

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0310b.12
(1485 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
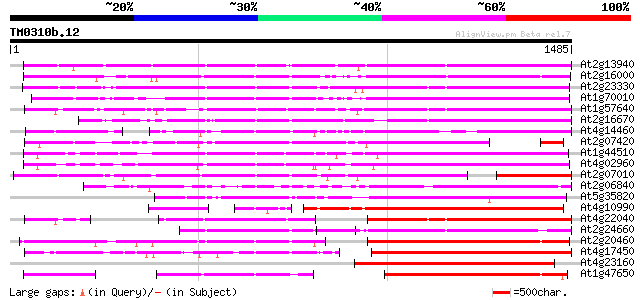
Score E
Sequences producing significant alignments: (bits) Value
At2g13940 putative retroelement pol polyprotein 874 0.0
At2g16000 putative retroelement pol polyprotein 847 0.0
At2g23330 putative retroelement pol polyprotein 844 0.0
At1g70010 hypothetical protein 838 0.0
At1g57640 824 0.0
At2g16670 putative retroelement pol polyprotein 701 0.0
At4g14460 retrovirus-related like polyprotein 683 0.0
At2g07420 putative retroelement pol polyprotein 667 0.0
At1g44510 polyprotein, putative 660 0.0
At4g02960 putative polyprotein of LTR transposon 643 0.0
At2g07010 putative retroelement pol polyprotein 625 e-179
At2g06840 putative retroelement pol polyprotein 560 e-159
At5g35820 copia-like retrotransposable element 557 e-158
At4g10990 putative retrotransposon polyprotein 528 e-150
At4g22040 LTR retrotransposon like protein 513 e-145
At2g24660 putative retroelement pol polyprotein 489 e-138
At2g20460 putative retroelement pol polyprotein 488 e-138
At4g17450 retrotransposon like protein 485 e-137
At4g23160 putative protein 457 e-128
At1g47650 hypothetical protein 452 e-127
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 874 bits (2257), Expect = 0.0
Identities = 533/1493 (35%), Positives = 816/1493 (53%), Gaps = 81/1493 (5%)
Query: 36 SFRLDGRNYLQWSELVRHTLISRQKISYIEETAPAE--TDPQYDTWREENSLIMTWFWYS 93
S L+G NY QW+ + + L +++K +I T P DP Y+ W NS+I+ W S
Sbjct: 46 SVELNGDNYNQWATEMLNALQAKRKTGFINGTIPRPPPNDPNYENWTAVNSMIVGWIRTS 105
Query: 94 MIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLN 153
+ P++ F A +W +L Q +S + +++ ++ + +Q +V +YYG L+
Sbjct: 106 IEPKVKATVTFISDAHLLWKDLKQRFSVGNKVRI-HQIRAQLSSCRQDGQAVIEYYGRLS 164
Query: 154 GLWIELDQYQNLKM------KCDSDSTTLNKFIEKARIFKFLSGLN-SEFDPIRVQILGK 206
LW E + Y+ + + +C + S K E+ +I +F+ GL+ S F + ++
Sbjct: 165 NLWEEYNIYKPVTVCTCGLCRCGATSEP-TKEREEEKIHQFVLGLDESRFGGLCATLINM 223
Query: 207 EQFPSLSEVFS-IVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRD 265
+ PSL E++S ++R E+ +V V ++ + + + + +R D +
Sbjct: 224 DPLPSLGEIYSRVIREEQRLASVHVREQKEEAVGFLARREQLD------HHSRVDASSSR 277
Query: 266 DRFNGDDRFNRDDR----CTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQR-RANHT 320
G R N + C+ C R+GH K+ C+++ G + + + +G+ R R
Sbjct: 278 SEHTGGSRSNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERNGGRGSNGRGRGGRG 337
Query: 321 ASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNA 380
++ G+V S+ E + L K + T S L+
Sbjct: 338 SNGGRGQGQVMAAHATSSNSSVFPEFTEEHMRVLSQLVKEKSNSGSTSNNNSDR---LSG 394
Query: 381 LSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPL 440
+ D I+D GA+ HMT S L++ P + A+GS+ G + LS ++ L
Sbjct: 395 KTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCP-VGFADGSKAFALSVGVLTLSNTVSL 453
Query: 441 KRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQH- 499
VL VP L+ L+S+ KL + C TF + QD ++ +IG +E+ G+Y L
Sbjct: 454 TNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFLQDRSSKTLIGSGEERGGVYYLTDV 513
Query: 500 EESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVS--VESFHCDVCQFSK 557
+K H + +S +W H+RLGHP FS+L ++ LFSK S V S CDVC +K
Sbjct: 514 TPAKIHTANVDS--DQALW--HQRLGHPSFSVLSSL--PLFSKTSSTVTSHSCDVCFRAK 567
Query: 558 HHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSE 617
R F S NK + F L+H DVWGP + GA +F+T +DD +R W +L+ +KSE
Sbjct: 568 QTREVFPESINKTEECFSLIHCDVWGPYRVPASCGAVYFLTIVDDYSRAVWTYLLLEKSE 627
Query: 618 VFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGV 677
V Q+ NF QFGK +K +RSDNG E + S + +NG++H+ +CV PQQNG
Sbjct: 628 VRQVLTNFLKYAEKQFGKTVKMVRSDNGTEFMC--LSSYFRENGIIHQTSCVGTPQQNGR 685
Query: 678 AERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPS 737
ERK+RH+L + RALLFQ ++P +WGE++LTAAYLINR PS +LS +P V+ S
Sbjct: 686 VERKHRHILNVARALLFQASLPIKFWGESILTAAYLINRTPSSILSGRTPYEVLHG---S 742
Query: 738 VPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYIS 797
P+ S Q RVFG + +VH + K R+ CIF+GY KKG+K Y +S
Sbjct: 743 KPVYS--QLRVFGSACYVHRVTRDKDKFGQRSRSCIFVGYPFGKKGWKVYDIERNEFLVS 800
Query: 798 MDVTFQESESFYPNPQLQGGDTQEAESPELS-----VIPLLQ-----EPLMPSSIPITED 847
DV F+E +P + P +S IP L+ + + + T D
Sbjct: 801 RDVIFREE--VFPYAGVNSSTLASTSLPTVSEDDDWAIPPLEVRGSIDSVETERVVCTTD 858
Query: 848 SD--DDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKV-KPVLI 904
D ++PNQ E VPD + P V + + V P+ +
Sbjct: 859 EVVLDTSVSDSEIPNQ------EFVPDDTPPSSPLSVSPSGSPNTPTTPIVVPVASPIPV 912
Query: 905 QQQDQPSDPEVSVPNPE------------PSNSYEHNLDDLPIALRKGKRSCAKYPISQY 952
Q + P P+ PS+ + D + GK + +P++ Y
Sbjct: 913 SPPKQRKSKRATHPPPKLNDYVLYNAMYTPSSIHALPADPSQSSTVPGK---SLFPLTDY 969
Query: 953 VSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPND 1012
VS S H+++++AI P +E ++ K W AM E+ AL+ N T IV+ P
Sbjct: 970 VSDAAFSSSHRAYLAAITDNVEPKHFKEAVQIKVWNDAMFTEVDALEINKTWDIVDLPPG 1029
Query: 1013 KRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLA 1072
K +G +W++ KY SDGT++RYKARLV +G Q G DY ETFAPV +M TVR +L
Sbjct: 1030 KVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGEDYKETFAPVVRMTTVRTLLRNV 1089
Query: 1073 AHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWF 1132
A WE+ Q DV NAFLHGDLEEEVYM++PPG+ S +KVC+L+K+LYGLKQ+PR WF
Sbjct: 1090 AANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRH-SHPDKVCRLRKSLYGLKQAPRCWF 1148
Query: 1133 GRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAV 1192
+ +++++ G+ QS D++LF +TRN +L+YV+D++I GND Q + L+
Sbjct: 1149 KKLSDSLLRFGFVQSYEDYSLF-SYTRNNIELRVLIYVDDLLICGNDGYMLQKFKDYLSR 1207
Query: 1193 QFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGS 1252
F MKDLGKLKYFLG+EV+ +GIF+SQRKY LD++ ++G LG +P P+EQNH + S
Sbjct: 1208 CFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDVIADSGNLGSRPAHTPLEQNHHLAS 1267
Query: 1253 SEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKA 1312
+ L Y+RLVG+L+YL HTRP+++Y V V++QFM +P+E H A R+++YLK
Sbjct: 1268 DDGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFMQNPREAHFDAALRVVRYLKG 1327
Query: 1313 SPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSS 1372
SPG+G+L L++EVY D+D+ + RRS + Y + LGG+ ++W++KKQ+ V+ SS
Sbjct: 1328 SPGQGILLNADPDLTLEVYCDSDWQSCPLTRRSISAYVVLLGGSPISWKTKKQDTVSHSS 1387
Query: 1373 AEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEI 1432
AEAE+RAM+ + E+ ++ +L +L I+ P RL+CD+K+AI IA NPV H+RTKHIE
Sbjct: 1388 AEAEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSKAAIHIAANPVFHERTKHIES 1447
Query: 1433 DRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
D H +++ + +I+TQ+V + QLADV TK L +F L KLG+ ++H+P
Sbjct: 1448 DCHSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQFLYLMSKLGVQNLHTP 1500
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 847 bits (2188), Expect = 0.0
Identities = 523/1489 (35%), Positives = 801/1489 (53%), Gaps = 153/1489 (10%)
Query: 36 SFRLDGRNYLQWSELVRHTLISRQKISYIEETA--PAETDPQYDTWREENSLIMTWFWYS 93
S RLD NY WS + +L ++ K +I+ T P E+D + W NS++ +W S
Sbjct: 76 SHRLDETNYGDWSVAMLISLDAKNKTGFIDGTLSRPLESDLNFRLWSRCNSMVKSWLLNS 135
Query: 94 MIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLN 153
+ P+I R+ + A +IW +L ++ +L Y L ++ + +QGTLS+++YY L
Sbjct: 136 VSPQIYRSILRMNDASDIWRDLNSRFNVT-NLPRTYNLTQEIQDFRQGTLSLSEYYTRLK 194
Query: 154 GLWIELDQYQNLKMKCD-SDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGKEQFPSL 212
LW +LD + L C + L + E+A+I KFL+GLN + +R QI+ K+ PSL
Sbjct: 195 TLWDQLDSTEALDEPCTCGKAMRLQQKAEQAKIVKFLAGLNESYAIVRRQIIAKKALPSL 254
Query: 213 SEVFSIVRGEETRRTV--------------MVEDKSVDGSALASGKGPIKGSTSFGRPNR 258
EV+ I+ + ++++ + + S+D + GP KG
Sbjct: 255 GEVYHILDQDNSQQSFSNVVAPPAAFQVSEITQSPSMDPTVCYVQNGPNKG--------- 305
Query: 259 DDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRRAN 318
RP C++ R GH E C+K +G K+ G ++
Sbjct: 306 --RPI----------------CSFYNRVGHIAERCYKKHGFPPGFTPKGKA-GEKLQKPK 346
Query: 319 HTASDMENDGEVPPVPPAEDVPSLSKAELER-LRVFMDSLSK-PSGSCSLTMTGKS---- 372
A+++ EV + V +LSK +L++ + +F L P + + T +S
Sbjct: 347 PLAANVAESSEVNTSLESM-VGNLSKEQLQQFIAMFSSQLQNTPPSTYATASTSQSDNLG 405
Query: 373 -----STFLSLNALSTEND------WIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANG 421
ST+ + L+ W+ID GAT H++ S SS T + + G
Sbjct: 406 ICFSPSTYSFIGILTVARHTLSSATWVIDSGATHHVSHDRSLFSSLDTSV-LSAVNLPTG 464
Query: 422 SQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLAT 481
+I+G G + L+ + LK VL +P+ NL+SI LT D+ V F + QDL
Sbjct: 465 PTVKISGVGTLKLNDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIK 524
Query: 482 GQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFS 541
G+++G + LY L + + I + H+RLGH L + L +
Sbjct: 525 GRMLGQGRRVANLYLLDVGDQSISVNA-----VVDISMWHRRLGHASLQRLDAISDSLGT 579
Query: 542 K--VSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTF 599
+ S C VC +K + +F SN C + FDL+H DVWGP S+ V G K+F+T
Sbjct: 580 TRHKNKGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTI 639
Query: 600 IDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTK 659
+DD +R TW++L+K KSEV +F F + Q+ +K +RSDN E F+ F +
Sbjct: 640 VDDHSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPE---LKFTSFYAE 696
Query: 660 NGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPS 719
G+V +C + P+QN V ERK++H+L + RAL+FQ VP WG+ VLTA +LINR PS
Sbjct: 697 KGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPS 756
Query: 720 RVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYAS 779
++L N +P ++T P Q R FGC + R K PR+ C+F+GY S
Sbjct: 757 QLLMNKTPYEILTGTAPVYE-----QLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPS 811
Query: 780 DKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLMP 839
KGYK S V+IS +V F E +P + G ++ L P++P
Sbjct: 812 GYKGYKLMDLESNTVFISRNVQFHEE--VFPLAKNPGSESSLK----------LFTPMVP 859
Query: 840 SSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKV 899
S I D+ + P +P+Q D +P Q I QR
Sbjct: 860 VSSGIISDTT---HSPSSLPSQISD-----------------LPPQ-------ISSQRVR 892
Query: 900 KPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLS 959
KP P N Y N +S KYPIS +S +S
Sbjct: 893 KP------------------PAHLNDYHCNT----------MQSDHKYPISSTISYSKIS 924
Query: 960 VQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCK 1019
H +I+ I I +PT+ E K+W +A++ E+ A++K T +I P K+ VGCK
Sbjct: 925 PSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCK 984
Query: 1020 WIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWEL 1079
W++T+K+ +DG L+RYKARLVAKGYTQ G+DY +TF+PVAKM T++++L ++A W L
Sbjct: 985 WVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWFL 1044
Query: 1080 QQFDVKNAFLHGDLEEEVYMEIPPGYDPTSG----RNKVCKLKKALYGLKQSPRAWFGRF 1135
+Q DV NAFL+G+LEEE++M+IP GY G N V +LK+++YGLKQ+ R WF +F
Sbjct: 1045 KQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKKF 1104
Query: 1136 TNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFE 1195
+++++SLG+K++ GDHTLF+K +G+ ++LVYV+D++IA E L +EL +F+
Sbjct: 1105 SSSLLSLGFKKTHGDHTLFLK-MYDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQRFK 1163
Query: 1196 MKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEE 1255
++DLG LKYFLG+EVA + GI I QRKY L+LL+ TG L CKP+ VP+ N K+ +
Sbjct: 1164 LRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNLKMRKDDG 1223
Query: 1256 DLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPG 1315
DL QY+R+VGKL+YL+ TRPDI + V+ + QF P+ HL A R+LQY+K + G
Sbjct: 1224 DLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVG 1283
Query: 1316 RGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEA 1375
+GL + S L+++ + D+D+A RRSTT + MF+G +L++WRSKKQ+ V+RSSAEA
Sbjct: 1284 QGLFYSASSDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEA 1343
Query: 1376 EFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRH 1435
E+RA+A CE++ + +L L+ P+ L+ D+ +AI IA NPV H+RTKHI++D H
Sbjct: 1344 EYRALALATCEMVWLFTLLVSLQASPPVPI-LYSDSTAAIYIATNPVFHERTKHIKLDCH 1402
Query: 1436 FIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHS 1484
++E+L++ + +V + Q+AD+LTK L +F L K+ +++I S
Sbjct: 1403 TVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSILNIFS 1451
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 844 bits (2181), Expect = 0.0
