

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0314.4
(199 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
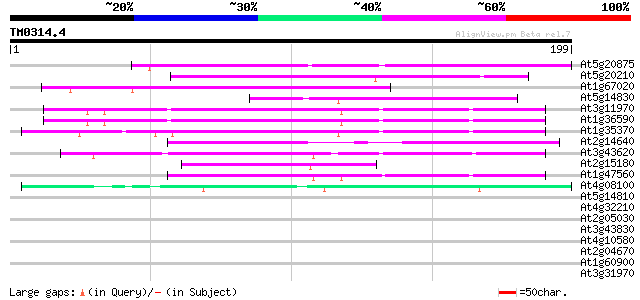
Score E
Sequences producing significant alignments: (bits) Value
At5g20875 unknown protein 85 3e-17
At5g20210 putative protein 80 5e-16
At1g67020 hypothetical protein 74 4e-14
At5g14830 putative protein 59 1e-09
At3g11970 hypothetical protein 54 4e-08
At1g36590 hypothetical protein 54 4e-08
At1g35370 hypothetical protein 54 7e-08
At2g14640 putative retroelement pol polyprotein 50 6e-07
At3g43620 putative protein 46 1e-05
At2g15180 hypothetical protein 45 2e-05
At1g47560 hypothetical protein 45 2e-05
At4g08100 putative polyprotein 44 6e-05
At5g14810 putative protein 39 0.002
At4g32210 putative protein 39 0.002
At2g05030 hypothetical protein 39 0.002
At3g43830 putative protein 35 0.020
At4g10580 putative reverse-transcriptase -like protein 35 0.026
At2g04670 putative retroelement pol polyprotein 34 0.045
At1g60900 putative U2 snRNP auxiliary factor 34 0.045
At3g31970 hypothetical protein 33 0.13
>At5g20875 unknown protein
Length = 272
Score = 84.7 bits (208), Expect = 3e-17
Identities = 45/157 (28%), Positives = 90/157 (56%), Gaps = 4/157 (2%)
Query: 44 HPHRV-HLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKAL 102
HPHR+ + ++++P+F+G + W+ R ER+F+ + E++++V +++EG+AL
Sbjct: 99 HPHRMSRYIQLMKPKIEMPVFEGPNVNSWLTRAERYFEFGSFTDAEKIQLVYMSVEGRAL 158
Query: 103 GWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAP 162
WF E + P W FK +++RF ++ LL++KQ SV++Y FE+ +
Sbjct: 159 CWFN-LENRNPFVDWNDFKARVLQRFGDP--RSAMERLLTLKQVDSVISYLGEFEDLSTQ 215
Query: 163 MRGADREILKGVFLNGLQEEIRAEMKLYPAADLAGLM 199
+L+ VF+ GL+EEI+ ++++ L+ ++
Sbjct: 216 CLDDLDTVLEVVFVKGLKEEIQEMLRVFQPKGLSDII 252
>At5g20210 putative protein
Length = 256
Score = 80.5 bits (197), Expect = 5e-16
Identities = 45/128 (35%), Positives = 68/128 (52%), Gaps = 2/128 (1%)
Query: 58 VDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAW 117
+++P FDG + W+ +E +F L E +++ +MEG AL W Q ++ P W
Sbjct: 65 IEMPSFDGDHLFIWLEIIEVYFTLQLAPEVCKVDTAFRSMEGDALLWLQCLRQENPTLTW 124
Query: 118 EPFKHALIRRF-QPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFL 176
K LI F + +P L S++Q GSV Y F AA + D +++KG+FL
Sbjct: 125 SQLKTELIEEFGGGDITASPEEQLASLRQTGSVDEYANEFRARAAQIGNKD-QLVKGMFL 183
Query: 177 NGLQEEIR 184
NGL+EEIR
Sbjct: 184 NGLKEEIR 191
>At1g67020 hypothetical protein
Length = 659
Score = 74.3 bits (181), Expect = 4e-14
Identities = 42/127 (33%), Positives = 66/127 (51%), Gaps = 3/127 (2%)
Query: 12 RQASRERDR-EASSHSGSRTGSRFSDHSRERF--DHPHRVHLRAVTGRRVDIPMFDGSDA 68
R RE R E HS FS +R + P + R+ RR+++P+FDGS
Sbjct: 61 RMEIREGKRVEQGEHSCQSPRHSFSSSMPQRGAGESPILLDNRSSLIRRIEMPVFDGSGV 120
Query: 69 YGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRF 128
Y W ++VERFF++ R + ++L++V +++EG AL WF R W F+ L+ RF
Sbjct: 121 YEWFSKVERFFRVGRYQDSDKLDLVALSLEGVALKWFLREMSTLEFRDWNSFEQRLLARF 180
Query: 129 QPTLVQN 135
P + +
Sbjct: 181 DPVKIHS 187
