

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
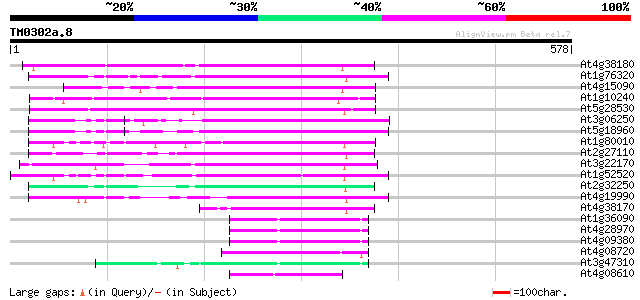
Query= TM0302a.8
(578 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 209 4e-54
At1g76320 putative phytochrome A signaling protein 162 5e-40
At4g15090 unknown protein 156 3e-38
At1g10240 unknown protein 152 5e-37
At5g28530 far-red impaired response protein (FAR1) - like 149 3e-36
At3g06250 unknown protein 149 3e-36
At5g18960 FAR1 - like protein 147 2e-35
At1g80010 hypothetical protein 144 1e-34
At2g27110 Mutator-like transposase 143 2e-34
At3g22170 far-red impaired response protein, putative 137 2e-32
At1g52520 F6D8.26 132 4e-31
At2g32250 Mutator-like transposase 119 5e-27
At4g19990 putative protein 112 4e-25
At4g38170 hypothetical protein 99 5e-21
At1g36090 hypothetical protein 64 2e-10
At4g28970 putative protein 63 5e-10
At4g09380 putative protein 62 7e-10
At4g08720 putative protein 61 1e-09
At3g47310 putative protein 61 2e-09
At4g08610 predicted transposon protein 59 6e-09
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 209 bits (531), Expect = 4e-54
Identities = 121/373 (32%), Positives = 193/373 (51%), Gaps = 15/373 (4%)
Query: 14 LDLMKYDFLD---SEVAYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEGLRE 69
LDL YD L+ E A FY+ Y GF+ R + ++G +IQR FVC KEG R
Sbjct: 69 LDLEPYDGLEFESEEAAKAFYNSYARRIGFSTRVSSSRRSRRDGAIIQRQFVCAKEGFRN 128
Query: 70 -DRGLTMDKRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIA 128
+ T D+ + RT R GC A V + + +G+W F +HNHEL +
Sbjct: 129 MNEKRTKDREIKRPRTITRVGCKASLSVKM-QDSGKWLVSGFVKDHNHELVPPDQVHCLR 187
Query: 129 AHRKMNESDIMHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKK 188
+HR+++ ++ + +G+ +I + + GG KV F++ + N R R K
Sbjct: 188 SHRQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYM-RNNRQKSI 246
Query: 189 EPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFD 248
E E + + Y+R + + +P + ++D ++ +FW+D + +++ FGD V FD
Sbjct: 247 EGE---IQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFD 303
Query: 249 ATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVI 308
TY NRY+ PF F GVNHH + +F A + NE ++VWL +L AM P+ +
Sbjct: 304 TTYRSNRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSAHPPVSIT 363
Query: 309 TDGDKAMKAAIKQVFPNAHHRLCAWHILDNARK---HVHIR--AFNTELRKCMFMELDVS 363
TD D ++AAI VFP A HR C WHIL ++ HV ++ +F ++ KC+ + V
Sbjct: 364 TDHDAVIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHPSFESDFHKCVNLTESVE 423
Query: 364 EFEEHWAATVAKF 376
+FE W + + K+
Sbjct: 424 DFERCWFSLLDKY 436
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 162 bits (410), Expect = 5e-40
Identities = 106/377 (28%), Positives = 177/377 (46%), Gaps = 21/377 (5%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGL-VIQRNFVCHKEGLREDRGLTMDKR 78
+F E AYLFY Y GF K ++ I F C + G ++ ++ R
Sbjct: 2 EFETHEDAYLFYKDYAKSVGFGTAKLSSRRSRASKEFIDAKFSCIRYGSKQQSDDAINPR 61
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDI 138
+ GC A V + +G+WY F EHNH+L ++ +HR N +
Sbjct: 62 ASP-----KIGCKASMHVK-RRPDGKWYVYSFVKEHNHDLLPEQ-AHYFRSHR--NTELV 112
Query: 139 MHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFA 198
++ R +TP T Y + F G M N + RR+ + DAE
Sbjct: 113 KSNDSRLRRKKNTPL---TDCKHLSAYHDLDFIDGYMRNQHDKGRRLVL---DTGDAEIL 166
Query: 199 ISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKC 258
