

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0304.15
(697 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
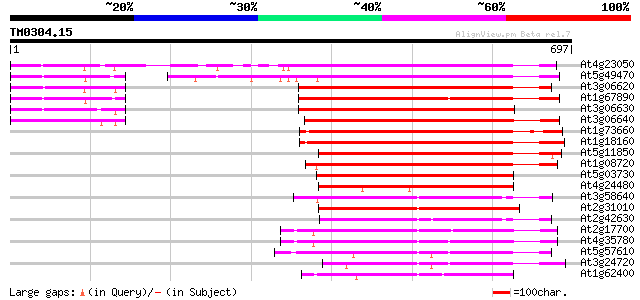
Score E
Sequences producing significant alignments: (bits) Value
At4g23050 putative serine/threonine kinase 511 e-145
At5g49470 protein kinase like protein 364 e-100
At3g06620 protein kinase, putative 358 4e-99
At1g67890 putative protein kinase (At1g67890) 358 7e-99
At3g06630 protein kinase, putative 348 8e-96
At3g06640 protein kinase, putative 342 3e-94
At1g73660 putative protein kinase 330 2e-90
At1g18160 MAP kinase, putative 325 5e-89
At5g11850 MAP3K like protein kinase 315 4e-86
At1g08720 MAP kinase kinase kinase EDR1 314 9e-86
At5g03730 SERINE/THREONINE-PROTEIN KINASE CTR1 308 5e-84
At4g24480 putative protein kinase 283 2e-76
At3g58640 unknown protein 226 3e-59
At2g31010 putative protein kinase 214 1e-55
At2g42630 putative protein kinase 201 9e-52
At2g17700 putative protein kinase 196 5e-50
At4g35780 unknown protein 189 5e-48
At5g57610 putative protein 179 5e-45
At3g24720 protein kinase, putative 170 2e-42
At1g62400 hypothetical protein 167 2e-41
>At4g23050 putative serine/threonine kinase
Length = 736
Score = 511 bits (1317), Expect = e-145
Identities = 311/711 (43%), Positives = 414/711 (57%), Gaps = 133/711 (18%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
+ SAE LY W E++G R L+TEEY L I R + GETW+G+FP++KK GE+
Sbjct: 117 SRSAENLYHWYAEEVVGYRTIDVLVTEEYRNSLTGIRNR-VCRGETWTGQFPFQKKTGEL 175
Query: 61 FMVMVTKAPLYEDGVLVGVISVSSEAAKFNT---IDSEFQRTCQSGASGQPGVQKLNSKG 117
FM +VTK+P+YE+G LVGV++VSS+A FN + +E Q+ +S + ++K
Sbjct: 176 FMALVTKSPVYENGELVGVVTVSSDATLFNRMHPLSNEHQQQARSNNRHESNLRK----- 230
Query: 118 NQWPL-RPLIAP------MPQIASSV-SNL-ASKILPNRHMDDTVSRNTSTDANDEKLEK 168
+QW L RP IA +PQ +S+V SNL ASK+LP R+ DD+ + N ++ + DE +
Sbjct: 231 HQWHLPRPQIAAASQVPVVPQYSSAVASNLKASKLLPQRNGDDSFNGNHNSRSRDENVPV 290
Query: 169 CSIYETKLHSRHHHKEQNATIREASEKDESTTEFGRASKIAARFMAKLQIGGTGKWG-KD 227
+ +T F + +A +F+ KLQ TG G +D
Sbjct: 291 VA----------------------------STTFEKYGSLADKFLGKLQRKITGSQGTED 322
Query: 228 TQSVKTNCRVDNPESDGVNNGNHSSGGSA------ALTSHQDVANGVDREENLHKCDSLF 281
+ + N G+N SGGS+ T+ +D NG + + D
Sbjct: 323 NEPILRN---------GINKSACGSGGSSKASNAVTCTAFRDNGNGKPKRAEVRISD--- 370
Query: 282 SMKTAHTNVYARRTSGVFEVSSAKPFSRECYECLGSLVPQDPLLRLHRQFDPKKLEP--- 338
VY G+ + + ++ +G+L P P+ LE
Sbjct: 371 --------VYGNGAEGLIH-------NGDRFQYIGNLGQSKP---------PRGLESGLV 406
Query: 339 ---EATHMA-----IEDEVQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVD--CEIHWE 388
T M+ IED + LP + G +S ++ +N V D CEI WE
Sbjct: 407 SGMRGTKMSDLNGEIEDAWNTRLSVDPLPILGVNSGRQQSPVNQRNNRLVTDSSCEIRWE 466
Query: 389 DLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVL 448
DL L +E+G+GS+A V+ G+WNGSDVAIKVYF Y TL + KKEI+IMK+LRHPNVL
Sbjct: 467 DLQLGEEVGRGSFAAVHRGVWNGSDVAIKVYFDGDYNAMTLTECKKEINIMKKLRHPNVL 526
Query: 449 LFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPP 508
LFMGAV + E+ AI+ E +PRGSLFK LH +NQ LD +RRLRMALD+ARGMNYLH RNPP
Sbjct: 527 LFMGAVCTEEKSAIIMEYMPRGSLFKILHNTNQPLDKKRRLRMALDVARGMNYLHRRNPP 586
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEK 568
IVHRDLKSSNLLVDKNW VKVGDFGLS+ K+ T L+TKSG+GTPQWMAPE+LR+EPSNEK
Sbjct: 587 IVHRDLKSSNLLVDKNWNVKVGDFGLSKWKNATFLSTKSGKGTPQWMAPEVLRSEPSNEK 646
Query: 569 SDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRC 628
DV+S+GV+LWELMT +PW+ LNS+QVVGVVGFMDRRLDLPEGL+P +ASII DCW+
Sbjct: 647 CDVFSFGVILWELMTTLVPWDRLNSIQVVGVVGFMDRRLDLPEGLNPRIASIIQDCWQ-- 704
Query: 629 FYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNR 679
+DP +RPSFEELI +M+ + +
Sbjct: 705 -----------------------------TDPAKRPSFEELISQMMSLFRK 726
>At5g49470 protein kinase like protein
Length = 770
Score = 364 bits (934), Expect = e-100
Identities = 226/558 (40%), Positives = 299/558 (53%), Gaps = 104/558 (18%)
Query: 197 ESTTEFGRA-SKIAARFMAKLQIGGTGKWGKDTQ--------------SVKTNCRVDNPE 241
E T F R S +A++ + K G D+Q + K +V +