>At5g14830 putative protein
Length = 757
Score = 59.3 bits (142), Expect = 1e-09
Identities = 33/97 (34%), Positives = 50/97 (51%), Gaps = 4/97 (4%)
Query: 86 EEERLEMVMIAMEGKALGWFQWWEEQAPER--AWEPFKHALIRRFQPTLVQNPFGPLLSI 143
E ++LE + A+ G A+ W W Q PE+ W+ F+ RF+P L +I
Sbjct: 620 ETQKLEQAITALTGTAINW--WQIAQYPEKISTWKDFRDKFKVRFKPARGAATIDQLFTI 677
Query: 144 KQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQ 180
Q+GSV YR FEE A + ++L+ FLNG++
Sbjct: 678 FQRGSVEEYRNRFEEIAVELPHVTDDVLESAFLNGVE 714
>At3g11970 hypothetical protein
Length = 1499
Score = 54.3 bits (129), Expect = 4e-08
Identities = 44/187 (23%), Positives = 85/187 (44%), Gaps = 12/187 (6%)
Query: 13 QASRERDREASSHS-GSRTGS----RFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGSD 67
Q++ +R SS S +R+G R + H + R DH + + G+ +D P FDG+
Sbjct: 63 QSTLDRSAPRSSQSMENRSGYPDPYRDARHQQVRSDHFNAYNNLTRLGK-IDFPRFDGTR 121
Query: 68 AYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA----WEPFKHA 123
W+ +VE FF + E+ +++M I + A W Q + + W+ +
Sbjct: 122 LKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTWHQSFIQSGVGLEVLYDWKGYVKL 181
Query: 124 LIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEI 183
L RF+ +P L +++ ++ Y + FE + E L V+L GL+ +
Sbjct: 182 LKERFEDD-CDDPMAELKHLQETDGIIDYHQKFELIKTRV-NLSEEYLVSVYLAGLRTDT 239
Query: 184 RAEMKLY 190
+ ++++
Sbjct: 240 QMHVRMF 246
>At1g36590 hypothetical protein
Length = 1499
Score = 54.3 bits (129), Expect = 4e-08
Identities = 44/187 (23%), Positives = 85/187 (44%), Gaps = 12/187 (6%)
Query: 13 QASRERDREASSHS-GSRTGS----RFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGSD 67
Q++ +R SS S +R+G R + H + R DH + + G+ +D P FDG+
Sbjct: 63 QSTLDRSAPRSSQSMENRSGYLDPYRDARHQQVRSDHFNAYNNLTRLGK-IDFPRFDGTR 121
Query: 68 AYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA----WEPFKHA 123
W+ +VE FF + E+ +++M I + A W Q + + W+ +
Sbjct: 122 LKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTWHQSFIQSGVGLEVLYDWKGYVKL 181
Query: 124 LIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEI 183
L RF+ +P L +++ ++ Y + FE + E L V+L GL+ +
Sbjct: 182 LKERFEDD-CDDPMAELKHLQETDGIIDYHQKFELIKTRV-NLSEEYLVSVYLAGLRTDT 239
Query: 184 RAEMKLY 190
+ ++++
Sbjct: 240 QMHVRMF 246
>At1g35370 hypothetical protein
Length = 1447
Score = 53.5 bits (127), Expect = 7e-08
Identities = 46/201 (22%), Positives = 84/201 (40%), Gaps = 18/201 (8%)
Query: 5 RSNTPRFRQASRERDREAS-SHSGSRTGSRFSDHSRERFDHPHRVHL--RAVTGR----- 56
R+ R + + DR S + S S F +H R+ F R + + V +
Sbjct: 54 RTTAKRVTEGEKVLDRSVPRSSTSSMARSGFVEHQRD-FQRDFRPEMVRQEVNNQYGNLT 112
Query: 57 ---RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAP 113
++D P FDGS W+ +VE FF + EE +++MV I + A W + +
Sbjct: 113 RLGKIDFPRFDGSRINEWLFKVEEFFGVDFTPEEMKVKMVAIHFDSHAATWHHSFIQSGI 172
Query: 114 ER----AWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADRE 169
W + L RF+ +P L +++ ++ Y + FE + E
Sbjct: 173 GLDVFFNWPEYVKLLKDRFEDA-CDDPMAELKKLQETDGIVEYHQQFELIKVRL-NLSEE 230
Query: 170 ILKGVFLNGLQEEIRAEMKLY 190
L V+L GL+ + + ++++
