

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.6
(561 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
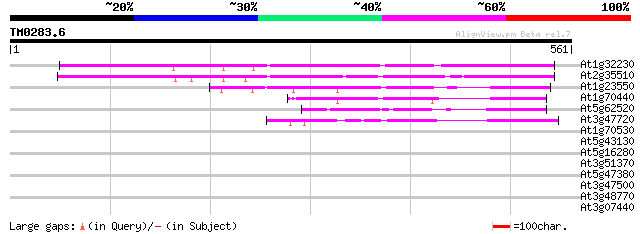
Score E
Sequences producing significant alignments: (bits) Value
At1g32230 ATP8 325 3e-89
At2g35510 unknown protein 304 1e-82
At1g23550 unknown protein 132 5e-31
At1g70440 hypothetical protein 107 2e-23
At5g62520 unknown protein 97 3e-20
At3g47720 unknown protein 89 5e-18
At1g70530 putative protein kinase 38 0.017
At5g43130 unkown protein 31 1.6
At5g16280 putative protein 31 1.6
At3g51370 protein phosphatase 2C like protein 31 1.6
At5g47380 putative protein 31 2.1
At3g47500 H-protein promoter binding factor-2a 30 3.6
At3g48770 hypothetical protein 29 6.1
At3g07440 unknown protein 29 6.1
>At1g32230 ATP8
Length = 588
Score = 325 bits (834), Expect = 3e-89
Identities = 188/517 (36%), Positives = 289/517 (55%), Gaps = 34/517 (6%)
Query: 50 KRKMNILGSGSGQSYLRFYLNYKKSARPERMMFYNHGEWVDYPKDIVDLVKKDFEIKNAA 109
+ K++ + SG++ +R+Y +KK+ +R+M Y +GEW D P+ ++ ++ + E K+AA
Sbjct: 62 ENKLSAYENRSGKALVRYYTYFKKTGIAKRVMMYENGEWNDLPEHVICAIQNELEEKSAA 121
Query: 110 IGIRLNGRDLMLDFLHACQMDLKTGFIQPIAWIDKVGGYFFPEVDVASGEESY------- 162
I +L G +LDFLH ++D++TG P+AWID G FFPE+ + +Y
Sbjct: 122 IEFKLCGHSFILDFLHMQRLDMETGAKTPLAWIDNAGKCFFPEIYESDERTNYCHHKCVE 181
Query: 163 ----NFGEQEGMKFHVEIKMNMVDEFQLRECIGES-NALVKDAQVKVRNMMSIMD----- 212
N ++ +++ L EC ES + ++ D + R+ D
Sbjct: 182 DPKQNAPHDIKLRLEIDVNGGETPRLNLEECSDESGDNMMDDVPLAQRSSNEHYDEATED 241
Query: 213 -CGY-IGEEIAKNEELGLSAYTGY-VHGNLVLD--SIRDLFCNGMSIIGNNDFEIVEIYR 267
C + ++K +E +G + G+ VLD +++ +F G + +G+ ++++ R
Sbjct: 242 SCSRKLEAAVSKWDETDAIVVSGAKLTGSEVLDKDAVKKMFAVGTASLGH--VPVLDVGR 299
Query: 268 CSCASMQVQSKLFLTQAEITKELHGDANVRYAWLSFSKGELSTMMEHGLGHCELFTTKCT 327
S + + LF Q EITK+ GDANVRYAWL + LS +M GLG F K
Sbjct: 300 FSSEIAEARLALFQKQVEITKKHRGDANVRYAWLPAKREVLSAVMMQGLGVGGAFIRKSI 359
Query: 328 YGVGVHLAAISCPYASARYCDIDGNGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSE 387
YGVG+HL A CPY SARYCD+D NGVR+++LCR+IMGNMELL QF E
Sbjct: 360 YGVGIHLTAADCPYFSARYCDVDENGVRYMVLCRVIMGNMELL----RGDKAQFFSGGEE 415
Query: 388 YDNGVDDIQCPRYYTIWNMNISTYICPAFVVSFKVSRGVEGYFCGTVDKNNVSRDNSSGA 447
YDNGVDDI+ P+ Y +WN+N++T+I P FVV FK+S + N +++ ++SG
Sbjct: 416 YDNGVDDIESPKNYIVWNINMNTHIFPEFVVRFKLSN------LPNAEGNLIAKRDNSGV 469
