

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0288.8
(846 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
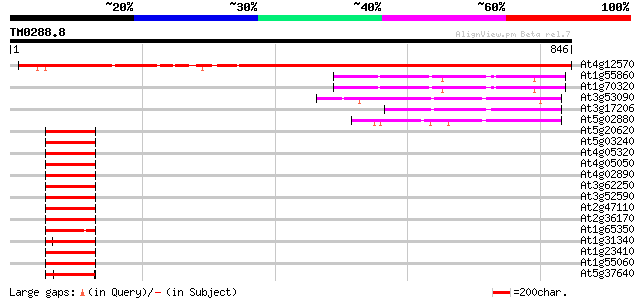
Score E
Sequences producing significant alignments: (bits) Value
At4g12570 polyubiquitin-like protein 767 0.0
At1g55860 ubiquitin-protein ligase 1, putative 173 4e-43
At1g70320 hypothetical protein 171 2e-42
At3g53090 putative protein 124 2e-28
At3g17206 putative ubiquitin ligase, 5'partial 115 1e-25
At5g02880 putative protein 86 9e-17
At5g20620 polyubiquitin 4 UBQ4 62 1e-09
At5g03240 polyubiquitin (ubq3) 62 1e-09
At4g05320 polyubiquitin (ubq10) 62 1e-09
At4g05050 62 1e-09
At4g02890 polyubiquitin 62 1e-09
At3g62250 ubiquitin extension protein (UBQ5) 62 1e-09
At3g52590 ubiquitin / ribosomal protein CEP52 62 1e-09
At2g47110 putative ubiquitin extension protein (UBQ6) 62 1e-09
At2g36170 putative ubiquitin/ribosomal protein CEP52 62 1e-09
At1g65350 62 1e-09
At1g31340 putative ubiquitin (AtRUB1) 62 1e-09
At1g23410 ubiquitin extension protein, putative 62 1e-09
At1g55060 ubiquitin-like (UBQ12) 62 2e-09
At5g37640 polyubiquitin 4 61 3e-09
>At4g12570 polyubiquitin-like protein
Length = 873
Score = 767 bits (1981), Expect = 0.0
Identities = 428/870 (49%), Positives = 561/870 (64%), Gaps = 61/870 (7%)
Query: 14 KRKHDEIDDDEGGVSD-IDPIRMKKDDA----AKAVYSAVHHRS---------------- 52
KRK DE D GV++ ++ ++ ++ DA A A + + RS
Sbjct: 28 KRKRDEDSSDYVGVAESLEMLKKQEIDADHMAASAQQTLISWRSGENSRSLSSSGECSSS 87
Query: 53 -----IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQS 107
LQ FVRMM G TIV+ A DTV+ +H+RI+ IP EQR+IY GKQLQ
Sbjct: 88 NRPESTRLQIFVRMMSGGKTIVIHAEKYDTVEKLHQRIEWKTKIPALEQRVIYKGKQLQR 147
Query: 108 EQTLAECSIGKDASLQLVGRMRSTGHPHAWQVINDMVSLVIRLCHNEPVHDANKTIKGLI 167
E +L SI +DASLQLV RM+ST HP AWQ I+D++ + R+ E + I I
Sbjct: 148 ENSLTYYSIEQDASLQLVARMQSTEHPVAWQTIDDIMYTISRMYKGE---NLQSNINEKI 204
Query: 168 TSYLNLSPKNDNDSASGYFQIFMSAT--AVLVSLYLSPYTGNKDCADSSIRHFLTTCKTT 225
++ + P ++S + Y IF +++ A LV LY S NK CA SS++ FL+ C
Sbjct: 205 VTFFAMIPVESDESIAKYLNIFSNSSVPAALVMLYASSLERNKSCAKSSVKLFLSNC--- 261
Query: 226 VSPPLHTQ--CARVVLDFCTLLKKRVSSEDPLYGFCRSTFGSLLETARVYCADSEGDNVK 283
V+ P + + C +VL+FC LL+K V + LY CR+T GS+LET DN
Sbjct: 262 VALPKNQKNYCLPIVLEFCKLLRK-VCPDQKLYVTCRNTLGSMLETF---------DNPH 311
Query: 284 GSILVQ------EIFPFVSELAKSLLVDLDLSIDSPSNAGTFLINVADFTAFLVPLRARI 337
G Q EIFPF +EL LL +L N+G + ++F LR I
Sbjct: 312 GVYNDQYETFGVEIFPFFTELTGLLLNEL------AQNSGPSFCDFQKVSSFWQQLRKVI 365
Query: 338 MEQQALRGSMPTQKRHKEALLVEEIEGLHLLYNDLLNKTDQCLQKLEQTWAGQGMMQGED 397
+ A +P + L EI LH L+ LL D C+ ++E + A + + E
Sbjct: 366 ELKVAF--PIPIVLPMQSTALEAEIRHLHRLFGSLLTTMDLCMCRVESSLADKEVGNSET 423
Query: 398 IYPAWSHYLSILKELYLISKLYDGAEEKLWKVLTRQRSVVCLLIVRYAKRTDEHQWILEH 457
+ +WS YLSILK + +S +Y GA+ +L +L + + L+V++AKR D+HQWI E+
Sbjct: 424 MSSSWSQYLSILKIINSMSNIYQGAKGQLAVMLNKNKVSFSALVVKFAKRGDDHQWIFEY 483
Query: 458 KCVTNFESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLFME 517
K TNFE+RRHLAM++FP+VKED+EE+HEMLIDRS LLSESFEYI A +LH GLFME
Sbjct: 484 KEATNFEARRHLAMLLFPDVKEDFEEMHEMLIDRSNLLSESFEYIVGASPEALHGGLFME 543
Query: 518 FKNEEATGPGVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFS 577
