

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296b.2
(1291 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
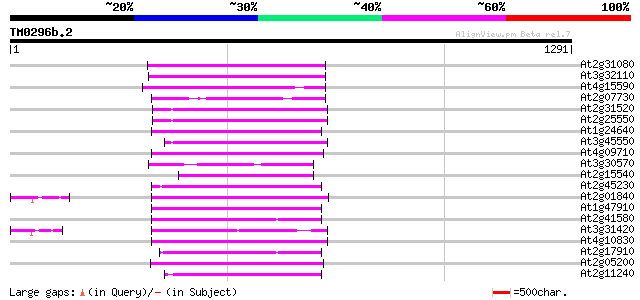
Score E
Sequences producing significant alignments: (bits) Value
At2g31080 putative non-LTR retroelement reverse transcriptase 327 3e-89
At3g32110 non-LTR reverse transcriptase, putative 323 4e-88
At4g15590 reverse transcriptase like protein 309 6e-84
At2g07730 putative non-LTR retroelement reverse transcriptase 236 8e-62
At2g31520 putative non-LTR retroelement reverse transcriptase 223 6e-58
At2g25550 putative non-LTR retroelement reverse transcriptase 223 7e-58
At1g24640 hypothetical protein 223 7e-58
At3g45550 putative protein 222 9e-58
At4g09710 RNA-directed DNA polymerase -like protein 220 4e-57
At3g30570 putative reverse transcriptase 220 4e-57
At2g15540 putative non-LTR retroelement reverse transcriptase 220 4e-57
At2g45230 putative non-LTR retroelement reverse transcriptase 219 6e-57
At2g01840 putative non-LTR retroelement reverse transcriptase 219 1e-56
At1g47910 reverse transcriptase, putative 215 2e-55
At2g41580 putative non-LTR retroelement reverse transcriptase 212 1e-54
At3g31420 hypothetical protein 211 2e-54
At4g10830 putative protein 210 4e-54
At2g17910 putative non-LTR retroelement reverse transcriptase 205 1e-52
At2g05200 putative non-LTR retroelement reverse transcriptase 202 1e-51
At2g11240 pseudogene 201 3e-51
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 327 bits (838), Expect = 3e-89
Identities = 172/411 (41%), Positives = 249/411 (59%), Gaps = 1/411 (0%)
Query: 317 TLTNTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEAS-QTPFPGD 375
T +T +I R+RN + LK D W +D +L+ AL Y++ L++ + + P
Sbjct: 216 TFFHTSTIIRRRRNRIESLKGDDDRWVTDKVELEAMALTYYKRLYSLEDVSEVRNMLPTG 275
Query: 376 NVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVV 435
+ E + L+Q +K EV A+ SM F APGPDG+QP F++Q WE VG +V + V
Sbjct: 276 GFASISEAEKAALLQAFTKAEVVSAVKSMGRFKAPGPDGYQPVFYQQCWETVGPSVTRFV 335
Query: 436 SEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLND 495
E FE+G + + L+VLI KV P I++FRP+SLCNV +K+ITK++V R++ ++
Sbjct: 336 LEFFETGVLPASTNDALLVLIAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISK 395
Query: 496 IVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRET 555
++GP Q SF+PGR ++DN L E +H M R RKG + K+DLEKAYD + W FL+ET
Sbjct: 396 LIGPAQASFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQET 455
Query: 556 LELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLS 615
LE IM V+ +S+LWNG SF P RGLRQGDP SPY FVLC+ERL
Sbjct: 456 LEAAGLSEGWTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLC 515
Query: 616 IHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVL 675
I+ V WKP+ + GG+ +SH+ FA D++LF +ASVAQ+ ++ L+ CE S
Sbjct: 516 HLIEASVGKREWKPIAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQ 575
Query: 676 RINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSE 726
+++++KSK S V+ E+ I + I +LGKYLG P+ +R+N E
Sbjct: 576 KVSLEKSKIFFSHNVSREMEQLISEESGIGCTKELGKYLGMPILQKRMNKE 626
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 323 bits (828), Expect = 4e-88
Identities = 168/408 (41%), Positives = 244/408 (59%), Gaps = 1/408 (0%)
Query: 320 NTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFA-GNQEASQTPFPGDNVP 378
+T +I R+RN + L+ +DG W S+ +L+ A+ Y++ L++ + +A P +
Sbjct: 895 HTSTIIRRRRNQIEMLQDNDGRWLSNAQELETHAIDYYKRLYSLDDLDAVVEQLPQEGFT 954
Query: 379 KLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEA 438
L E L +P S EV A+ SM + APGPDGFQP F++Q WE VG++V K V +
Sbjct: 955 ALSEADFSSLTKPFSPLEVEGAIRSMGKYKAPGPDGFQPVFYQQGWEVVGESVTKFVMDF 1014
Query: 439 FESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVG 498
F SG + ++LVVLI KV P I +FRPISLCNV +K ITK++V R++ +N ++G
Sbjct: 1015 FSSGSFPQETNDVLVVLIAKVLKPEKITQFRPISLCNVLFKTITKVMVGRLKGVINKLIG 1074
Query: 499 PMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLEL 558
P Q SF+PGR + DN + E++H M R KG + K+DLEKAYD I W L +TL+
Sbjct: 1075 PAQTSFIPGRLSTDNIVVVQEVVHSMRRKKGVKGWMLLKLDLEKAYDRIRWDLLEDTLKA 1134
Query: 559 YEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHI 618
P + IM V + LLWNG +FKP RGLRQGDP SPY FVLC+ERL I
Sbjct: 1135 AGLPGTWVQWIMKCVEGPSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLI 1194
Query: 619 QNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRIN 678
++ + +WKP+ + G +SH+ FA D++LF +AS+ Q+ VL L+ C S +++
Sbjct: 1195 ESSIAAKKWKPIKISQSGPRLSHICFADDLILFAEASIDQIRVLRGVLEKFCGASGQKVS 1254
Query: 679 VQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSE 726
++KSK S V ++ I + + DLGKYLG P+ +R+N E