+ ++ + ++P + ++D L +FW D +Y+ F DVV+F+ +Y ++YK
Sbjct: 167 LEFLMRMQEENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKV 226
Query: 259 PFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAA 318
P V+F GVNHH + + L++++ + TYVWL++ +L AM G+ P ++TD + A+KAA
Sbjct: 227 PLVLFVGVNHHVQPVLLGCGLLADDTVYTYVWLMQSWLVAMGGQKPKVMLTDQNNAIKAA 286
Query: 319 IKQVFPNAHHRLCAWHILDNARKHVHI-----RAFNTELRKCMFMELDVSEFEEHWAATV 373
I V P H C WH+LD +++ F +L KC++ EF+ W +
Sbjct: 287 IAAVLPETRHCYCLWHVLDQLPRNLDYWSMWQDTFMKKLFKCIYRSWSEEEFDRRWLKLI 346
Query: 374 AKFGWKKILGLRNYMKE 390
KF + + +R+ +E
Sbjct: 347 DKFHLRDVPWMRSLYEE 363
>At4g15090 unknown protein
Length = 768
Score = 156 bits (395), Expect = 3e-38
Identities = 100/340 (29%), Positives = 169/340 (49%), Gaps = 37/340 (10%)
Query: 56 IQRNFVCHKEGL--REDRGLTMDKRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDE 113
I F C + G+ + + +R +TD + K R +G+W F +
Sbjct: 34 IDAKFACSRYGVTPESESSGSSSRRSTVKKTDCKASMHVKRR-----PDGKWIIHEFVKD 88
Query: 114 HNHELEDDKMCGMIAAHRKM---------NESDIMHMNNMSRSGISTPQIHNTFASQTGG 164
HNHEL +A H ++ N DI+H + T +++ + Q+GG
Sbjct: 89 HNHEL-----LPALAYHFRIQRNVKLAEKNNIDILHAVSER-----TKKMYVEMSRQSGG 138
Query: 165 YQKVRFSKGNMY--NVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDG 222
Y+ + G++ +V + + + E+ D++ + Y + + ++P + N+D
Sbjct: 139 YKNI----GSLLQTDVSSQVDKGRYLALEEGDSQVLLEYFKRIKKENPKFFYAIDLNEDQ 194
Query: 223 NLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSN 282
L LFW+D S+ +Y F DVV+FD TY K K P +F GVNHH++ + ALV++
Sbjct: 195 RLRNLFWADAKSRDDYLSFNDVVSFDTTYVKFNDKLPLALFIGVNHHSQPMLLGCALVAD 254
Query: 283 EKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARK- 341
E +ET+VWL++ +L AM G+AP ++TD DK + +A+ ++ PN H WH+L+ +
Sbjct: 255 ESMETFVWLIKTWLRAMGGRAPKVILTDQDKFLMSAVSELLPNTRHCFALWHVLEKIPEY 314
Query: 342 --HVHIRAFNTELR--KCMFMELDVSEFEEHWAATVAKFG 377
HV R N L+ KC+F EF+ W V++FG
Sbjct: 315 FSHVMKRHENFLLKFNKCIFRSWTDDEFDMRWWKMVSQFG 354
>At1g10240 unknown protein
Length = 680
Score = 152 bits (384), Expect = 5e-37
Identities = 102/368 (27%), Positives = 173/368 (46%), Gaps = 17/368 (4%)
Query: 21 FLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGL---VIQRNFVCHKEGLREDRGLTMDK 77
FL + AY FY + GF++R++ + K+G+ + +R FVCH+ G + L+ K
Sbjct: 54 FLTHDTAYEFYSTFAKRCGFSIRRH-RTEGKDGVGKGLTRRYFVCHRAGNTPIKTLSEGK 112
Query: 78 RQRESRTDFRCGCTAKFRVH--IDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNE 135
QR R+ RCGC A R+ + + W F++ HNHEL + + A+R +++
Sbjct: 113 PQRNRRSS-RCGCQAYLRISKLTELGSTEWRVTGFANHHNHELLEPNQVRFLPAYRSISD 171
Query: 136 SDIMHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDA 195
+D + S++GIS Q+ + + F +V+ + KK +PE +
Sbjct: 172 ADKSRILMFSKTGISVQQMMRLLELEK--CVEPGFLPFTEKDVRNLLQSFKKLDPEDENI 229
Query: 196 EFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNR 255
+F + + + KDP + + L+ + WS S +Y++FGD V FD T+ +
Sbjct: 230 DF-LRMCQSIKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYELFGDAVVFDTTHRLSA 288
Query: 256 YKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAM 315
+ P ++ GVN++ F L+ +E + ++ W L+ F M GKAP ++TD + +
Sbjct: 289 VEMPLGIWVGVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFMNGKAPQTILTDHNMCL 348
Query: 316 KAAIKQVFPNAHHRLCAWHILD------NARKHVHIRAFNTELRKCMFMELDVSEFEEHW 369
K AI P H LC W ++ NA + E + +E V EFE W