Query: 448 NSDSHCSGDPVQSASSAYNEIAGNGVACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLIK 507
+ P ++ A + N V + PKSPWMP L AAI+ KV NDM LI
Sbjct: 470 TLEGPKDLPPQLESNGARGSGSANSVGSSTTRPKSPWMPFPTLFAAISHKVAENDMLLIN 529
Query: 508 ENFELYKAKQISRDDFVKEMRLIVGDTLLRDTITTLQ 544
+++ + K+++R +FV+++R+IVGD LLR TITTLQ
Sbjct: 530 ADYQQLRDKKMTRAEFVRKLRVIVGDDLLRSTITTLQ 566
>At2g35510 unknown protein
Length = 568
Score = 304 bits (778), Expect = 1e-82
Identities = 192/519 (36%), Positives = 281/519 (53%), Gaps = 35/519 (6%)
Query: 48 DYKRKMNILGSGSGQSYLRFYLNYKKSARPERMMFYNHGEWVDYPKDIVDLVKKDFEIKN 107
D + K+ + + +S +R++ YKK+ P+R+MF+ +GEW+D P I+ ++ D E K
Sbjct: 59 DGENKVIVSENHVEKSLVRYFSYYKKTGVPKRVMFHENGEWIDLPDHILCDIRNDLEAKR 118
Query: 108 AAIGIRLNGRDLMLDFLHACQMDLKTGFIQPIAWIDKVGGYFFPEV-DVASGEESYNF-- 164
A I GR +LDFLH ++DL+TG +AWID G FFPE D + ++
Sbjct: 119 ATIEFNWCGRHFLLDFLHMYRLDLETGVKTQLAWIDIAGKCFFPETFDTLERDGCHHIRG 178
Query: 165 -----GEQEGMKFHVEIKMNM--VDEFQLRECIGESNALVKDAQVKVRNMMSIMD----- 212
+Q +K H+EI +N + L ES + D Q R+ D
Sbjct: 179 EDPEQHDQREIKLHIEIDVNSGELPRLNLNVVTDESGDNMDDFQAVQRSSNGPNDEASED 238
Query: 213 -CG-YIGEEIAKNEELGLSAYTGY--VHGNLVLDSIRDLFCNGMSIIGNNDFEIVEIYRC 268
C + + + K ++ ++G L D+++ +F G + +G+ E +++Y+
Sbjct: 239 SCSRELDDAVEKWDKTETDRFSGVKPAEEELDKDAVKQMFALGAATLGH--VESLDVYQF 296
Query: 269 SCASMQVQSKLFLTQAEITKELHGDANVRYAWLSFSKGELSTMMEHGLGHCELFTTKCTY 328
S + + LF QA+ITK+ GDAN+RYAW+ K LS +M HGLG F K Y
Sbjct: 297 SSEIAKARLSLFQKQADITKKHRGDANIRYAWVPAKKEVLSAVMMHGLGVGGAFIKKSMY 356
Query: 329 GVGVHLAAISCPYASARYCDIDGNGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEY 388
GVGVH A +CPY SARYCDID NGVRH++LCR+IMGNME + N Q+ EY
Sbjct: 357 GVGVH--AANCPYFSARYCDIDDNGVRHMVLCRVIMGNME----PLRGDNTQYFTGGEEY 410
Query: 389 DNGVDDIQCPRYYTIWNMNISTYICPAFVVSFKVS-RGVEGYFCGTVDKNNVSRDNSSGA 447
DNGVDD++ P++Y IWNMN++T+I P FVVSFK+S EG T SR SSG
Sbjct: 411 DNGVDDVESPKHYLIWNMNMNTHIYPEFVVSFKLSIPNAEGNILPTTQ----SRHESSGL 466
Query: 448 NSDSHCSGDPVQSASSAYNEIAGNGV-ACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLI 506
+ G P N +G+ + + + P+SP MP +L AI+ K+ DM LI
Sbjct: 467 TLEGP-KGSPSNEPGRVSNGGSGSEKNSSSSRRPRSPIMPFPLLFKAISSKIARKDMDLI 525
Query: 507 KENFELYKAKQISRDDFVKEMRLIVG-DTLLRDTITTLQ 544
++ + K++SR +F K + +IVG D LL TIT LQ
Sbjct: 526 IAGYQELREKKVSRKEFYKTLSMIVGDDDLLISTITGLQ 564