Sbjct: 1255 LEKSKIYFSKNVLRDLGKRISEESGMKATKDLGKYLGVPILQKRINKE 1302
Score = 39.3 bits (90), Expect = 0.015
Identities = 32/119 (26%), Positives = 56/119 (46%), Gaps = 14/119 (11%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSFRLVD-----L 55
V+ + E + R LGF +F ++AVG SGG+W+L +VD +
Sbjct: 596 VLAIFETHAGGDQASRICQGLGFENSFRVDAVGHSGGLWLLWRTGIG-EVSVVDSTDQFI 654
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYL---IRVREDVSALGSCWGTLTK 111
HA+ V +D+++ V VY +P ++R +W L IR + +G + T+ +
Sbjct: 655 HAKDVN---GKDNVNLV--VVYAAPTASRRSGLWDRLGDVIRSMDGPVVIGGDFNTIVR 708
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 309 bits (792), Expect = 6e-84
Identities = 171/421 (40%), Positives = 248/421 (58%), Gaps = 18/421 (4%)
Query: 307 KLERTGSDWGTLTNTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFA-GNQ 365
KL G T +T VI R+RN + LK S+ W ++ L++ A+ Y++ L++ +
Sbjct: 158 KLLALGDRNTTFFHTSTVIRRRRNRIEMLKDSEDRWVTEKEALEKLAMDYYRKLYSLEDV 217
Query: 366 EASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWE 425
+ P + P+L + L +P +++EV A+ SM F APGPDG+QP F++Q WE
Sbjct: 218 SVVRGTLPTEGFPRLTREEKNNLNRPFTRDEVVVAVRSMGRFKAPGPDGYQPVFYQQCWE 277
Query: 426 QVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLL 485
VG++V K V E FESG + + ++L+VL+ KV P I +FRP+SLCNV +K+ITK++
Sbjct: 278 TVGESVSKFVMEFFESGVLPKSTNDVLLVLLAKVAKPERITQFRPVSLCNVLFKIITKMM 337
Query: 486 VNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYD 545
V R++ ++ ++GP Q SF+PGR + DN + E +H M R RKG + K+DLEKAYD
Sbjct: 338 VIRLKNVISKLIGPAQASFIPGRLSFDNIVVVQEAVHSMRRKKGRKGWMLLKLDLEKAYD 397
Query: 546 SISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPY 605
I W FL ETLE I IM V+ ++SLLWNG SF P RGLRQGDP SPY
Sbjct: 398 RIRWDFLAETLEAAGLSEGWIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPY 457
Query: 606 FFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDA 665
FVLC+ERL I+ V G WK + + GG +SH+ FA D++LF +ASVAQ
Sbjct: 458 LFVLCIERLCHQIETAVGRGDWKSISISQGGPKVSHVCFADDLILFAEASVAQ------- 510
Query: 666 LQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNS 725
+++++KSK S V+ ++ I I +LGKYLG P+ +R+N
Sbjct: 511 ----------KVSLEKSKIFFSNNVSRDLEGLITAETGIGSTRELGKYLGMPVLQKRINK 560
Query: 726 E 726
+
Sbjct: 561 D 561
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 236 bits (601), Expect = 8e-62
Identities = 145/401 (36%), Positives = 208/401 (51%), Gaps = 51/401 (12%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFA-GNQEASQTPFPGDNVPKLKEVVR 385
+KRN + K D W +D +L+ AL Y+ L++ + A P L + +
Sbjct: 108 KKRNRIESPKDGDNRWMTDKVELETMALNYYTRLYSLEDVSAVWNMLPIGGFATLTDGEK 167
Query: 386 FLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVD 445
L++P + EEV A+ M F A GP + ++A
Sbjct: 168 AALLKPFTSEEVVVAVKGMGRFKASGP-------------------YAATNDA------- 201
Query: 446 ERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFL 505
L+VLI KV P I +FRP+SLCNV +K+ITK++VNR++ + ++GP Q +F+
Sbjct: 202 ------LLVLIAKVAKPERITQFRPMSLCNVLFKIITKMIVNRLKKVILKLIGPAQANFI 255
Query: 506 PGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
PGR ++DN L E +H M R RKG + K+DLEKAYD I W+FL+ETL V
Sbjct: 256 PGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRIRWNFLQETLVATGLSEVW 315
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
IM V+ +S+LWNG SF P RGLRQGDP SPY FVLC+ERL I+ V
Sbjct: 316 THRIMAGVTDPSMSVLWNGGKTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGTW 375
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
WKP+ L C+ASVAQ+ ++ L+ CE S ++++KSK +
Sbjct: 376 DWKPM------------------ALSCEASVAQILIIRRVLEQFCEASGQNVSLEKSKIV 417
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSE 726
S V+ ++ I + I +LGKYLG P+ +R+N E
Sbjct: 418 FSNNVSRDMERLISGESGIGCTRELGKYLGMPILQKRMNKE 458
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 223 bits (568), Expect = 6e-58
Identities = 142/404 (35%), Positives = 202/404 (49%), Gaps = 3/404 (0%)
Query: 329 RNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQ-EASQTPFPGDNVPKLKEVVRFL 387
+N V+ + G + ++ A +F +F+ N + S F D + V
Sbjct: 521 QNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIFSTNGIKVSPIDF-ADFKSTVTNTVNLD 579
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + S E+ A+ + APGPDG F+K W+ VG V V + FE+ +
Sbjct: 580 LTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPS 639
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
+ + +IPK+ +PT++ ++RPI+LCNV YK+I+K LVNR++ LN IV Q +F+PG
Sbjct: 640 INHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPG 699
Query: 508 RGTMDNAFLA*EIIHQMH-RSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DN +A E++H + R K +A K D+ KAYD + W FL T+ L+ F I