Sbjct: 229 EGETSFSRVTSSVASKLGLDSKEAVVSKLGLDSQQPIQVAIASKISDLASKVGNKVKSKM 288
Query: 242 SDGVNNGNHSSGGSAALTSHQDVANGVDREENLHKCDSLFSMKTAHTNVYARRTSGVF-- 299
G NN + GGS SHQ D + D+ S + + GVF
Sbjct: 289 RAGDNNAANLEGGSG--DSHQSDQGFFDAAFADRREDAATSGADTPRGDFIQSPFGVFLR 346
Query: 300 --EVSSAKPFSRECYECLGSLVPQDPLLRLHRQFDPKK-------------LEPEATHMA 344
E +S KPF E G+ V L ++ KK LE +H
Sbjct: 347 SDEKASTKPFRDSSDESDGNSVVPKTLTSKAEEWMVKKGLSWPWKGNEREGLEGRRSHSV 406
Query: 345 ---IEDEVQKQQGR-------------------------LQLPSSRESVGSHESSSSKGD 376
+ +E QKQQ + SS +V S SSSS G
Sbjct: 407 WPWVRNEQQKQQAYQSNSNHSVKSESQACESIKASSNEPMGYWSSSVNVNSTSSSSSCGS 466
Query: 377 NDSVV-----------DCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYT 425
S V D EI WEDL + ++IGQGS VYHG+W GSDVA+KV+ Y+
Sbjct: 467 TSSSVMNKVDMDSDCLDYEILWEDLTIGEQIGQGSCGTVYHGLWFGSDVAVKVFSKQEYS 526
Query: 426 EETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDI 485
EE + +++E+ +MKRLRHPNVLLFMGAV S +RL IVTE LPRGSLF+ L ++ LD
Sbjct: 527 EEIITSFRQEVSLMKRLRHPNVLLFMGAVTSPQRLCIVTEFLPRGSLFRLLQRNTSKLDW 586
Query: 486 RRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTT 545
RRR+ MA DIARGMNYLHH PPI+HRDLKSSNLLVDKNWTVKV DFGLSR+K T LTT
Sbjct: 587 RRRIHMASDIARGMNYLHHCTPPIIHRDLKSSNLLVDKNWTVKVADFGLSRIKHETYLTT 646
Query: 546 KSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDR 605
K+GRGTPQWMAPE+LRNE ++EKSDVYS+GV+LWEL+T+ IPWE+LN++QV+G VGFM++
Sbjct: 647 KTGRGTPQWMAPEVLRNEAADEKSDVYSFGVILWELVTEKIPWESLNAMQVIGAVGFMNQ 706
Query: 606 RLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPS 665
RL++P+ +DP S++ CW S+P+ RPS
Sbjct: 707 RLEVPKNVDPQWISLMESCWH-------------------------------SEPQDRPS 735
Query: 666 FEELIQRMLFVLNRVTAE 683
F+E+++++ + + T +
Sbjct: 736 FQEIMEKLRELQRKYTIQ 753
Score = 70.9 bits (172), Expect = 2e-12
Identities = 52/159 (32%), Positives = 81/159 (50%), Gaps = 20/159 (12%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE LYG+ E +GK L+ + + Q I R ++GE+W+G+FP K K GE
Sbjct: 126 NSMAEKLYGFSASEALGKDPIDILVDVQDASVAQNITRRC-SSGESWTGEFPVKNKAGER 184
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNTI-------------DSEFQRTCQSGAS- 105
F V+ T +P Y +DG L+G+I +++++A F ++ F R S AS
Sbjct: 185 FSVVTTMSPSYDDDGCLIGIICITNDSALFQDPRGSPAKTRRGQEGETSFSRVTSSVASK 244
Query: 106 -GQPGVQKLNSKGNQWPLRPLIAPMPQIASSVSNLASKI 143
G + + SK +P+ IAS +S+LASK+
Sbjct: 245 LGLDSKEAVVSKLGLDSQQPI---QVAIASKISDLASKV 280
>At3g06620 protein kinase, putative
Length = 773
Score = 358 bits (920), Expect = 4e-99
Identities = 179/316 (56%), Positives = 227/316 (71%), Gaps = 33/316 (10%)
Query: 360 SSRESVGSHESS-SSKGDNDSV-VDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
SS S GS SS +K D DS ++ EI W+DL + +++GQGS VYHG+W GSDVA+K
Sbjct: 462 SSASSCGSTSSSVMNKVDTDSEGLEYEILWDDLTIGEQVGQGSCGTVYHGLWFGSDVAVK 521
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+ E ++ +K+E+ +MKRLRHPNVLLFMGAV S +RL IV+E LPRGSLF+ L
Sbjct: 522 VFSKQEYSAEVIESFKQEVLLMKRLRHPNVLLFMGAVTSPQRLCIVSEFLPRGSLFRLLQ 581
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
KS LD RRR+ MALDIARGMNYLHH +PPI+HRDLKSSNLLVDKNWTVKV DFGLSR+
Sbjct: 582 KSTSKLDWRRRIHMALDIARGMNYLHHCSPPIIHRDLKSSNLLVDKNWTVKVADFGLSRI 641
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LT+KSG+GTPQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWE LNS+QV+
Sbjct: 642 KHETYLTSKSGKGTPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWETLNSMQVI 701
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMS 657
G VGFMD+RL++P+ +DP S++ CW
Sbjct: 702 GAVGFMDQRLEIPKDIDPRWISLMESCWH------------------------------- 730
Query: 658 SDPEQRPSFEELIQRM 673
SD + RP+F+EL+ ++
Sbjct: 731 SDTKLRPTFQELMDKL 746
Score = 58.2 bits (139), Expect = 2e-08
Identities = 44/156 (28%), Positives = 72/156 (45%), Gaps = 14/156 (8%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE +YG+ E +G+ + + A I R + GE+W+G+FP K K G+
Sbjct: 129 NAMAEKVYGYSAAEALGENPINVIADDRDAAFAMNIARRCVR-GESWTGEFPVKSKSGDR 187
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNT---------IDSEFQRTCQSGASGQPGV 110