Sbjct: 349 KEAIAGEMPATKHALCIWMVVGKFPSWFNAGLGERYNDWKAEFYRLYHLE-SVEEFELGW 407
Query: 370 AATVAKFG 377
V FG
Sbjct: 408 RDMVNSFG 415
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 149 bits (377), Expect = 3e-36
Identities = 96/372 (25%), Positives = 179/372 (47%), Gaps = 17/372 (4%)
Query: 21 FLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKRQR 80
F + A+ +Y + +GF++RK ++ V +R+FVC++ G + R + R
Sbjct: 76 FTTDDEAFEYYSTFARKSGFSIRKARSTESQNLGVYRRDFVCYRSGFNQPRKKANVEHPR 135
Query: 81 ESRTDFRCGCTAKFRVHIDKTNG--RWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDI 138
E R RCGC K + + +G WY FS+ HNHEL +D ++ A+RK+ +SD
Sbjct: 136 E-RKSVRCGCDGKLYLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLPAYRKIQQSDQ 194
Query: 139 MHMNNMSRSGISTPQIHNTFASQTGGYQ-KVRFSKGNMYN-VQGRQRRMKKK-----EPE 191
+ +S++G +I + G ++ F + ++ N V+ ++ +++ E
Sbjct: 195 ERILLLSKAGFPVNRIVKLLELEKGVVSGQLPFIEKDVRNFVRACKKSVQENDAFMTEKR 254
Query: 192 KTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATY 251
++D + +GL +D ++++ ++ + W+ G S Y +FGDVV FD +Y
Sbjct: 255 ESDTLELLECCKGLAERDMDFVYDCTSDENQKVENIAWAYGDSVRGYSLFGDVVVFDTSY 314
Query: 252 GKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDG 311
Y VF+G++++ K+ + L+ +E ++ W L+ F+ M+G+ P ++TD
Sbjct: 315 RSVPYGLLLGVFFGIDNNGKAMLLGCVLLQDESCRSFTWALQTFVRFMRGRHPQTILTDI 374
Query: 312 DKAMKAAIKQVFPNAHHRLCAWHILDNARKHV------HIRAFNTELRKCMFMELDVSEF 365
D +K AI + PN +H + HI+ H F + +V EF
Sbjct: 375 DTGLKDAIGREMPNTNHVVFMSHIVSKLASWFSQTLGSHYEEFRAGF-DMLCRAGNVDEF 433
Query: 366 EEHWAATVAKFG 377
E+ W V +FG
Sbjct: 434 EQQWDLLVTRFG 445
>At3g06250 unknown protein
Length = 764
Score = 149 bits (377), Expect = 3e-36
Identities = 107/384 (27%), Positives = 164/384 (41%), Gaps = 65/384 (16%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNK-EGLVIQRNFVCHKEGLREDRGLTMDKR 78
+F + A FY Y V GF VR +K +G + R FVC KEG +
Sbjct: 195 EFNSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSKEGFQHPS------- 247
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNES-- 136
R GC A R+ + G W + +HNH+LE K A +K+ +
Sbjct: 248 --------RMGCGAYMRIKRQDSGG-WIVDRLNKDHNHDLEPGKKN---AGMKKITDDVT 295
Query: 137 ------DIMHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEP 190
D++ +N++S ST + NT + Y V
Sbjct: 296 GGLDSVDLIELNDLSNHISSTRE--NTIGKE-------------WYPV------------ 328
Query: 191 EKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDAT 250
+ Y + ++D + + +G+ +FW+D S+ FGD V FD +
Sbjct: 329 -------LLDYFQSKQAEDMGFFYAIELDSNGSCMSIFWADSRSRFACSQFGDAVVFDTS 381
Query: 251 YGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITD 310
Y K Y PF F G NHH + + ALV++E E + WL + +L AM G+ P ++ D
Sbjct: 382 YRKGDYSVPFATFIGFNHHRQPVLLGGALVADESKEAFSWLFQTWLRAMSGRRPRSMVAD 441
Query: 311 GDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVHI--RAFNTELRKCMFMELDVSEFEEH 368
D ++ A+ QVFP HHR AW I R+++ F E KC++ EF+
Sbjct: 442 QDLPIQQAVAQVFPGTHHRFSAWQIRSKERENLRSFPNEFKYEYEKCLYQSQTTVEFDTM 501
Query: 369 WAATVAKFGWKKILGLRN-YMKEE 391
W++ V K+G + + LR Y K E
Sbjct: 502 WSSLVNKYGLRDNMWLREIYEKRE 525
Score = 42.7 bits (99), Expect = 5e-04
Identities = 34/100 (34%), Positives = 44/100 (44%), Gaps = 17/100 (17%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNK-EGLVIQRNFVCHKEGLREDRGLTMDKR 78
+F +E A +Y+ Y GF VR ++ +G V R FVC KEG
Sbjct: 33 EFDTAEEARDYYNSYATRTGFKVRTGQLYRSRTDGTVSSRRFVCSKEGF----------- 81