Identities = 531/1503 (35%), Positives = 790/1503 (52%), Gaps = 98/1503 (6%)
Query: 33 LQGSFRLDGRNYLQWSELVRHTLISRQKISYIEETAPAET-DPQYDTWREENSLIMTWFW 91
L S L NY +WSE +++ L ++QK+ +I+ + P DP+ W NS+I+ W
Sbjct: 33 LISSVVLKENNYAEWSEELQNFLRAKQKLGFIDGSIPKPAADPELSLWIAINSMIVGWIR 92
Query: 92 YSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGT 151
S+ P I F A ++W NL + +S + L++++ Q V YYG
Sbjct: 93 TSIDPTIRSTVGFVSEASQLWENLRRRFSVGNGVRKTL-LKDEIAACTQDGQPVLAYYGR 151
Query: 152 LNGLWIELDQYQN-LKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGKEQFP 210
L LW EL Y++ + KC++ S + K E R+ KFL GL+S F IR I E P
Sbjct: 152 LIKLWEELQNYKSGRECKCEAASD-IEKEREDDRVHKFLLGLDSRFSSIRSSITDIEPLP 210
Query: 211 SLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNG 270
L +V+S V EE V A+ ++ ST+ RF
Sbjct: 211 DLYQVYSRVVREEQNLNASRTKDVVKTEAIGFS---VQSSTT-------------PRFRD 254
Query: 271 DDRFNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRR---ANHTASDMEND 327
CT+C R GH CF ++G + + + P R +N S
Sbjct: 255 KSTLF----CTHCNRKGHEVTQCFLVHGYPDWWLEQNPQENQPSTRGRGSNGRGSSSGRG 310
Query: 328 GEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGK-SSTFLSLNALSTEND 386
G P + A+ V D + + SL + SS+ L+ + D
Sbjct: 311 GNRSSAPTTRGRGRANNAQAAAPTVSGDGNDQIAQLISLLQAQRPSSSSERLSGNTCLTD 370
Query: 387 WIIDFGATDHMTPHPSHLSS-YSTYPGKHYITVANGSQNQITGCGNIHLSPSLPLKRVLH 445
+ID GA+ HMT S L + P +T +G +Q T CG + L S L VL
Sbjct: 371 GVIDTGASHHMTGDCSILVDVFDITPSP--VTKPDGKASQATKCGTLLLHDSYKLHDVLF 428
Query: 446 VPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEES-KC 504
VP L+S+ KL + + F + QD +IG +E++G+Y + +
Sbjct: 429 VPDFDCTLISVSKLLKQTSSIAIFTDTFCFLQDRFLRTLIGAGEEREGVYYFTGVLAPRV 488
Query: 505 HQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFH-CDVCQFSKHHRSTF 563
H+ SS+ + +W H+RLGHP S+L ++ S + CD C SK R F
Sbjct: 489 HKASSDFAISGDLW--HRRLGHPSTSVLLSLPECNRSSQGFDKIDSCDTCFRSKQTREVF 546
Query: 564 LPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFA 623
SNNK + F L+H DVWGP + +GA +F+T +DD +R W +LM K+EV QL
Sbjct: 547 PISNNKTMECFSLIHGDVWGPYRTPSTTGAVYFLTLVDDYSRSVWTYLMSSKTEVSQLIK 606
Query: 624 NFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNR 683
NF M QFGK +K R+DNG E + + + +G++H+ +CVD PQQNG ERK+R
Sbjct: 607 NFCAMSERQFGKQVKAFRTDNGTEFMC--LTPYFQTHGILHQTSCVDTPQQNGRVERKHR 664
Query: 684 HLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSG 743
H+L + RA LFQ N+P +WGE++LTA +LINR PS VL +P ++ PS +
Sbjct: 665 HILNVARACLFQGNLPVKFWGESILTATHLINRTPSAVLKGKTPYELLFGERPSYDML-- 722
Query: 744 LQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQ 803
R FGC + H+ ++ K R+ KC+FIGY KK ++ Y + +++ S DV F
Sbjct: 723 ---RSFGCLCYAHIRPRNKDKFTSRSRKCVFIGYPHGKKAWRVYDLETGKIFASRDVRFH 779
Query: 804 ESESFYPNPQLQGGDTQEAESPELSVIPLLQEP-LMPSSIPITEDSDDDDNGPEQVPNQN 862
E YP TQ P++ + +P S + DS + D+ P Q+
Sbjct: 780 ED--IYPYATA----TQSNVPLPPPTPPMVNDDWFLPISTQV--DSTNVDSSSSSSPAQS 831
Query: 863 DD-----NGLELVPDQNDDNGPE----LVPDQN---NVDRFRIKYQRKVKPVLIQ--QQD 908
++ P + + PE +VP + ++DR L+ +
Sbjct: 832 GSIDQPPRSIDQSPSTSTNPVPEEIGSIVPSSSPSRSIDRSTSDLSASDTTELLSTGESS 891
Query: 909 QPSDP--------------------EVSVPNPEPSNSYEHNLDD--------LPIALRKG 940
PS P + + HN+ LP +L
Sbjct: 892 TPSSPGLPELLGKGCREKKKSVLLKDFVTNTTSKKKTASHNIHSPSQVLPSGLPTSLSAD 951
Query: 941 KRSCAK-YPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALD 999
S YP+S +++ S H +F++AI P ++ + K+W +AM++E+ AL+
Sbjct: 952 SVSGKTLYPLSDFLTNSGYSANHIAFMAAILDSNEPKHFKDAILIKEWCEAMSKEIDALE 1011
Query: 1000 KNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPV 1059
N T I + P+ K+ + KW+Y +KY SDGTL+R+KARLV G Q G+D+ ETFAPV
Sbjct: 1012 ANHTWDITDLPHGKKAISSKWVYKLKYNSDGTLERHKARLVVMGNHQKEGVDFKETFAPV 1071
Query: 1060 AKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKK 1119
AK+ TVR IL++AA DWE+ Q DV NAFLHGDLEEEVYM +PPG+ S +KVC+L+K
Sbjct: 1072 AKLTTVRTILAVAAAKDWEVHQMDVHNAFLHGDLEEEVYMRLPPGFK-CSDPSKVCRLRK 1130
Query: 1120 ALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTL-LLVYVNDMIIAGN 1178
+LYGLKQ+PR WF + + A+ ++G+ QS D++LF +NG + +LVYV+D+I+AGN
Sbjct: 1131 SLYGLKQAPRCWFSKLSTALRNIGFTQSYEDYSLF--SLKNGDTIIHVLVYVDDLIVAGN 1188
Query: 1179 DELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCK 1238
+ + +L F MKDLGKLKYFLG+EV+ G +SQRKY LD++KETG LGCK
Sbjct: 1189 NLDAIDRFKSQLHKCFHMKDLGKLKYFLGLEVSRGPDGFCLSQRKYALDIVKETGLLGCK 1248
Query: 1239 PMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQER 1298
P VPI NHK+ S + + QY+RLVG+ IYL+ TRPD++Y V ++SQFM P
Sbjct: 1249 PSAVPIALNHKLASITGPVFTNPEQYRRLVGRFIYLTITRPDLSYAVHILSQFMQAPLVA 1308
Query: 1299 HLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLV 1358
H +A R+++YLK SP +G+ + L + Y D+DY + RRS + Y ++LG + +
Sbjct: 1309 HWEAALRLVRYLKGSPAQGIFLRSDSSLIINAYCDSDYNACPLTRRSLSAYVVYLGDSPI 1368
Query: 1359 TWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIA 1418
+W++KKQ+ V+ SSAEAE+RAMA + EL +K +L DL + + +PM+L CD+++AI IA
Sbjct: 1369 SWKTKKQDTVSYSSAEAEYRAMAYTLKELKWLKALLKDLGVHHSSPMKLHCDSEAAIHIA 1428
Query: 1419 HNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLG 1478
NPV H+RTKHIE D H +++ + LI+T+++ + Q+AD+LTK LP F L LG
Sbjct: 1429 ANPVFHERTKHIESDCHKVRDAVLDKLITTEHIYTEDQVADLLTKSLPRPTFERLLSTLG 1488
Query: 1479 MID 1481
+ D
Sbjct: 1489 VTD 1491
>At1g70010 hypothetical protein
Length = 1315
Score = 838 bits (2166), Expect = 0.0
Identities = 501/1449 (34%), Positives = 774/1449 (52%), Gaps = 167/1449 (11%)
Query: 57 SRQKISYIEETAPA--ETDPQYDTWREENSLIMTWFWYSMIPEISRNYMFFPTAKEIWNN 114
++ K+ +++ + P + DP WR NS++ +W S+ EI + ++FPTA IW +
Sbjct: 7 AKNKLGFVDGSIPKPDDDDPYCKIWRRCNSMVKSWLLNSVSKEIYTSILYFPTAAAIWKD 66
Query: 115 LAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLWIELDQYQNLKMKCDSDST 174
L + K L Y+L ++ + +QG L ++ Y+ LW EL Q + T
Sbjct: 67 LYTRFHKSS-LPRLYKLRQQIHSLRQGNLDLSSYHTRTQTLWEELTSLQAVPR------T 119
Query: 175 TLNKFIEKA--RIFKFLSGLNSEFDPIRVQILGKEQFPSLSEVFSIVRGEETRRTVMVED 232
+ IE+ R+ FL GLN +D +R QIL K+ PSLSEVF+++ +ET+R+ +
Sbjct: 120 VEDLLIERETNRVIDFLMGLNDCYDTVRSQILMKKTLPSLSEVFNMIDQDETQRSARIST 179
Query: 233 KSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFNRDDR--CTYCKRSGHTK 290
+ S P+ +S NGD + + +R C+YC R GH +
Sbjct: 180 TP----GMTSSVFPVSNQSS------------QSALNGDT-YQKKERPVCSYCSRPGHVE 222
Query: 291 EYCFKLYGRENVLARMSKS-KGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELER 349
+ C+K +G K K + A + ++ N+ V L+ +++++
Sbjct: 223 DTCYKKHGYPTSFKSKQKFVKPSISANAAIGSEEVVNNTSV-------STGDLTTSQIQQ 275
Query: 350 LRVFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYST 409
L F+ S +P TP
Sbjct: 276 LVSFLSSKLQPPS-----------------------------------TP---------V 291
Query: 410 YPGKHYITVANGSQNQITGC---GNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCA 466
P H I+V++ + T C G++HL L L VL +P+ NLLS+ LT+ + C
Sbjct: 292 QPEVHSISVSSDPSSSSTVCPISGSVHLGRHLILNDVLFIPQFKFNLLSVSSLTKSMGCR 351
Query: 467 VTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSE----SWNTSQIWLRHK 522
+ F + V QD ++G+ K+ LY + + T S S + +W HK
Sbjct: 352 IWFDETSCVLQDATRELMVGMGKQVANLYIVDLDSLSHPGTDSSITVASVTSHDLW--HK 409
Query: 523 RLGHPPFSILRTMFPHL-FSKVSVES-FHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSD 580
RLGHP L+ M L F K + FHC VC SK F+ NNK ++PFDL+H D
Sbjct: 410 RLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHID 469
Query: 581 VWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRL 640
WGP S+ G ++F+T +DD +R TWV+L+++KS+V + F M+ QF IK +
Sbjct: 470 TWGPFSVQTHDGYRYFLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGV 529
Query: 641 RSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPK 700
RSDN E NF++F G+V +C + PQQN V ERK++H+L + R+L FQ ++P
Sbjct: 530 RSDNAPE---LNFTQFYHSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPI 586
Query: 701 FYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGH 760
YWG+ +LTA YLINRLP+ +L + P V+T P+ +VFGC +
Sbjct: 587 SYWGDCILTAVYLINRLPAPILEDKCPFEVLTKTVPTYD-----HIKVFGCLCYASTSPK 641
Query: 761 HRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQ 820
R K PRA C FIGY S KGYK + + +S V F E +P G D
Sbjct: 642 DRHKFSPRAKACAFIGYPSGFKGYKLLDLETHSIIVSRHVVFHEE--LFP---FLGSDLS 696
Query: 821 EAES---PELSVIPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDN 877
+ E P+L+ P P+ S D N + + +E++P N N
Sbjct: 697 QEEQNFFPDLNPTP-----------PMQRQSSDHVNPSDS------SSSVEILPSANPTN 739
Query: 878 GPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIAL 937
VP+ + + +++ KP +Q Y H++
Sbjct: 740 N---VPEPS----VQTSHRKAKKPAYLQDY------------------YCHSVV------ 768
Query: 938 RKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHA 997
S + I +++S ++ + +F++ +D + P++ E K + W AM E
Sbjct: 769 -----SSTPHEIRKFLSYDRINDPYLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDF 823
Query: 998 LDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFA 1057
L+ T ++ P DKR +GC+WI+ +KY SDG+++RYKARLVA+GYTQ GIDY+ETF+
Sbjct: 824 LEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFS 883
Query: 1058 PVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGR----NK 1113
PVAK+N+V+++L +AA F L Q D+ NAFL+GDL+EE+YM +P GY G N
Sbjct: 884 PVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNA 943
Query: 1114 VCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDM 1173
VC+LKK+LYGLKQ+ R W+ +F++ ++ LG+ QS DHT F+K + +G +LVY++D+
Sbjct: 944 VCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKIS-DGIFLCVLVYIDDI 1002
Query: 1174 IIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETG 1233
IIA N++ L+ ++ F+++DLG+LKYFLG+E+ S +GI ISQRKY LDLL ETG
Sbjct: 1003 IIASNNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQRKYALDLLDETG 1062
Query: 1234 KLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMH 1293
+LGCKP +P++ + V Y+RL+G+L+YL+ TRPDI + V+ ++QF
Sbjct: 1063 QLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSM 1122
Query: 1294 DPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFL 1353
P++ HLQAV +ILQY+K + G+GL + + +L ++VY +ADY RRST+GYCMFL
Sbjct: 1123 APRKAHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRDSRRSTSGYCMFL 1182
Query: 1354 GGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKS 1413
G +L+ W+S+KQ+VV++SSAEAE+R+++ EL+ + L +L++ P LFCDN++
Sbjct: 1183 GDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEA 1242
Query: 1414 AISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFREL 1473
AI IA+N V H+RTKHIE D H ++E+L L ++ + LQ+AD TK L F L
Sbjct: 1243 AIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKPLYPSHFHRL 1302
Query: 1474 TCKLGMIDI 1482
K+G+++I
Sbjct: 1303 ISKMGLLNI 1311
>At1g57640
Length = 1444
Score = 824 bits (2129), Expect = 0.0
Identities = 513/1499 (34%), Positives = 792/1499 (52%), Gaps = 145/1499 (9%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYIEETAPAETD--PQYDTWREENSLIMTWFWYSMIP 96
L NY +W+ + L SR+K +++ T P D P + W N+L+++W ++
Sbjct: 38 LKTNNYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLEDWLTINALLVSWMKMTIDS 97
Query: 97 EISRNYMFFPTAKEIWNNLAQAYS---------KKQDLSACYELENKVFNSKQGTLSVTD 147
E+ N A+++W + + +S K DL+ C KQ ++V
Sbjct: 98 ELLTNISHRDVARDLWEQIRKRFSVSNGPKNQKMKADLATC----------KQEGMTVEG 147
Query: 148 YYGTLNGLWIELDQYQNLKM-KCD----SDSTTLNKFIEKARIFKFLSGLN-SEFDPIRV 201
YYG LN +W ++ Y+ L++ KC + T K+ E + ++L GLN ++F IR
Sbjct: 148 YYGKLNKIWDNINSYRPLRICKCGRCICNLGTDQEKYREDDMVHQYLYGLNETKFHTIRS 207
Query: 202 QILGKEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDR 261
+ + P L EV++IVR EE MV ++S + T+F R
Sbjct: 208 SLTSRVPLPGLEEVYNIVRQEED----MVNNRSSNEERT--------DVTAFAVQMRPRS 255
Query: 262 PNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYG-----------RENVLARMSKSK 310
++F ++ CT+C R GH+ E CF L G + N S+ +
Sbjct: 256 EVISEKFANSEKLQNKKLCTHCNRGGHSPENCFVLIGYPEWWGDRPRGKSNSNGSTSRGR 315
Query: 311 G--APQRRANHTASDMENDGEVPPVPPAEDVPS-LSKAELERLRVFMDSLSKPSGSCSLT 367
G P N P P +E V ++ ++ + + D + G L
Sbjct: 316 GRFGPGFNGGQPRPTYVNVVMTGPFPSSEHVNRVITDSDRDAVSGLTDEQWR--GVVKLL 373
Query: 368 MTGKSSTFLSLNALSTEN-------DWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVAN 420
G+S NA T++ WI+D GA+ HMT + LS + I +A+
Sbjct: 374 NAGRSDN--KSNAHETQSGTCSLFTSWILDTGASHHMTGNLELLSDMRSM-SPVLIILAD 430
Query: 421 GSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLA 480
G++ G + L L LK V +V +L ++L+S+ ++ + +C