FKNEEATGPGVLREWF LVCQ +FNP+N LF+ +D RRF PN ASKV PLH ++F F+
Sbjct: 544 FKNEEATGPGVLREWFYLVCQEIFNPKNTLFLRSADDFRRFSPNPASKVDPLHPDFFEFT 603
Query: 578 GRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDS 637
GRVIALALMHKVQVG++ DRVFFLQLAG I+LED+KD D +Y+SCKQILEMD +F DS
Sbjct: 604 GRVIALALMHKVQVGVLFDRVFFLQLAGLKISLEDIKDTDRIMYNSCKQILEMDPEFFDS 663
Query: 638 DA-LGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHF 696
+A LGLTFV E EELG +ELCP GK VNSKNR++Y+DLLI+ RF T I EQV F
Sbjct: 664 NAGLGLTFVLETEELGKRDTIELCPDGKLKAVNSKNRKQYVDLLIERRFATPILEQVKQF 723
Query: 697 SKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITW 756
S+GFTD+LS S + FF+ L LEDLD ML G E+ IS++DWKAHTEYNG+KETD QI W
Sbjct: 724 SRGFTDMLSHSVPPRSFFKRLYLEDLDGMLRGGENPISIDDWKAHTEYNGFKETDRQIDW 783
Query: 757 FWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYIYKSPEPEDRLPSSHTCFYRM 816
FW+I+ +MT E+++ +LFFWTS K++PVEGF GL+S+LYIY+ E DRLP SHTCFYR+
Sbjct: 784 FWKILKKMTEEEQRSILFFWTSNKFVPVEGFRGLSSKLYIYRLYEANDRLPLSHTCFYRL 843
Query: 817 CFPAYSSMTVMQERLEVITQEHIGCSFGTW 846
C P Y ++T+M++RL +I Q+H+ SFG W
Sbjct: 844 CIPRYPTITLMEQRLRLIAQDHVSSSFGKW 873
>At1g55860 ubiquitin-protein ligase 1, putative
Length = 3891
Score = 173 bits (438), Expect = 4e-43
Identities = 117/362 (32%), Positives = 177/362 (48%), Gaps = 24/362 (6%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ +L +S+ + L L ++F+ EE G L REW+ L+ + +F+ L
Sbjct: 3533 VRRAYVLEDSYNQLRMRSPQDLKGRLNVQFQGEEGIDAGGLTREWYQLLSRVIFDKGALL 3592
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F ND F PN S HL YF F GR++A AL + + R F+ + G
Sbjct: 3593 FTTVGNDAT-FQPNPNSVYQTEHLSYFKFVGRMVAKALFDGQLLDVYFTRSFYKHILGVK 3651
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEE-----LGDTKVV--ELC 660
+T D++ DP Y + K +LE D SD L LTF + +E T+V EL
Sbjct: 3652 VTYHDIEAVDPDYYKNLKWLLENDV----SDILDLTFSMDADEEKHILYEKTEVTDYELK 3707
Query: 661 PGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELE 720
PGG+N+ V + + +Y+DL+ +I Q++ F +GF +++ F + LEL
Sbjct: 3708 PGGRNIRVTEETKHEYVDLVAGHILTNAIRPQINAFLEGFNELIPRELVSIFNDKELEL- 3766
Query: 721 DLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVK 780
++ G + I +D KA+TEY Y I WFWE+V + E L F T
Sbjct: 3767 ----LISGLPE-IDFDDLKANTEYTSYTAGSPVIHWFWEVVKAFSKEDMARFLQFVTGTS 3821
Query: 781 YLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVIT 835
+P+EGF L RL I+K+ +RLPS+HTCF ++ P Y S +QERL +
Sbjct: 3822 KVPLEGFKALQGISGPQRLQIHKAYGAPERLPSAHTCFNQLDLPEYQSKEQLQERLLLAI 3881
Query: 836 QE 837
E
Sbjct: 3882 HE 3883
>At1g70320 hypothetical protein
Length = 3658
Score = 171 bits (433), Expect = 2e-42
Identities = 116/362 (32%), Positives = 177/362 (48%), Gaps = 24/362 (6%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ +L +S+ + L L ++F+ EE G L REW+ L+ + +F+ L
Sbjct: 3300 VRRAYVLEDSYNQLRMRSPQDLKGRLNVQFQGEEGIDAGGLTREWYQLLSRVIFDKGALL 3359
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F ND F PN S HL YF F GR++A AL + + R F+ + G
Sbjct: 3360 FTTVGNDAT-FQPNPNSVYQTEHLSYFKFVGRMVAKALFDGQLLDVYFTRSFYKHILGVK 3418
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEE-----LGDTKVV--ELC 660
+T D++ DP Y + K +LE D SD L LTF + +E T+V EL