F + T +P Y +DG L+G+I ++S A + E + + +
Sbjct: 188 FSAVTTCSPFYDDDGALMGIICITSNTAPYLNPRISLAKLKAQEEGETSSIPARNSFASK 247
Query: 111 QKLNSKGNQWPLRPLIAPMP---QIASSVSNLASKI 143
L+S+G L + P IAS +S+LASK+
Sbjct: 248 LGLDSRGAVISKLGLDSDQPIQVAIASKISDLASKV 283
>At1g67890 putative protein kinase (At1g67890)
Length = 765
Score = 358 bits (918), Expect = 7e-99
Identities = 181/326 (55%), Positives = 233/326 (70%), Gaps = 34/326 (10%)
Query: 360 SSRESVGSHESS-SSKGDNDS-VVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
SS S GS SS +K D DS +D EI WEDL + ++IGQGS VYHG+W GSDVA+K
Sbjct: 455 SSASSCGSTSSSVMNKVDMDSDCLDYEILWEDLTIGEQIGQGSCGTVYHGLWFGSDVAVK 514
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+EE + +K+E+ +MKRLRHPNVLLFMGAV S +RL IVTE LPRGSLF+ L
Sbjct: 515 VFSKQEYSEEIITSFKQEVSLMKRLRHPNVLLFMGAVASPQRLCIVTEFLPRGSLFRLLQ 574
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
++ LD+RRR+ MA DIARGMNYLHH +PPI+HRDLKSSNLLVD+NWTVKV DFGLSR+
Sbjct: 575 RNKSKLDLRRRIHMASDIARGMNYLHHCSPPIIHRDLKSSNLLVDRNWTVKVADFGLSRI 634
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LTT +GRGTPQWMAPE+LRNE ++EKSDVYS+GVVLWEL+T+ IPWENLN++QV+
Sbjct: 635 KHETYLTT-NGRGTPQWMAPEVLRNEAADEKSDVYSFGVVLWELVTEKIPWENLNAMQVI 693
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMS 657
G VGFM++RL++P+ +DP +++ CW
Sbjct: 694 GAVGFMNQRLEVPKDVDPQWIALMESCWH------------------------------- 722
Query: 658 SDPEQRPSFEELIQRMLFVLNRVTAE 683
S+P+ RPSF+EL+ ++ + + T +
Sbjct: 723 SEPQCRPSFQELMDKLRELQRKYTIQ 748
Score = 72.8 bits (177), Expect = 6e-13
Identities = 54/158 (34%), Positives = 83/158 (52%), Gaps = 19/158 (12%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE LYG+ E +GK L+ + A + I +R ++GE+W+G+FP K K GE
Sbjct: 127 NSMAEKLYGFSAAEALGKDSINILVDGQDAAVAKNIFQRC-SSGESWTGEFPVKNKMGER 185
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNTI------------DSEFQRTCQSGAS-- 105
F V+ T +P Y +DG+L+G+I +++++A F DS F R AS
Sbjct: 186 FSVVTTISPFYDDDGLLIGIICITNDSALFQRPRVPPAKNRWQEGDSSFCRGTNGVASRL 245
Query: 106 GQPGVQKLNSKGNQWPLRPLIAPMPQIASSVSNLASKI 143
G + + SK +P+ A IAS +S+LASK+
Sbjct: 246 GFDSKEAVVSKLGLDSQQPIQA---AIASKISDLASKV 280
>At3g06630 protein kinase, putative
Length = 671
Score = 348 bits (892), Expect = 8e-96
Identities = 169/270 (62%), Positives = 212/270 (77%), Gaps = 2/270 (0%)
Query: 360 SSRESVGSHESS-SSKGDNDS-VVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
+S S GS S K D DS ++ EI W+DL + ++IG+GS VYHGIW GSDVA+K
Sbjct: 402 NSASSCGSTSRSVMDKVDIDSDPLEHEILWDDLTIGEQIGRGSCGTVYHGIWFGSDVAVK 461
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+E ++ ++KE+ +MKRLRHPNVLLFMGAV S +RL IV+E +PRGSLF+ L
Sbjct: 462 VFSKQEYSESVIKSFEKEVSLMKRLRHPNVLLFMGAVTSPQRLCIVSEFVPRGSLFRLLQ 521
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
+S LD RRR+ MALDIARGMNYLH +PPI+HRDLKSSNLLVD+NWTVKV DFGLSR+
Sbjct: 522 RSMSKLDWRRRINMALDIARGMNYLHCCSPPIIHRDLKSSNLLVDRNWTVKVADFGLSRI 581
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LT+KSG+GTPQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWENLNS+QV+
Sbjct: 582 KHQTYLTSKSGKGTPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWENLNSMQVI 641
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
G VGFM++RL++P+ DP S+I CW R
Sbjct: 642 GAVGFMNQRLEIPKDTDPDWISLIESCWHR 671
Score = 69.7 bits (169), Expect = 5e-12
Identities = 46/147 (31%), Positives = 76/147 (51%), Gaps = 8/147 (5%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N +E LYG+ E++G+ ++ ++ A + R GE+W+G+FP K K G++
Sbjct: 90 NAMSEKLYGYSAAEVVGRNPVHVIVDDQNAAFALNVARRC-ANGESWTGEFPVKTKSGKI 148
Query: 61 FMVMVTKAPLYED-GVLVGVISVSSEAAKFNTIDSEFQRTCQSGASGQPGVQKLNSKGNQ 119
F + T +P Y+D G +VG+IS++S+ A + R + G L+SKG
Sbjct: 149 FSAVTTCSPFYDDNGTVVGIISITSDIAPYLNPRLSLPRLKPQEPERKLG---LDSKGAV 205
Query: 120 WPLRPLIAPMP---QIASSVSNLASKI 143
L + P IAS +S+LASK+
Sbjct: 206 ISKPGLDSDQPIQVSIASKISSLASKL 232
>At3g06640 protein kinase, putative
Length = 763
Score = 342 bits (878), Expect = 3e-94