Sbjct: 700 RIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWI 759
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
IM V S S+L NG+P P RG+RQGDP SPY F+LC + LS I G
Sbjct: 760 GWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGD 819
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ + G I+HL FA D L FC+A+V L D S +INVQKS
Sbjct: 820 LRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITF 879
Query: 687 SIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSERCEY 730
V ++++ IP GKYLG P Q R E EY
Sbjct: 880 GSRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEY 923
Score = 36.6 bits (83), Expect = 0.096
Identities = 29/102 (28%), Positives = 49/102 (47%), Gaps = 12/102 (11%)
Query: 5 MEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDLHAQV 59
+E Q +V + LGF G SGG+ ++ N+ +LS RL+D H
Sbjct: 190 LETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKNNVSLSLISQDERLIDSH--- 246
Query: 60 VTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSA 101
VTF ++ S+ S VY P ++R +W+ L + ++ +A
Sbjct: 247 VTF----NNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNA 284
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 223 bits (567), Expect = 7e-58
Identities = 142/404 (35%), Positives = 201/404 (49%), Gaps = 3/404 (0%)
Query: 329 RNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQ-EASQTPFPGDNVPKLKEVVRFL 387
+N V+ + G + ++ A +F +F+ N + S F D + V
Sbjct: 747 QNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIFSTNGIKVSPIDF-ADFKSTVTNTVNLD 805
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + S E+ A+ + APGPDG F+K W+ VG V V + FE+ +
Sbjct: 806 LTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPS 865
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
+ + +IPK+ +PT++ ++RPI+LCNV YK+I+K LVNR++ LN IV Q +F+PG
Sbjct: 866 INHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPG 925
Query: 508 RGTMDNAFLA*EIIHQMH-RSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DN +A E++H + R K +A K D+ KAYD + W FL T+ L+ F I
Sbjct: 926 RIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWI 985
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
IM V S S+L NG+P P RG+RQGDP SPY F+LC + LS I G
Sbjct: 986 GWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGD 1045
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ + G I+HL FA D L FC+A+V L D S +INVQKS
Sbjct: 1046 LRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITF 1105
Query: 687 SIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSERCEY 730
V ++ + IP GKYLG P Q R E EY
Sbjct: 1106 GSRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEY 1149
Score = 39.3 bits (90), Expect = 0.015
Identities = 30/106 (28%), Positives = 52/106 (48%), Gaps = 12/106 (11%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDL 55
++ L+E Q +V + LGF G SGG+ ++ N+ +LS RL+D
Sbjct: 412 ILFLVETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKNNVSLSLISQDERLIDS 471
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSA 101
H VTF ++ S+ S VY P ++R +W+ L + ++ +A
Sbjct: 472 H---VTF----NNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNA 510
>At1g24640 hypothetical protein
Length = 1270
Score = 223 bits (567), Expect = 7e-58
Identities = 139/393 (35%), Positives = 205/393 (51%), Gaps = 3/393 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R+R + +LK +G+ + + E A YF LF + + T + VP++ EV+
Sbjct: 326 RQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNLFKSSNPSGFTDWFSGLVPRVSEVMNE 385
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LV VS +E+++A+ S+K +APGPDG FF+ YW VG+ V V + F G +
Sbjct: 386 SLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFFQHYWSTVGNQVTSEVKKFFADGIMPA 445
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
+ LIPK HPT + + RPISLC+V YK+I+K++ R++P+L +IV Q++F+
Sbjct: 446 EWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIVSDTQSAFVS 505
Query: 507 GRGTMDNAFLA*EIIHQM--HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLV 564
R DN +A E++H + H ++ + +A K D+ KAYD + WS+LR L F L
Sbjct: 506 ERLITDNILVAHELVHSLKVHPRISSEF-MAVKSDMSKAYDRVEWSYLRSLLLSLGFHLK 564
Query: 565 IIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP 624
++ IM VSS S+L N P RGLRQGDP SP+ FVLC E L+ +
Sbjct: 565 WVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQWE 624
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
G + + + G + HL FA D L CKAS Q VL L+ + IN+ KS
Sbjct: 625 GALEGIQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKSSI 684
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
V +++ I I G YLG P
Sbjct: 685 TFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLP 717