Sbjct: 231 YLVSVYLAGLRTDTQMHVRMF 251
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 50.4 bits (119), Expect = 6e-07
Identities = 33/139 (23%), Positives = 59/139 (41%), Gaps = 28/139 (20%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P F G D W+++ +++F+ E+ + ++G A GW+Q
Sbjct: 105 KLDFPRFHGGDLTKWLSKAKQYFEYHEGPVEQAVRFTTFHLDGVANGWWQ---------- 154
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFL 176
A +RR +I Q GS+ Y+ FE + G +E+L G F+
Sbjct: 155 ------ATLRR------------SATIGQTGSLSEYQHEFEWLQNKVYGWTQEVLVGAFM 196
Query: 177 NGLQEEIRAEMKLYPAADL 195
NGL I ++++ L
Sbjct: 197 NGLYYSISNGIRMFQPRSL 215
>At3g43620 putative protein
Length = 221
Score = 45.8 bits (107), Expect = 1e-05
Identities = 39/181 (21%), Positives = 76/181 (41%), Gaps = 15/181 (8%)
Query: 19 DREASSHSGS---RTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGSDAYGWINRV 75
DR +S GS ++G +H H LR + +VD P F+G W+ ++
Sbjct: 29 DRSSSMVLGSPESQSGCSDPNHDVRSDSQYHYRRLRRLG--KVDFPRFNGDGIKDWLFQI 86
Query: 76 ERFFQLSRVGEEERLEMVMIAMEGKALGWFQ------WWEEQAPERAWEPFKHALIRRFQ 129
E+FF + EE ++++ I + W Q W + W +K L R+
Sbjct: 87 EQFFLIDHTPEELKVDIASIHFDDIDATWHQSIVQSIMWRHVRHD--WWNYKLLLQVRYN 144
Query: 130 PTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKL 189
V + L +++ + Y FE ++ D + L ++L GL+ + + +++
Sbjct: 145 KH-VDDSIAKLKLLQETEGIEVYHARFESICTRVK-LDEDFLVSLYLTGLKTDTQVNIRM 202
Query: 190 Y 190
+
Sbjct: 203 F 203
>At2g15180 hypothetical protein
Length = 474
Score = 45.4 bits (106), Expect = 2e-05
Identities = 19/75 (25%), Positives = 38/75 (50%), Gaps = 6/75 (8%)
Query: 62 MFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWF------QWWEEQAPER 115
+F G W + + +F+ +E++L + + ++G AL W+ +W+E +AP R
Sbjct: 113 LFSGLGYLQWESNMNYYFEFHSTAQEDKLSIALGQLKGSALWWWDQDEYNRWYERRAPIR 172
Query: 116 AWEPFKHALIRRFQP 130
WE K + ++ P
Sbjct: 173 TWERLKWNMCAKYSP 187
>At1g47560 hypothetical protein
Length = 1564
Score = 45.4 bits (106), Expect = 2e-05
Identities = 31/138 (22%), Positives = 60/138 (43%), Gaps = 6/138 (4%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQ-WWEEQAPER 115
++D F G W+ ++E+FF L EE ++ + +G A W++ +E +
Sbjct: 967 KLDFHRFSGDRVRDWLFKLEQFFSLDFTPEELKVSIASTLCDGAAAKWYKSLFESDFGVK 1026
Query: 116 A---WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILK 172
W +K L F L +P L +K+ + Y + FE A + + L
Sbjct: 1027 LLSNWNMYKLLLEEHFAEVL-DDPISELKQLKETNGIEEYHKKFELLRARV-NLSEDYLV 1084
Query: 173 GVFLNGLQEEIRAEMKLY 190
V+L+GL + + + ++
Sbjct: 1085 RVYLDGLHPDTQMNVNMF 1102
>At4g08100 putative polyprotein
Length = 1054
Score = 43.9 bits (102), Expect = 6e-05
Identities = 49/211 (23%), Positives = 77/211 (36%), Gaps = 30/211 (14%)
Query: 5 RSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFD 64
R PR + +R ASSHS R H RER P R L G ++ IP F
Sbjct: 52 RQEEPREDEEARNYYSHASSHSSQRR------HRRER--PPPRDPL---GGLKLKIPAFH 100
Query: 65 GSD----AYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEE---------Q 111
G+D W ++E F + ++++ AL WW++