Identities = 167/317 (52%), Positives = 222/317 (69%), Gaps = 31/317 (9%)
Query: 367 SHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTE 426
SH + + N + ++ EI W+DL + ++IGQGS VYHG+W GSDVA+K+ Y+E
Sbjct: 423 SHSTMNKVDTNSNCLEYEILWDDLTIGEQIGQGSCGTVYHGLWFGSDVAVKLISKQEYSE 482
Query: 427 ETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIR 486
E +Q +++E+ +M+RLRHPNVLLFMGAV + L IV+E LPRGSLF+ L ++ LD R
Sbjct: 483 EVIQSFRQEVSLMQRLRHPNVLLFMGAVTLPQGLCIVSEFLPRGSLFRLLQRNMSKLDWR 542
Query: 487 RRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTK 546
RR+ MALDIARGMNYLH +PPI+HRDLKSSNLLVDKN TVKV DFGLSR+K T LT+K
Sbjct: 543 RRINMALDIARGMNYLHRCSPPIIHRDLKSSNLLVDKNLTVKVADFGLSRIKHHTYLTSK 602
Query: 547 SGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRR 606
SG+G PQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWENLNS+QV+G VGFM++R
Sbjct: 603 SGKGMPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWENLNSMQVIGAVGFMNQR 662
Query: 607 LDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSF 666
L++P+ +DP S+I CW R D + RP+F
Sbjct: 663 LEIPKDIDPDWISLIESCWHR-------------------------------DAKLRPTF 691
Query: 667 EELIQRMLFVLNRVTAE 683
+EL++R+ + + T +
Sbjct: 692 QELMERLRDLQRKYTIQ 708
Score = 72.0 bits (175), Expect = 1e-12
Identities = 48/155 (30%), Positives = 81/155 (51%), Gaps = 13/155 (8%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE +YG+ E +G+ ++ ++ AP + +L + GE+W+GKFP K++ GE
Sbjct: 90 NAMAEKVYGYSAAEAVGQNPIDVMV-DDRDAPFAMTIAQLCSNGESWTGKFPVKRRTGEK 148
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKF--NTIDSEFQRTCQSGASGQPGVQK----- 112
F + T +P Y +DG L+G++S++S+ A + TI + + S P
Sbjct: 149 FSAVTTCSPFYADDGSLIGIVSITSDVAPYLNPTISLAKLKASEVETSSTPARNSFAFKL 208
Query: 113 -LNSKGNQWPLRPLIAPMP---QIASSVSNLASKI 143
L++KG L + P IAS +S+LASK+
Sbjct: 209 GLDTKGAVVSKLGLDSDQPIQVAIASKISDLASKV 243
>At1g73660 putative protein kinase
Length = 1030
Score = 330 bits (846), Expect = 2e-90
Identities = 169/327 (51%), Positives = 218/327 (65%), Gaps = 34/327 (10%)
Query: 361 SRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYF 420
S +S+G+ SSK D D V DCEI WE++ + + IG GSY VY G W+G++VA+K +
Sbjct: 722 SDKSIGNE---SSKSDCDDVSDCEILWEEITVGERIGLGSYGEVYRGDWHGTEVAVKKFL 778
Query: 421 GNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN 480
T E L++++ E+ IMK+LRHPN++LFMGAV L+IVTE LPRGSL++ +H+ N
Sbjct: 779 DQDLTGEALEEFRSEVRIMKKLRHPNIVLFMGAVTRPPNLSIVTEFLPRGSLYRLIHRPN 838
Query: 481 QTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDT 540
LD RRRLRMALD ARGMNYLH NP IVHRDLKS NLLVDKNW VKV DFGLSR+K +
Sbjct: 839 NQLDERRRLRMALDAARGMNYLHSCNPMIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKHS 898
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVV 600
T L++KS GT +WMAPE+LRNEP++EK DVYSYGV+LWEL T PW +N +QVVG V
Sbjct: 899 TYLSSKSTAGTAEWMAPEVLRNEPADEKCDVYSYGVILWELFTLQQPWGKMNPMQVVGAV 958
Query: 601 GFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDP 660
GF RRLD+P+ +DP +A +I+ CW QT+S L
Sbjct: 959 GFQHRRLDIPDFVDPAIADLISKCW---------------------QTDSKL-------- 989
Query: 661 EQRPSFEELIQRMLFVLNRVTAESVKR 687
RPSF E++ + + VT ++ R
Sbjct: 990 --RPSFAEIMASLKRLQKPVTGSNIPR 1014
>At1g18160 MAP kinase, putative
Length = 992
Score = 325 bits (833), Expect = 5e-89
Identities = 166/329 (50%), Positives = 214/329 (64%), Gaps = 32/329 (9%)
Query: 361 SRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYF 420
S S+G+ ESS S D V +CEI WE++ + + IG GSY VY G W+G+ VA+K +
Sbjct: 687 SDRSIGN-ESSKSDAAIDDVAECEILWEEITVAERIGLGSYGEVYRGDWHGTAVAVKKFI 745
Query: 421 GNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN 480
T E L++++ E+ +M+RLRHPN++LFMGAV L+IVTE LPRGSL++ +H+ N
Sbjct: 746 DQDITGEALEEFRSEVRMMRRLRHPNIVLFMGAVTRPPNLSIVTEFLPRGSLYRLIHRPN 805
Query: 481 QTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDT 540
LD R+RLRMALD ARGMNYLH NP IVHRDLKS NLLVDKNW VKV DFGLSR+K +
Sbjct: 806 NQLDERKRLRMALDAARGMNYLHSCNPVIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKVS 865
