVTTHELKDLKHQM 797
Query: 219 EEKPEEIRDYEGSSKEVERHIAKM 242
+K E IR E + +E + +A++
Sbjct: 798 LKKEESIRRLESNLQEAAKEMARL 821
>At3g54670 structural maintenance of chromosomes (SMC) - like
protein
Length = 1265
Score = 33.9 bits (76), Expect = 0.12
Identities = 26/94 (27%), Positives = 43/94 (45%), Gaps = 5/94 (5%)
Query: 186 KTSKQERNALRREKEILKESKHSSFIRTLMN---DIEEKPEEIRDYEGSSKEVERHIAKM 242
K K+E R +E LK K F+ L N DIE+ E++ + + K+V R + K
Sbjct: 210 KAQKEEAEKHLRLQEELKALKRERFLWQLYNIENDIEKANEDVDSEKSNRKDVMRELEKF 269
Query: 243 EERAWQEEELFNRVPLTKHERKREKYLKKSRNGL 276
E A + + + K +REK + + + L
Sbjct: 270 EREAGKRK--VEQAKYLKEIAQREKKIAEKSSKL 301
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 33.9 bits (76), Expect = 0.12
Identities = 47/264 (17%), Positives = 111/264 (41%), Gaps = 35/264 (13%)
Query: 3 ETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKV-KEGQYPTADGFSYLE 61
E NH + + + + S L A ++ L+ + I+ LT ++ EG+ + S+ E
Sbjct: 1017 ELANHGSEANELQTKLSALEAEKEQTANELEASKTTIEDLTKQLTSEGEKLQSQISSHTE 1076
Query: 62 AKNLLLLNY------CQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPID----- 110
N + + QS++ L + S + +V I ++R + ++
Sbjct: 1077 ENNQVNAMFQSTKEELQSVIAKLEEQLTVESSKADTLVSEIEKLRAVAAEKSVLESHFEE 1136
Query: 111 -KKQQYQIQKLMKVG-ENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEG 168
+K +++ +K ENAAT AS K +++ + + D+ ++
Sbjct: 1137 LEKTLSEVKAQLKENVENAAT----------ASVKVAELTSKLQEHEHIAGERDVLNEQV 1186
Query: 169 HDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDY 228
+ ++ A +S+D+ K + + K+S+ S ++ +IE K + + ++
Sbjct: 1187 LQLQKELQAAQSSIDEQKQAHSQ-----------KQSELESALKKSQEEIEAKKKAVTEF 1235
Query: 229 EGSSKEVERHIAKMEERAWQEEEL 252
E K++E+ + + + + E +
Sbjct: 1236 ESMVKDLEQKVQLADAKTKETEAM 1259
Score = 27.7 bits (60), Expect = 8.6
Identities = 53/268 (19%), Positives = 108/268 (39%), Gaps = 36/268 (13%)
Query: 8 LNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAK---VKEGQYPTAD--------- 55
L +HD+K+KE L E+KE D + +++S K ++EG +A+
Sbjct: 169 LQSHDAKDKE-------LTEVKEAFDALGIELESSRKKLIELEEGLKRSAEEAQKFEELH 221
Query: 56 --GFSYLEAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQ 113
S+ ++++ L + + L+ AK + + + I E+ + + ++
Sbjct: 222 KQSASHADSESQKALEFSE-LLKSTKESAKEMEEKMASLQQEIKELNEKMSENEKVEAAL 280
Query: 114 QYQIQKLMKVGENAA--------TSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAP 165
+ +L V E A T S E + + ++++ + + + +E+L+
Sbjct: 281 KSSAGELAAVQEELALSKSRLLETEQKVSSTEALIDELTQELEQKKASESRF--KEELSV 338
Query: 166 QEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILK--ESKHSSFIRTLMNDIEEKPE 223
+ DA A S + SK +EKE+L+ +RT + E +
Sbjct: 339 LQDLDAQTKGLQAKLSEQEGINSKLAEEL--KEKELLESLSKDQEEKLRTANEKLAEVLK 396
Query: 224 EIRDYEGSSKEVERHIAKMEERAWQEEE 251
E E + EV ++A + E + EE
Sbjct: 397 EKEALEANVAEVTSNVATVTEVCNELEE 424
>At1g65470 hypothetical protein, 3' partial
Length = 618
Score = 33.9 bits (76), Expect = 0.12
Identities = 29/151 (19%), Positives = 70/151 (46%), Gaps = 11/151 (7%)
Query: 130 SDLPSKKEPVASDKSE-DVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTS 188
SDL E + SE D+ + N + +++ + D+ R K +++++
Sbjct: 203 SDLSKAAEKLGKILSEVDIRSFMDN----MMQKNSSEMAEKDSKREEKLLLKQLEKNRC- 257
Query: 189 KQERNALRREKEILKES-----KHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKME 243
+ E+ R E+++LKE + + ++++ ++ EE + K+ + + +
Sbjct: 258 EAEKEKKRMERQVLKEKLQQEKEQKLLQKAIVDENNKEKEETESRKRIKKQQDESEKEQK 317
Query: 244 ERAWQEEELFNRVPLTKHERKREKYLKKSRN 274
R ++ EL ++ + K E++LKKS++
Sbjct: 318 RREKEQAELKKQLQVQKQASIMERFLKKSKD 348
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.130 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,357,109
Number of Sequences: 26719
Number of extensions: 332952
Number of successful extensions: 1548
Number of sequences better than 10.0: 143
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 115
Number of HSP's that attempted gapping in prelim test: 1328
Number of HSP's gapped (non-prelim): 207
length of query: 318
length of database: 11,318,596
effective HSP length: 99
effective length of query: 219
effective length of database: 8,673,415
effective search space: 1899477885
effective search space used: 1899477885
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0316.5