Sbjct: 101 GTDNPDTYLEWEQKIELVFLCQECLQSNKVKIAATKFYNYAL---SWWDQLVTSRRRTRD 157
Query: 112 APERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRG---ADR 168
P + W K + +RF P+ L +GS E +R D
Sbjct: 158 YPIKTWNQLKFVMRKRFVPSYYHRELHQRLRNLVQGSKTVEEYFLEMETLMLRADLQEDG 217
Query: 169 EILKGVFLNGLQEEIRAEMKLYPAADLAGLM 199
E + F+ GL EI+ ++ +L ++
Sbjct: 218 EAVMSRFMGGLNREIQDRLETQHYVELEEML 248
>At5g14810 putative protein
Length = 716
Score = 38.9 bits (89), Expect = 0.002
Identities = 34/157 (21%), Positives = 65/157 (40%), Gaps = 21/157 (13%)
Query: 40 ERFDHPHRVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEG 99
ER D HLR F+G W+ ++E+FF + R EE ++ + +
Sbjct: 99 ERSDFARLEHLR-----------FNGDRITEWLFQIEQFFLIDRTPEELKVGFASLHFDD 147
Query: 100 KALGWFQ------WWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYR 153
A Q WW+ + W +K L R+ V + L +++ + Y
Sbjct: 148 TAATLHQSIVQSMWWKHVRHD--WWSYKLLLQVRYDEH-VNDSIAKLTQLQETEGIEEYH 204
Query: 154 ESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKLY 190
FE + + A+ + L V+L GL+ + + ++++
Sbjct: 205 ARFELISTRLNFAE-DYLVSVYLAGLRTDTQLNVRMF 240
>At4g32210 putative protein
Length = 1086
Score = 38.9 bits (89), Expect = 0.002
Identities = 34/157 (21%), Positives = 65/157 (40%), Gaps = 21/157 (13%)
Query: 40 ERFDHPHRVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEG 99
ER D HLR F+G W+ ++E+FF + R EE ++ + +
Sbjct: 99 ERSDFARLEHLR-----------FNGDRITEWLFQIEQFFLIDRTPEELKVGFASLHFDD 147
Query: 100 KALGWFQ------WWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYR 153
A Q WW+ + W +K L R+ V + L +++ + Y
Sbjct: 148 TAATLHQSIVQSMWWKHVRHD--WWSYKLLLQVRYDEH-VNDSIAKLTQLQETEGIEEYH 204
Query: 154 ESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKLY 190
FE + + A+ + L V+L GL+ + + ++++
Sbjct: 205 ARFELISTRLNFAE-DYLVSVYLAGLRTDTQLNVRMF 240
>At2g05030 hypothetical protein
Length = 205
Score = 38.5 bits (88), Expect = 0.002
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 6/75 (8%)
Query: 36 DHSRE-RFDH----PHRVHLRAVTGR-RVDIPMFDGSDAYGWINRVERFFQLSRVGEEER 89
DH R+ R D P + H +T ++D+P FDG+ W+ +VE FF + + +
Sbjct: 106 DHRRDNRLDQVRYEPQQQHYGNLTRLGKIDLPRFDGTGIKEWLFKVEEFFGIDFTPADLK 165
Query: 90 LEMVMIAMEGKALGW 104
++MV I + A W
Sbjct: 166 VKMVAIHFDSHAATW 180
>At3g43830 putative protein
Length = 329
Score = 35.4 bits (80), Expect = 0.020
Identities = 39/144 (27%), Positives = 57/144 (39%), Gaps = 20/144 (13%)
Query: 56 RRVDIPMFDGSD----AYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQ 111
R +DI F+G AY W N +E F+ R E ++ + G+A +WWE
Sbjct: 109 RDLDIETFNGGTDPVKAYNWRNMLECKFKSMRCPVEFWRDLASCYLRGEAQ---EWWERV 165
Query: 112 APER------AWEPFKHALIRRFQPTLVQNPFGPLLSIKQKG--SVMAYRESF---EEAA 160
W FK RR+ P + Q+G +V Y + F E
Sbjct: 166 KQREQVGCVDQWSFFKEEFTRRYLPEETIDDLEMKFLRLQQGTKTVRKYEKEFHSLERFE 225
Query: 161 APMRGADREILKGVFLNGLQEEIR 184
RG I K F++GL+ +IR
Sbjct: 226 RRKRGEHELIHK--FISGLRVDIR 247
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 35.0 bits (79), Expect = 0.026
Identities = 36/153 (23%), Positives = 58/153 (37%), Gaps = 35/153 (22%)