Sbjct: 3419 VTYHDIEAVDPDYYKNLKWLLENDV----SDILDLTFSMDADEEKHILYEKTEVTDYELK 3474
Query: 661 PGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELE 720
PGG+N+ V + + +Y+DL+ ++I Q++ F +G +++ F + LEL
Sbjct: 3475 PGGRNIRVTEETKHEYVDLVADHILTSAIRPQINAFLEGLNELIPRELVSIFNDKELEL- 3533
Query: 721 DLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVK 780
++ G + I +D KA+TEY Y I WFWE+V + E L F T
Sbjct: 3534 ----LISGLPE-IDFDDLKANTEYTSYTVGSPVIRWFWEVVKAFSKEDMARFLQFVTGTS 3588
Query: 781 YLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVIT 835
+P+EGF L RL I+K+ +RLPS+HTCF ++ P Y S +QERL +
Sbjct: 3589 KVPLEGFKALQGISGPQRLQIHKAYGSPERLPSAHTCFNQLDLPEYQSKEQVQERLLLAI 3648
Query: 836 QE 837
E
Sbjct: 3649 HE 3650
>At3g53090 putative protein
Length = 1142
Score = 124 bits (311), Expect = 2e-28
Identities = 94/387 (24%), Positives = 173/387 (44%), Gaps = 27/387 (6%)
Query: 463 FESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLFMEFKNEE 522
F S+ + M EV E+++ R ++ + F+ + + + L + + + F NE
Sbjct: 752 FISKDKASRKMAGEVDAPGARSIEIVVRRGHVVEDGFQQL-NSIGSRLKSSIHVSFVNES 810
Query: 523 ATGP------GVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGF 576
G+ +E+ + +A F + LF P R P+ +++ ++ F
Sbjct: 811 GLPEAGLDYGGLSKEFLTDITKAAFATEYGLFSQTPTSDRLLVPSPSARHLENGIQMIEF 870
Query: 577 SGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFID 636
GR++ AL + + VF +L G++ ++++ DP LY + + D D +
Sbjct: 871 LGRIVGKALYEGILLDYSFSHVFIQKLLGRYSFIDELSGLDPELYRNLMYVKHYDGDLKE 930
Query: 637 SDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHF 696
L L F E G ++EL PGGK+ V ++N+ +YI + + I + F
Sbjct: 931 ---LCLDFTVTEEFCGKMSIIELKPGGKDTSVTNENKMQYIHAMADYKLNRQIVPFSNAF 987
Query: 697 SKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQIT 755
+G TD++S + + F + + +L G I V+D + +T+Y GY ++ I
Sbjct: 988 YRGLTDLISPAWLKLF-----NAHEFNQLLSGGNHDIDVDDLRRNTKYTGGYSDSSRTIK 1042
Query: 756 WFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYIYK-----------SPEPED 804
FWE++ +R +LL F TS P+ GF L I+K + +
Sbjct: 1043 IFWEVMKGFEPSERCLLLKFVTSCSRAPLLGFKYLQPTFIIHKVSCDTSLWAAIGGQDVE 1102
Query: 805 RLPSSHTCFYRMCFPAYSSMTVMQERL 831
RLPS+ TC+ + P Y + M+E+L
Sbjct: 1103 RLPSASTCYNTLKLPTYKRASTMREKL 1129
>At3g17206 putative ubiquitin ligase, 5'partial
Length = 276
Score = 115 bits (288), Expect = 1e-25
Identities = 81/268 (30%), Positives = 125/268 (46%), Gaps = 10/268 (3%)
Query: 566 VHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCK 625
+H HL++F F G ++A A+ + V I F +L K+ L D+ DP LY
Sbjct: 4 IHEQHLQFFHFLGSLLAKAMFEGILVDIPFATFFLSKLKQKYNYLNDLPSLDPELYRHLI 63
Query: 626 QILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRF 685
+ D D L L FV E G+ EL PGG+++ V ++N +I L+ R
Sbjct: 64 FLKRYKGDISD---LELYFVILNNEYGERTEEELLPGGQDMRVTNENVITFIHLVSNHRL 120
Query: 686 VTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEY- 744
I +Q SHF +GF ++ F L++ ++ GS D++ ++D + +T Y
Sbjct: 121 NFQIRQQSSHFLRGFQQLIPKEWIDMFNEHELQV-----LISGSVDSLDIDDLRNNTNYA 175
Query: 745 NGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYIYK-SPEPE 803
GY I FWE++ + E +K L F T P+ GF L I S E
Sbjct: 176 GGYHAGHYVIDMFWEVMKSFSTENQKKFLKFVTGCSRGPLLGFKYLEPAFCIQSASNESV 235