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
Q SRT GC A RV + G+W EHNH+L
Sbjct: 82 QLNSRT----GCPAFIRVQ-RRDTGKWVLDQIQKEHNHDL 116
>At5g18960 FAR1 - like protein
Length = 788
Score = 147 bits (371), Expect = 2e-35
Identities = 100/377 (26%), Positives = 168/377 (44%), Gaps = 51/377 (13%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNK-EGLVIQRNFVCHKEGLREDRGLTMDKR 78
+F + A FY Y V GF VR +K +G + R FVC +EG +
Sbjct: 216 EFGSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSREGFQHPS------- 268
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDI 138
R GC A R+ + G W + +HNH+LE K N++ +
Sbjct: 269 --------RMGCGAYMRIKRQDSGG-WIVDRLNKDHNHDLEPGKK----------NDAGM 309
Query: 139 MHMNNMSRSGISTPQIH--NTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAE 196
+ + G+ + + N F GN + + R+ R+ K+
Sbjct: 310 KKIPDDGTGGLDSVDLIELNDF--------------GNNHIKKTRENRIGKEW-----YP 350
Query: 197 FAISYIRGLGSKDP-LLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNR 255
+ Y + ++D Y + ++G+ +FW+D ++ FGD V FD +Y K
Sbjct: 351 LLLDYFQSRQTEDMGFFYAVELDVNNGSCMSIFWADSRARFACSQFGDSVVFDTSYRKGS 410
Query: 256 YKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAM 315
Y PF G NHH + + A+V++E E ++WL + +L AM G+ P ++ D D +
Sbjct: 411 YSVPFATIIGFNHHRQPVLLGCAMVADESKEAFLWLFQTWLRAMSGRRPRSIVADQDLPI 470
Query: 316 KAAIKQVFPNAHHRLCAWHILDNARKHV--HIRAFNTELRKCMFMELDVSEFEEHWAATV 373
+ A+ QVFP AHHR AW I + R+++ F E KC++ + EF+ W+A +
Sbjct: 471 QQALVQVFPGAHHRYSAWQIREKERENLIPFPSEFKYEYEKCIYQTQTIVEFDSVWSALI 530
Query: 374 AKFGWKKILGLRNYMKE 390
K+G + + LR ++
Sbjct: 531 NKYGLRDDVWLREIYEQ 547
Score = 47.8 bits (112), Expect = 2e-05
Identities = 37/100 (37%), Positives = 45/100 (45%), Gaps = 17/100 (17%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNK-EGLVIQRNFVCHKEGLREDRGLTMDKR 78
+F +E A FY+ Y GF VR ++ +G V R FVC KEG
Sbjct: 48 EFDTAEEAREFYNAYAARTGFKVRTGQLYRSRTDGTVSSRRFVCSKEGF----------- 96
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
Q SRT GCTA RV + G+W EHNHEL
Sbjct: 97 QLNSRT----GCTAFIRVQ-RRDTGKWVLDQIQKEHNHEL 131
>At1g80010 hypothetical protein
Length = 696
Score = 144 bits (364), Expect = 1e-34
Identities = 113/378 (29%), Positives = 170/378 (44%), Gaps = 39/378 (10%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVR---KYIKIHNKE--GLVIQRNFVCHKEGLREDRGLT 74
+F + AY FY+ Y GFA+R + K ++KE G V+ C+ +G + L
Sbjct: 71 EFESYDDAYSFYNSYARELGFAIRVKSSWTKRNSKEKRGAVL----CCNCQGFK----LL 122
Query: 75 MDKRQRESRTDFRCGCTAKFR---VHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHR 131
D R T R GC A R +H D RW +HNH D + +H+
Sbjct: 123 KDAHSRRKET--RTGCQAMIRLRLIHFD----RWKVDQVKLDHNHSF-DPQRAHNSKSHK 175
Query: 132 KMNESDIMHMNNMSRSG----ISTPQIHNTFASQTGGYQKVRFSKGNMYNVQ----GRQR 183
K + S + T +++ T A T S G ++ R
Sbjct: 176 KSSSSASPATKTNPEPPPHVQVRTIKLYRTLALDTPPALGTSLSSGETSDLSLDHFQSSR 235
Query: 184 RMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGD 243
R++ + + +F + + S LY +A DDG+L +FW D ++ Y FGD
Sbjct: 236 RLELRGGFRALQDF---FFQIQLSSPNFLYLMDLA-DDGSLRNVFWIDARARAAYSHFGD 291
Query: 244 VVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKA 303
V+ FD T N Y+ P V F G+NHH + + L++++ ETYVWL +L M G+
Sbjct: 292 VLLFDTTCLSNAYELPLVAFVGINHHGDTILLGCGLLADQSFETYVWLFRAWLTCMLGRP 351
Query: 304 PLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHV----HIRAFNTELRKCMFME 359