Query: 56 RRVDIPMFDGS----DAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQ 111
+R+D F G +A W +RV R F SR E R+++ + +EG A WW
Sbjct: 145 QRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLEGDA---HLWWRSV 201
Query: 112 APER-----AWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGA 166
R +W F ++ P +P+ + E+ A MR
Sbjct: 202 TARRRQTDMSWADFVAEFKAKYFPQEALDPYA--------------GQGMEDDQAQMRR- 246
Query: 167 DREILKGVFLNGLQEEIRAEMKLYPAADLAGLM 199
FL GL+ ++R ++ A A L+
Sbjct: 247 --------FLRGLRPDLRVRCRVSQYATKAALV 271
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 34.3 bits (77), Expect = 0.045
Identities = 36/142 (25%), Positives = 58/142 (40%), Gaps = 12/142 (8%)
Query: 67 DAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPER-----AWEPFK 121
+A W +RV+R F SR E R+++ + +EG A WW R +W F
Sbjct: 146 EADSWRSRVQRNFGSSRCPAEYRVDLAVHFLEGDA---HLWWRSVTARRRQADMSWADFV 202
Query: 122 HALIRRFQP-TLVQNPFGPLLSIKQ-KGSVMAYRESFEE--AAAPMRGADREILKGVFLN 177
++ P + L + Q + SV Y F A A D + FL
Sbjct: 203 AEFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRLLAYAGRGMEDDQAQMRRFLR 262
Query: 178 GLQEEIRAEMKLYPAADLAGLM 199
GL+ ++R + ++ A A L+
Sbjct: 263 GLRPDLRVQCRVSQYATKAALV 284
>At1g60900 putative U2 snRNP auxiliary factor
Length = 589
Score = 34.3 bits (77), Expect = 0.045
Identities = 18/48 (37%), Positives = 25/48 (51%)
Query: 5 RSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRA 52
R + R R+ SR+RDRE SR+ SR R R +H H+ R+
Sbjct: 128 RRSRDRDRRRSRDRDREVRHRRRSRSRSRSRSERRSRSEHRHKSEHRS 175
Score = 26.9 bits (58), Expect = 7.1
Identities = 19/52 (36%), Positives = 25/52 (47%), Gaps = 2/52 (3%)
Query: 10 RFRQASRERDREASSHSGSRTGSRFSDHSRERFDHP-HRVHLRAVTGRRVDI 60
R R+ SR+RDRE S R R D R+R HR R + +R D+
Sbjct: 72 RDREKSRDRDRE-KSRDRDRDRERSKDRQRDRHHRDRHRDRSRERSEKRDDL 122
>At3g31970 hypothetical protein
Length = 1329
Score = 32.7 bits (73), Expect = 0.13
Identities = 35/144 (24%), Positives = 58/144 (39%), Gaps = 16/144 (11%)
Query: 67 DAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPER-----AWEPFK 121
+A W +RVE F SR E R+++ + +EG A WW R +W F
Sbjct: 166 EADSWKSRVEHNFGSSRCPAEYRVDLAVHFLEGDA---HLWWRSVTARRRQAHMSWADFV 222
Query: 122 HALIRRFQP-TLVQNPFGPLLSIKQ-KGSVMAYRESFEE----AAAPMRGADREILKGVF 175
++ P + L + Q + SV Y F A M+ ++ + F
Sbjct: 223 AEFNAKYFPQEALDRMEARFLELTQGERSVREYEREFNRLLVYAGRGMQDDQAQMRR--F 280
Query: 176 LNGLQEEIRAEMKLYPAADLAGLM 199
L GL+ ++R ++ A A L+
Sbjct: 281 LRGLRPDLRVRCRVLQYATKAALV 304
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,288,755
Number of Sequences: 26719
Number of extensions: 174562
Number of successful extensions: 563
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 511
Number of HSP's gapped (non-prelim): 62
length of query: 199
length of database: 11,318,596
effective HSP length: 94
effective length of query: 105
effective length of database: 8,807,010
effective search space: 924736050
effective search space used: 924736050
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0314.4