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVV 600
T L++KS GT +WMAPE+LRNEP++EK DVYSYGV+LWEL T PW +N +QVVG V
Sbjct: 866 TYLSSKSTAGTAEWMAPEVLRNEPADEKCDVYSYGVILWELFTLQQPWGKMNPMQVVGAV 925
Query: 601 GFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDP 660
GF RRLD+PE +DP +A II CW+ +DP
Sbjct: 926 GFQHRRLDIPEFVDPGIADIIRKCWQ-------------------------------TDP 954
Query: 661 EQRPSFEELIQRMLFVLNRVTAESVKRSS 689
RPSF E++ + + + +V SS
Sbjct: 955 RLRPSFGEIMDSLKQLQKPIQRAAVPSSS 983
>At5g11850 MAP3K like protein kinase
Length = 880
Score = 315 bits (808), Expect = 4e-86
Identities = 158/308 (51%), Positives = 207/308 (66%), Gaps = 37/308 (12%)
Query: 384 EIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLR 443
EI WEDL + + IG GSY VY WNG++VA+K + ++ + L +K EI+IM RLR
Sbjct: 603 EIMWEDLQIGERIGIGSYGEVYRAEWNGTEVAVKKFLDQDFSGDALTQFKSEIEIMLRLR 662
Query: 444 HPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLH 503
HPNV+LFMGAV +I+TE LPRGSL++ LH+ N LD +RR+RMALD+A+GMNYLH
Sbjct: 663 HPNVVLFMGAVTRPPNFSILTEFLPRGSLYRLLHRPNHQLDEKRRMRMALDVAKGMNYLH 722
Query: 504 HRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNE 563
+P +VHRDLKS NLLVDKNW VKV DFGLSR+K T L++KS GTP+WMAPE+LRNE
Sbjct: 723 TSHPTVVHRDLKSPNLLVDKNWVVKVCDFGLSRMKHHTYLSSKSTAGTPEWMAPEVLRNE 782
Query: 564 PSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIIND 623
P+NEK DVYS+GV+LWEL T +PW+ LN +QVVG VGF +RRL++P+ +D VA II +
Sbjct: 783 PANEKCDVYSFGVILWELATSRVPWKGLNPMQVVGAVGFQNRRLEIPDDIDLTVAQIIRE 842
Query: 624 CWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRM-----LFVLN 678
CW+ ++P RPSF +L+Q + L + N
Sbjct: 843 CWQ-------------------------------TEPHLRPSFTQLMQSLKRLQGLNISN 871
Query: 679 RV-TAESV 685
R T+ES+
Sbjct: 872 RANTSESL 879
>At1g08720 MAP kinase kinase kinase EDR1
Length = 933
Score = 314 bits (805), Expect = 9e-86
Identities = 161/325 (49%), Positives = 209/325 (63%), Gaps = 44/325 (13%)
Query: 368 HESSSSKGDNDS------------VVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVA 415
HES+SS D+ S V +CEI W DL + + IG GSY VYH W+G++VA
Sbjct: 635 HESTSSSLDSTSYRNDPQVLDDADVGECEIPWNDLVIAERIGLGSYGEVYHADWHGTEVA 694
Query: 416 IKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKT 475
+K + ++ L +++ E+ IM+RLRHPNV+ F+GAV L+IVTE LPRGSL++
Sbjct: 695 VKKFLDQDFSGAALAEFRSEVRIMRRLRHPNVVFFLGAVTRPPNLSIVTEFLPRGSLYRI 754
Query: 476 LHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLS 535
LH+ +D RRR++MALD+A GMN LH P IVHRDLK+ NLLVD NW VKVGDFGLS
Sbjct: 755 LHRPKSHIDERRRIKMALDVAMGMNCLHTSTPTIVHRDLKTPNLLVDNNWNVKVGDFGLS 814
Query: 536 RLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQ 595
RLK T L++KS GTP+WMAPE+LRNEPSNEK DVYS+GV+LWEL T +PW +N +Q
Sbjct: 815 RLKHNTFLSSKSTAGTPEWMAPEVLRNEPSNEKCDVYSFGVILWELATLRLPWRGMNPMQ 874
Query: 596 VVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLN 655
VVG VGF +RRL++P+ LDP V II +CW+
Sbjct: 875 VVGAVGFQNRRLEIPKELDPVVGRIILECWQ----------------------------- 905
Query: 656 MSSDPEQRPSFEELIQRMLFVLNRV 680
+DP RPSF +L + +L LNR+
Sbjct: 906 --TDPNLRPSFAQLTE-VLKPLNRL 927
>At5g03730 SERINE/THREONINE-PROTEIN KINASE CTR1
Length = 821
Score = 308 bits (790), Expect = 5e-84
Identities = 148/246 (60%), Positives = 188/246 (76%), Gaps = 2/246 (0%)
Query: 382 DCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKR 441
D +I W DL+++++IG GS+ V+ W+GSDVA+K+ + E + ++ +E+ IMKR
Sbjct: 543 DMDIPWCDLNIKEKIGAGSFGTVHRAEWHGSDVAVKILMEQDFHAERVNEFLREVAIMKR 602
Query: 442 LRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN--QTLDIRRRLRMALDIARGM 499
LRHPN++LFMGAV L+IVTE L RGSL++ LHKS + LD RRRL MA D+A+GM
Sbjct: 603 LRHPNIVLFMGAVTQPPNLSIVTEYLSRGSLYRLLHKSGAREQLDERRRLSMAYDVAKGM 662
Query: 500 NYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEI 559
NYLH+RNPPIVHRDLKS NLLVDK +TVKV DFGLSRLK +T L++KS GTP+WMAPE+
Sbjct: 663 NYLHNRNPPIVHRDLKSPNLLVDKKYTVKVCDFGLSRLKASTFLSSKSAAGTPEWMAPEV 722
Query: 560 LRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVAS 619
LR+EPSNEKSDVYS+GV+LWEL T PW NLN QVV VGF +RL++P L+P VA+