P IT+ KAM+ A+ +VFP AHHRL H+L N + V F L + ++
Sbjct: 352 PQIFITEQCKAMRTAVSEVFPRAHHRLSLTHVLHNICQSVVQLQDSDLFPMALNRVVYGC 411
Query: 360 LDVSEFEEHWAATVAKFG 377
L V EFE W + +FG
Sbjct: 412 LKVEEFETAWEEMIIRFG 429
>At2g27110 Mutator-like transposase
Length = 851
Score = 143 bits (361), Expect = 2e-34
Identities = 94/362 (25%), Positives = 157/362 (42%), Gaps = 37/362 (10%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKRQ 79
+F + A FY Y GF + + +G V R FVC R R L+
Sbjct: 54 EFNSEKEAKSFYDEYSRQLGFTSKL---LPRTDGSVSVREFVCSSSSKRSKRRLSES--- 107
Query: 80 RESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDIM 139
C A R+ + + + +W F EH H L M + R S+
Sbjct: 108 ----------CDAMVRIEL-QGHEKWVVTKFVKEHTHGLASSNMLHCLRPRRHFANSE-- 154
Query: 140 HMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAI 199
+ + G++ P G V N + K DA +
Sbjct: 155 --KSSYQEGVNVPS----------GMMYVSMD-ANSRGARNASMATNTKRTIGRDAHNLL 201
Query: 200 SYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCP 259
Y + + +++P + ++D + +FW+D S++ Y FGD V D Y N+++ P
Sbjct: 202 EYFKRMQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVP 261
Query: 260 FVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAI 319
F F GVNHH ++ +F AL+ +E +++WL + FL AM+ + P+ ++TD D+A++ A
Sbjct: 262 FAPFTGVNHHGQAILFGCALILDESDTSFIWLFKTFLTAMRDQPPVSLVTDQDRAIQIAA 321
Query: 320 KQVFPNAHHRLCAWHIL-DNARKHVHI----RAFNTELRKCMFMELDVSEFEEHWAATVA 374
QVFP A H + W +L + K H+ +F EL C+ + EFE W++ +
Sbjct: 322 GQVFPGARHCINKWDVLREGQEKLAHVCLAYPSFQVELYNCINFTETIEEFESSWSSVID 381
Query: 375 KF 376
K+
Sbjct: 382 KY 383
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 137 bits (345), Expect = 2e-32
Identities = 104/387 (26%), Positives = 169/387 (42%), Gaps = 48/387 (12%)
Query: 11 LVPLDLMKYDFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGL-VIQRNFVCHKEGLRE 69
L PL+ M+++ AY FY Y GF +K I F C + G +
Sbjct: 68 LEPLNGMEFE--SHGEAYSFYQEYSRAMGFNTAIQNSRRSKTTREFIDAKFACSRYGTKR 125
Query: 70 DRGLTMDK-RQRESRTDFR-------CGCT-AKFRVHIDKT-NGRWYAKLFSDEHNHELE 119
+ + ++ R R+S+ D C T K +H+ + +G+W F EHNHEL
Sbjct: 126 EYDKSFNRPRARQSKQDPENMAGRRTCAKTDCKASMHVKRRPDGKWVIHSFVREHNHEL- 184
Query: 120 DDKMCGMIAAHRKMNESDIMHMNNMSRSGIS--TPQIHNTFASQTGGYQKVRFSKGNMYN 177
+ +S T +I+ A Q Y+ V K + +
Sbjct: 185 ------------------------LPAQAVSEQTRKIYAAMAKQFAEYKTVISLKSDSKS 220
Query: 178 VQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLN 237
+ R + E D + + ++ + S + + DD + +FW D S+ N
Sbjct: 221 SFEKGRTLSV---ETGDFKILLDFLSRMQSLNSNFFYAVDLGDDQRVKNVFWVDAKSRHN 277
Query: 238 YQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLE 297
Y F DVV+ D TY +N+YK P +F GVN H + + AL+S+E TY WL+E +L
Sbjct: 278 YGSFCDVVSLDTTYVRNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWLMETWLR 337
Query: 298 AMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHV-----HIRAFNTEL 352
A+ G+AP +IT+ D M + + ++FPN H L WH+L +++ F +
Sbjct: 338 AIGGQAPKVLITELDVVMNSIVPEIFPNTRHCLFLWHVLMKVSENLGQVVKQHDNFMPKF 397
Query: 353 RKCMFMELDVSEFEEHWAATVAKFGWK 379
KC++ +F W +A+FG K
Sbjct: 398 EKCIYKSGKDEDFARKWYKNLARFGLK 424
>At1g52520 F6D8.26
Length = 703
Score = 132 bits (333), Expect = 4e-31
Identities = 110/400 (27%), Positives = 183/400 (45%), Gaps = 34/400 (8%)
Query: 1 QDILGIDLKKLVPLDLMKYDFLDSEVAYLFYHWYG*VNGFAVR---KYIKIHNKE--GLV 55