Sbjct: 431 GNKRVAVSEGTVRLGSHLILKSVFYVKELESDLISVGQMMDENHCV-------------- 476
Query: 481 TGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLF 540
+ H + ++ +W H+RLGH I+ + L
Sbjct: 477 -------------------NAAAVHTSVKAPFD---LW--HRRLGHASDKIVNLLPRELL 512
Query: 541 S--KVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVT 598
S K +E+ CD C +K R TF S+N+ F L+H DVWGP + SGA++F+T
Sbjct: 513 SSGKEILENV-CDTCMRAKQTRDTFPLSDNRSMDSFQLIHCDVWGPYRAPSYSGARYFLT 571
Query: 599 FIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLT 658
+DD +R WV+LM DKSE + +F ++ QF IK +RSDNG E + E+
Sbjct: 572 IVDDYSRGVWVYLMTDKSETQKHLKDFIALVERQFDTEIKIVRSDNGTEFLCMR--EYFL 629
Query: 659 KNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLP 718
G+ HE +CV P QNG ERK+RH+L I RAL FQ +P +WGE +L+AAYLINR P
Sbjct: 630 HKGIAHETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQFWGECILSAAYLINRTP 689
Query: 719 SRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYA 778
S +L SP ++ + + P S L RVFG + H H K R+ +C+F+GY
Sbjct: 690 SMLLQGKSPYEML---YKTAPKYSHL--RVFGSLCYAHNQNHKGDKFAARSRRCVFVGYP 744
Query: 779 SDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLM 838
+KG++ + ++ ++S DV FQE+E +P ++ + E + P ++E +
Sbjct: 745 HGQKGWRLFDLEEQKFFVSRDVIFQETE--FPYSKMSCNEEDERVLVDCVGPPFIEEAIG 802
Query: 839 PSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNG-PELVPDQNNVDRFRIKYQR 897
P +I I + + GP ++P+ N ++ P +++D F
Sbjct: 803 PRTI-IGRNIGEATVGPNVATGP-------IIPEINQESSSPSEFVSLSSLDPF------ 848
Query: 898 KVKPVLIQQQDQPSDPEVSVP-----------NPEPSNSYEHNLDDLPIALRKGKRSCAK 946
+ +Q D P P P ++ N + ++ S +
Sbjct: 849 -LASSTVQTADLPLSSTTPAPIQLRRSSRQTQKPMKLKNFVTNTVSVE-SISPEASSSSL 906
Query: 947 YPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKI 1006
YPI +YV + H++F++A+ + PT+ E + DK W +AM+ E+ +L N T I
Sbjct: 907 YPIEKYVDCHRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFSI 966
Query: 1007 VEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVR 1066
V P KR +G KW+Y +KYRSDG ++RYKARLV G Q G+DYDETFAPVAKM+TVR
Sbjct: 967 VNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTVR 1026
Query: 1067 IILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQ 1126
+ L +AA DW + Q DV NAFLHGDL+EEVYM++P G+ +KVC+L K+LYGLKQ
Sbjct: 1027 LFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQ-CDDPSKVCRLHKSLYGLKQ 1085
Query: 1127 SPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNL 1186
+PR WF + ++A+ G+ QS D++LF + +G +LVYV+D+II+G+
Sbjct: 1086 APRCWFSKLSSALKQYGFTQSLSDYSLF-SYNNDGIFVHVLVYVDDLIISGSCPDAVAQF 1144
Query: 1187 RKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQ 1246
+ L F MKDLG LKYFLG+EV+ + QG ++SQRKYVLD++ E G LG +P P+EQ
Sbjct: 1145 KSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQ 1204
Query: 1247 NHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRI 1306
NHK+ S L ++Y+RLVG+LIYL TRP+++Y V ++QFM +P++ H A R+
Sbjct: 1205 NHKLSLSTSPLLSDSSRYRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRV 1264
Query: 1307 LQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQN 1366
++YLK++PG+G+L + L + + D+DYA + RRS TGY + LG ++W++KKQ
Sbjct: 1265 VRYLKSNPGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQP 1324
Query: 1367 VVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDR 1426
V+RSSAEAE+RAMA EL+ +K +L DL + + MR+F D+KSAI+++ NPVQH+R
Sbjct: 1325 TVSRSSAEAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHER 1384
Query: 1427 TKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
TKH+E+D HFI++ + +I+T +VPS QLAD+LTK L + R KLG++D+H+P
Sbjct: 1385 TKHVEVDCHFIRDAILDGIIATSFVPSHKQLADILTKALGEKEVRYFLRKLGILDVHAP 1443
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 701 bits (1810), Expect = 0.0
Identities = 462/1332 (34%), Positives = 699/1332 (51%), Gaps = 119/1332 (8%)
Query: 181 EKARIFKFLSGLNSE-FDPIRVQILGKEQFPSLSEVFSIVRGEET--RRTVMVEDKSVDG 237
E+ ++ +FLSGL+ F +R ++ + L EV++IVR EE R V D +
Sbjct: 93 EEDKVHEFLSGLDDALFRTVRSSLVSRIPVQPLEEVYNIVRQEEDLLRNGANVLDDQREV 152
Query: 238 SALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLY 297
+A A+ P GR GD++ ++ C +C RSGH E C+ +
Sbjct: 153 NAFAAQMRP---KLYQGR--------------GDEK-DKSMVCKHCNRSGHASESCYAVI 194
Query: 298 GRENVLARMSKSKGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSL 357
G +S+ R T S VP+ E
Sbjct: 195 GYPEWWGDRPRSRSLQTRGRGGTNSSGGRGRGAAAYANRVTVPNHDTYE----------- 243
Query: 358 SKPSGSCSLTMTGKSSTFLSLNALSTENDW-----IIDFG---ATDHMTPHPSHLSSYST 409
+ +LT + +N L T++ W I++ G AT+ +T L
Sbjct: 244 ---QANYALTDEDRDG----VNGL-TDSQWRTIKSILNSGKDAATEKLTDMDPIL----- 290
Query: 410 YPGKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTF 469
I +A+G + G + L +L + V +V + ++L+SI +L + C +
Sbjct: 291 ------IVLADGRERISVKEGTVRLGSNLVMISVFYVEEFQSDLISIGQLMDENRCVLQM 344
Query: 470 FHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPF 529
V QD + ++G + G + + E T E N ++W H R+GHP
Sbjct: 345 SDRFLVVQDRTSRMVMGAGRRVGGTFHFRSTEIAASVTVKEEKNY-ELW--HSRMGHPAA 401
Query: 530 SILRTMFPHLFSKVSVESFH----CDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPS 585
++ ++ P S VSV S H CDVC +K R++F S NK + F+L++ D+WGP
Sbjct: 402 RVV-SLIPE--SSVSVSSTHLNKACDVCHRAKQTRNSFPLSINKTLRIFELIYCDLWGPY 458
Query: 586 SISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNG 645
+ +GA++F+T IDD +R W++L+ DKSE NFF M QF IK +RSDNG
Sbjct: 459 RTPSHTGARYFLTIIDDYSRGVWLYLLNDKSEAPCHLKNFFAMTDRQFNVKIKTVRSDNG 518
Query: 646 KE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGE 705
E + ++F + GV+HE +CV P++N ERK+RHLL + RAL FQ N+P +WGE
Sbjct: 519 TEFLC--LTKFFQEQGVIHERSCVATPERNDRVERKHRHLLNVARALRFQANLPIQFWGE 576
Query: 706 AVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKL 765
VLTAAYLINR PS VL++ +P + P RVFG + H K
Sbjct: 577 CVLTAAYLINRTPSSVLNDSTPYERLHKKQPRFD-----HLRVFGSLCYAHNRNRGGDKF 631
Query: 766 DPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESP 825
R+ +C+F+GY +KG++ + ++S DV F E E + Q +E E+
Sbjct: 632 AERSRRCVFVGYPHGQKGWRLFDLEQNEFFVSRDVVFSELEFPFRISHEQNVIEEEEEAL 691
Query: 826 ELSVIP-LLQEPL-----MPSSIPITEDSDDDDNGPEQVPNQNDDNGL--ELVPDQNDDN 877
++ L++E + + PI S + + + L E+VP
Sbjct: 692 WAPIVDGLIEEEVHLGQNAGPTPPICVSSPISPSATSSRSEHSTSSPLDTEVVPTPATST 751
Query: 878 GPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDP---EVSVPNPEPSNSYEHNLDDLP 934
P ++ + + KP Q P+ P S N P + + + +
Sbjct: 752 TSASSP--SSPTNLQFLPLSRAKPTTAQAVAPPAVPPPRRQSTRNKAPPVTLKDFVVNTT 809
Query: 935 IALRK-GKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNE 993
+ K + Y + + T+ S H +++ AID
Sbjct: 810 VCQESPSKLNSILYQLQKRDDTRRFSASHTTYV-AID----------------------- 845
Query: 994 EMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYD 1053
A ++N T I + P KR +G +W+Y VK+ SDG+++RYKARLVA G Q G DY
Sbjct: 846 ---AQEENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARLVALGNKQKEGEDYG 902
Query: 1054 ETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNK 1113
ETFAPVAKM TVR+ L +A +WE+ Q DV NAFLHGDL EEVYM++PPG++ S NK
Sbjct: 903 ETFAPVAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYMKLPPGFE-ASHPNK 961
Query: 1114 VCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDM 1173
VC+L+KALYGLKQ+PR WF + T A+ G++QS D++LF + ++ +L +YV+D+
Sbjct: 962 VCRLRKALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGSVRIKIL-IYVDDL 1020
Query: 1174 IIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETG 1233
II GN + Q ++ LA F MKDLG LKYFLG+EVA S GI+I QRKY LD++ ETG
Sbjct: 1021 IITGNSQRATQQFKEYLASCFHMKDLGPLKYFLGIEVARSTTGIYICQRKYALDIISETG 1080
Query: 1234 KLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMH 1293
LG KP P+EQNHK+G S L +Y+RLVG+LIYL+ TR D+A+ V ++++FM
Sbjct: 1081 LLGVKPANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTRLDLAFSVHILARFMQ 1140
Query: 1294 DPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFL 1353
+P+E H A R+++YLKA PG+G+ ++S + + D+D+AG + RRS TGY +
Sbjct: 1141 EPREDHWAAALRVVRYLKADPGQGVFLRRSGDFQITGWCDSDWAGDPMSRRSVTGYFVQF 1200
Query: 1354 GGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKS 1413
G + ++W++KKQ+ V++SSAEAE+RAM+ ELL +K +L L + + PM + CD+KS
Sbjct: 1201 GDSPISWKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLGVSHVQPMIMCCDSKS 1260
Query: 1414 AISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFREL 1473
AI IA NPV H+RTKHIEID HF++++ +I+ ++V + QLAD+ TK L + F
Sbjct: 1261 AIYIATNPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLADIFTKPLGRDCFSAF 1320
Query: 1474 TCKLGMIDIHSP 1485
KLG+ ++++P
Sbjct: 1321 RIKLGIRNLYAP 1332
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 683 bits (1763), Expect = 0.0
Identities = 397/1140 (34%), Positives = 608/1140 (52%), Gaps = 147/1140 (12%)
Query: 371 KSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCG 430
++ T +L + WIID GA+ H+ S L+ + G
Sbjct: 466 QNHTLSALQKFLPSDAWIIDSGASSHVC---SDLAMFRELKSVS---------------G 507
Query: 431 NIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKE 490
+H++ L L VLHVP NL+S+ L + ++C+ F+ + Q+L+ G +IG +
Sbjct: 508 TVHITQKLILHNVLHVPDFKFNLMSVSSLVKTISCSAHFYVDCCLIQELSQGLMIGRGRL 567
Query: 491 QDGLYCLQHEESK--------CHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSK 542
LY L+ E + C T S N +W H+RLGHP +L+
Sbjct: 568 YHNLYILETENTSPSTSTPAACLFTGSVL-NDGHLW--HQRLGHPSSVVLQ--------- 615
Query: 543 VSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDD 602
K R ++ NN + PFDLVH D+WGP SI ++ G ++F+T +DD
Sbjct: 616 --------------KLKRLAYISHNNLASNPFDLVHLDIWGPFSIESIEGFRYFLTVVDD 661
Query: 603 CTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGV 662
CTR TWV+++++K +V +F F ++ TQF IK +RSDN E F+E + ++G+
Sbjct: 662 CTRTTWVYMLRNKKDVSSVFPEFIKLVSTQFNAKIKAIRSDNAPE---LGFTEIVKEHGM 718
Query: 663 VHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVL 722
+H +C PQQN V ERK++H+L + RALLFQ N+P YW + V TA +LINRLPS +L
Sbjct: 719 LHHFSCAYTPQQNSVVERKHQHILNVARALLFQSNIPMQYWSDCVTTAVFLINRLPSPLL 778
Query: 723 SNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKK 782
+N SP ++ + P + FGC FV + H R K PRA C+F+GY S K
Sbjct: 779 NNKSPYELILNKQPDYSLLKN-----FGCLCFVSTNAHERTKFTPRARACVFLGYPSGYK 833
Query: 783 GYKCYHPPSRRVYISMDVTFQE-------------SESFYPNPQLQGGDTQEAESPELSV 829
GYK S V +S +V F+E + +PN L A +
Sbjct: 834 GYKVLDLESHSVTVSRNVVFKEHVFPFKTSELLNKAVDMFPNSILP----LPAPLHFVET 889
Query: 830 IPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVD 889
+PL+ E S IP T DS DN + ++P ++ ++ D N V
Sbjct: 890 MPLIDED---SLIPTTTDSRTADNHASSSSSALPS----IIPPSSNTETQDI--DSNAVP 940
Query: 890 RFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPI 949
R K + P + + P +S P S+ H L ++ A + YPI
Sbjct: 941 ITRSKRTTRA-PSYLSEYHCSLVPSISTLPPTDSSIPIHPLPEIFTA--SSPKKTTPYPI 997
Query: 950 SQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEK 1009
S VS + QS+I A ++ P + + +K +KWI+ EE+ A++ N T +
Sbjct: 998 STVVSYDKYTPLCQSYIFAYNTETEPKTFSQAMKSEKWIRVAVEELQAMELNKTWSVESL 1057
Query: 1010 PNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIIL 1069
P DK VGCKW++T+KY DGT++RYKARLVA+G+TQ GID+ +TF+PVAK+ + +++L
Sbjct: 1058 PPDKNVVGCKWVFTIKYNPDGTVERYKARLVAQGFTQQEGIDFLDTFSPVAKLTSAKMML 1117
Query: 1070 SLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGR----NKVCKLKKALYGLK 1125
LAA W L Q DV +AFLHGDL+EE++M +P GY P +G N VC+L K++YGLK
Sbjct: 1118 GLAAITGWTLTQMDVSDAFLHGDLDEEIFMSLPQGYTPPAGTILPPNPVCRLLKSIYGLK 1177
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQN 1185
Q+ R W+ RF A LVY++D++IA N++ E +N
Sbjct: 1178 QASRQWYKRFVAA----------------------------LVYIDDIMIASNNDAEVEN 1209
Query: 1186 LRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIE 1245
L+ L +F++KDLG ++FLG+ LGCKP +P++
Sbjct: 1210 LKALLRSEFKIKDLGPARFFLGL--------------------------LGCKPSSIPMD 1243
Query: 1246 QNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDR 1305
+ + Y++L+G+L+YL+ TRPDI Y V +SQF+ P + HLQA +
Sbjct: 1244 PTLHLVRDMGTPLPNPTAYRKLIGRLLYLTITRPDITYAVHQLSQFISAPSDIHLQAAHK 1303
Query: 1306 ILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQ 1365
+L+Y+KA+PG+GL++ ++ + ++DAD+A RRS +G+C++LG +L++W+SKKQ
Sbjct: 1304 VLRYIKANPGQGLMYSADYEICLNGFSDADWAACKDTRRSISGFCIYLGTSLISWKSKKQ 1363
Query: 1366 NVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHD 1425
V +RSS E+E+R+MAQ CE++ ++ +L DL I P +LFCDNKSA+ + NPV H+