Sbjct: 723 LRDEPSNEKSDVYSFGVILWELATLQQPWGNLNPAQVVAAVGFKCKRLEIPRNLNPQVAA 782
Query: 620 IINDCW 625
II CW
Sbjct: 783 IIEGCW 788
>At4g24480 putative protein kinase
Length = 963
Score = 283 bits (725), Expect = 2e-76
Identities = 138/256 (53%), Positives = 181/256 (69%), Gaps = 14/256 (5%)
Query: 384 EIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEI-----DI 438
E+ W +LH+++ +G GS+ V+ W+GSDVA+K+ + ++ +++ +E+ I
Sbjct: 663 EVSWNELHIKERVGAGSFGTVHRAEWHGSDVAVKILSIQDFHDDQFREFLREVCKQAVAI 722
Query: 439 MKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHK--SNQTLDIRRRLRMALDI- 495
MKR+RHPNV+LFMGAV RL+I+TE LPRGSLF+ +H+ S + LD RRRLRMALD+
Sbjct: 723 MKRVRHPNVVLFMGAVTERPRLSIITEYLPRGSLFRLIHRPASGELLDQRRRLRMALDVV 782
Query: 496 ------ARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGR 549
A+G+NYLH NPP+VH DLKS NLLVDKNWTVKV DFGLSR K T + +KS
Sbjct: 783 CAIPHYAKGLNYLHCLNPPVVHWDLKSPNLLVDKNWTVKVCDFGLSRFKANTFIPSKSVA 842
Query: 550 GTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDL 609
GTP+WMAPE LR EP+NEKSDVYS+GVVLWEL+T PW L+ QVVG V F +RRL +
Sbjct: 843 GTPEWMAPEFLRGEPTNEKSDVYSFGVVLWELITLQQPWNGLSPAQVVGAVAFQNRRLII 902
Query: 610 PEGLDPHVASIINDCW 625
P P + S++ CW
Sbjct: 903 PPNTSPVLVSLMEACW 918
>At3g58640 unknown protein
Length = 809
Score = 226 bits (577), Expect = 3e-59
Identities = 130/344 (37%), Positives = 187/344 (53%), Gaps = 59/344 (17%)
Query: 353 QGRLQLPSSRESVGS-HESSSSKGDNDSVV-------------------DCEIHWEDLHL 392
Q + LPSS ++ S +E S S N S + + I + +L +
Sbjct: 496 QKAISLPSSPQNYRSQYEQSGSSHRNISHIWDKVLGSPMFQNKPLLPYEEWNIDFSELTV 555
Query: 393 RDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMG 452
+G G + V+ GIWNG+DVAIKV+ T E ++D+ EI I+ RLRHPNV+LF+G
Sbjct: 556 GTRVGIGFFGEVFRGIWNGTDVAIKVFLEQDLTAENMEDFCNEISILSRLRHPNVILFLG 615
Query: 453 AVYSLERLAIVTELLPRGSLFKTLHKSNQT--LDIRRRLRMALDIARGMNYLHHRNPPIV 510
A RL+++TE + GSL+ LH S Q L RR+L+M DI RG+ +H IV
Sbjct: 616 ACTKPPRLSLITEYMEMGSLYYLLHLSGQKKRLSWRRKLKMLRDICRGLMCIHRMG--IV 673
Query: 511 HRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSD 570
HRD+KS+N L+ WTVK+ DFGLSR+ T + GTP+WMAPE++RNEP +EK D
Sbjct: 674 HRDIKSANCLLSNKWTVKICDFGLSRIMTGTTMRDTVSAGTPEWMAPELIRNEPFSEKCD 733
Query: 571 VYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFY 630
++S GV++WEL T + PWE + +VV + + RL++PEG + +I DCW
Sbjct: 734 IFSLGVIMWELCTLTRPWEGVPPERVVYAIAYEGARLEIPEG---PLGKLIADCW----- 785
Query: 631 MYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRML 674
++PEQRPS E++ R+L
Sbjct: 786 ---------------------------TEPEQRPSCNEILSRLL 802
>At2g31010 putative protein kinase
Length = 375
Score = 214 bits (546), Expect = 1e-55
Identities = 111/255 (43%), Positives = 160/255 (62%), Gaps = 7/255 (2%)
Query: 384 EIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLR 443
+I + +L + +G G + V+ G+WNG+DVAIK++ T E ++D+ EI I+ R+R
Sbjct: 50 DIDFSELTVGTRVGIGFFGEVFRGVWNGTDVAIKLFLEQDLTAENMEDFCNEISILSRVR 109
Query: 444 HPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQT--LDIRRRLRMALDIARGMNY 501
HPNV+LF+GA RL+++TE + GSL+ +H S Q L RRLRM DI RG+
Sbjct: 110 HPNVVLFLGACTKPPRLSMITEYMELGSLYYLIHMSGQKKKLSWHRRLRMLRDICRGLMC 169
Query: 502 LHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILR 561
+H IVHRDLKS+N LVDK+WTVK+ DFGLSR+ + S GTP+WMAPE++R
Sbjct: 170 IHRMK--IVHRDLKSANCLVDKHWTVKICDFGLSRIMTDENMKDTSSAGTPEWMAPELIR 227
Query: 562 NEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEG-LDPHVASI 620
N P EK D++S GV++WEL T PWE + +VV V RL++P+G L +A +
Sbjct: 228 NRPFTEKCDIFSLGVIMWELSTLRKPWEGVPPEKVVFAVAHEGSRLEIPDGPLSKLIAGL 287
Query: 621 I--NDCWRRCFYMYS 633
+ N + F ++S
Sbjct: 288 VLLNTMFLELFCLFS 302
>At2g42630 putative protein kinase
Length = 357
Score = 201 bits (512), Expect = 9e-52
Identities = 108/289 (37%), Positives = 165/289 (56%), Gaps = 38/289 (13%)
Query: 385 IHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRH 444
I + L + +G G+ VV G+WN ++VAIK++ G T E ++ + EI I+ RL+H
Sbjct: 99 IDFSKLKVGASVGSGTSGVVCRGVWNKTEVAIKIFLGQQLTAENMKVFCNEISILSRLQH 158
