

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0284.4
(247 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
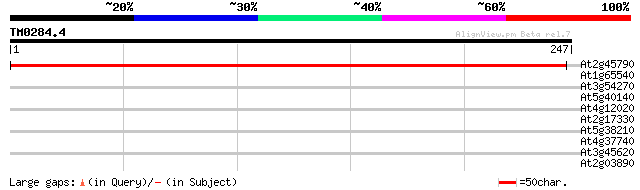
Score E
Sequences producing significant alignments: (bits) Value
At2g45790 putative phosphomannomutase 419 e-118
At1g65540 hypothetical protein 29 2.0
At3g54270 sucrose-phosphatase (SPP3) 29 2.7
At5g40140 putative protein 28 3.5
At4g12020 putative disease resistance protein 28 3.5
At2g17330 putative obtusifoliol 14-alpha demethylase 28 3.5
At5g38210 protein kinase - like protein 28 4.5
At4g37740 transcription activator (GRL2) 28 4.5
At3g45620 unknown protein 28 5.9
At2g03890 unknown protein 27 7.7
>At2g45790 putative phosphomannomutase
Length = 246
Score = 419 bits (1078), Expect = e-118
Identities = 201/245 (82%), Positives = 225/245 (91%)
Query: 1 MAVAKPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGST 60
MA PGVIALFDVDGTLTAPRK A E+L F++ELRKVVT+GVVGGSDL KISEQLG T
Sbjct: 1 MAAKIPGVIALFDVDGTLTAPRKEATPELLDFIRELRKVVTIGVVGGSDLSKISEQLGKT 60
Query: 61 VTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTF 120
VT+DYDY FSENGLVAHK GK IG +SLK LGD+KLKE INFTLHYIADLDIPIKRGTF
Sbjct: 61 VTNDYDYCFSENGLVAHKDGKSIGIQSLKLHLGDDKLKELINFTLHYIADLDIPIKRGTF 120
Query: 121 IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQIS 180
IEFR+GMLNVSPIGRNCSQEERDEFE+YDKVQNIRPKMV+ LRE+FAHLNLTFSIGGQIS
Sbjct: 121 IEFRNGMLNVSPIGRNCSQEERDEFERYDKVQNIRPKMVAELRERFAHLNLTFSIGGQIS 180
Query: 181 FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
FDVFP+GWDKTYCL+YL+ F+EIHFFGDKTY+GGND+EIYES +TIGH+VTSP+DT+ +C
Sbjct: 181 FDVFPKGWDKTYCLQYLEDFSEIHFFGDKTYEGGNDYEIYESPKTIGHSVTSPDDTVAKC 240
Query: 241 KSLFL 245
K+LF+
Sbjct: 241 KALFM 245
>At1g65540 hypothetical protein
Length = 398
Score = 29.3 bits (64), Expect = 2.0
Identities = 25/93 (26%), Positives = 48/93 (50%), Gaps = 7/93 (7%)
Query: 13 DVDGTLTAPRK---VANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTV--THDY-D 66
D+DG +T R+ V+N E+LGF + +T+ + S L+ + + +G + T Y
Sbjct: 266 DLDGFMTKVRRGVGVSNDEILGFAKLFNDELTLDNINRSRLVNMCKYMGISPFGTDAYLR 325
Query: 67 YVFSENGLVAHKQGKLIGTESLKTFLGDEKLKE 99
Y+ + K KLI E +++ L + +L++
Sbjct: 326 YMLRKRLQEIKKDDKLIKAEGVES-LSEAELRQ 357
>At3g54270 sucrose-phosphatase (SPP3)
Length = 425
Score = 28.9 bits (63), Expect = 2.7
Identities = 33/141 (23%), Positives = 57/141 (40%), Gaps = 13/141 (9%)
Query: 107 YIADLDIPIKRGTFIEFRSGMLNVSPIGRNCSQEERDEF--EKYDKVQNIRPKMVSVLRE 164
+ A LD R +E + P +E + F + D V+ I + +L E
Sbjct: 96 WTARLDYKWNRDIVVEETLKFPKLEPQPDKSQEEHKVSFFVGREDAVE-IMKVLPGILEE 154
Query: 165 KFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYK------GGNDHE 218
+ + L +S G +FDV P+G K L YL +++ G + GND E
Sbjct: 155 RGVDVKLVYSNG--YAFDVLPRGAGKQGALTYL--LDKLDIEGKQPSNTLVCGDSGNDAE 210
Query: 219 IYESERTIGHTVTSPEDTIKQ 239
++ G V++ + + Q
Sbjct: 211 LFNISDVYGVMVSNSHEELLQ 231
>At5g40140 putative protein
Length = 550
Score = 28.5 bits (62), Expect = 3.5
Identities = 20/83 (24%), Positives = 40/83 (48%), Gaps = 2/83 (2%)
Query: 42 VGVVGGSDLIKISEQLGSTVT-HDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEF 100