Sbjct: 1364 AVASRSSTESEYRSMAQATCEIIWLQQLLKDLHIPLTCPAKLFCDNKSALHSSLNPVFHE 1423
Query: 1426 RTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
RTKHIEID H +++++ + + +VP+ Q AD+LTK L F L ++ + + P
Sbjct: 1424 RTKHIEIDCHTVRDQIKAGNLKALHVPTENQHADILTKALHPGPFHHLLRQMSLSSLFLP 1483
Score = 107 bits (268), Expect = 4e-23
Identities = 69/263 (26%), Positives = 124/263 (46%), Gaps = 21/263 (7%)
Query: 43 NYLQWSELVRHTLISRQKISYIEETA--PAETDPQYDTWREENSLIMTWFWYSMIPEISR 100
++ W + L R K+ +I T P E + W N ++ TW S+ +I +
Sbjct: 56 DFHSWRRSILMALNVRNKLGFINGTITKPPEDHRDFGAWSRCNDIVSTWLMNSVDKKIGQ 115
Query: 101 NYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLWIELD 160
+ ++ T + IWNNL + K+ D +++E K+ +QG++ ++ YY L LW E
Sbjct: 116 SLLYIATVQGIWNNLLSRF-KQDDAPRIFDIEQKLSKIEQGSMDISTYYTALLTLWEEHR 174
Query: 161 QYQNLKM----KCDSDSTTLNKFI-EKARIFKFLSGLNSEFDPIRVQILGKEQFPSLSEV 215
Y L + +C+ D+ + + +++R+ KFL LN FD R IL + P++ E
Sbjct: 175 NYVELPVCTCGRCECDAAVKWEHLQQRSRVTKFLKELNEGFDQTRRHILMLKPIPTIKEA 234
Query: 216 FSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFN 275
F++V +E +R V + VD A + + ++ RP N
Sbjct: 235 FNMVTQDERQRNVKPLTR-VDSVAFQNTSMINEDENAYVAAYNTVRP------------N 281
Query: 276 RDDRCTYCKRSGHTKEYCFKLYG 298
+ CT+C + GHT + C+K++G
Sbjct: 282 QKPICTHCGKVGHTIQKCYKVHG 304
>At2g07420 putative retroelement pol polyprotein
Length = 1664
Score = 667 bits (1720), Expect = 0.0
Identities = 425/1275 (33%), Positives = 657/1275 (51%), Gaps = 145/1275 (11%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYI-EETAPAETDPQY---------DTWREENSLIMT 88
L G NYL W+ + TL SR ++I AP+E + + W +E+ ++
Sbjct: 13 LKGVNYLLWARTTKTTLCSRGLWAHILTSEAPSEATIREGMEIVHVGEEKWFQEDQSVLA 72
Query: 89 WFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDY 148
S+ + Y + TAKE+W L + + +LS +E++ + + QG + T +
Sbjct: 73 LLQNSLEASLLEAYSYCETAKELWETLFNVFGNQSNLSRVFEVKKAINDLSQGDMEFTQH 132
Query: 149 YGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGKEQ 208
+G LW EL+ + + D L + E+ ++F L L+S ++ + +L ++
Sbjct: 133 FGKFRSLWAELEMLRPNTL----DPKVLIERREQDKVFGLLLTLSSTYNDLIKHLLRADK 188
Query: 209 FPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRF 268
P+L EV S ++ E+ R
Sbjct: 189 LPNLEEVCSQIQKEQANR------------------------------------------ 206
Query: 269 NGDDRFNRDDR---CTYCKRSGHTKEYCFKLY-----GRENVLARMSKSKGAPQRRANHT 320
G+ +++ + + C +CKRSGHTKE C+ L+ GR A + + + T
Sbjct: 207 -GNYKYDNNKKALWCEHCKRSGHTKEKCWTLHPHLRPGRREPRANQVTGENFGTQEQSGT 265
Query: 321 ASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNA 380
S+ G + + D+ S L+ + +L + SG S +A
Sbjct: 266 -SNQHLGGNGAAMAASSDLVRRSD-----LKALIKALKESSGK-------------SYHA 306
Query: 381 LSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPL 440
LS+ IID GA+ HM +S+ P + +ANG + + G G++ L
Sbjct: 307 LSSLKPLIIDSGASHHMISDSKLISNIE--PALGNVVIANGDRIPVKGVGDLDLFDKS-- 362
Query: 441 KRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQ-- 498
+ ++P ++NLLS+ K T DLNC F + FQD+ T +++G +DGLY L+
Sbjct: 363 SKAFYMPTFTSNLLSVKKATTDLNCYAIFGPNEVHFQDIETSRVLGQGVTKDGLYVLEDT 422
Query: 499 --------HEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFHC 550
H S +SESW H RLGHP L+ + P S ++ C
Sbjct: 423 KPSVPLSSHFSSILGNANSESW--------HARLGHPHSRALKLLLP----STSFKNDEC 470
Query: 551 DVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVF 610
+ C KH +S F S+ + FDL+HSDVW +S K+FVTFID+ ++ TW
Sbjct: 471 EACILGKHCKSVFPKSSTIYEKCFDLIHSDVWTSPCLSR-ENHKYFVTFIDEKSKFTWFT 529
Query: 611 LMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVD 670
L+ K V + F NF + + IK LRSDN E + F + L K+G++H+ +C
Sbjct: 530 LLPSKDRVLEAFTNFQTYVTNHYDAKIKILRSDNRGEYTSHAFKQHLNKHGIIHQTSCPY 589
Query: 671 APQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLV 730
PQQNGVAERKNRHL+E+ R ++F NVPK +W + V++A YLIN+ P+++L + SP V
Sbjct: 590 TPQQNGVAERKNRHLMEVRRVMMFHTNVPKHFWIDGVVSACYLINQTPTKILLDSSPFEV 649
Query: 731 MTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPP 790
+ P + RVFGC FV + G R KL P++ K +FIGY+ ++KGYKCY
Sbjct: 650 LNKVKPFIN-----HLRVFGCVCFVLISGEQRNKLQPKSTKGMFIGYSINQKGYKCYVLE 704
Query: 791 SRRVYISMDVTFQESESFYPNPQLQG-GDTQEAESPELSVIPLLQEPLMPSSIPITEDSD 849
+R+V IS DV F ES+S+Y + D ++ S + + ++ E L S+I T+ +
Sbjct: 705 TRKVLISRDVKFLESKSYYDKKNWEDIQDLTDSPSDRATNLRIILERLGVSNIQ-TQTTP 763
Query: 850 DDDNGPE---QVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQ 906
N PE Q N ++ E ++ EL+ + K Q K +L
Sbjct: 764 RTSN-PETITQPENMEEEEEEEEEEEEKQGKEQELITLEETESS---KVQEKDTSLLNDD 819
Query: 907 QDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKR----------SCAKYPISQYVSTQ 956
++ E E SNS E K KR + ++PI +
Sbjct: 820 NGHTNNQE------EDSNSREEPRIPRRSEHLKDKRVYYNNQVYFDNVVEHPIQVVCTLA 873
Query: 957 NLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTV 1016
+L +HQ F +D +P + +E + + W A+ E A++ N T E P K+ V
Sbjct: 874 HLPEEHQVFFGKVDQHWIPQTYEEAITHQVWRDAIAAEKQAMENNHTWDEDELPRGKKVV 933
Query: 1017 GCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFD 1076
KW++ +KY+SDG ++RYKARLVA+G+TQTYG DY +TFAPVAK++TVR++LSL + +
Sbjct: 934 TSKWVFAIKYKSDGEIERYKARLVARGFTQTYGEDYLDTFAPVAKLHTVRVVLSLTTNLE 993
Query: 1077 WELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFT 1136
W+L Q DVKNAFL G+LEE+VYM+ PPG + + NKV KLKKA+YGLKQSPRAW+ + +
Sbjct: 994 WDLWQMDVKNAFLQGELEEKVYMKPPPGLEDINAPNKVFKLKKAIYGLKQSPRAWYHKLS 1053
Query: 1137 NAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEM 1196
++ G+K+S+ D+TLF ++ G + ++LVYV+D+II+GND++ Q+ + L F++
Sbjct: 1054 TTLMGRGFKRSEADNTLFTLPSQKG-IVVILVYVDDIIISGNDKVGIQDTKTFLKSVFDI 1112
Query: 1197 KDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKI---GSS 1253
KDLG+LKYFLG+EV SK+G+F+SQRKY LDLL + GKLG KP P+E ++K G
Sbjct: 1113 KDLGELKYFLGIEVCRSKEGLFLSQRKYTLDLLAQVGKLGVKPAKTPLEDDYKAKRKGEH 1172
Query: 1254 EEDLRVHKAQYQRLV 1268
+ +Y+RLV
Sbjct: 1173 DNKPFEDATRYRRLV 1187
Score = 64.3 bits (155), Expect = 5e-10
Identities = 30/61 (49%), Positives = 41/61 (67%)
Query: 1404 PMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTK 1463
P+ + CDN++AI IA N V H+RTKHIE+D H ++E++ +I Y QLADV TK
Sbjct: 1196 PIPMHCDNQAAIHIASNSVFHERTKHIEVDCHKVREQVQLGVILPHYTEGEEQLADVFTK 1255
Query: 1464 G 1464
G
Sbjct: 1256 G 1256
>At1g44510 polyprotein, putative
Length = 1459
Score = 660 bits (1703), Expect = 0.0
Identities = 455/1512 (30%), Positives = 713/1512 (47%), Gaps = 160/1512 (10%)
Query: 38 RLDGRNYLQWSELVRHTLISRQKISYIEETA--PAET---------DPQYDTWREENSLI 86
+L NYL WS + L Y++ + P ET +P + W+ ++ LI
Sbjct: 32 KLTSTNYLMWSIQIHALLDGYDLAGYLDNSVVIPPETTTINSVVSANPSFTLWKRQDKLI 91
Query: 87 MTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVT 146
+ ++ P + + +IW+ L Y+K +L ++ +GT ++
Sbjct: 92 FSALIGAISPAVQSLVSRATNSSQIWSTLNNTYAKPS-YGHIKQLRQQIQRLTKGTKTID 150
Query: 147 DYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGK 206
+Y + LDQ L + + ++ L GL E+ + QI GK
Sbjct: 151 EY---VQSHTTRLDQLAILGKPMEHEE----------QVEHILKGLPEEYKTVVDQIEGK 197
Query: 207 EQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDD 266
+ P+++E+ + E++ ++ D+ S+ ++ N++ NR
Sbjct: 198 DNTPTITEIHERLINHESK---LLSDEVPPSSSFPMSANAVQQRNFNNNCNQNQHKNRYQ 254
Query: 267 RFNGDDRFNRDDRCTYCKRSGHTKEYCFKLY-GRENVLARMSKS-KGAPQRRANHTASDM 324
++ N + + + +SG + FK Y G+ + + S + PQ +A +
Sbjct: 255 GNTHNNNTNTNSQPSTYNKSG---QRTFKPYLGKCQICSVQGHSARRCPQLQAMQLPASS 311
Query: 325 ENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTE 384
P P + L++ +
Sbjct: 312 SAHSPFTPWQPRAN-------------------------------------LAIGSPYAA 334
Query: 385 NDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHL---SPSLPLK 441
N W++D GAT H+T + LS + Y G Y+ +A+G+ I G+ L + L L
Sbjct: 335 NPWLLDSGATHHITSDLNALSLHQPYNGGEYVMIADGTGLTIKQTGSTFLPSQNRDLALH 394
Query: 442 RVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLY--CLQH 499
+VL+VP + NL+S+++L +V FF + +DL TG ++ + +D LY + +
Sbjct: 395 KVLYVPDIRKNLISVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDDLYEWPVTN 454
Query: 500 EESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFH---CDVCQFS 556
+ TS T W H RLGHP SIL T+ VSV S + C C +
Sbjct: 455 PPATALFTSPSPKTTLSSW--HSRLGHPSASILNTLLSKFSLPVSVASSNKTSCSDCLIN 512
Query: 557 KHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKS 616
K H+ F S+ + P + + +DVW IS+ K+++ +D TR TW++ ++ KS
Sbjct: 513 KSHKLPFATSSIHSSSPLEYIFTDVWTSPIISH-DNYKYYLVLVDHYTRYTWLYPLQQKS 571
Query: 617 EVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNG 676
+V F F ++ +F I+ L SDNG E + +FL NG+ H + P+ NG
Sbjct: 572 QVKATFIAFKALVENRFQAKIRTLYSDNGGEFIA--LRDFLVSNGISHLTSPPHTPEHNG 629
Query: 677 VAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFP 736
++ERK+RH++E LL Q +VP+ YW A TA YLINR+P+ VL SP F
Sbjct: 630 LSERKHRHIVETGLTLLTQASVPREYWTYAFATAVYLINRMPTPVLCLQSP---FQKLFG 686
Query: 737 SVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYI 796
S P L RVFGC F + + R KL+ R+ +C+F+GY+ + Y C + R+Y
Sbjct: 687 SSPNYQRL--RVFGCLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDVDNNRLYT 744
Query: 797 SMDVTFQESESFYP-----NPQLQGG-----DTQEAESPELSVIPL----LQEPLMPSSI 842
S V F ES YP Q Q ++ + SP S P LQ P P+S
Sbjct: 745 SRHVMFDEST--YPFAASIREQSQSSLVTPPESSSSSSPANSGFPCSVLRLQSP--PASS 800
Query: 843 PITEDSDDDDNGPEQVPNQNDD---------NGLELVPDQNDDNGPELVPDQNNVDRFRI 893
P T N P Q L P + N P +N +
Sbjct: 801 PETPSPPQQQNDSPVSPRQTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHENGPEP--- 857
Query: 894 KYQRKVKPVLIQQQDQPSDPEVS-VPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQ- 951
+ Q P+ P + +PNP P + +++ P+ K + P +Q
Sbjct: 858 -----------EAQSNPNSPFIGPLPNPNPETNPSSSIEQRPV----DKSTTTALPPNQT 902
Query: 952 -YVSTQNLSVQ-----HQSFISAIDSI------------------RVPTSVQEVLKDKKW 987
+T N Q HQ + ++I P +V + LKDKKW
Sbjct: 903 TIAATSNSRSQPPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSEPNTVTQALKDKKW 962
Query: 988 IQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQT 1047
AM++E A +N T +V + VGC+W++ +KY +G +D+YKARLVAKG+ Q
Sbjct: 963 RFAMSDEFDAQQRNHTWDLVPPNPTQHLVGCRWVFKLKYLPNGLIDKYKARLVAKGFNQQ 1022
Query: 1048 YGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDP 1107
YG+DY ETF+PV K T+R++L +A +W L+Q DV NAFL G L EEVYM PPG+
Sbjct: 1023 YGVDYAETFSPVIKATTIRVVLDVAVKKNWPLKQLDVNNAFLQGTLTEEVYMAQPPGFVD 1082
Query: 1108 TSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLL 1167
+ VC+L+KA+YGLKQ+PRAW+ ++++G+ S D +LFI ++ L LL
Sbjct: 1083 KDRPSHVCRLRKAIYGLKQAPRAWYMELKQHLLNIGFVNSLADTSLFI-YSHGTTLLYLL 1141
Query: 1168 VYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLD 1227
VYV+D+I+ G+D + LA +F +KD L YFLG+E + G+ + QRKY+ D
Sbjct: 1142 VYVDDIIVTGSDHKSVSAVLSSLAERFSIKDPTDLHYFLGIEATRTNTGLHLMQRKYMTD 1201
Query: 1228 LLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSV 1287
LL + L KP+ P+ + K+ ++Y+ +VG L YL+ TRPDIA+ V+
Sbjct: 1202 LLAKHNMLDAKPVATPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLAFTRPDIAFAVNR 1261
Query: 1288 VSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTT 1347
+SQFMH P H QA R+L+YL + G+ S + + ++DAD+AG D ST
Sbjct: 1262 LSQFMHQPTSDHWQAAKRVLRYLAGTTTHGIFLNSSSPIHLHAFSDADWAGDSADYVSTN 1321
Query: 1348 GYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRL 1407
Y ++LG N ++W SKKQ V+RSS E+E+RA+A E+ + +L +L I+ +
Sbjct: 1322 AYVIYLGRNPISWSSKKQRGVSRSSTESEYRAVANAASEIRWLCSLLTELHIRLPHGPTI 1381
Query: 1408 FCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPL 1467
FCDN A I NPV H R KHI +D HF++ + S + +V + QLAD LTK L
Sbjct: 1382 FCDNIGATYICANPVFHSRMKHIALDYHFVRGMIQSRALRVSHVSTNDQLADALTKSLSR 1441
Query: 1468 ERFRELTCKLGM 1479
F K+G+
Sbjct: 1442 PHFLSARSKIGV 1453
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 643 bits (1659), Expect = 0.0
Identities = 450/1529 (29%), Positives = 714/1529 (46%), Gaps = 191/1529 (12%)
Query: 38 RLDGRNYLQWSELVRHTLISRQKISYIEETAPA-----------ETDPQYDTWREENSLI 86
+L NYL WS V + +++ + P +P Y WR ++ LI
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTPMPPATIGTDAVPRVNPDYTRWRRQDKLI 84
Query: 87 MTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVT 146
+ ++ + TA +IW L + Y+
Sbjct: 85 YSAILGAISMSVQPAVSRATTAAQIWETLRKIYANPS----------------------- 121
Query: 147 DYYGTLNGLWI--ELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQIL 204
YG + L DQ L D D ++ + L L ++ P+ QI
Sbjct: 122 --YGHVTQLRFITRFDQLALLGKPMDHDE----------QVERVLENLPDDYKPVIDQIA 169