Query: 445 PNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHH 504
PNV+L +GA +L++VTE + GSL+ + + L +R+L++ +I RG+ Y+H
Sbjct: 159 PNVILLLGACTKPPQLSLVTEYMSTGSLYDVIRTRKKELSWQRKLKILAEICRGLMYIHK 218
Query: 505 RNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEP 564
IVHRDL S+N L++K+ VK+ DFGLSR T + GTP+WMAPE++RNEP
Sbjct: 219 MG--IVHRDLTSANCLLNKS-IVKICDFGLSRRMTGTAVKDTEAAGTPEWMAPELIRNEP 275
Query: 565 SNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDC 624
EKSD++S+GV++WEL T S PW+ + +V+ +V RL +PEG + +I DC
Sbjct: 276 VTEKSDIFSFGVIMWELSTLSKPWKGVPKEKVIHIVANEGARLKIPEG---PLQKLIADC 332
Query: 625 WRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRM 673
W S+PEQRPS +E++ R+
Sbjct: 333 W--------------------------------SEPEQRPSCKEILHRL 349
>At2g17700 putative protein kinase
Length = 546
Score = 196 bits (497), Expect = 5e-50
Identities = 116/354 (32%), Positives = 189/354 (52%), Gaps = 47/354 (13%)
Query: 337 EPEATHMAIEDEVQKQQGRLQLPSSRESVGSHESSSSKGD---------NDSVVDCEIHW 387
E + A+ E+ K + Q S ++S+ E S + D + EI
Sbjct: 226 ETDGLRDALSKEILKLKD--QPGSKQKSISFFEHDKSSNELIPACIEIPTDGTDEWEIDV 283
Query: 388 EDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNV 447
L + ++ GSY ++ G + +VAIK + E L+++ +E+ IM+++RH NV
Sbjct: 284 TQLKIEKKVASGSYGDLHRGTYCSQEVAIKFLKPDRVNNEMLREFSQEVFIMRKVRHKNV 343
Query: 448 LLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNP 507
+ F+GA L IVTE + RGS++ LHK ++ L++ALD+A+GM+YLH N
Sbjct: 344 VQFLGACTRSPTLCIVTEFMARGSIYDFLHKQKCAFKLQTLLKVALDVAKGMSYLHQNN- 402
Query: 508 PIVHRDLKSSNLLVDKNWTVKVGDFGLSRLK-DTTLLTTKSGRGTPQWMAPEILRNEPSN 566
I+HRDLK++NLL+D++ VKV DFG++R++ ++ ++T ++G T +WMAPE++ ++P N
Sbjct: 403 -IIHRDLKTANLLMDEHGLVKVADFGVARVQIESGVMTAETG--TYRWMAPEVIEHKPYN 459
Query: 567 EKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWR 626
K+DV+SY +VLWEL+T IP+ L LQ V R +P+ P V ++ CW
Sbjct: 460 HKADVFSYAIVLWELLTGDIPYAFLTPLQAAVGVVQKGLRPKIPKKTHPKVKGLLERCWH 519
Query: 627 RCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRV 680
+ DPEQRP FEE+I+ + ++ V
Sbjct: 520 Q-------------------------------DPEQRPLFEEIIEMLQQIMKEV 542
>At4g35780 unknown protein
Length = 570
Score = 189 bits (480), Expect = 5e-48
Identities = 111/353 (31%), Positives = 188/353 (52%), Gaps = 45/353 (12%)
Query: 337 EPEATHMAIEDEVQKQQGRLQLPSSRESVGSHESSSSKGD---------NDSVVDCEIHW 387
E E A++ E++K + Q S ++S+ E S + D + EI
Sbjct: 232 ETEGLKDALKKEIRKFKD--QPCSKQKSITFFEHDKSTNELLPACVEIPTDGTDEWEIDM 289
Query: 388 EDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNV 447
+ L + ++ GSY ++ G + +VAIK+ E L+++ +E+ IM+++RH NV
Sbjct: 290 KQLKIEKKVACGSYGELFRGTYCSQEVAIKILKPERVNAEMLREFSQEVYIMRKVRHKNV 349
Query: 448 LLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNP 507
+ F+GA L IVTE + RGS++ LHK I+ L++ALD+++GMNYLH N
Sbjct: 350 VQFIGACTRSPNLCIVTEFMTRGSIYDFLHKHKGVFKIQSLLKVALDVSKGMNYLHQNN- 408
Query: 508 PIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNE 567
I+HRDLK++NLL+D++ VKV DFG++R++ + + T + GT +WMAPE++ ++P +
Sbjct: 409 -IIHRDLKTANLLMDEHEVVKVADFGVARVQTESGVMT-AETGTYRWMAPEVIEHKPYDH 466
Query: 568 KSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
++DV+SY +VLWEL+T +P+ L LQ V R +P+ P + ++ CW++
Sbjct: 467 RADVFSYAIVLWELLTGELPYSYLTPLQAAVGVVQKGLRPKIPKETHPKLTELLEKCWQQ 526
Query: 628 CFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRV 680
DP RP+F E+I+ + ++ V
Sbjct: 527 -------------------------------DPALRPNFAEIIEMLNQLIREV 548
>At5g57610 putative protein
Length = 1054
Score = 179 bits (454), Expect = 5e-45
Identities = 121/358 (33%), Positives = 184/358 (50%), Gaps = 53/358 (14%)
Query: 330 QFDPKKLEPEATHMAIEDEVQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWED 389
Q P+ +E E+ HM +DE + + S + E + ++ + S I +D
Sbjct: 726 QVRPELVENESEHMNAQDEPE-----IDSDSDNPNNFKIEQTKAEAEAKSRGLQSIRNDD 780
Query: 390 LHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYT------EETLQDYKKEIDIMKRLR 443
L E+G G+Y VYHG W GSDVAIK + + E ++D+ KE ++ L