+GV+GG + + ++G+ +T HD LV +GKL+ +++ LG L +
Sbjct: 348 IGVLGGLEPLLHLIRVGTELTRHDSALALYHLSLVQSNRGKLVKLGAVQMLLGMVSLGQM 407
Query: 101 INFTLHYIADL-DIPIKRGTFIE 122
I L + ++ P+ R ++
Sbjct: 408 IGRVLLILCNMASCPVSRPALLD 430
>At4g12020 putative disease resistance protein
Length = 1895
Score = 28.5 bits (62), Expect = 3.5
Identities = 18/53 (33%), Positives = 25/53 (46%), Gaps = 1/53 (1%)
Query: 146 EKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLD 198
EK D + K + K HL T ++ G IS + FP + CLR+LD
Sbjct: 1375 EKLDLENSRHLKNLPTSIYKLKHLE-TLNLSGCISLERFPDSSRRMKCLRFLD 1426
>At2g17330 putative obtusifoliol 14-alpha demethylase
Length = 473
Score = 28.5 bits (62), Expect = 3.5
Identities = 13/29 (44%), Positives = 17/29 (57%)
Query: 156 PKMVSVLREKFAHLNLTFSIGGQISFDVF 184
PK+ SV K H N+TF IG ++S F
Sbjct: 69 PKLGSVFTVKLLHKNITFLIGPEVSSHFF 97
>At5g38210 protein kinase - like protein
Length = 686
Score = 28.1 bits (61), Expect = 4.5
Identities = 21/92 (22%), Positives = 41/92 (43%), Gaps = 4/92 (4%)
Query: 63 HDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIE 122
+ + V SE L++ K+ I L + + + N +H +ADL + R +
Sbjct: 542 YSFGVVLSE--LISSKEAVDITRHRHDINLANMAISKIQNDAVHELADLSLGFARDPSV- 598
Query: 123 FRSGMLNVSPIGRNCSQEERDEFEKYDKVQNI 154
+ M +V+ + C Q+ERD D++ +
Sbjct: 599 -KKMMSSVAELAFRCLQQERDVRPSMDEIVEV 629
>At4g37740 transcription activator (GRL2)
Length = 535
Score = 28.1 bits (61), Expect = 4.5
Identities = 21/85 (24%), Positives = 34/85 (39%), Gaps = 5/85 (5%)
Query: 148 YDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLDGFNEIHFFG 207
Y N++PK V+ +K + N F G S + KTY +LD +
Sbjct: 334 YPSTVNLQPKESPVIHQKHRNNNNPFEFGHISSDSLLNPNTAKTYGSSFLDFSS-----N 388
Query: 208 DKTYKGGNDHEIYESERTIGHTVTS 232
+ + G ++H + E T T S
Sbjct: 389 QEKHSGNHNHNSWPEELTSDWTQLS 413
>At3g45620 unknown protein
Length = 481
Score = 27.7 bits (60), Expect = 5.9
Identities = 17/52 (32%), Positives = 23/52 (43%), Gaps = 6/52 (11%)
Query: 183 VFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGG------NDHEIYESERTIGH 228
V P T+C R+L N +H G K G ND IY E+ +G+
Sbjct: 242 VLPDAPVNTFCPRHLRETNSVHITGLAYSKAGELLVSYNDELIYLFEKNMGY 293
>At2g03890 unknown protein
Length = 650
Score = 27.3 bits (59), Expect = 7.7
Identities = 11/37 (29%), Positives = 21/37 (56%)
Query: 123 FRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMV 159
F +G ++ + + +EE DE E+ DK +N P ++
Sbjct: 502 FSNGRSSLGKLEESIKEEEEDEEEEEDKTENTVPMII 538
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.140 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,639,081
Number of Sequences: 26719
Number of extensions: 247508
Number of successful extensions: 503
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 500
Number of HSP's gapped (non-prelim): 10
length of query: 247
length of database: 11,318,596
effective HSP length: 97
effective length of query: 150
effective length of database: 8,726,853
effective search space: 1309027950
effective search space used: 1309027950
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0284.4