Query: 205 GKEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNR 264
K+ PSL+E+ + E++ LA + T+ +R+ NR
Sbjct: 170 AKDTPPSLTEIHERLINRESK-------------LLALNSAEVVPITANVVTHRNTNTNR 216
Query: 265 DDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRRANHTASDM 324
+ GD+R Y N + + + R N
Sbjct: 217 NQNNRGDNRN----------------------YNNNNNRSNSWQPSSSGSRSDNRQPKPY 254
Query: 325 ENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTE 384
++ V S +L F + ++ + T + L++N+
Sbjct: 255 LGRCQIC------SVQGHSAKRCPQLHQFQSTTNQQQSTSPFTPWQPRAN-LAVNSPYNA 307
Query: 385 NDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHL---SPSLPLK 441
N+W++D GAT H+T ++LS + Y G + +A+GS IT G+ L S SL L
Sbjct: 308 NNWLLDSGATHHITSDFNNLSFHQPYTGGDDVMIADGSTIPITHTGSASLPTSSRSLDLN 367
Query: 442 RVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLY------ 495
+VL+VP + NL+S+++L +V FF + +DL TG + K +D LY
Sbjct: 368 KVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWPIAS 427
Query: 496 --CLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKV---SVESFHC 550
+ S C + + SW H RLGHP +IL ++ + V S + C
Sbjct: 428 SQAVSMFASPCSKATHSSW--------HSRLGHPSLAILNSVISNHSLPVLNPSHKLLSC 479
Query: 551 DVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVF 610
C +K H+ F S ++P + ++SDVW S I ++ +++V F+D TR TW++
Sbjct: 480 SDCFINKSHKVPFSNSTITSSKPLEYIYSDVWS-SPILSIDNYRYYVIFVDHFTRYTWLY 538
Query: 611 LMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVD 670
+K KS+V F F ++ +F I L SDNG E V ++L+++G+ H +
Sbjct: 539 PLKQKSQVKDTFIIFKSLVENRFQTRIGTLYSDNGGEFVV--LRDYLSQHGISHFTSPPH 596
Query: 671 APQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLV 730
P+ NG++ERK+RH++E+ LL +VPK YW A A YLINRLP+ +L SP
Sbjct: 597 TPEHNGLSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSP--- 653
Query: 731 MTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPP 790
F P L +VFGC+ + + ++R KL+ ++ +C F+GY+ + Y C H P
Sbjct: 654 FQKLFGQPPNYEKL--KVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIP 711
Query: 791 SRRVYISMDVTF-------------------QESES---------------FYPNPQLQG 816
+ R+Y S V F Q S+S P P G
Sbjct: 712 TGRLYTSRHVQFDERCFPFSTTNFGVSTSQEQRSDSAPNWPSHTTLPTTPLVLPAPPCLG 771
Query: 817 GDTQEAESPELSVIPLLQEPLMPSSIP--------ITEDSDDDDNGPEQV----PNQNDD 864
+ P S PL + S++P +E + NGP+ QN +
Sbjct: 772 PHLDTSPRPPSSPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSN 831
Query: 865 NGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSN 924
+ ++ + N ++ P+QN+ P+ P P S EP++
Sbjct: 832 SNSPILNNPNPNSPSPNSPNQNS-------------PLPQSPISSPHIPTPSTSISEPNS 878
Query: 925 SYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSV----------QHQSFISAIDSIRV 974
+ P+ + V+T +++ Q S+ +++ +
Sbjct: 879 PSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSLAANSE 938
Query: 975 PTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRT-VGCKWIYTVKYRSDGTLD 1033
P + + +KD +W QAM E++A N T +V P T VGC+WI+T K+ SDG+L+
Sbjct: 939 PRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSDGSLN 998
Query: 1034 RYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDL 1093
RYKARLVAKGY Q G+DY ETF+PV K ++RI+L +A W ++Q DV NAFL G L
Sbjct: 999 RYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTL 1058
Query: 1094 EEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTL 1153
+EVYM PPG+ + VC+L+KA+YGLKQ+PRAW+ ++++G+ S D +L
Sbjct: 1059 TDEVYMSQPPGFVDKDRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSISDTSL 1118
Query: 1154 FIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYS 1213
F+ R + +LVYV+D++I GND + ++ L+ +F +K+ L YFLG+E
Sbjct: 1119 FVLQ-RGRSIIYMLVYVDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEAKRV 1177
Query: 1214 KQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIY 1273
QG+ +SQR+Y LDLL T L KP+ P+ + K+ +Y+ +VG L Y
Sbjct: 1178 PQGLHLSQRRYTLDLLARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGSLQY 1237
Query: 1274 LSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTD 1333
L+ TRPD++Y V+ +SQ+MH P + H A+ R+L+YL +P G+ KK LS+ Y+D
Sbjct: 1238 LAFTRPDLSYAVNRLSQYMHMPTDDHWNALKRVLRYLAGTPDHGIFLKKGNTLSLHAYSD 1297
Query: 1334 ADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKII 1393
AD+AG D ST GY ++LG + ++W SKKQ V RSS EAE+R++A EL + +
Sbjct: 1298 ADWAGDTDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWICSL 1357
Query: 1394 LDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPS 1453
L +L I+ P ++CDN A + NPV H R KHI +D HFI+ ++ S + +V +
Sbjct: 1358 LTELGIQLSHPPVIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVHVST 1417
Query: 1454 GLQLADVLTKGLPLERFRELTCKLGMIDI 1482
QLAD LTK L F+ + K+G+I +
Sbjct: 1418 HDQLADTLTKPLSRVAFQNFSRKIGVIKV 1446
>At2g07010 putative retroelement pol polyprotein
Length = 1413
Score = 625 bits (1611), Expect = e-179
Identities = 415/1250 (33%), Positives = 632/1250 (50%), Gaps = 93/1250 (7%)
Query: 9 PKEKLTSGVKGVSLPPLTHKDLP-VLQGSFRLDGRNYLQWSELVRHTLISRQKISYIEET 67
P+ V VS L D P + S L G NY +WS + + L +++K +I +
Sbjct: 13 PRTNQQPDVTKVSPYTLASSDNPGAMISSVMLTGDNYNEWSTKMLNALQAKRKTGFINGS 72
Query: 68 A--PAETDPQYDTWREENSLIMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDL 125
P +P Y+ W+ NS+I+ W S+ P++ F A ++W+ L Q +S +
Sbjct: 73 ISKPPLDNPDYENWQAVNSMIVGWIRASIEPKVKSTVTFICDAHQLWSELKQRFSVGNKV 132
Query: 126 SACYELENKVFNSKQGTLSVTDYYGTLNGLWIELDQYQNLKM-KCD--SDSTTL--NKFI 180
++++ ++ +Q V DYYG L LW E Y+ + + KC + TL +K
Sbjct: 133 HV-HQIKTQLAACRQDGQPVIDYYGRLCKLWEEFQIYKPITVCKCGLCTCGATLEPSKER 191
Query: 181 EKARIFKFLSGLN-SEFDPIRVQILGKEQFPSLSEVFS-IVRGEETRRTVMVEDKSVDGS 238
E+ +I +F+ GL+ S F + ++ + FPSL E++S +VR E+ +V + ++
Sbjct: 192 EEEKIHQFVLGLDDSRFGGLSATLIAMDPFPSLGEIYSRVVREEQRLASVQIREQQQSAI 251
Query: 239 ALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYG 298
+ + + T+ GR + +RD R C++C RSGH K+ C+++ G
Sbjct: 252 GFLTRQSEV---TADGRTDSSIIKSRD----------RSVLCSHCGRSGHEKKDCWQIVG 298
Query: 299 -------RENVLARMSKSKGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELERLR 351
R N R S S+G R + S ++ + ++LR
Sbjct: 299 FPDWWTERTNGGGRGSSSRGRGGRSSGSNNSGRGRGQVTAAHATTSNLSPFPEFTPDQLR 358
Query: 352 VFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYP 411
V + + S ++GK D I+D GA+ HMT S L++ T P
Sbjct: 359 VITQMIQNKNNGTSDKLSGKMKL----------GDVILDTGASHHMTGQLSLLTNIVTIP 408
Query: 412 GKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFH 471
+ A+G + G LS ++ L VL+VP L+ +L+S+ KL + + C F
Sbjct: 409 SCS-VGFADGRKTFAISMGTFKLSETVSLSNVLYVPALNCSLISVSKLVKQIKCLALFTD 467
Query: 472 SHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSI 531
+ V QD + +IG +E+DG+Y L + T + T+ L H+RLGHP FS+
Sbjct: 468 TICVLQDRFSRTLIGTGEERDGVYYLTDAATT---TVHKVDITTDHALWHQRLGHPSFSV 524
Query: 532 LRTMFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVS 591
L ++ S SV S CDVC +K R F S+NK F L+H DVWGP + +
Sbjct: 525 LSSLPLFSGSSCSVSSRSCDVCFRAKQTREVFPDSSNKSTDCFSLIHCDVWGPYRVPSSC 584
Query: 592 GAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNK 651
GA +F+T +DD +R W +L+ KSEV + NF QFGK +K +RSDNG E +
Sbjct: 585 GAVYFLTIVDDFSRSVWTYLLLAKSEVRSVLTNFLAYTEKQFGKSVKIIRSDNGTEFMC- 643
Query: 652 NFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAA 711
S + + G+VH+ +CV PQQNG ERK+RH+L ++RALLFQ ++P +WGEAV+TAA
Sbjct: 644 -LSSYFKEQGIVHQTSCVGTPQQNGRVERKHRHILNVSRALLFQASLPIKFWGEAVMTAA 702
Query: 712 YLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIK 771
YLINR PS + + +SP ++ P Q RVFG + + H + K R+
Sbjct: 703 YLINRTPSSIHNGLSPYELLHGCKPDYD-----QLRVFGSACYAHRVTRDKDKFGERSRL 757
Query: 772 CIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIP 831
CIF+GY +KG+K Y + +S DV F+E+ Y + GDT +P ++
Sbjct: 758 CIFVGYPFGQKGWKVYDLSTNEFIVSRDVVFRENVFPYATNE---GDT--IYTPPVTCPI 812
Query: 832 LLQEPLMP--------------SSIPI----TEDSDDDDNGPEQVPNQNDDN---GLELV 870
E +P S P+ +SD + + P+ +P DD +
Sbjct: 813 TYDEDWLPFTTLEDRGSDENYLSDPPVCVTNVSESDTEHDTPQSLPTPVDDPLSPSTSVT 872
Query: 871 PDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVP---------NPE 921
P Q N NV Q+ P++ + +V P N
Sbjct: 873 PTQTPTNSSSSTSPSTNVS----PPQQDTTPIIENTPPRQGKRQVQQPARLKDYILYNAS 928
Query: 922 PSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEV 981
+ + H L + +YP++ Y+S + S H+ F++AI + P +E
Sbjct: 929 CTPNTPHVLSPSTSQSSSSIQGNLQYPLTDYISDECFSAGHKVFLAAITANDEPKHFKED 988
Query: 982 LKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVA 1041
+K K W AM +E+ AL+ N T IV+ P K +G +W+Y K+ +DGT++RYKARLV
Sbjct: 989 VKVKVWNDAMYKEVDALEVNKTWDIVDLPTGKVAIGSQWVYKTKFNADGTVERYKARLVV 1048
Query: 1042 KGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEI 1101
+G Q G DY ETFAPV KM TVR +L L A WE+ Q DV NAFLHGDLEEEVYM++
Sbjct: 1049 QGNNQIEGEDYTETFAPVVKMTTVRTLLRLVAANQWEVYQMDVHNAFLHGDLEEEVYMKL 1108
Query: 1102 PPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNG 1161
PPG+ S +KVC+L+K+LYGLKQ+PR WF + ++A+ G+ Q D++ F ++ G
Sbjct: 1109 PPGF-RHSHPDKVCRLRKSLYGLKQAPRCWFKKLSDALKRFGFIQGYEDYS-FFSYSCKG 1166
Query: 1162 KLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVA 1211
+LVYV+D+II GNDE Q ++ L F MKDLGKLKYFLG+EV+
Sbjct: 1167 IELRVLVYVDDLIICGNDEYMVQKFKEYLGRCFSMKDLGKLKYFLGIEVS 1216
Score = 181 bits (460), Expect = 2e-45
Identities = 90/198 (45%), Positives = 137/198 (68%)
Query: 1288 VSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTT 1347
VS+ P+E HL+A RI++YLK SPG+G+L ++ L++EVY D+D+ + RRS +
Sbjct: 1215 VSRGPDAPREAHLEAAMRIVRYLKGSPGQGILLSANKDLTLEVYCDSDFQSCPLTRRSLS 1274
Query: 1348 GYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRL 1407
Y + LGG+ ++W++KKQ+ V+ SSAEAE+RAM+ + E+ + +L +L I AP RL
Sbjct: 1275 AYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSVALKEIKWLNKLLKELGITLAAPTRL 1334
Query: 1408 FCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPL 1467
FCD+K+AISIA NPV H+RTKHIE D H +++ + +I+T +V + QLAD+ TK L
Sbjct: 1335 FCDSKAAISIAANPVFHERTKHIERDCHSVRDAVRDGIITTHHVRTSEQLADIFTKALGR 1394
Query: 1468 ERFRELTCKLGMIDIHSP 1485
+F L KLG+ ++H+P
Sbjct: 1395 NQFIYLMSKLGIQNLHTP 1412
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 560 bits (1443), Expect = e-159
Identities = 415/1307 (31%), Positives = 617/1307 (46%), Gaps = 237/1307 (18%)
Query: 196 FDPIRVQILGKEQFPSLSEVFS-IVRGEET---RRTVMVEDKSVDGSALASGKGPIKGST 251
F PIR +I ++ PS + V+S ++RG++ R+ + ++ + P +
Sbjct: 15 FAPIRSKITDEDPLPSHNRVYSRVIRGQQNLDVARSKETTNSEAINFSVKTPSAPQVAAV 74
Query: 252 SFGRPNRDDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYG-RENVLARMSKSK 310
+P RD CT+C R GH CF ++G E + S+
Sbjct: 75 YAPKP-------------------RDRSCTHCHRQGHDVTDCFLVHGFPEWYYEQKGGSR 115
Query: 311 GAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELE---RLRVFMDSLSKPSGSCSLT 367
+ R S +EN PA+ SK R+ LS +GS +T
Sbjct: 116 VSSDNR--EVVSRLENK-------PAKREGRSSKGNGRGRGRVNSARAPLSSSNGSDQIT 166
Query: 368 MT-------GKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSS-YSTYPGKHYITVA 419
ST L+ + D IID GA+ HMT S L + P +T
Sbjct: 167 QLISLLQAQRPKSTSERLSGNTCLTDVIIDSGASHHMTGDCSILVDVFDIIPSA--VTKP 224
Query: 420 NGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDL 479
+G + T C + LS S L+ VL VP L+S+ KL + G F
Sbjct: 225 DGKASCATKCVTLLLSSSYKLQDVLFVPDFDCTLISVSKLLKQT----------GPF--- 271
Query: 480 ATGQIIGIAKEQDGLYCLQHE-ESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPH 538
+ +IG + ++ +Y + H+TSS+S ++ +W H+RLGHP S L ++
Sbjct: 272 -SRTLIGAGEVRERVYYFTGVLVASAHKTSSDSTSSGALW--HRRLGHPSTSFLLSL--- 325
Query: 539 LFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVT 598
C H S L + C
Sbjct: 326 ------------PEC----HQSSKDLGKIDSC---------------------------- 341
Query: 599 FIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLT 658
D C+R K + L NF M QF KP++ +RSDNG E + + +
Sbjct: 342 --DTCSRA--------KQTLPSLIRNFCAMADRQFRKPVRSIRSDNGTEFMCH--TSYFQ 389
Query: 659 KNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLP 718
++G++HE +CVD PQQN ERK+RH+L + R LFQ N P P
Sbjct: 390 EHGILHETSCVDTPQQNARVERKHRHILNVARTCLFQGNFPT-----------------P 432
Query: 719 SRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYA 778
S VL +P V+ PS + R FGC + H+ + K R+ KCIFIGY
Sbjct: 433 SPVLKGKTPYEVLFGKQPSYDML-----RTFGCLCYAHIRPRDKDKFASRSRKCIFIGYP 487
Query: 779 SDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLM 838