Sbjct: 781 LEEIRELGHGTYGSVYHGKWKGSDVAIKRIKASCFAGKPSERERLIEDFWKEALLLSSLH 840
Query: 444 HPNVLLFMGAVYSLE--RLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNY 501
HPNV+ F G V LA V E + GSL + L K ++T+D R+RL +A+D A GM Y
Sbjct: 841 HPNVVSFYGIVRDGPDGSLATVAEFMVNGSLKQFLQKKDRTIDRRKRLIIAMDTAFGMEY 900
Query: 502 LHHRNPPIVHRDLKSSNLLVD----KNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAP 557
LH +N IVH DLK NLLV+ + K+GD GLS++K TL++ RGT WMAP
Sbjct: 901 LHGKN--IVHFDLKCENLLVNMRDPQRPICKIGDLGLSKVKQKTLVSG-GVRGTLPWMAP 957
Query: 558 EILRNEPS--NEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDP 615
E+L + + +EK DVYS+G+V+WEL+T P+ +++ ++G + R +P+ DP
Sbjct: 958 ELLSGKSNMVSEKIDVYSFGIVMWELLTGEEPYADMHCASIIGGIVNNALRPKIPQWCDP 1017
Query: 616 HVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRM 673
++ CW +S+P +RPSF E+ Q++
Sbjct: 1018 EWKGLMESCW-------------------------------TSEPTERPSFTEISQKL 1044
>At3g24720 protein kinase, putative
Length = 297
Score = 170 bits (431), Expect = 2e-42
Identities = 112/316 (35%), Positives = 162/316 (50%), Gaps = 48/316 (15%)
Query: 389 DLHLRDEIGQGSYAVVYHGIWNGSDVAIK-----VYFGNGYTEETL-QDYKKEIDIMKRL 442
DL E+G G+Y VYHG W G+DVAIK + G +E L +D+ +E I+ L
Sbjct: 15 DLEDLTELGSGTYGTVYHGTWRGTDVAIKRIRNSCFAGRSSEQERLTKDFWREAQILSNL 74
Query: 443 RHPNVLLFMGAVYSLE--RLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMN 500
HPNV+ F G V LA VTE + GSL L K ++ LD R+++ +A+D A GM
Sbjct: 75 HHPNVVAFYGIVPDGTGGTLATVTEFMVNGSLRHALLKKDRLLDTRKKIIIAMDAAFGME 134
Query: 501 YLHHRNPPIVHRDLKSSNLLVD----KNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMA 556
YLH +N IVH DLK NLLV+ + KVGD GLSR+K TL++ RGT WMA
Sbjct: 135 YLHSKN--IVHFDLKCENLLVNLRDPQRPICKVGDLGLSRIKRNTLVSG-GVRGTLPWMA 191
Query: 557 PEILRNEPS--NEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLD 614
PE+L + +EK DV+SYG+ LWE++T P+ +++ ++G + R +P+
Sbjct: 192 PELLNGSSTRVSEKVDVFSYGISLWEILTGEEPYADMHCGAIIGGIVKNTLRPPIPKSCS 251
Query: 615 PHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRML 674
P ++ CW S DP+ RP F E+ R+
Sbjct: 252 PEWKKLMEQCW-------------------------------SVDPDSRPPFTEITCRLR 280
Query: 675 FVLNRVTAESVKRSSE 690
+ V +S +R ++
Sbjct: 281 SMSMEVVTKSKRRENK 296
>At1g62400 hypothetical protein
Length = 345
Score = 167 bits (422), Expect = 2e-41
Identities = 97/268 (36%), Positives = 149/268 (55%), Gaps = 11/268 (4%)
Query: 363 ESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGN 422
ES +SKG+ + + L + ++ G+++ +Y GI+ VA+K+
Sbjct: 17 ESENVETWEASKGERE---EWTADLSQLFIGNKFASGAHSRIYRGIYKQRAVAVKMVRIP 73
Query: 423 GYTEETL----QDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHK 478
+ EET Q +K E+ ++ RL HPN++ F+ A I+TE + +G+L L+K
Sbjct: 74 THKEETRAKLEQQFKSEVALLSRLFHPNIVQFIAACKKPPVYCIITEYMSQGNLRMYLNK 133
Query: 479 SNQ-TLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
+L I LR+ALDI+RGM YLH + ++HRDLKS+NLL++ VKV DFG S L
Sbjct: 134 KEPYSLSIETVLRLALDISRGMEYLHSQG--VIHRDLKSNNLLLNDEMRVKVADFGTSCL 191
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
+T K GT +WMAPE+++ +P K DVYS+G+VLWEL T +P++ + +Q
Sbjct: 192 -ETQCREAKGNMGTYRWMAPEMIKEKPYTRKVDVYSFGIVLWELTTALLPFQGMTPVQAA 250
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCW 625
V + R LP P +A +I CW
Sbjct: 251 FAVAEKNERPPLPASCQPALAHLIKRCW 278
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,115,901
Number of Sequences: 26719
Number of extensions: 725382
Number of successful extensions: 5325
Number of sequences better than 10.0: 1007
Number of HSP's better than 10.0 without gapping: 858
Number of HSP's successfully gapped in prelim test: 149
Number of HSP's that attempted gapping in prelim test: 2097
Number of HSP's gapped (non-prelim): 1214
length of query: 697
length of database: 11,318,596
effective HSP length: 106
effective length of query: 591
effective length of database: 8,486,382
effective search space: 5015451762
effective search space used: 5015451762
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0304.15