Q L +D +K + +F + AY +Y+ Y GF VR + K +KE G V
Sbjct: 71 QSGLLVDERKEFDAPAVGMEFESYDDAYNYYNCYASEVGFRVRVKNSWFKRRSKEKYGAV 130
Query: 56 IQRNFVCHKEGLREDRGLTMDKRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHN 115
+ C +G + R +++ ++E+RT GC A R+ + RW + +HN
Sbjct: 131 L----CCSSQGFK--RINDVNRVRKETRT----GCPAMIRMR-QVDSKRWRVVEVTLDHN 179
Query: 116 HELEDDKMCGMIAAHRKMNESDIMHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNM 175
H L K+ + RK S + S T +++ G +
Sbjct: 180 HLL-GCKLYKSVKRKRKCVSSPV--------SDAKTIKLYRACVVDNGSNVNPNSTLNKK 230
Query: 176 Y-NVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGIS 234
+ N G + K + D+ +Y + +P + ND+G L +FW+D S
Sbjct: 231 FQNSTGSPDLLNLK---RGDSAAIYNYFCRMQLTNPNFFYLMDVNDEGQLRNVFWADAFS 287
Query: 235 QLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLER 294
+++ FGDV+ D++Y +++ P V F GVNHH K+T+ S ++ E +E+Y WLL+
Sbjct: 288 KVSCSYFGDVIFIDSSYISGKFEIPLVTFTGVNHHGKTTLLSCGFLAGETMESYHWLLKV 347
Query: 295 FLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHV----HIRAFNT 350
+L MK ++P ++TD K ++AAI QVFP +H R HI+ + + + A
Sbjct: 348 WLSVMK-RSPQTIVTDRCKPLEAAISQVFPRSHQRFSLTHIMRKIPEKLGGLHNYDAVRK 406
Query: 351 ELRKCMFMELDVSEFEEHWAATVAKFGWKKILGLRNYMKE 390
K ++ L V EFE W V FG + LR+ +E
Sbjct: 407 AFTKAVYETLKVVEFEAAWGFMVHNFGVIENEWLRSLYEE 446
>At2g32250 Mutator-like transposase
Length = 684
Score = 119 bits (298), Expect = 5e-27
Identities = 87/363 (23%), Positives = 139/363 (37%), Gaps = 55/363 (15%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKR 78
DF E AY FY Y GF + K + + G I C + G + ++ ++ R
Sbjct: 43 DFESKEAAYYFYREYARSVGFGITIKASRRSKRSGKFIDVKIACSRFGTKREKATAINPR 102
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDI 138
+ GC A + K + +W F EHNHE+ D + K
Sbjct: 103 SCP-----KTGCKAGLHMK-RKEDEKWVIYNFVKEHNHEICPDDFYVSVRGKNK------ 150
Query: 139 MHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFA 198
G + +G Q + E+ D +
Sbjct: 151 --------------------------------PAGALAIKKGLQLAL-----EEEDLKLL 173
Query: 199 ISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKC 258
+ + + K P + + D + +FW D ++ +Y F DVV FD Y +N Y+
Sbjct: 174 LEHFMEMQDKQPGFFYAVDFDSDKRVRNVFWLDAKAKHDYCSFSDVVLFDTFYVRNGYRI 233
Query: 259 PFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAA 318
PF F GV+HH + + AL+ TY WL +L+A+ G+AP +ITD DK +
Sbjct: 234 PFAPFIGVSHHRQYVLLGCALIGEVSESTYSWLFRTWLKAVGGQAPGVMITDQDKLLSDI 293
Query: 319 IKQVFPNAHHRLCAWHILDNARKHVH-----IRAFNTELRKCMFMELDVSEFEEHWAATV 373
+ +VFP+ H C W +L + ++ F C+ FE W+ +
Sbjct: 294 VVEVFPDVRHIFCLWSVLSKISEMLNPFVSQDDGFMESFGNCVASSWTDEHFERRWSNMI 353
Query: 374 AKF 376
KF
Sbjct: 354 GKF 356
>At4g19990 putative protein
Length = 672
Score = 112 bits (281), Expect = 4e-25
Identities = 96/389 (24%), Positives = 161/389 (40%), Gaps = 74/389 (19%)
Query: 20 DFLDSEVAYLFYHWYG*VNGFA-VRKYIKIHNKEGLVIQRNFVCHKEGLRE---DRGLTM 75
+F E A+ FY Y GF + K + G I FVC + G ++ D GL
Sbjct: 26 EFESKEEAFEFYKEYANSVGFTTIIKASRRSRMTGKFIDAKFVCTRYGSKKEDIDTGLGT 85
Query: 76 D--------KRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMI 127
D KR R +R+ + C A V + +GRW + EHNHE+ G
Sbjct: 86 DGFNIPQARKRGRINRSSSKTDCKAFLHVK-RRQDGRWVVRSLVKEHNHEI----FTGQA 140
Query: 128 AAHRKMNESDIMHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKK 187
+ R+++ G +K+ G + + +K