+ + + S+D T S+ E +P V P ++P
Sbjct: 488 --------HETATPNTHDSIDPTSTSSD--------------ENNTPPEPVTPQAEQPHS 525
Query: 839 PSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRK 898
PSSI + P V N+ + L D +D + P L
Sbjct: 526 PSSI----------SSPHIVHNKGSVHSRHLNED-HDSSSPGL----------------- 557
Query: 899 VKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNL 958
P L+ + +P P V + + H + P G +S + +
Sbjct: 558 --PELLGKGHRPKHPPVYL-----KDYVAHKVHSSPHTSSPG--------LSDSNVSPTV 602
Query: 959 SVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGC 1018
S H +F++AI ++ + K+W AM +E+ AL+ N T + + P+ K+ +
Sbjct: 603 SANHIAFMAAILDSNEQNHFKDDVLIKEWCDAMQKEIEALEANHTWDVTDLPHGKKAISS 662
Query: 1019 KWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWE 1078
KW+Y +K+ SDGTL+R+KARLV G Q GID+ ETFAPVAKM TVR++L++AA DW+
Sbjct: 663 KWVYKLKFNSDGTLERHKARLVVMGNHQKEGIDFKETFAPVAKMTTVRLLLAVAAAKDWD 722
Query: 1079 LQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNA 1138
+ Q DV NAFLHGDLE+E +LYGLKQ+PR WF + + A
Sbjct: 723 VFQMDVHNAFLHGDLEQE-----------------------SLYGLKQAPRCWFAKLSTA 759
Query: 1139 MVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKD 1198
+ LG+ QS D++LF + R+G + LVYV+D II GN+ + ++ L F MKD
Sbjct: 760 LRKLGFTQSYEDYSLFSLN-RDGTVIHFLVYVDDFIIVGNNLKAIDHFKEHLHKCFHMKD 818
Query: 1199 LGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLR 1258
LGKLKYFLG+EV+ G +SQ+KY LD++ E G LG KP VP+E +HK+GS +
Sbjct: 819 LGKLKYFLGLEVSRGADGFCLSQQKYALDIINEAGLLGYKPSAVPMELHHKLGSISSPVF 878
Query: 1259 VHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGL 1318
+ AQY+RLV + IYL+ TRPD++Y V ++SQFM P E H A R+++YLK SP +G+
Sbjct: 879 DNPAQYRRLVDRFIYLTITRPDLSYAVHILSQFMQTPLEAHWHATLRLVRYLKGSPDQGI 938
Query: 1319 LFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFR 1378
L + LS+ Y D+DY RRS + Y ++LG ++W++KKQ+ V+ SSAEAE+R
Sbjct: 939 LLRSDRALSLTAYCDSDYNPCPRTRRSLSAYVLYLGDTPISWKTKKQDTVSSSSAEAEYR 998
Query: 1379 AMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIK 1438
AMA + E+ +K ++ L + + P+ LFCD+++AI IA NPV H+RTKHIE D H ++
Sbjct: 999 AMAYTLKEIKWLKALMTTLGVDHTQPILLFCDSQAAIHIAANPVFHERTKHIEKDCHQVR 1058
Query: 1439 EKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
+ + +IST ++ + D+LTK LP F L LG + P
Sbjct: 1059 DAVTDKVISTPHIST----TDLLTKALPRPTFERLLSTLGTCNYDLP 1101
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 557 bits (1435), Expect = e-158
Identities = 361/1111 (32%), Positives = 567/1111 (50%), Gaps = 107/1111 (9%)
Query: 383 TENDWIIDFGATDHMTPHPSHLSSYS-TYPGKHYITVANGSQNQITGCGNIHLS----PS 437
T + WI+D G + HMT + + T GK + + N + +++ G G++ + +
Sbjct: 308 TPDTWILDTGCSFHMTCRKDWIIDFKETASGK--VRMGNDTYSEVKGIGDVRIKNEDGST 365
Query: 438 LPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCL 497
+ L V ++P++S NL+S+ L D C F G+ + K++ LY L
Sbjct: 366 ILLTDVRYIPEMSKNLISLGTL-EDKGC--WFESKKGILTIFKNDLTVLTGKKESTLYFL 422
Query: 498 QHEE--SKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFHCDVCQF 555
Q + + E TS +W H RLGH L+ + V H D
Sbjct: 423 QGTTLAGEANVIDKEKDETS-LW--HSRLGHIGAKGLQVL---------VSKGHLD---- 466
Query: 556 SKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSIS-NVSGAKWFVTFIDDCTRITWVFLMKD 614
K+ +F + + D VHSD+WG +++ ++ ++F+TFIDD TR TW++ ++
Sbjct: 467 -KNIMISFGAAKHVTKDKLDYVHSDLWGSTNVPFSIGKCQYFITFIDDFTRRTWIYFIRT 525
Query: 615 KSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQ 674
K E F F + I Q K +K L +DNG E N+ F F K GV+ TC PQQ
Sbjct: 526 KDEAFSKFVEWKTQIENQQDKKLKILITDNGLEFCNQEFDSFCRKEGVIRHRTCAYTPQQ 585
Query: 675 NGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSF 734
NGVAER NR ++ R +L + + K +W EA TA +LIN+ PS + P T
Sbjct: 586 NGVAERMNRTIMNKVRCMLSESGLGKQFWAEAASTAVFLINKSPSSSIEFDIPEEKWTGH 645
Query: 735 FPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRV 794
P I + FG A++H +GKL+PRA K IF+GY K +K + R+
Sbjct: 646 PPDYKI-----LKKFGSVAYIH---SDQGKLNPRAKKGIFLGYPDGVKRFKVWLLEDRKC 697
Query: 795 YISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDDNG 854
+S D+ FQE++ + +LQ D E + V L I + S DD+N
Sbjct: 698 VVSRDIVFQENQMY---KELQKNDMSEEDKQLTEVERTL--------IELKNLSADDENQ 746
Query: 855 PEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPE 914
E N N + D E D + ++ + + R + + Q
Sbjct: 747 SEGGDNSNQEQASTTRSASKDKQVEETDSDDDCLENYLLARDRIRRQIRAPQ-------- 798
Query: 915 VSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRV 974
++V + V ++ +
Sbjct: 799 ------------------------------------RFVEEDDSLVGFALTMTEDGEVYE 822
Query: 975 PTSVQEVLKD---KKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGT 1031
P + +E ++ +KW QA EEM ++ KN T +++KP KR +GCKWI+ K G
Sbjct: 823 PETYEEAMRSPECEKWKQATIEEMDSMKKNDTWDVIDKPEGKRVIGCKWIFKRKAGIPGV 882
Query: 1032 -LDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLH 1090
RYKARLVAKG++Q GIDY E F+PV K ++R +LS+ FD EL+Q DVK AFLH
Sbjct: 883 EPPRYKARLVAKGFSQREGIDYQEIFSPVVKHVSIRYLLSIVVQFDMELEQLDVKTAFLH 942
Query: 1091 GDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGD 1150
G+L+E + M P GY+ KVC LKK+LYGLKQSPR W RF + M++ GY++S+ +
Sbjct: 943 GNLDEYILMSQPEGYEDEDSTEKVCLLKKSLYGLKQSPRQWNQRFDSFMINSGYQRSKYN 1002
Query: 1151 HTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEV 1210
++ + +G LL+YV+DM+IA ++ + Q L++ L +FEMKDLG + LGME+
Sbjct: 1003 PCVYTQQLNDGSYIYLLLYVDDMLIASQNKDQIQKLKESLNREFEMKDLGPARKILGMEI 1062
Query: 1211 AYSK-QGIF-ISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLV 1268
++ QGI +SQ +YV +L+ G K P+ + K+ ++ E A+Y +LV
Sbjct: 1063 TRNREQGILDLSQSEYVAGVLRAFGMDQSKVSQTPLGAHFKLRAANEKTLARDAEYMKLV 1122
Query: 1269 ------GKLIY-LSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFK 1321
G ++Y + +RPD+AYPV VVS+FM P + H QAV +++Y+K + L FK
Sbjct: 1123 PYPNAIGSIMYSMIGSRPDLAYPVGVVSRFMSKPSKEHWQAVKWVMRYMKGTQDTCLRFK 1182
Query: 1322 KSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMA 1381
K ++ + Y D+DYA + RRS TG+ GGN ++W+S Q VVA S+ EAE+ A+A
Sbjct: 1183 KDDKFEIRGYCDSDYATDLDRRRSITGFVFTAGGNTISWKSGLQRVVALSTTEAEYMALA 1242
Query: 1382 QGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKL 1441
+ V E + ++ + ++ + +A + + CD++SAI+++ N V H+RTKHI++ HFI+EK+
Sbjct: 1243 EAVKEAIWLRGLAAEMGFEQDA-VEVMCDSQSAIALSKNSVHHERTKHIDVRYHFIREKI 1301
Query: 1442 NSSLISTQYVPSGLQLADVLTKGLPLERFRE 1472
I + + AD+ TK +P+ + +E
Sbjct: 1302 ADGEIQVVKISTTWNPADIFTKTVPVSKLQE 1332
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 528 bits (1360), Expect = e-150
Identities = 276/694 (39%), Positives = 424/694 (60%), Gaps = 15/694 (2%)
Query: 777 YASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEP 836
Y S KGYK S + I+ +V F E++ + + + + S++PL
Sbjct: 335 YPSGYKGYKVLDLESHSISITRNVVFHETKFPFKTSKFL---KESVDMFPNSILPLPAPL 391
Query: 837 LMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQ-NDDNGPELVPDQNNVDRFRIKY 895
S+P+ +D DDN + + + + +P N N L D N+V R K
Sbjct: 392 HFVESMPLDDDLRADDNNASTSNSASSASSIPPLPSTVNTQNTDALDIDTNSVPIARPKR 451
Query: 896 QRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVST 955
K P + + S P +S +P S S E +P K+ YP+S +S
Sbjct: 452 NAKA-PAYLSEYHCNSVPFLSSLSPTTSTSIETPSSSIP-----PKKITTPYPMSTAISY 505
Query: 956 QNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRT 1015
L+ S+I A + P + + +K +KW +A NEE+HAL++N T + K
Sbjct: 506 DKLTPLFHSYICAYNVETEPKAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNV 565
Query: 1016 VGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHF 1075
VGCKW++T+KY DG+++RYKARLVA+G+TQ GIDY ETF+PVAK +V+++L LAA
Sbjct: 566 VGCKWVFTIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAAT 625
Query: 1076 DWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSG----RNKVCKLKKALYGLKQSPRAW 1131
W L Q DV NAFLHG+L+EE+YM +P GY P +G VC+L K+LYGLKQ+ R W
Sbjct: 626 GWSLTQMDVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQASRQW 685
Query: 1132 FGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELA 1191
+ R ++ + + QS D+T+F+K + + ++LVYV+D++IA ND +NL++ L
Sbjct: 686 YKRLSSVFLGANFIQSPADNTMFVKVSCTS-IIVVLVYVDDLMIASNDSSAVENLKELLR 744
Query: 1192 VQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIG 1251
+F++KDLG ++FLG+E+A S +GI + QRKY +LL++ G GCKP +P++ N +
Sbjct: 745 SEFKIKDLGPARFFLGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLT 804
Query: 1252 SSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLK 1311
L + Y+ LVG+L+YL TRPDI + V +SQF+ P + H+QA ++L+YLK
Sbjct: 805 KEMGTLLPNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLK 864
Query: 1312 ASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARS 1371
+PG+GL++ S +L + ++DAD+ RRS TG+C++LG +L+TW+SKKQ+VV+RS
Sbjct: 865 GNPGQGLMYSASSELCLNGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRS 924
Query: 1372 SAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIE 1431
S E+E+R++AQ CE++ ++ +L DL + P +LFCDNKSA+ +A NPV H+RTKHIE
Sbjct: 925 STESEYRSLAQATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIE 984
Query: 1432 IDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGL 1465
ID H +++++ + + T +VP+G QLAD+LTK L
Sbjct: 985 IDCHTVRDQIKAGKLKTLHVPTGNQLADILTKPL 1018
Score = 91.3 bits (225), Expect = 4e-18
Identities = 54/164 (32%), Positives = 88/164 (52%), Gaps = 22/164 (13%)
Query: 595 WFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFS 654
+F+T +DDCTR TWV++MK+KSEV +F F +I TQ+ IK +RSDN KE F+
Sbjct: 252 YFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKE---LAFT 308
Query: 655 EFLTKNGVVHELTCVDAPQQNGVAER------------KNRHLLEITRALLFQMNVPKFY 702
+F+ + G++H+ +C PQQN V ER H + ITR ++F F
Sbjct: 309 KFVKEQGMIHQFSCAYTPQQNSVVERYPSGYKGYKVLDLESHSISITRNVVFHETKFPFK 368
Query: 703 WGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQS 746
+ + + ++ P+ +L +P+ F S+P+ L++
Sbjct: 369 TSKFLKES---VDMFPNSILPLPAPL----HFVESMPLDDDLRA 405
Score = 61.6 bits (148), Expect = 3e-09
Identities = 50/163 (30%), Positives = 78/163 (47%), Gaps = 6/163 (3%)
Query: 368 MTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQIT 427
+T ++ T SL + + + WIID GA+ H+ + G +T+ NG++ IT
Sbjct: 80 LTFQNHTLSSLQNVLSSDAWIIDSGASSHVCSDLTMFRELIHVSGVT-VTLPNGTRVAIT 138
Query: 428 GCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGI 487
G I ++ +L L VL VP NL+S+ L + L+ + FF Q+L G +IG
Sbjct: 139 HTGTICITSTLILHNVLLVPDFKFNLISVCCLVKTLSYSAHFFADCCYIQELTRGLMIGR 198
Query: 488 AKEQDGLYCLQHEE---SKCHQTSSESWNTSQ--IWLRHKRLG 525
K + LY L+ + S +S T Q L H+RLG
Sbjct: 199 GKTYNNLYILETQRTSFSPSLPAASSFTGTVQDDCLLWHQRLG 241
>At4g22040 LTR retrotransposon like protein
Length = 1109
Score = 513 bits (1322), Expect = e-145
Identities = 254/540 (47%), Positives = 372/540 (68%), Gaps = 3/540 (0%)
Query: 946 KYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRK 1005
++P S+ +S + H++F++A+ + PT+ E + DK W +AM+ E+ +L N T
Sbjct: 572 EFPYSK-MSCNRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFS 630
Query: 1006 IVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTV 1065
IV P KR +G KW+Y +KYRSDG ++RYKARLV G Q G+DYDETFAPVAKM+TV
Sbjct: 631 IVNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTV 690
Query: 1066 RIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLK 1125
R+ L +AA DW + Q DV NAFLHGDL+EEVYM++P G+ +KVC+L K+LYGLK
Sbjct: 691 RLFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQ-CDDPSKVCRLHKSLYGLK 749
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQN 1185
Q+PR WF + ++A+ G+ QS D++LF + +G +LVYV+D+II+G+
Sbjct: 750 QAPRCWFSKLSSALKQYGFTQSLSDYSLF-SYNNDGVFVHVLVYVDDLIISGSCPDAVAQ 808
Query: 1186 LRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIE 1245
+ L F MKDLG LKYFLG+EV+ + QG ++SQRKYVLD++ E G LG +P P+E
Sbjct: 809 FKSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLE 868
Query: 1246 QNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDR 1305
QNHK+ S L ++Y+RLVG+LIYL+ TRP+++Y V ++QFM +P++ H A R
Sbjct: 869 QNHKLSLSTSPLLSDSSRYRRLVGRLIYLAVTRPELSYSVHTLAQFMQNPRQDHWNAAIR 928
Query: 1306 ILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQ 1365
+++YLK++PG+G+L + L + + D+DYA + RRS TGY + LG ++W++KKQ
Sbjct: 929 VVRYLKSNPGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQ 988
Query: 1366 NVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHD 1425
++RSSAEAE+RAMA EL+ +K +L DL + + MR+F D+KSAI+++ NPVQH+
Sbjct: 989 PTISRSSAEAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHE 1048