>At3g45550 putative protein
Length = 851
Score = 222 bits (566), Expect = 9e-58
Identities = 139/377 (36%), Positives = 192/377 (50%), Gaps = 3/377 (0%)
Query: 356 YFQALFAGNQ-EASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDG 414
+F +F N + S F D + ++ L Q E+ +A+ + APGPDG
Sbjct: 13 FFTDIFTTNGIQVSPIDF-ADFPSSVTNIINSELTQDFRDSEIFEAICQIGDDKAPGPDG 71
Query: 415 FQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLC 474
F+KQ W+ VG+ V K V FES + + + +IPK+ +P ++ ++RPI+LC
Sbjct: 72 LTARFYKQCWDIVGNDVIKEVKLFFESSHMKTSVNHTNICMIPKIQNPQTLSDYRPIALC 131
Query: 475 NVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMH-RSMARKGN 533
NV YK+I+K +VNR++ LN IV Q +F+PGR DN +A EI+H + R K
Sbjct: 132 NVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSKTY 191
Query: 534 LAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPG 593
+A K D+ KAYD + W FL T+ L+ F I IM V S S+L NG+P P
Sbjct: 192 MAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYISPT 251
Query: 594 RGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCK 653
RG+RQGDP SPY F+LC + LS I+ G + + G I+HL FA D L FC+
Sbjct: 252 RGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGNGAPAITHLQFADDSLFFCQ 311
Query: 654 ASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKY 713
A+V L D S +INVQKS V +T + IP GKY
Sbjct: 312 ANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGGKY 371
Query: 714 LGFPLQGRRLNSERCEY 730
LG P Q R E Y
Sbjct: 372 LGLPEQFGRKKKEMFNY 388
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 220 bits (561), Expect = 4e-57
Identities = 134/398 (33%), Positives = 203/398 (50%), Gaps = 2/398 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R N++ ++ G + ++ YFQ +F + + P +
Sbjct: 304 RMLNNLSVIEDGSGQEFHEEEQIASTISSYFQNIFTTSNNSDLQVVQEALSPIISSHCNE 363
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L++ S E+++AL S+ + APGPDGF FF YW+ + V + + F +
Sbjct: 364 ELIKISSLLEIKEALFSISADKAPGPDGFSASFFHAYWDIIEADVSRDIRSFFVDSCLSP 423
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
RL E V LIPK++ P + ++RPI+LCNV YK++ K+L R++P+L++++ Q++F+P
Sbjct: 424 RLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVP 483
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARK-GNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
GR DN + EI+H + S A+K ++A K D+ KAYD I W+FL+E L F
Sbjct: 484 GRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDMSKAYDRIKWNFLQEVLMRLGFHDKW 543
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
I +M V + S L NG+P S P RGLRQGDP SPY F+LC E LS + E G
Sbjct: 544 IRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLSGLCRKAQEKG 603
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
+ G ++HL FA D + FCK + L + L+ S IN+ KS
Sbjct: 604 VMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSAIT 663
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQ-GRR 722
S +++ + L I +GKYLG P GRR
Sbjct: 664 FSSKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEHFGRR 701
>At3g30570 putative reverse transcriptase
Length = 1099
Score = 220 bits (561), Expect = 4e-57
Identities = 133/380 (35%), Positives = 196/380 (51%), Gaps = 43/380 (11%)
Query: 320 NTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFA-GNQEASQTPFPGDNVP 378
+T +I R+ N + LK G W S+T++L++ A+ Y++ L++ + + P
Sbjct: 409 HTSTIIRRRHNRIEMLKNEKGGWISNTNELEKLAIDYYKRLYSLDDVDEVVERLPAGRFL 468
Query: 379 KLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEA 438
L + + L +P S EV AL + + V +
Sbjct: 469 GLNQEEQIRLNKPFSAMEVESALK----------------------------ITRFVLDF 500
Query: 439 FESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVG 498
F SG + + + +VVLI KV P I +FRPISLCNV ++IT +V R++ + ++G
Sbjct: 501 FLSGNLPQETNDAMVVLIAKVGKPEKITQFRPISLCNVLLEIITNTMVERLKSVITKLIG 560
Query: 499 PMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLEL 558
P Q SF+PGR + DN + E +H M KG + K DLEKAYD + W FL +TLE+
Sbjct: 561 PSQTSFIPGRLSTDNIEIVEEDVHSMRCKKGHKGCMLLKFDLEKAYDRVRWDFLEDTLEI 620
Query: 559 YEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHI 618
E P +SLLWNG FK +GLRQGD S Y FVLCMERL I
Sbjct: 621 AEGP--------------SMSLLWNGEQKELFKARQGLRQGDLLSLYLFVLCMERLCHLI 666
Query: 619 QNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRIN 678
+ RWKP+ + G +SH+ FA D++LF +ASVAQV V+ L+ C S +++
Sbjct: 667 DGAIAGKRWKPICLSQSGPKLSHICFAYDLILFAEASVAQVRVIQSVLEKCCIASGQKVS 726
Query: 679 VQKSKAITSIGVAPEVRTEI 698
++KSK S V+ E+ +I
Sbjct: 727 LEKSKIYFSDNVSRELGQQI 746
Score = 37.7 bits (86), Expect = 0.043
Identities = 26/84 (30%), Positives = 38/84 (44%), Gaps = 9/84 (10%)
Query: 16 RFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSN----LSFRLVDLHAQVVTFEMWRDSLSW 71