Sbjct: 141 DSLRELS-----------------------------GRRKLEKLNGAIV------KEVKS 165
Query: 188 KEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAF 247
++ E D E +++ + S L +FW D + +Y F DVV+
Sbjct: 166 RKLEDGDVERLLNFFTDMQS----------------LRNIFWVDAKGRFDYTCFSDVVSI 209
Query: 248 DATYGKNRYKCPFVVFYGVNHHNKSTIFSTA-LVSNEKIETYVWLLERFLEAMKGKAPLF 306
D T+ KN YK P V F GVNHH + + L+++E +VWL +L+AM G P
Sbjct: 210 DTTFIKNEYKLPLVAFTGVNHHGQFLLLGFGLLLTDESKSGFVWLFRAWLKAMHGCRPRV 269
Query: 307 VITDGDKAMKAAIKQVFPNAHHRLCAWHILDN-ARKHVHI----RAFNTELRKCMFMELD 361
++T D+ +K A+ +VFP++ H W L K H+ + E+ ++
Sbjct: 270 ILTKHDQMLKEAVLEVFPSSRHCFYMWDTLGQMPEKLGHVIRLEKKLVDEINDAIYGSCQ 329
Query: 362 VSEFEEHWAATVAKFGWKKILGLRNYMKE 390
+FE++W V +F + + L++ ++
Sbjct: 330 SEDFEKNWWEVVDRFHMRDNVWLQSLYED 358
>At4g38170 hypothetical protein
Length = 531
Score = 99.4 bits (246), Expect = 5e-21
Identities = 57/188 (30%), Positives = 94/188 (49%), Gaps = 12/188 (6%)
Query: 196 EFAISYIRGLGSKDP-LLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKN 254
E ++Y++ ++P LY + +D GN+ FW+D +LNY FGD + FD TY +
Sbjct: 5 EHVLNYLKRRQLENPGFLYA--IEDDCGNV---FWADPTCRLNYTYFGDTLVFDTTYRRG 59
Query: 255 -RYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDK 313
RY+ PF F G NHH + +F AL+ NE ++ WL + +L+AM P + + D+
Sbjct: 60 KRYQVPFAAFTGFNHHGQPVLFGCALILNESESSFAWLFQTWLQAMSAPPPPSITVEPDR 119
Query: 314 AMKAAIKQVFPNAHHRLCAWHILDNA-RKHVHI----RAFNTELRKCMFMELDVSEFEEH 368
++ A+ +VF R I + K H+ F +E C+ +EFE
Sbjct: 120 LIQVAVSRVFSQTRLRFSQPLIFEETEEKLAHVFQAHPTFESEFINCVTETETAAEFEAS 179
Query: 369 WAATVAKF 376
W + V ++
Sbjct: 180 WDSIVRRY 187
>At1g36090 hypothetical protein
Length = 645
Score = 64.3 bits (155), Expect = 2e-10
Identities = 43/147 (29%), Positives = 69/147 (46%), Gaps = 7/147 (4%)
Query: 227 LFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVF--YGVNHHNKSTIFSTALVSNEK 284
LF + G ++ V+ DAT+ K Y V+ + NHH+ F ++ +EK
Sbjct: 353 LFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLVIATAHDPNHHHYPLAFG--IIDSEK 410
Query: 285 IETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVH 344
+++W LE+ L I+D +++K A+K V+PNA H C WH+ N R V
Sbjct: 411 DVSWIWFLEKLKTVYSDVPRLVFISDRHQSIKKAVKTVYPNALHAACIWHLCQNMRDRVK 470
Query: 345 I--RAFNTELRKCMFMELDVSEFEEHW 369
I + R C + SEFE+ +
Sbjct: 471 IDKDGAAVKFRDCAHAYTE-SEFEKEF 496
>At4g28970 putative protein
Length = 914
Score = 62.8 bits (151), Expect = 5e-10
Identities = 43/147 (29%), Positives = 68/147 (46%), Gaps = 7/147 (4%)
Query: 227 LFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGV--NHHNKSTIFSTALVSNEK 284
LF + G ++ V+ DAT+ K Y V+ NHH+ F ++ +EK
Sbjct: 393 LFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPLAFG--IIDSEK 450
Query: 285 IETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVH 344
+++W LE+ L I+D +++K A+K V+PNA H C WH+ N R V
Sbjct: 451 DVSWIWFLEKLKTVYSDVPGLVFISDRHQSIKKAVKTVYPNALHAACIWHLCQNMRDRVT 510
Query: 345 I--RAFNTELRKCMFMELDVSEFEEHW 369
I + R C + SEFE+ +
Sbjct: 511 IDKDGAAVKFRDCAHAYTE-SEFEKEF 536
>At4g09380 putative protein
Length = 960
Score = 62.4 bits (150), Expect = 7e-10
Identities = 43/147 (29%), Positives = 67/147 (45%), Gaps = 7/147 (4%)
Query: 227 LFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGV--NHHNKSTIFSTALVSNEK 284
LF + G ++ V+ DAT+ K Y V+ NHH+ F ++ +EK
Sbjct: 393 LFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPLAFG--IIDSEK 450