Query: 1426 RTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
RTKH+E+D HFI++ + +I+T +VPS QLAD+LTK L + R KLG++D+H+P
Sbjct: 1049 RTKHVEVDCHFIRDAILDGIIATSFVPSHKQLADILTKALGEKEVRYFLRKLGILDVHAP 1108
Score = 253 bits (645), Expect = 8e-67
Identities = 150/419 (35%), Positives = 226/419 (53%), Gaps = 14/419 (3%)
Query: 393 ATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNN 452
A+ HMT + LS + I +A+G++ G + L L LK V +V +L ++
Sbjct: 169 ASHHMTGNLELLSGMRSM-SPVLIILADGNKRVAVSEGTVRLGSHLILKSVFYVKELESD 227
Query: 453 LLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESW 512
L+S+ ++ + +C V V QD T + GI K ++G +C + E+ +S
Sbjct: 228 LISVGQMMDENHCVVQLADHFLVIQDRTTRMVTGIGKRENGSFCFRGMENAAAVHTSVK- 286
Query: 513 NTSQIWLRHKRLGHPPFSILRTMFPHLFS--KVSVESFHCDVCQFSKHHRSTFLPSNNKC 570
+ L H+RLGH I+ + L S K +E+ CD C +K R TF +N+
Sbjct: 287 --APFDLWHRRLGHASDKIVNLLPRELLSSGKEILENV-CDTCMRAKQTRDTFPLRDNRS 343
Query: 571 AQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIR 630
F L+H DVWGP + SGA++F+T +DD +R WV+LM DKSE + +F ++
Sbjct: 344 MDSFQLIHCDVWGPYRTPSYSGARYFLTIVDDYSRGVWVYLMTDKSETQKHLKDFMALVE 403
Query: 631 TQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITR 690
QF IK +RSDNG E + E+ G+ HE +CV P QNG ERK+RH+L I R
Sbjct: 404 RQFDTEIKTVRSDNGTEFL--CMREYFLHKGIAHETSCVGTPHQNGRVERKHRHILNIAR 461
Query: 691 ALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFG 750
AL FQ +P +WGE +L+AAYLINR PS +L SP ++ + + P S L RVFG
Sbjct: 462 ALRFQSYLPIQFWGECILSAAYLINRTPSMLLQGKSPYEML---YKTAPKYSHL--RVFG 516
Query: 751 CSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFY 809
+ H H K R+ +C+F+GY +KG++ + ++ ++S DV FQE+E Y
Sbjct: 517 SLCYAHNQNHKGDKFAARSRRCVFVGYPHGQKGWRLFDLEEQKFFVSRDVIFQETEFPY 575
Score = 63.9 bits (154), Expect = 7e-10
Identities = 47/187 (25%), Positives = 85/187 (45%), Gaps = 31/187 (16%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYIEETAPAETD--PQYDTWREENSLIMTWFWYSMIP 96
L NY +W+ + L SR+K +++ T P D P + W N+L+++W ++
Sbjct: 38 LKTNNYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLEDWLTINALLVSWMKMTIDS 97
Query: 97 EISRNYMFFPTAKEIWNNLAQAYS---------KKQDLSACYELENKVFNSKQGTLSVTD 147
E+ N A+++W + + +S K DL+ C KQ ++V
Sbjct: 98 ELLTNISHRDVARDLWEQIRKRFSVSNGPKNQKMKADLATC----------KQEGMTVEG 147
Query: 148 YYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGKE 207
YYG LN +W ++ Y+ L++ C S T N + LSG+ S P+ + +
Sbjct: 148 YYGKLNKIWDNINSYRPLRI-CASHHMTGN--------LELLSGMRS-MSPVLIILADGN 197
Query: 208 QFPSLSE 214
+ ++SE
Sbjct: 198 KRVAVSE 204
>At2g24660 putative retroelement pol polyprotein
Length = 1156
Score = 489 bits (1260), Expect = e-138
Identities = 272/678 (40%), Positives = 405/678 (59%), Gaps = 34/678 (5%)
Query: 811 NPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELV 870
N LQ E +S + L S + ++ S D N D+ ++V
Sbjct: 480 NSALQTDSVFEPDSDSDVAPASIASDLQNSRVDSSDSSTPDKNLASGDTLAQIDDSPDIV 539
Query: 871 PDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPS-DPEVSVPNPE--PSNSYE 927
N +N P V D V+ + +++ ++ QD + VS NP P +S +
Sbjct: 540 STPNRNNQPLFVVDSPFVEATSPRQRKRQIRQSVRLQDYVLYNATVSPINPHALPDSSSQ 599
Query: 928 HNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKW 987
+ ++ +G S YP+S YVS S H++F++AI + P +E ++ K W
Sbjct: 600 SS------SMVQGTSSL--YPLSDYVSDDCFSAGHKAFLAAITANDEPKHFKEAVRIKVW 651
Query: 988 IQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQT 1047
AM +E+ AL+ N T IV+ P K +G +W+Y KY +DG+++RYKARLV +G Q
Sbjct: 652 NDAMFKEVDALEINKTWDIVDLPPGKVAIGSQWVYKTKYNADGSIERYKARLVVQGNKQV 711
Query: 1048 YGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDP 1107
G DY+ETFAPV KM TVR +L L A WE+ Q DV NAFLHGDL+EEVYM++PPG+
Sbjct: 712 EGEDYNETFAPVVKMTTVRTLLRLVAANQWEVYQMDVNNAFLHGDLDEEVYMKLPPGFRH 771
Query: 1108 TSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLL 1167
S +KVC+L+K+LYGLKQ+PR WF + ++A++ G+ Q D++ F +TRNG +L
Sbjct: 772 -SHPDKVCRLRKSLYGLKQAPRCWFKKLSDALLRFGFVQGHEDYSFF-SYTRNGIELRVL 829
Query: 1168 VYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLD 1227
VYV+D++I GND Q ++ L F MKDLGKLKYFLG+EV+ +GIF+SQRKY LD
Sbjct: 830 VYVDDLLICGNDGYMLQKFKEYLGRCFSMKDLGKLKYFLGIEVSRGSEGIFLSQRKYALD 889
Query: 1228 LLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSV 1287
++ ++G LGC+P P+EQNH + + + L Y+RLVG+L+YL HTRP+++Y V V
Sbjct: 890 IITDSGNLGCRPALTPLEQNHHLATDDGPLLADAKPYRRLVGRLLYLLHTRPELSYSVHV 949
Query: 1288 VSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTT 1347
+SQFM P+E HL A RI+++LK SPG+G+L +L ++VY D+D+ RRS +
Sbjct: 950 LSQFMQTPREAHLAAAMRIVRFLKGSPGQGILLNADTELQLDVYCDSDFQTCPKTRRSLS 1009
Query: 1348 GYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRL 1407
Y + LGG+ ++W++KKQ+ V+ SSAEAE+RAM+ + E+ ++ +L +L
Sbjct: 1010 AYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSVALREIKWLRKLLKELA--------- 1060
Query: 1408 FCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPL 1467
NPV H+RTKHIE D H +++ + +I+T +V + QLAD+ TK L
Sbjct: 1061 ------------NPVFHERTKHIESDCHSVRDAVRDGIITTHHVRTNEQLADIFTKALGR 1108
Query: 1468 ERFRELTCKLGMIDIHSP 1485
+F L KLG+ D+H+P
Sbjct: 1109 SQFLYLMSKLGIQDLHTP 1126
Score = 242 bits (617), Expect = 1e-63
Identities = 160/469 (34%), Positives = 238/469 (50%), Gaps = 28/469 (5%)
Query: 450 SNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSS 509
SN SI R + +T + D + +IG +E+DG+Y L + T+
Sbjct: 24 SNKEESISSEHRLVTLTLTHENLKACKLDRFSRTLIGSGEERDGVYYLTDVATAKIHTAK 83
Query: 510 ESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNK 569
S + +W H+RLGHP FS+L ++ S +SV S CDVC +K R F S NK
Sbjct: 84 VS-SDQALW--HQRLGHPSFSVLSSLPVLTSSSLSVGSRSCDVCFRAKQTREVFPVSTNK 140
Query: 570 CAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMI 629
+ F L+H DVWGP + + GA +F+T +DD +R W +L+ KSEV + NF
Sbjct: 141 SIECFSLIHCDVWGPYRVPSSCGAVYFLTIVDDFSRAVWTYLLLAKSEVRTVLTNFLVYT 200
Query: 630 RTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEIT 689
QFGK +K LRSDNG E + + + ++G+VH+ +CV PQQNG ERK+RH+L +
Sbjct: 201 EKQFGKSVKVLRSDNGTEFM--CLASYFREHGIVHQTSCVGTPQQNGRVERKHRHILNVA 258
Query: 690 RALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVF 749
RA+LFQ ++P +WGEAVLTAAYLINR P+ + + +SP ++ + P+ RVF
Sbjct: 259 RAILFQASLPIQFWGEAVLTAAYLINRTPTSLHNGLSPYEILHNSKPNYE-----HLRVF 313
Query: 750 GCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFY 809
G + +VH + K R+ C+FIGY +KG+K + + +S DV F+ E +
Sbjct: 314 GSACYVHRASRDKDKFGERSRLCVFIGYPFAQKGWKVFDMEKKEFLVSRDVVFR--EDVF 371
Query: 810 PNPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDDNGPEQV--PNQNDDNGL 867
P + +P L+ E M S+ + D G V N+N D
Sbjct: 372 PYAATNTDHVSASPAP-----LLVDEDWMVSTTTL-------DRGSSAVLSANKNTDPKP 419
Query: 868 ELVPDQNDDNGPELV--PDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPE 914
V +D PE V PD K +PV + + D ++P+
Sbjct: 420 NSVTKPDDFAEPESVTKPDDFAEPESVTKPDDFAEPVSVTKPDDFAEPD 468
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 488 bits (1257), Expect = e-138
Identities = 243/540 (45%), Positives = 366/540 (67%), Gaps = 6/540 (1%)
Query: 947 YPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKI 1006
+PIS +S +S H +I+ I I +P S E K+W A+++E+ A+++ T +I
Sbjct: 919 HPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIGAMERTDTWEI 978
Query: 1007 VEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVR 1066
P K+ VGCKW++TVK+ +DG+L+R+KAR+VAKGYTQ G+DY ETF+PVAKM TV+
Sbjct: 979 TSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVK 1038
Query: 1067 IILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGR----NKVCKLKKALY 1122
++L ++A W L Q D+ NAFL+GDLEE +YM++P GY G N VC+LKK++Y
Sbjct: 1039 LLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIY 1098
Query: 1123 GLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELE 1182
GLKQ+ R WF +F+N++++LG+++ GDHTLF++ + +LLVYV+D++IA E
Sbjct: 1099 GLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCI-GSEFIVLLVYVDDIVIASTTEQA 1157
Query: 1183 KQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGV 1242
Q+L + L F++++LG LKYFLG+EVA + +GI +SQRKY L+LL L CKP +
Sbjct: 1158 AQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSADMLDCKPSSI 1217
Query: 1243 PIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQA 1302
P+ N ++ ++ L K Y+RLVGKL+YL+ TRPDI + V+ + QF P+ HL A
Sbjct: 1218 PMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAA 1277
Query: 1303 VDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRS 1362
V ++LQY+K + G+GL + + L+++ YTDAD+ RRSTTG+ MF+G +L++WRS
Sbjct: 1278 VYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISWRS 1337
Query: 1363 KKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPV 1422
KKQ V+RSSAEAE+RA+A CE+ + +L L++ P+ L+ D+ +A+ IA NPV
Sbjct: 1338 KKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVHSGVPI-LYSDSTAAVYIATNPV 1396
Query: 1423 QHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDI 1482
H+RTKHIEID H ++EKL++ + +V + Q+AD+LTK L +F L K+ + +I
Sbjct: 1397 FHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAHLLSKMSIQNI 1456
Score = 387 bits (993), Expect = e-107
Identities = 266/860 (30%), Positives = 424/860 (48%), Gaps = 92/860 (10%)
Query: 27 HKDLPVLQGSFRLDGRNYLQWSELVRHTLISRQKISYIEETAPA--ETDPQYDTWREENS 84
H L ++ S RLD Y WS +R +L ++ K+ +++ + P E+DP + W NS
Sbjct: 73 HPGLSII--SHRLDETTYGDWSVAMRISLDAKNKLGFVDGSLPRPLESDPNFRLWSRCNS 130
Query: 85 LIMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLS 144
++ +W S+ P+I R+ + A +IW +L ++ +L Y L ++ + +QGT+S
Sbjct: 131 MVKSWLLNSVSPQIYRSILRLNDATDIWRDLFDRFNLT-NLPRTYNLTQEIQDLRQGTMS 189
Query: 145 VTDYYGTLNGLWIELDQYQNLKMKCD-SDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQI 203
+++YY L LW +LD + L C + L + EKA+I KFL+GLN + +R QI
Sbjct: 190 LSEYYTLLKTLWDQLDSTEALDDPCTCGKAVRLYQKAEKAKIMKFLAGLNESYAIVRRQI 249
Query: 204 LGKEQFPSLSEVFSIVRGEETRR--------TVMVEDKSVDGSALASGKGP-IKGSTSFG 254
+ K+ PSL+EV+ I+ + +++ + V S + S + ++ + G
Sbjct: 250 IAKKALPSLAEVYHILDQDNSQKGFFNVVAPPAAFQVSEVSHSPITSPEIMYVQSGPNKG 309
Query: 255 RPNRDDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQ 314
RP C++C R GH E C+K +G KS P
Sbjct: 310 RPT----------------------CSFCNRVGHIAERCYKKHGFPPGFTPKGKSSDKPP 347
Query: 315 R----RANHTASDMENDGEVPPVP------PAEDVPSLSKAELERLRVFMDSLS---KPS 361
+ A T S + G++ + +++ +L ++L+ V + S + S
Sbjct: 348 KPQAVAAQVTLSPDKMTGQLETLAGNFSPDQIQNLIALFSSQLQPQIVSPQTASSQHEAS 407
Query: 362 GSCSLTMTG---KSSTF-------LSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYP 411
S S+ +G ST+ +S N+LS++ W+ID GAT H++ H L
Sbjct: 408 SSQSVAPSGILFSPSTYCFIGILAVSHNSLSSDT-WVIDSGATHHVS-HDRKLFQTLDTS 465
Query: 412 GKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFH 471
++ + G +I+G G + ++ + L+ VL +P+ NL+SI LT DL V F
Sbjct: 466 IVSFVNLPTGPNVRISGVGTVLINKDIILQNVLFIPEFRLNLISISSLTTDLGTRVIFDP 525
Query: 472 SHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSI 531
S QDL G +G K LY L +++ S + +W HKRLGHP FS
Sbjct: 526 SCCQIQDLTKGLTLGEGKRIGNLYVL---DTQSPAISVNAVVDVSVW--HKRLGHPSFSR 580
Query: 532 LRTMFPHLFSK--VSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISN 589
L ++ L + + +S +C VC +K + +F +NN C F+L+H DVWGP S+
Sbjct: 581 LDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFSVET 640
Query: 590 VSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*V 649
V G K+F+T +DD +R TW++L+K KS+V +F F ++ Q+ +K +RSDN KE
Sbjct: 641 VEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKE-- 698
Query: 650 NKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLT 709
F+EF G+V +C + P+QN V ERK++H+L + RAL+FQ N+ YWG+ VLT
Sbjct: 699 -LAFTEFYKAKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLT 757
Query: 710 AAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRA 769
A +LINR PS +LSN +P V+T P Q + FGC + R K PR+
Sbjct: 758 AVFLINRTPSALLSNKTPFEVLTGKLPDYS-----QLKTFGCLCYSSTSSKQRHKFLPRS 812
Query: 770 IKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQE---------------SESFYPNPQL 814
C+F+GY KGYK S V+IS +V F E S+ F P L
Sbjct: 813 RACVFLGYPFGFKGYKLLDLESNVVHISRNVEFHEELFPLASSQQSATTASDVFTPMDPL 872
Query: 815 QGGDTQEAESPELSVIPLLQ 834
G++ + P + P Q
Sbjct: 873 SSGNSITSHLPSPQISPSTQ 892
>At4g17450 retrotransposon like protein
Length = 1433
Score = 485 bits (1248), Expect = e-137