R L F +F ++A G SGG+WVL + F +HA+VV + D +
Sbjct: 95 RICQGLDFENSFWVDAEGQSGGLWVLWRTCIGDVVIVEFAYQFIHARVVNGDAAVDLI-- 152
Query: 72 VCSAVYTSPIPAQRESMWRYLIRV 95
VY P P++R +W L V
Sbjct: 153 ---VVYAPPTPSRRSKLWDTLSAV 173
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 220 bits (561), Expect = 4e-57
Identities = 121/312 (38%), Positives = 175/312 (55%), Gaps = 1/312 (0%)
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + +S EEV++AL S+ APGPDG FF++ YW+ G + K+V +G DER
Sbjct: 371 LTKVISPEEVKRALFSLNPDKAPGPDGMTAFFYQHYWDLTGPDLIKLVQNFHSTGFFDER 430
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
L E + LIPK P + EFRPISLCNV+YK+I+K+L +R++ L +++ Q++F+
Sbjct: 431 LNETNICLIPKTERPRKMAEFRPISLCNVSYKVISKVLSSRLKRLLPELISETQSAFVAE 490
Query: 508 RGTMDNAFLA*EIIHQMHRSMA-RKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DN +A E H + + A +K +A K D+ KAYD + WSFLR + F +
Sbjct: 491 RLITDNILIAQENFHALRTNPACKKKYMAIKTDMSKAYDRVEWSFLRALMLKMGFAQKWV 550
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
D I+ +SS +L NG+P KP RG+RQGDP SP+ F+LC E L +++ GR
Sbjct: 551 DWIIFCISSVSYKILLNGSPKGFIKPSRGIRQGDPISPFLFILCTEALVAKLKDAEWHGR 610
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ L + SHL FA D L FCKA Q ++D L+ E S ++N KS +
Sbjct: 611 IQGLQISRASPSTSHLLFADDSLFFCKADPLQGKEIIDILRLYGEASGQQLNPDKSSVMF 670
Query: 687 SIGVAPEVRTEI 698
V +R I
Sbjct: 671 GHEVDNSIRNTI 682
Score = 34.3 bits (77), Expect = 0.48
Identities = 25/99 (25%), Positives = 43/99 (43%), Gaps = 12/99 (12%)
Query: 2 VILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDLH 56
+ L E K + ++ SLGF +E +G SGG+ + + + F RL+D+
Sbjct: 32 LFLSETKNDLVYLQNVQVSLGFDCLKTVEPIGNSGGLALFYSRDYPVKFIYVCDRLIDIE 91
Query: 57 AQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRV 95
+ D + VY P+ RE +W+ L R+
Sbjct: 92 TII-------DGNRVFITFVYGDPVVQYRELVWKRLTRI 123
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 219 bits (559), Expect = 6e-57
Identities = 132/393 (33%), Positives = 201/393 (50%), Gaps = 3/393 (0%)
Query: 327 RKRNSVHRLKLSDG-SWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVR 385
R +N + +L +G W SD L A YF+ LFA + P + + +
Sbjct: 363 RAQNRIQKLIDEEGREWTSDED-LGRVAEAYFKKLFASEDVGYTVEELENLTPLVSDQMN 421
Query: 386 FLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVD 445
L+ P++KEEV++A S+ PGPDG F ++Q+WE +GD + ++V F SG ++
Sbjct: 422 NNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMGDQITEMVQAFFRSGSIE 481
Query: 446 ERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFL 505
E + + + LIPK+ + +FRPISLCNV YK+I KL+ NR++ L ++ Q +F+
Sbjct: 482 EGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFV 541
Query: 506 PGRGTMDNAFLA*EIIHQM-HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLV 564
GR DN +A E++H + + + +A K D+ KAYD + W FL + + F
Sbjct: 542 KGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADH 601
Query: 565 IIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP 624
I LIM V S + +L NG P P RGLRQGDP SPY FV+C E L +Q+ +
Sbjct: 602 WIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQK 661
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
+ L G ISHL FA D + +CK + + ++ ++ S R+N KS
Sbjct: 662 NQITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSI 721
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
++ E R + + I G YLG P
Sbjct: 722 YFGKHISEERRCLVKRKLGIEREGGEGVYLGLP 754
Score = 35.4 bits (80), Expect = 0.21
Identities = 23/95 (24%), Positives = 43/95 (45%), Gaps = 2/95 (2%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSFRLVDLHAQVV 60
V+ L E K + + LGF +E +G SGG+ ++ +S + D +
Sbjct: 31 VIFLCETKKRRNYLENVVGHLGFFDLHTVEPIGKSGGLALMWKDSVQIKVLQSDKRL-ID 89
Query: 61 TFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRV 95
+W+D + + +Y P+ A+R +W L R+
Sbjct: 90 ALLIWQDK-EFYLTCIYGEPVQAERGELWERLTRL 123
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 219 bits (557), Expect = 1e-56
Identities = 135/408 (33%), Positives = 204/408 (49%), Gaps = 1/408 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
+ RN + + + G + + A YF LF Q + PK+ E +
Sbjct: 725 KSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTTTQTSDWEEIISGIAPKVTEQMNH 784
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L+Q V+ +EVR A+ ++ + APG DGF F+ W+ +G+ V +V FES +D
Sbjct: 785 ELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWDLIGNDVCLMVRHFFESDVMDN 844
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
++ + + LIPK+ P + ++RPISLC +YK+I+K+L+ R++ L D++ Q +F+P
Sbjct: 845 QINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVP 904