Query: 285 IETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVH 344
+++W LE L I+D +++K A+K V+PNA H C WH+ N R V
Sbjct: 451 DVSWIWFLENLKTVYSDVPGLVFISDRHQSIKKAVKTVYPNALHAACIWHLCQNMRDRVK 510
Query: 345 I--RAFNTELRKCMFMELDVSEFEEHW 369
I + R C + SEFE+ +
Sbjct: 511 IDKDGAAVKFRDCAHAYTE-SEFEKEF 536
>At4g08720 putative protein
Length = 848
Score = 61.2 bits (147), Expect = 1e-09
Identities = 39/155 (25%), Positives = 66/155 (42%), Gaps = 6/155 (3%)
Query: 219 NDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTA 278
++ G LF + G S +Q V+ DAT+ KN Y V + + I +
Sbjct: 365 DESGKFQYLFIALGASIEGFQAMRKVIIVDATHLKNGYGGVLVFASARDPNRHHYIIAVG 424
Query: 279 LVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDN 338
++ E ++ W E+ L + L ++D + ++ I+ + AHH C WH+ N
Sbjct: 425 VLDGENDASWGWFFEKLLSVVPDTPELVFMSDRNSSLIKGIRNAYTAAHHGYCVWHLSQN 484
Query: 339 ARKHVHIRAFNTELRKCMFMELD----VSEFEEHW 369
+ H N ++ FMEL SEFE +
Sbjct: 485 VK--AHSTNINRDVLAWRFMELSRIYTWSEFEREY 517
>At3g47310 putative protein
Length = 735
Score = 60.8 bits (146), Expect = 2e-09
Identities = 67/290 (23%), Positives = 108/290 (37%), Gaps = 21/290 (7%)
Query: 89 GCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRSG 148
GCT RV K + W ++ H + I + ++H +
Sbjct: 208 GCTWYLRVARTKKSHFWSVRVHRKMHTCSRSVETTSNSIQRGTPRLIASVLHCDYPGNLE 267
Query: 149 ISTPQIHNTFASQTGGYQKVRFS-----KGNMYNVQGRQRRMKKKEPEKTDAEFAISYIR 203
TP N S G V S +G M +V + PE++ SY+
Sbjct: 268 TPTP---NNIMSIVRGRLGVHCSYSTALRGKMLHVSD-----VRGTPERSYT-MLFSYLY 318
Query: 204 GLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVF 263
L +P + LF + G ++ V+ DAT+ K Y V+
Sbjct: 319 MLEKVNPGTVTYVELEGEKKFKYLFIALGACIEGFRTMRKVIVVDATHLKTVYGGMLVIA 378
Query: 264 YGV--NHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQ 321
NHH+ F ++ +E +++W LE+ L I+D +++K +K
Sbjct: 379 TAQDPNHHHYPLAFG--IIDSENDVSWIWFLEKLKTVYSDVPGLVFISDRHQSIKKVVKT 436
Query: 322 VFPNAHHRLCAWHILDNARKHVHI--RAFNTELRKCMFMELDVSEFEEHW 369
V+PNA H C WH+ N R V I + R C + SEFE+ +
Sbjct: 437 VYPNALHAACIWHLCQNMRDRVKIDKDGAAVKFRDCAHAYTE-SEFEKEF 485
>At4g08610 predicted transposon protein
Length = 907
Score = 59.3 bits (142), Expect = 6e-09
Identities = 35/119 (29%), Positives = 56/119 (46%), Gaps = 4/119 (3%)
Query: 227 LFWSDGISQLNYQVFGDVVAFDATYGKNRY--KCPFVVFYGVNHHNKSTIFSTALVSNEK 284
LF S G+ ++V VV DAT+ K Y F NHHN I ++A++ E
Sbjct: 395 LFVSLGVCIEGFKVMRKVVIVDATFLKTVYGDMLVFATAQDPNHHNY--IIASAVIDREN 452
Query: 285 IETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHV 343
++ W + + + L ++D +++ +I VFPNA H C WH+ N + V
Sbjct: 453 DASWSWFFNKLKTVIPDELGLVFVSDRHQSIIKSIMHVFPNARHGHCVWHLSQNVKVRV 511
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.340 0.149 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,478,561
Number of Sequences: 26719
Number of extensions: 518878
Number of successful extensions: 1731
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 77
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 1587
Number of HSP's gapped (non-prelim): 111
length of query: 578
length of database: 11,318,596
effective HSP length: 105
effective length of query: 473
effective length of database: 8,513,101
effective search space: 4026696773
effective search space used: 4026696773
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0302a.8


