

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0299.14
(88 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
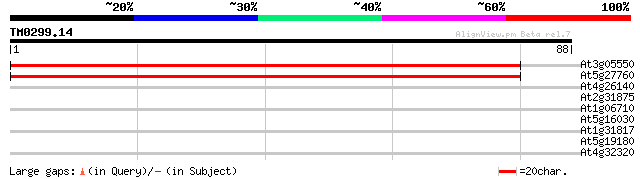
At3g05550 unknown protein 146 2e-36
At5g27760 unknown protein 139 3e-34
At4g26140 putative beta-galactosidase 28 0.61
At2g31875 poly(ADP-ribose) like glycohydrolase 28 0.80
At1g06710 hypothetical protein 26 3.0
At5g16030 unknown protein 26 4.0
At1g31817 30S ribosomal like protein S11 25 6.8
At5g19180 putative ubiquitin activating enzyme E1 (ECR1) 25 8.8
At4g32320 L-ascorbate peroxidase - like protein 25 8.8
>At3g05550 unknown protein
Length = 97
Score = 146 bits (368), Expect = 2e-36
Identities = 70/80 (87%), Positives = 74/80 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M E+KT+ E IRKWV DHKLRTVGCLWLSGITGSIAYNWS+P MKTSVKIIHARLHAQAL
Sbjct: 1 MVESKTKFEEIRKWVSDHKLRTVGCLWLSGITGSIAYNWSQPAMKTSVKIIHARLHAQAL 60
Query: 61 TLAALAGAAVVEYYDHRAEA 80
TLAALAGAAVVEYYDH+ EA
Sbjct: 61 TLAALAGAAVVEYYDHKTEA 80
>At5g27760 unknown protein
Length = 96
Score = 139 bits (349), Expect = 3e-34
Identities = 64/80 (80%), Positives = 74/80 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAE KT+V IR+W+++HKLRTVGCLWLSGI+GSIAYNWS+P MKTSV+IIHARLHAQAL
Sbjct: 1 MAEPKTKVAEIREWIIEHKLRTVGCLWLSGISGSIAYNWSKPAMKTSVRIIHARLHAQAL 60
Query: 61 TLAALAGAAVVEYYDHRAEA 80
TLAALAGAA VEYYDH++ A
Sbjct: 61 TLAALAGAAAVEYYDHKSGA 80
>At4g26140 putative beta-galactosidase
Length = 729
Score = 28.5 bits (62), Expect = 0.61
Identities = 18/62 (29%), Positives = 30/62 (48%), Gaps = 8/62 (12%)
Query: 27 WLSGITGSIAYN------WSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRAEA 80
W +G+ G + N W K S K I + +AL++ LAG++ VE+ + A
Sbjct: 558 WNTGVLGPVTLNGVNSGTWDMTKWKWSYKQIGTK--GEALSVHTLAGSSTVEWKEGSLVA 615
Query: 81 KR 82
K+
Sbjct: 616 KK 617
>At2g31875 poly(ADP-ribose) like glycohydrolase
Length = 548
Score = 28.1 bits (61), Expect = 0.80
Identities = 18/59 (30%), Positives = 28/59 (46%), Gaps = 11/59 (18%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLW----------LSGITGSIAYNWSRPNMKTSVKII 51
EA ++ + KW++ HK TVG LW L T ++W P++ T+ K I
Sbjct: 488 EALRNLDQVTKWILSHKW-TVGDLWNMMLEYSAQRLYKQTSVGFFSWLLPSLATTNKAI 545
>At1g06710 hypothetical protein
Length = 946
Score = 26.2 bits (56), Expect = 3.0
Identities = 17/56 (30%), Positives = 26/56 (46%), Gaps = 17/56 (30%)
Query: 8 VESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLA 63
+E RKW +++R VGC PN+ T +IHA L A+ ++ A
Sbjct: 493 IEQARKWF--NEMREVGCT---------------PNVVTYTALIHAYLKAKKVSYA 531
>At5g16030 unknown protein
Length = 339
Score = 25.8 bits (55), Expect = 4.0
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 2/41 (4%)
Query: 31 ITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVV 71
+ +IA N +PN +V++IH L+A L ALA VV
Sbjct: 157 LMNNIAQN--KPNNNPNVRVIHEDLYAPDPELLALAEKKVV 195
>At1g31817 30S ribosomal like protein S11
Length = 314
Score = 25.0 bits (53), Expect = 6.8
Identities = 13/50 (26%), Positives = 26/50 (52%), Gaps = 10/50 (20%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARL 55
+ ++ +R+ + D RT G I ++ RPN++T+ IIH ++
Sbjct: 161 SDLDFVREVIEDEGRRTAG----------IFSHFQRPNLETNADIIHIKM 200
>At5g19180 putative ubiquitin activating enzyme E1 (ECR1)
Length = 454
Score = 24.6 bits (52), Expect = 8.8
Identities = 15/64 (23%), Positives = 27/64 (41%), Gaps = 3/64 (4%)
Query: 13 KWVVDHKLRTVGCLWLSGITGSIAYNWSR---PNMKTSVKIIHARLHAQALTLAALAGAA 69
KWV D +R + G+T S+ + P + ++ II A + L + +
Sbjct: 254 KWVYDEAIRRAELFGIPGVTYSLTQGVVKNIIPAIASTNAIISAACALETLKIVSACSKT 313
Query: 70 VVEY 73
+V Y
Sbjct: 314 LVNY 317
>At4g32320 L-ascorbate peroxidase - like protein
Length = 329
Score = 24.6 bits (52), Expect = 8.8
Identities = 12/25 (48%), Positives = 16/25 (64%), Gaps = 3/25 (12%)
Query: 30 GITGSIAYNWSRP---NMKTSVKII 51
GI GSIAY RP +K S+K++
Sbjct: 136 GINGSIAYELERPENIGLKKSLKVL 160
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,689,696
Number of Sequences: 26719
Number of extensions: 47721
Number of successful extensions: 113
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 109
Number of HSP's gapped (non-prelim): 9
length of query: 88
length of database: 11,318,596
effective HSP length: 64
effective length of query: 24
effective length of database: 9,608,580
effective search space: 230605920
effective search space used: 230605920
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0299.14