Query: 507 GRGTMDNAFLA*EIIHQM-HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
G+ DN +A E++H + R + G +A K D+ KAYD + W+FL + + F
Sbjct: 905 GQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRW 964
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
+ IM V+S +L NG+P P RG+RQGDP SPY F+ C E LS ++
Sbjct: 965 VKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNK 1024
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
+ + T ISHL FA D L FC+AS + L + E S +IN KS I
Sbjct: 1025 QIHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSII 1084
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSERCEYNFT 733
+ R + I V GKYLG P Q R E EY T
Sbjct: 1085 FGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVT 1132
Score = 43.5 bits (101), Expect = 8e-04
Identities = 36/143 (25%), Positives = 60/143 (41%), Gaps = 14/143 (9%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDL 55
++ L+E K Q R +GF ++ G SGG+ V ++ RLVDL
Sbjct: 393 MLFLIETKQQDNYTRDLGVKMGFEDMCIISPRGLSGGLVVYWKKHLSIQVISHDVRLVDL 452
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSALGSCWGTLTKSYNL 115
+ + F + S +Y PIP++R +W L RV S G + NL
Sbjct: 453 YVEYKNFNFY-------LSCIYGHPIPSERHHLWEKLQRVSAHRSGPWMMCGDFNEILNL 505
Query: 116 MRSEEELSYQAEQQNLQRFLMLV 138
+E++ + +LQ F ++
Sbjct: 506 --NEKKGGRRRSIGSLQNFTNMI 526
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 215 bits (547), Expect = 2e-55
Identities = 131/392 (33%), Positives = 195/392 (49%), Gaps = 1/392 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R RN + L +G W + +Q A+ YFQ LF G+ + + +
Sbjct: 147 RARNRITGLHDENGIWSIEDDDIQNIAVSYFQNLFTTANPQVFDEALGEVQVLITDRIND 206
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LL ++ EVR AL + APGPDG FF++ W + + +V+ + G D+
Sbjct: 207 LLTADATECEVRAALFMIHPEKAPGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDK 266
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
RL + LIPK PT + E RPISLCNV YK+I+K+L R++ L +++ Q++F+
Sbjct: 267 RLNTTNICLIPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVD 326
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARKGN-LAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
GR DN +A E+ H + + + K +A K D+ KAYD + W+F+ L F
Sbjct: 327 GRLISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKW 386
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
I IM +++ Q +L NG P P RGLRQGDP SPY F+LC E L +I+
Sbjct: 387 ISWIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQN 446
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
+ +SHL FA D L FCKA+ Q ++++ L+ S +IN KS
Sbjct: 447 LITGIKVATPSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQ 506
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
V ++ +I L I + +G YLG P
Sbjct: 507 FGHKVEDSIKADIKLILGIHNLGGMGSYLGLP 538
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 212 bits (540), Expect = 1e-54
Identities = 137/393 (34%), Positives = 205/393 (51%), Gaps = 3/393 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R NS+ L +G+ + + + A+ +F+ LF+ + +S ++ E +
Sbjct: 92 RVSNSLRFLVDENGNEQTVNREKGKIAVTFFEDLFSSSYPSSMDSVLEGFNKRVTEDMNQ 151
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L + V+++E+ +A+ S+ + +APGPDGF FF++ W V + + + F++G + E
Sbjct: 152 DLTKKVNEQEIYKAVFSINAESAPGPDGFTALFFQRQWPLVKNQIISDIELFFQTGILPE 211
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
+ LIPK+ P + + RPISLC+V YK+I+K+L R++ +L IV P Q++F+
Sbjct: 212 DWNHTHLCLIPKITKPARMADIRPISLCSVMYKIISKILSARLKKYLPVIVSPTQSAFVA 271
Query: 507 GRGTMDNAFLA*EIIHQMHRS-MARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
R DN LA EI+H + + K + FK D+ KAYD + W FL+ L F
Sbjct: 272 ERLVSDNIILAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKGILLALGFNSTW 331
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP- 624
I+ +M VSS S+L NG P P RGLRQGDP SP+ FVLC E L IHI N E
Sbjct: 332 INWMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEAL-IHILNQAEKI 390
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
G+ + G ++HL FA D LL CKAS + ++ L S IN +KS
Sbjct: 391 GKISGIQFNGTGPSVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQMINSEKSAI 450
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
V E + I R+ I GKYLG P
Sbjct: 451 TFGAKVNEETKQWIMNRSGIQTEGGTGKYLGLP 483
>At3g31420 hypothetical protein
Length = 1491
Score = 211 bits (537), Expect = 2e-54
Identities = 137/411 (33%), Positives = 208/411 (50%), Gaps = 38/411 (9%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEAS---QTPFPGDNVPKLKEV 383
R RN + ++ SDG C + + A+ YF L+ S F G K+ +
Sbjct: 546 RCRNRLLSVQDSDGDICRGDENIAKVAINYFDDLYKSTPNTSLRYADVFQGFQ-QKITDE 604
Query: 384 VRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGK 443