Identities = 233/533 (43%), Positives = 351/533 (65%), Gaps = 5/533 (0%)
Query: 957 NLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTV 1016
N + +FI+ I + +P E K W AM EE+ A+ + T +V P +K+ +
Sbjct: 899 NTTEPFHAFINNITNAVIPQRYSEAKDFKAWCDAMKEEIGAMVRTNTWSVVSLPPNKKAI 958
Query: 1017 GCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFD 1076
GCKW++T+K+ +DG+++RYKARLVAKGYTQ G+DY+ETF+PVAK+ +VR++L LAA
Sbjct: 959 GCKWVFTIKHNADGSIERYKARLVAKGYTQEEGLDYEETFSPVAKLTSVRMMLLLAAKMK 1018
Query: 1077 WELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGR----NKVCKLKKALYGLKQSPRAWF 1132
W + Q D+ NAFL+GDL+EE+YM+IPPGY G + +C+L K++YGLKQ+ R W+
Sbjct: 1019 WSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVGEALPPHAICRLHKSIYGLKQASRQWY 1078
Query: 1133 GRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAV 1192
+ +N + +G+++S DHTLFIK+ NG L +LVYV+D++I N + EL
Sbjct: 1079 LKLSNTLKGMGFQKSNADHTLFIKYA-NGVLMGVLVYVDDIMIVSNSDDAVAQFTAELKS 1137
Query: 1193 QFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGS 1252
F+++DLG KYFLG+E+A S++GI I QRKY+L+LL TG LG KP +P++ + K+
Sbjct: 1138 YFKLRDLGAAKYFLGIEIARSEKGISICQRKYILELLSTTGFLGSKPSSIPLDPSVKLNK 1197
Query: 1253 SEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKA 1312
+ Y++LVGKL+YL TRPDIAY V+ + QF H P HL AV ++L+YLK
Sbjct: 1198 EDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVNTLCQFSHAPTSVHLSAVHKVLRYLKG 1257
Query: 1313 SPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSS 1372
+ G+GL + ++ + YTD+D+ RR YCMF+G LV+W+SKKQ+ V+ S+
Sbjct: 1258 TVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCVAAYCMFIGDYLVSWKSKKQDTVSMST 1317
Query: 1373 AEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEI 1432
AEAEFRAM+QG E++ + + DD K+ + P L+CDN +A+ I +N V H+RTK +E+
Sbjct: 1318 AEAEFRAMSQGTKEMIWLSRLFDDFKVPFIPPAYLYCDNTAALHIVNNSVFHERTKFVEL 1377
Query: 1433 DRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
D + +E + S + T +V +G Q+AD LTK + +F +L K+G+ +I +P
Sbjct: 1378 DCYKTREAVESGFLKTMFVETGEQVADPLTKAIHPAQFHKLIGKMGVCNIFAP 1430
Score = 315 bits (808), Expect = 1e-85
Identities = 252/888 (28%), Positives = 396/888 (44%), Gaps = 151/888 (17%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYIEETAPAE--TDPQYDTWREENSLIMTWFWYSMIP 96
LDG NY WS +R +L ++ K+S+++ + P +D + W NS++ TW
Sbjct: 85 LDGTNYNNWSIAMRMSLDAKNKLSFVDGSLPRPDVSDRMFKIWSRCNSMVKTWL------ 138
Query: 97 EISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLW 156
E+WN+L + + +L Y+LE + KQG L ++ YY LW
Sbjct: 139 --------LNVVTEMWNDLFSRF-RVSNLPRKYQLEQSIHTLKQGNLDLSTYYTKKKTLW 189
Query: 157 IELDQYQNLKM-KCDSDSTT-LNKFIEKARIFKFLSGLNSEFDPIRVQILGKEQFPSLSE 214
+L + L + KC+ + L + E +RI +FL GLN F IR QIL + P L+E
Sbjct: 190 EQLANTRVLTVRKCNCEHVKELLEEAETSRIIQFLMGLNDNFAHIRGQILNMKPRPGLTE 249
Query: 215 VFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRF 274
+++++ +E++R V S +A PI D + N
Sbjct: 250 IYNMLDQDESQRLVGNPTLSNPTAAFQVQASPII----------------DSQVNMAQGS 293
Query: 275 NRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRRANHTASDMENDGEVPPVP 334
+ +C+YC + GH + C+K +G +KG N ++ ++ E P
Sbjct: 294 YKKPKCSYCNKLGHLVDKCYKKHGYP---PGSKWTKGQTIGSTNLASTQLQPVNETPN-- 348
Query: 335 PAEDVPSLSKAELERLRVFMDSLSKP-----------------SGSCSLTMTGK-SSTFL 376
E S + ++++ + LS S S S+ M + S TFL
Sbjct: 349 --EKTDSYEEFSTDQIQTMISYLSTKLHIASASPMPTTSSASISASPSVPMISQISGTFL 406
Query: 377 SL----------NALSTE-----NDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANG 421
SL +++S E W+ID GAT H+T + ++ + ++ + N
Sbjct: 407 SLFSNAYYDMLISSVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENT-FVRLPND 465
Query: 422 SQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLAT 481
+I G G I LS ++ L VL++P+ NL+S +LT++L
Sbjct: 466 CTVKIAGIGFIQLSDAISLHNVLYIPEFKFNLIS--ELTKEL------------------ 505
Query: 482 GQIIGIAKEQDGLYCLQHEE-----------SKCHQTSSESWNTSQIWLRHKRLGHPPFS 530
+IG + LY L E S C + S S HKRLGHP +S
Sbjct: 506 --MIGRGSQVGNLYVLDFNENNHTVSLKGTTSMCPEFSVCSSVVVDSVTWHKRLGHPAYS 563
Query: 531 ILRTMFPHLFSKVS-VESFH------CDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWG 583
+ + L KV + H C VC SK +F N C+ FDLVH D WG
Sbjct: 564 KIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNMCSAAFDLVHIDTWG 623
Query: 584 PSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSD 643
P S+ TW++L+K+KS+V +F F +M+ TQ+ +K +RSD
Sbjct: 624 PFSVPTNDA--------------TWIYLLKNKSDVLHVFPAFINMVHTQYQTKLKSVRSD 669
Query: 644 NGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYW 703
N E F++ +G+V +C + P+QN V ERK++H+L + RALLFQ N+P +W
Sbjct: 670 NAHE---LKFTDLFAAHGIVAYHSCPETPEQNSVVERKHQHILNVARALLFQSNIPLEFW 726
Query: 704 GEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRG 763
G+ VLTA +LINRLP+ VL+N SP + + P+ + FGC + R
Sbjct: 727 GDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYE-----SLKTFGCLCYSSTSPKQRH 781
Query: 764 KLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAE 823
K +PRA C+F+GY KGYK + V IS V F E + + ++ D ++
Sbjct: 782 KFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPFISSTIK-DDIKD-- 838
Query: 824 SPELSVIPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVP 871
PLLQ P +P+ + S D + P+Q+ + LVP
Sbjct: 839 -----FFPLLQFPARTDDLPLEQTSIIDTH-----PHQDVSSSKALVP 876
>At4g23160 putative protein
Length = 1240
Score = 457 bits (1177), Expect = e-128
Identities = 236/538 (43%), Positives = 347/538 (63%), Gaps = 11/538 (2%)
Query: 913 PEVSVPN--PEPSNSYEHNLDDLPIALRK----GKRSCAKYPISQYVSTQNLSVQHQSFI 966
P ++ N PEPS H P L+ S + ISQ++S + +S + SF+
Sbjct: 18 PSANIQNDVPEPSVHTSHRRTRKPAYLQDYYCHSVASLTIHDISQFLSYEKVSPLYHSFL 77
Query: 967 SAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKY 1026
I + P++ E + W AM++E+ A++ T +I P +K+ +GCKW+Y +KY
Sbjct: 78 VCIAKAKEPSTYNEAKEFLVWCGAMDDEIGAMETTHTWEICTLPPNKKPIGCKWVYKIKY 137
Query: 1027 RSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKN 1086
SDGT++RYKARLVAKGYTQ GID+ ETF+PV K+ +V++IL+++A +++ L Q D+ N
Sbjct: 138 NSDGTIERYKARLVAKGYTQQEGIDFIETFSPVCKLTSVKLILAISAIYNFTLHQLDISN 197
Query: 1087 AFLHGDLEEEVYMEIPPGYDPTSGR----NKVCKLKKALYGLKQSPRAWFGRFTNAMVSL 1142
AFL+GDL+EE+YM++PPGY G N VC LKK++YGLKQ+ R WF +F+ ++
Sbjct: 198 AFLNGDLDEEIYMKLPPGYAARQGDSLPPNAVCYLKKSIYGLKQASRQWFLKFSVTLIGF 257
Query: 1143 GYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKL 1202
G+ QS DHT F+K T L +L VYV+D+II N++ L+ +L F+++DLG L
Sbjct: 258 GFVQSHSDHTYFLKITATLFLCVL-VYVDDIIICSNNDAAVDELKSQLKSCFKLRDLGPL 316
Query: 1203 KYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKA 1262
KYFLG+E+A S GI I QRKY LDLL ETG LGCKP VP++ + + V
Sbjct: 317 KYFLGLEIARSAAGINICQRKYALDLLDETGLLGCKPSSVPMDPSVTFSAHSGGDFVDAK 376
Query: 1263 QYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKK 1322
Y+RL+G+L+YL TR DI++ V+ +SQF P+ H QAV +IL Y+K + G+GL +
Sbjct: 377 AYRRLIGRLMYLQITRLDISFAVNKLSQFSEAPRLAHQQAVMKILHYIKGTVGQGLFYSS 436
Query: 1323 SEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQ 1382
++ ++V++DA + RRST GYCMFLG +L++W+SKKQ VV++SSAEAE+RA++
Sbjct: 437 QAEMQLQVFSDASFQSCKDTRRSTNGYCMFLGTSLISWKSKKQQVVSKSSAEAEYRALSF 496
Query: 1383 GVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEK 1440
E++ + +L++ P LFCDN +AI IA N V H+RTKHIE D H ++E+
Sbjct: 497 ATDEMMWLAQFFRELQLPLSKPTLLFCDNTAAIHIATNAVFHERTKHIESDCHSVRER 554
>At1g47650 hypothetical protein
Length = 1409
Score = 452 bits (1163), Expect = e-127
Identities = 230/496 (46%), Positives = 342/496 (68%), Gaps = 12/496 (2%)
Query: 991 MNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGI 1050
++ E+ A++ T +I P K+ VGCKW++++K+ +DG+L+RYKARLVAKGYTQ G+
Sbjct: 747 VSSEIGAMETTHTWEITTLPAGKKAVGCKWVFSLKFLADGSLERYKARLVAKGYTQKEGL 806
Query: 1051 DYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSG 1110
DY +TF+PVAKM T++++L ++A W L+Q DV NAFL+G+LEEE+YM++P GY G
Sbjct: 807 DYTDTFSPVAKMTTIKLLLKISASKKWFLKQLDVSNAFLNGELEEEIYMKLPEGYAARKG 866
Query: 1111 ----RNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLL 1166
N VC+LK+++YGLKQ+ R WF +F+ +++ LG+K++ GDHTLFIK +G+ ++
Sbjct: 867 ITLPPNSVCRLKRSIYGLKQASRQWFKKFSASLLDLGFKKTHGDHTLFIKEY-DGEFVIV 925
Query: 1167 LVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVL 1226
LVYV+D+ IA E L ++L +F+++DLG LKYFLG+E+A ++ GI I QRKY L
Sbjct: 926 LVYVDDIAIASTSEGAAIQLTQDLQQRFKLRDLGDLKYFLGLEIARTEAGISICQRKYAL 985
Query: 1227 DLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVS 1286
+LL TG + CKP+ VP+ N K+ ++ DL + QY+R+VGKL+YL+ TR DI + V+
Sbjct: 986 ELLASTGMVNCKPVSVPMIPNVKMMKTDGDLLDDREQYRRIVGKLMYLTITRLDITFAVN 1045
Query: 1287 VVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRST 1346
+ QF P+ HLQA R+LQY+K S G+GL + S L+++ + D+D+A RRST
Sbjct: 1046 KLCQFSSAPRTSHLQAAYRVLQYIKGSVGQGLFYSASADLTLKGFADSDWASCPDSRRST 1105
Query: 1347 TGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMR 1406
TG+ MF+G +L++WRSKKQ+VV+RSSAEAE+RA+A CEL+ + +L LK Y P+
Sbjct: 1106 TGFSMFVGDSLISWRSKKQHVVSRSSAEAEYRALALVTCELVWLHTLLMSLKASYLVPV- 1164
Query: 1407 LFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADV------ 1460
L+ D+ +AI IA NPV H+RTKHIE+D H ++EKL++ + +V + Q +
Sbjct: 1165 LYSDSTTAIYIATNPVFHERTKHIELDCHTVREKLDNGELKLLHVRTEDQPSTAAYPPPN 1224
Query: 1461 LTKGLPLERFRELTCK 1476
L + LP R T K
Sbjct: 1225 LAEPLPCPRCNSTTTK 1240
Score = 234 bits (596), Expect = 4e-61
Identities = 139/419 (33%), Positives = 212/419 (50%), Gaps = 30/419 (7%)
Query: 388 IIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVP 447
+ID GA H++ S + T + + G +I+G G + L+ + LK +L +P
Sbjct: 333 VIDSGAIHHVSHDKSLFLNLDTSVVSA-VNLPAGPTVRISGVGTLRLNDDILLKNILFIP 391
Query: 448 KLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQT 507
+ NL+SI LT D+ V F + QD +++G + LY L E+
Sbjct: 392 EFRLNLISISSLTDDIGSRVIFDKTSCEIQDPIKARMLGQGRRIANLYVLDIEDPIVSVN 451
Query: 508 SSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVE--SFHCDVCQFSKHHRSTFLP 565
+ I + H+RLGH L + L + S +C VC +K + F
Sbjct: 452 A-----VVDISMWHRRLGHASLQRLDVISDSLGTTKPKNKGSDYCHVCHLAKQRKLPFPS 506
Query: 566 SNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANF 625
N C + FDL+H D+WGP S+ V G K+F+T +DD +R TW++L+++KSEV +F F
Sbjct: 507 QNKVCNEIFDLLHIDIWGPFSVETVDGYKYFLTIVDDHSRATWIYLLRNKSEVLTVFPAF 566
Query: 626 FHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHL 685
+ Q+ +K +RSDN E F+ + G+V +C + P+QN V ERK++H+
Sbjct: 567 IEQVENQYKVRVKAVRSDNAPE---LKFTSLYQQKGIVSFHSCPETPEQNLVVERKHQHI 623
Query: 686 LEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQ 745
L ++RAL+FQ VP WG+ VLTA +LINR PS++LSN +P ++T P K
Sbjct: 624 LNVSRALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLSNKTPYEILTGTVPFYFAKK--- 680
Query: 746 SRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQE 804
R K PR+ C+F+GY + KGYK S V+IS +V F E
Sbjct: 681 ----------------RHKFQPRSKACVFLGYPAGYKGYKLMDLESHTVFISRNVVFHE 723
Score = 105 bits (262), Expect = 2e-22
Identities = 61/194 (31%), Positives = 107/194 (54%), Gaps = 4/194 (2%)
Query: 36 SFRLDGRNYLQWSELVRHTLISRQKISYIEETAPA--ETDPQYDTWREENSLIMTWFWYS 93
S RLD NY W+ + +L ++ K +I+ T P E+D + W S++ +W S
Sbjct: 79 SHRLDETNYGDWNVAMLISLDAKNKSGFIDGTLPRPLESDKNFRLWSRCKSMVKSWLLNS 138
Query: 94 MIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLN 153
+ P+I R+ + A +IW +L ++ +L Y L ++ + +QGTLS+++YY L
Sbjct: 139 VSPQIYRSILRMNDASDIWRDLHSRFNVT-NLPRTYNLTQEIQDFRQGTLSLSEYYTRLK 197
Query: 154 GLWIELDQYQNLKMKCD-SDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGKEQFPSL 212
LW +L+ + L C + L + E+A+I K L+GLN + IR QI+ K+ PSL
Sbjct: 198 TLWDQLESTEELDDPCICGKAMRLQQKAEQAKIVKLLAGLNDSYAIIRRQIIAKKALPSL 257
Query: 213 SEVFSIVRGEETRR 226
+EV+ I+ + +++
Sbjct: 258 AEVYHILDQDNSQQ 271
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,479,944
Number of Sequences: 26719
Number of extensions: 1641102
Number of successful extensions: 5758
Number of sequences better than 10.0: 169
Number of HSP's better than 10.0 without gapping: 137
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 4805
Number of HSP's gapped (non-prelim): 426
length of query: 1485
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1373
effective length of database: 8,326,068
effective search space: 11431691364
effective search space used: 11431691364
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0310b.12


