

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.2
(80 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
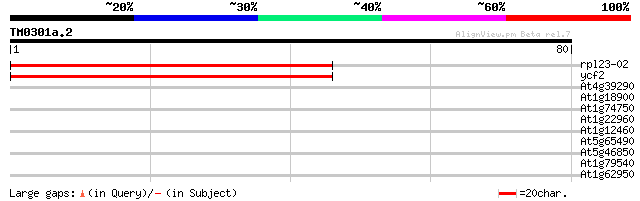
Score E
Sequences producing significant alignments: (bits) Value
rpl23-02 -chloroplast genome- 82 5e-17
ycf2 -chloroplast genome- 82 5e-17
At4g39290 putative protein 31 0.096
At1g18900 unknown protein 29 0.37
At1g74750 27 2.4
At1g22960 putative salt-inducible protein 26 4.1
At1g12460 unknown protein 26 4.1
At5g65490 unknown protein 25 5.3
At5g46850 unknown protein 25 6.9
At1g79540 unknown protein 25 9.0
At1g62950 protein kinase like protein 25 9.0
>rpl23-02 -chloroplast genome-
Length = 2294
Score = 82.0 bits (201), Expect = 5e-17
Identities = 39/46 (84%), Positives = 42/46 (90%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI RQRWLR N+SLSNGFFRSNT SESYQYLSN+F+SNGTLLD
Sbjct: 2227 FELLIQRQRWLRTNSSLSNGFFRSNTRSESYQYLSNLFISNGTLLD 2272
>ycf2 -chloroplast genome-
Length = 2294
Score = 82.0 bits (201), Expect = 5e-17
Identities = 39/46 (84%), Positives = 42/46 (90%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI RQRWLR N+SLSNGFFRSNT SESYQYLSN+F+SNGTLLD
Sbjct: 2227 FELLIQRQRWLRTNSSLSNGFFRSNTRSESYQYLSNLFISNGTLLD 2272
>At4g39290 putative protein
Length = 365
Score = 31.2 bits (69), Expect = 0.096
Identities = 14/29 (48%), Positives = 19/29 (65%)
Query: 5 IHRQRWLRPNNSLSNGFFRSNTLSESYQY 33
IH+ RW P+ SLS+GF SN + E+ Y
Sbjct: 235 IHKGRWREPHLSLSHGFHFSNCVIENVFY 263
>At1g18900 unknown protein
Length = 860
Score = 29.3 bits (64), Expect = 0.37
Identities = 15/31 (48%), Positives = 20/31 (64%), Gaps = 4/31 (12%)
Query: 11 LRPN----NSLSNGFFRSNTLSESYQYLSNM 37
LRPN NSL + F R N ++E+Y+ L NM
Sbjct: 605 LRPNVPTCNSLLSTFLRVNKIAEAYELLQNM 635
>At1g74750
Length = 855
Score = 26.6 bits (57), Expect = 2.4
Identities = 15/34 (44%), Positives = 20/34 (58%), Gaps = 4/34 (11%)
Query: 8 QRWLRPN----NSLSNGFFRSNTLSESYQYLSNM 37
Q LRPN NSL + F R + +SE+Y L +M
Sbjct: 597 QAGLRPNVPTCNSLLSTFLRVHRMSEAYNLLQSM 630
>At1g22960 putative salt-inducible protein
Length = 718
Score = 25.8 bits (55), Expect = 4.1
Identities = 12/35 (34%), Positives = 21/35 (59%), Gaps = 4/35 (11%)
Query: 7 RQRWLRPN----NSLSNGFFRSNTLSESYQYLSNM 37
++R +RPN N+L G ++ + E+Y+YL M
Sbjct: 612 KKRGVRPNVMTHNALLYGMCKAGNIDEAYRYLCKM 646
>At1g12460 unknown protein
Length = 882
Score = 25.8 bits (55), Expect = 4.1
Identities = 13/39 (33%), Positives = 23/39 (58%), Gaps = 4/39 (10%)
Query: 13 PNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLDNTKMK 51
PN S G++ S+T+ Q + + F+ NG+L DN ++
Sbjct: 648 PNLSSFQGYYFSSTM----QLILSEFVPNGSLYDNLHLR 682
>At5g65490 unknown protein
Length = 643
Score = 25.4 bits (54), Expect = 5.3
Identities = 13/30 (43%), Positives = 15/30 (49%), Gaps = 3/30 (10%)
Query: 1 FLLL---IHRQRWLRPNNSLSNGFFRSNTL 27
FLL+ H RWL P SL+ F R L
Sbjct: 123 FLLIEAAFHLPRWLNPETSLNRVFIRGGDL 152
>At5g46850 unknown protein
Length = 296
Score = 25.0 bits (53), Expect = 6.9
Identities = 11/34 (32%), Positives = 21/34 (61%)
Query: 18 SNGFFRSNTLSESYQYLSNMFLSNGTLLDNTKMK 51
S+ FF S+T+ S QY+ + + T LD+++ +
Sbjct: 31 SHEFFDSDTICISDQYIMESYFTTYTSLDDSEFR 64
>At1g79540 unknown protein
Length = 780
Score = 24.6 bits (52), Expect = 9.0
Identities = 10/31 (32%), Positives = 19/31 (61%)
Query: 11 LRPNNSLSNGFFRSNTLSESYQYLSNMFLSN 41
LR +SL +G FR+ +++++ +NM N
Sbjct: 303 LRGYSSLIDGLFRARRYTQAFELYANMLKKN 333
>At1g62950 protein kinase like protein
Length = 890
Score = 24.6 bits (52), Expect = 9.0
Identities = 12/35 (34%), Positives = 22/35 (62%), Gaps = 4/35 (11%)
Query: 13 PNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLDN 47
PN + G++ S+T+ Q + + F++NG+L DN
Sbjct: 655 PNLASFQGYYFSSTM----QLILSEFVTNGSLYDN 685
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.350 0.154 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,441,301
Number of Sequences: 26719
Number of extensions: 43557
Number of successful extensions: 269
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 262
Number of HSP's gapped (non-prelim): 11
length of query: 80
length of database: 11,318,596
effective HSP length: 56
effective length of query: 24
effective length of database: 9,822,332
effective search space: 235735968
effective search space used: 235735968
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0301a.2