+ L++PV++ E+ +++ S+ P PDGF F++Q+W + V V+ FE +
Sbjct: 605 INEDLIRPVTELEIEESVFSVAPSRTPDPDGFTADFYQQFWPDIKQKVIDEVTRFFERSE 664
Query: 444 VDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNS 503
+DER + LIPKV PT+I +FRPI+LCNV+YK+I+K+LVNR++ L + Q +
Sbjct: 665 LDERHNHTNLCLIPKVETPTTIAKFRPIALCNVSYKIISKILVNRLKKHLGGAITENQAA 724
Query: 504 FLPGRGTMDNAFLA*EIIHQMHRSMARKGN--LAFKIDLEKAYDSISWSFLRETLELYEF 561
F+PGR +NA +A E+ + + ++ R+ N +A K D+ KAYD + W FL ET+ F
Sbjct: 725 FVPGRLITNNAIIAHEVYYAL-KARKRQANSYMALKTDITKAYDRLEWDFLEETMRQMGF 783
Query: 562 PLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNL 621
I+ IM V+ + S+L NG+P + KP RG+R GDP SPY F+LC E LS I+
Sbjct: 784 NTKWIERIMICVTMVRFSVLINGSPHGTIKPERGIRHGDPLSPYLFILCAEVLSHMIKQA 843
Query: 622 VEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQK 681
+ K + + G ISHL FA D + F A+ T +
Sbjct: 844 EINKKLKGIRLSTQGPFISHLLFADDSIFFTLANQRSCTAI------------------- 884
Query: 682 SKAITSIGVAPEVRTEIGLRALIPFVND--LGKYLGFPLQGRRLNSERCEY 730
E T+ +R L+ N+ GKYLG P Q + E Y
Sbjct: 885 ----------KEPDTKRRMRHLLGIHNEGGEGKYLGLPEQFNKKKKELFNY 925
Score = 44.3 bits (103), Expect = 5e-04
Identities = 34/126 (26%), Positives = 57/126 (44%), Gaps = 14/126 (11%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNL-----SFRLVDL 55
++ L E K Q R+R LGF + G SGG+ V N+S ++ + LVDL
Sbjct: 211 IICLSETKQQDDRIRDVGAELGFCNHVTVPPDGLSGGLVVFWNSSVDIFLCFSNSNLVDL 270
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSALGSCWGTLTKSYNL 115
H + + S+ S VY P P+ R +W L R+ + + G+ W + +
Sbjct: 271 HVK-------SNEGSFYLSFVYGHPNPSHRHHLWERLERL--NTTRQGTAWFIMGDFNEI 321
Query: 116 MRSEEE 121
+ + E+
Sbjct: 322 LSNREK 327
>At4g10830 putative protein
Length = 1294
Score = 210 bits (535), Expect = 4e-54
Identities = 139/411 (33%), Positives = 213/411 (51%), Gaps = 10/411 (2%)
Query: 327 RKRNSVHRL---KLSDGSWCSDTSKLQEEALGYFQALFAGN-QEASQTPFPGDNVPKLKE 382
+ R SV+RL K +G ++ A +F ++ N + S F G P + E
Sbjct: 722 KTRFSVNRLVTIKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVSIIDFAGFK-PIVTE 780
Query: 383 VVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESG 442
+ L + +S E+ A+ + APGPDG F+K WE VG V K V F +
Sbjct: 781 QINDDLTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTS 840
Query: 443 KVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQN 502
+ + + + +IPK+ +P ++ ++RPI+LCNV YK+I+K LV R++ L+ IV Q
Sbjct: 841 YMKQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQA 900
Query: 503 SFLPGRGTMDNAFLA*EIIHQMH-RSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEF 561
+F+PGR DN +A E++H + R + +A K D+ KAYD + W+FL T+ L+ F
Sbjct: 901 AFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGF 960
Query: 562 PLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNL 621
I IM V S S+L NG P + +P RG+RQGDP SPY F+LC + L+ I+N
Sbjct: 961 SETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNR 1020
Query: 622 VEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQK 681
V G + + G G++HL FA D L FC+++V L D S +IN+
Sbjct: 1021 VAEGDIRGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINM-- 1078
Query: 682 SKAITSIGVAPEVRTEIGLRALIPFVN--DLGKYLGFPLQGRRLNSERCEY 730
SK++ + G T+ L+ ++ + GKYLG P Q R + Y
Sbjct: 1079 SKSMITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNY 1129
Score = 33.5 bits (75), Expect = 0.81
Identities = 27/106 (25%), Positives = 48/106 (44%), Gaps = 15/106 (14%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLS-----FRLVDL 55
V+ L+E + + + LGF G SGG+ +L +S LS R +D+
Sbjct: 392 VLFLIETLNKCEVISNLASVLGFPNVITQPPQGHSGGLALLWKDSVRLSNLYQDDRHIDV 451
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRY---LIRVRED 98
H + +++++ S VY P ++R S+W + L + R D
Sbjct: 452 HISI-------NNINFYLSRVYGHPCQSERHSLWTHFENLSKTRND 490
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 205 bits (522), Expect = 1e-52
Identities = 133/376 (35%), Positives = 190/376 (50%), Gaps = 6/376 (1%)
Query: 344 SDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMS 403
SD K+ A +F+ LF + K+ + L+Q V++ EV A+ S
Sbjct: 361 SDKGKI---ASSFFENLFTSTYILTHNNHLEGLQAKVTSEMNHNLIQEVTELEVYNAVFS 417
Query: 404 MKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPT 463
+ +APGPDGF FF+Q+W+ V + + FE+G + + + LIPK+ P
Sbjct: 418 INKESAPGPDGFTALFFQQHWDLVKHQILTEIFGFFETGVLPQDWNHTHICLIPKITSPQ 477
Query: 464 SIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQ 523
+ + RPISLC+V YK+I+K+L R++ L IV Q++F+P R DN +A E+IH
Sbjct: 478 RMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLISDNILVAHEMIHS 537
Query: 524 MH-RSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLW 582
+ K ++AFK D+ KAYD + W FL + F I IM V+S S+L