Query: 804 DRLPSSHTCFYRMCFPAYSSMTVMQERL 831
DRLP+S TC + P Y S +++ +L
Sbjct: 236 DRLPTSATCMNLLKLPPYQSKELLETKL 263
>At5g02880 putative protein
Length = 1502
Score = 85.9 bits (211), Expect = 9e-17
Identities = 89/355 (25%), Positives = 149/355 (41%), Gaps = 47/355 (13%)
Query: 516 MEFKNEEATGPGVLREWFLLVCQALFNPQNALF-------VACPNDRRR-------FYPN 561
+E+ E TG G E++ LV +A NP ++ V P + +P
Sbjct: 1142 VEYSEEVGTGLGPTLEFYTLVSRAFQNPDLGMWRNDCSFIVGKPVEHSGVLASSSGLFPR 1201
Query: 562 HASKVHPLH--LEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPF 619
S L+ F G V+A AL + + L + F+ + G+ ++ D+ DP
Sbjct: 1202 PWSGTSTTSDVLQKFVLLGTVVAKALQDGRVLDLPLSKAFYKLILGQELSSFDIHFVDPE 1261
Query: 620 LYSSCKQILEMDS----DFIDSDALGLTFVREVE-ELGDTKVVELC-------------- 660
L CK ++E+ + + ++A G + + + TK+ +LC
Sbjct: 1262 L---CKTLVELQALVRRKKLFAEAHGDSGAAKCDLSFHGTKIEDLCLEFALPGYTDYDLA 1318
Query: 661 PGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELE 720
P N +VN N E+YI ++ I +QV F GF + S + F E
Sbjct: 1319 PYSANDMVNLDNLEEYIKGIVNATVCNGIQKQVEAFRSGFNQVFSIEHLRIF-----NEE 1373
Query: 721 DLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSV 779
+L+ ML G D S+ + H ++ +GY + + + +I+ EQ++ L F T
Sbjct: 1374 ELETMLCGECDLFSMNEVLDHIKFDHGYTSSSPPVEYLLQILHEFDREQQRAFLQFVTGS 1433
Query: 780 KYLPVEGFCGLASRLYIYK---SPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERL 831
LP G L+ +L I + S + LPS TC + P YSS M+E+L
Sbjct: 1434 PRLPHGGLASLSPKLTIVRKHGSDSSDTDLPSVMTCANYLKLPPYSSKEKMKEKL 1488
>At5g20620 polyubiquitin 4 UBQ4
Length = 382
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 229 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 287
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 288 NIQKESTLHLVLRLR 302
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 305 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 363
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 364 NIQKESTLHLVLRLR 378
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 135
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 136 NIQKESTLHLVLRLR 150
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 153 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 211
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 212 NIQKESTLHLVLRLR 226
>At5g03240 polyubiquitin (ubq3)
Length = 306
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 135
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 136 NIQKESTLHLVLRLR 150
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 153 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 211
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 212 NIQKESTLHLVLRLR 226
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 229 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 287
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 288 NIQKESTLHLVLRLR 302
>At4g05320 polyubiquitin (ubq10)
Length = 464
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 153 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 211
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 212 NIQKESTLHLVLRLR 226
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 229 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 287