Sbjct: 538 LRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIMNCVTSVSYSVLI 597
Query: 583 NGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVE-PGRWKPLHRTAGGTGISH 641
NG P P RG+RQGDP SP FVLC E L IHI N E G+ + ++H
Sbjct: 598 NGQPYGHIIPTRGIRQGDPLSPALFVLCTEAL-IHILNKAEQAGKITGIQFQDKKVSVNH 656
Query: 642 LFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLR 701
L FA D LL CKA+ + L+ L + S IN+ KS V +++ I R
Sbjct: 657 LLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNVDIQIKDWIKSR 716
Query: 702 ALIPFVNDLGKYLGFP 717
+ I GKYLG P
Sbjct: 717 SGISLEGGTGKYLGLP 732
Score = 39.7 bits (91), Expect = 0.011
Identities = 23/92 (25%), Positives = 41/92 (44%), Gaps = 2/92 (2%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSFRLVDLHAQVV 60
++ LME K V + + LG+ F +E G SGG+ + + + F D ++
Sbjct: 9 ILFLMETKNSQDFVYKVFCWLGYDFIHTVEPEGRSGGLAIFWKSHLEIEFLYAD--KNLM 66
Query: 61 TFEMWRDSLSWVCSAVYTSPIPAQRESMWRYL 92
++ + W S VY P+ R +W +L
Sbjct: 67 DLQVSSRNKVWFISCVYGLPVTHMRPKLWEHL 98
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 202 bits (513), Expect = 1e-51
Identities = 125/403 (31%), Positives = 197/403 (48%), Gaps = 5/403 (1%)
Query: 325 IIRKRNSVHRLKLSD---GSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLK 381
+ R R + +RL + + G + ++ + GYFQ +F + + P +
Sbjct: 253 VTRNRRTQNRLTVMEDINGVAQHEEHQISQIISGYFQQIFTSESDGDFSVVDEAIEPMVS 312
Query: 382 EVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFES 441
+ L + + EEV+ A+ S+ + APGPDGF F+ YW + V + + F S
Sbjct: 313 QGDNDFLTRIPNDEEVKDAVFSINASKAPGPDGFTAGFYHSYWHIISTDVGREIRLFFTS 372
Query: 442 GKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQ 501
R+ E + LIPK P + ++RPI+LCN+ YK++ K++ R++ L ++ Q
Sbjct: 373 KNFPRRMNETHIRLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLISENQ 432
Query: 502 NSFLPGRGTMDNAFLA*EIIHQMHRSMARKG-NLAFKIDLEKAYDSISWSFLRETLELYE 560
++F+PGR DN + E++H + S A+K ++A K D+ KAYD + W FL++ L+ +
Sbjct: 433 SAFVPGRVISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFG 492
Query: 561 FPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQN 620
F + ID ++ V+S S L NG P P RGLRQGDP SP F+LC E LS
Sbjct: 493 FHSIWIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTR 552
Query: 621 LVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQ 680
+ + + G ++HL FA D + F K+ L + L + S IN
Sbjct: 553 AQRLRQLPGVRVSINGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFH 612
Query: 681 KSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQ-GRR 722
KS S V+ ++ I GKYLG P GRR
Sbjct: 613 KSSVTFSSKTPRSVKGQVKRILKIRKEGGTGKYLGLPEHFGRR 655
>At2g11240 pseudogene
Length = 1044
Score = 201 bits (510), Expect = 3e-51
Identities = 118/363 (32%), Positives = 182/363 (49%), Gaps = 9/363 (2%)
Query: 356 YFQALFAGNQEASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGF 415
YFQ LF+ N+ G +KE ++ + + EE++ A S+ + APGPDGF
Sbjct: 226 YFQKLFSANE--------GARAATIKEAIKPFISPEQNPEEIKSACFSIHADKAPGPDGF 277
Query: 416 QPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCN 475
FF+ W VG + + F S + + + + LIPK+ + ++RPI+LC
Sbjct: 278 SASFFQSNWMTVGPNIVLEIQSFFSSSTLQPTINKTHITLIPKIQSLKRMVDYRPIALCT 337
Query: 476 VTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKG-NL 534
V YK+I+KLL R++P L +I+ Q++F+P R + DN + E +H + A K +
Sbjct: 338 VFYKIISKLLSRRLQPILQEIISENQSAFVPKRASNDNVLITHEALHYLKSLGAEKRCFM 397
Query: 535 AFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGR 594
A K ++ KAYD I W F++ ++ F I I+ +++ S L NG+ + P R
Sbjct: 398 AVKTNMSKAYDRIEWDFIKLVMQEMGFHQTWISWILQCITTVSYSFLLNGSAQGAVTPER 457
Query: 595 GLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKA 654
GLRQGDP SP+ F++C E LS + G L + G ++HL FA D + FC++
Sbjct: 458 GLRQGDPLSPFLFIICSEVLSGLCRKAQLDGSLLGLRVSKGNPRVNHLLFADDTIFFCRS 517
Query: 655 SVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYL 714
+ + L+ E S IN KS S ++TE I V LGKYL
Sbjct: 518 DLKSCKTFLCILKKYEEASGQMINKSKSAITFSRKTPDHIKTEAQQILGIQLVGGLGKYL 577
Query: 715 GFP 717
G P
Sbjct: 578 GLP 580
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.340 0.149 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,180,764
Number of Sequences: 26719
Number of extensions: 1122478
Number of successful extensions: 3135
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 2904
Number of HSP's gapped (non-prelim): 138
length of query: 1291
length of database: 11,318,596
effective HSP length: 111
effective length of query: 1180
effective length of database: 8,352,787
effective search space: 9856288660
effective search space used: 9856288660
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0296b.2