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 288 NIQKESTLHLVLRLR 302
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 135
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 136 NIQKESTLHLVLRLR 150
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 305 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 363
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 364 NIQKESTLHLVLRLR 378
>At4g05050
Length = 229
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 153 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 211
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 212 NIQKESTLHLVLRLR 226
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 135
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 136 NIQKESTLHLVLRLR 150
>At4g02890 polyubiquitin
Length = 229
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 153 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 211
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 212 NIQKESTLHLVLRLR 226
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 135
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 136 NIQKESTLHLVLRLR 150
>At3g62250 ubiquitin extension protein (UBQ5)
Length = 157
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
>At3g52590 ubiquitin / ribosomal protein CEP52
Length = 128
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
>At2g47110 putative ubiquitin extension protein (UBQ6)
Length = 157
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
>At2g36170 putative ubiquitin/ribosomal protein CEP52
Length = 128
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
>At1g65350
Length = 294
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 58.9 bits (141), Expect = 1e-08
Identities = 30/75 (40%), Positives = 50/75 (66%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ + IP +QRLI+ GKQL+ +TLA+
Sbjct: 152 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEWIPPDQQRLIFAGKQLEDGRTLADY 210
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 211 NIQKESTLHLVLRLR 225
Score = 58.2 bits (139), Expect = 2e-08
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 2/75 (2%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLAD- 134
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 135 NIQKESTLHLVLRLR 149
Score = 42.4 bits (98), Expect = 0.001
Identities = 20/52 (38%), Positives = 33/52 (63%), Gaps = 1/52 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQ 106
+Q FV+ + G TI ++ T+ ++ +IQ +GIP +QRLI+ GKQL+
Sbjct: 228 MQIFVKTL-TGKTITLEVESSGTIDNVKAKIQDKEGIPPDQQRLIFAGKQLE 278
>At1g31340 putative ubiquitin (AtRUB1)
Length = 156
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
Score = 50.4 bits (119), Expect = 4e-06
Identities = 26/65 (40%), Positives = 38/65 (58%)
Query: 65 GNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAECSIGKDASLQL 124
G I + P DT+ I ER++ +GIP +QRLIY GKQL ++T + +I + L L
Sbjct: 86 GKEIEIDIEPTDTIDRIKERVEEKEGIPPVQQRLIYAGKQLADDKTAKDYNIEGGSVLHL 145
Query: 125 VGRMR 129
V +R
Sbjct: 146 VLALR 150
>At1g23410 ubiquitin extension protein, putative
Length = 156
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 60 NIQKESTLHLVLRLR 74
>At1g55060 ubiquitin-like (UBQ12)
Length = 230
Score = 61.6 bits (148), Expect = 2e-09
Identities = 31/75 (41%), Positives = 51/75 (67%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 77 MQIFVKTL-TGKTITLEVESSDTIDNLKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 135
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 136 NIQKESTLHLVLRLR 150
Score = 57.0 bits (136), Expect = 4e-08
Identities = 27/75 (36%), Positives = 50/75 (66%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q F++ + G T V++ DT+ ++ +IQ ++GIP + RLI+ GKQL+ +TLA+
Sbjct: 1 MQIFLKTL-TGKTKVLEVESSDTIDNVKAKIQDIEGIPPDQHRLIFAGKQLEDGRTLADY 59
Query: 115 SIGKDASLQLVGRMR 129
++ +D++L L+ R R
Sbjct: 60 NVQEDSTLHLLLRFR 74
Score = 56.2 bits (134), Expect = 7e-08
Identities = 29/75 (38%), Positives = 49/75 (64%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GI +QRLI+ GKQ + +TLA+
Sbjct: 153 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGISPDQQRLIFAGKQHEDGRTLADY 211
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 212 NIQKESTLHLVLRLR 226
>At5g37640 polyubiquitin 4
Length = 322
Score = 60.8 bits (146), Expect = 3e-09
Identities = 30/75 (40%), Positives = 50/75 (66%), Gaps = 1/75 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +G+P +QRLI+ GKQL +TLA+
Sbjct: 79 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGVPPDQQRLIFAGKQLDDGRTLADY 137
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 138 NIQKESTLHLVLRLR 152
Score = 57.0 bits (136), Expect = 4e-08
Identities = 30/74 (40%), Positives = 48/74 (64%), Gaps = 1/74 (1%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FVR + TI ++ DT ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 155 MQIFVRTLTR-KTIALEVESSDTTDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 213
Query: 115 SIGKDASLQLVGRM 128
+I K+++L LV R+
Sbjct: 214 NIQKESTLHLVLRL 227
Score = 54.7 bits (130), Expect = 2e-07
Identities = 27/63 (42%), Positives = 42/63 (65%)
Query: 67 TIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAECSIGKDASLQLVG 126
TI + DT+ ++ +IQ ++GIP+ +QRLI++GK L +TLA+ SI KD+ L L
Sbjct: 14 TITIDVVSSDTINNVKAKIQDIEGIPLDQQRLIFSGKLLDDGRTLADYSIQKDSILHLAL 73
Query: 127 RMR 129
R+R
Sbjct: 74 RLR 76
Score = 48.1 bits (113), Expect = 2e-05
Identities = 28/76 (36%), Positives = 47/76 (61%), Gaps = 2/76 (2%)
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQ-SEQTLAE 113
+Q FV + G TI ++ DT+ ++ +IQ + I +QRLI+ G+QL+ TLA+
Sbjct: 231 MQIFVNTL-TGKTITLEVESSDTIDNVKAKIQDKERIQPDQQRLIFAGEQLEDGYYTLAD 289
Query: 114 CSIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 290 YNIQKESTLHLVLRLR 305
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,091,185
Number of Sequences: 26719
Number of extensions: 827925
Number of successful extensions: 2097
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1990
Number of HSP's gapped (non-prelim): 86
length of query: 846
length of database: 11,318,596
effective HSP length: 108
effective length of query: 738
effective length of database: 8,432,944
effective search space: 6223512672
effective search space used: 6223512672
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0288.8