>At1g23550 unknown protein
Length = 323
Score = 132 bits (332), Expect = 5e-31
Identities = 94/359 (26%), Positives = 175/359 (48%), Gaps = 67/359 (18%)
Query: 200 AQVKVRNMMSI--MDCGYIGEEIAKNEELGLS-AYTGYVHGNLVL--------DSIRDLF 248
AQV++ + S+ +D G I + I+ + + +S A + + +L+L D I+
Sbjct: 3 AQVEIEDQTSVTNLDNGEIFDSISDDADSSVSHAGSSFSSSSLILLGEGNPEHDVIKTCL 62
Query: 249 CNGMSIIGNNDFEIVEIYRCSCASMQVQSKLFLT----QAEITKELHGDANVRYAWLSFS 304
+GM ++ ++D IV I + S FL + ++ GDANV+Y W + S
Sbjct: 63 LSGMGVV-SSDTTIVTISKNSSERGITTRAKFLAFRIFTDAVARKHGGDANVKYGWYAGS 121
Query: 305 KGELSTMMEHGLGHCELFTTKC---TYGVGVHLAAISCPYASARYCDIDGNGVRHLILCR 361
+ E+ ++ +G + ++ + ++G+G+HL C +A + D G+R+L+LCR
Sbjct: 122 RDEIQRIISYGFSNRDVGKFENDGGSHGIGIHLVPSKCSLLAASATEQDEEGLRYLLLCR 181
Query: 362 IIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICPAFVVSFK 421
+I+G E++ + + Q PSS+E+D+GVDD+ PR Y IW+ N+++ I P+++VSF+
Sbjct: 182 VILGKPEIII----SGSKQSYPSSAEFDSGVDDLHNPRNYVIWSCNMNSCILPSYIVSFR 237
Query: 422 VSRGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAGNGVACTQKIPK 481
R VSR + P
Sbjct: 238 SPR------------LRVSRGGFASR--------------------------------PS 253
Query: 482 SPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTI 540
SPW+ L++ ++ + P+ M+LI ++ ++ ++I RD V++MR + GD LL + I
Sbjct: 254 SPWVSFASLMSMLSTSMDPSRMNLIIRTYDDFRKRKIRRDQLVRKMREVAGDNLLAEII 312
>At1g70440 hypothetical protein
Length = 305
Score = 107 bits (266), Expect = 2e-23
Identities = 72/264 (27%), Positives = 135/264 (50%), Gaps = 61/264 (23%)
Query: 278 KLFLTQAEITKELHGDANVRYAWLSFSKGELSTMMEHGLGHCELFTTKC---TYGVGVHL 334
KLF T+A + ++ +G AN+RY W S SK E+ ++ +G + E+ + ++GVG+HL
Sbjct: 89 KLF-TEA-MKRKNNGYANIRYGWYSGSKEEIDRVITYGFSNREIKKVENDVGSHGVGIHL 146
Query: 335 AAISCPYASARYCDIDGNGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDD 394
A+A + D G+++++LCR+I+G E I T + Q PSS+++D+GVD+
Sbjct: 147 VHHRYSLAAALVGEGDEEGIKNILLCRVILGKPE----QIVTGSKQSYPSSNQFDSGVDN 202
Query: 395 IQCPRYYTIWNMNISTYICPAFVVSFK--VSRGVEGYFCGTVDKNNVSRDNSSGANSDSH 452
++ PR Y IW+ N+++YI P ++VSFK + RG+ G
Sbjct: 203 LENPRKYVIWSCNMNSYILPTYIVSFKSHLLRGLIGR----------------------- 239
Query: 453 CSGDPVQSASSAYNEIAGNGVACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLIKENFEL 512
+SP + +L++ ++ + M+LI +++
Sbjct: 240 ---------------------------ARSPCVSFSVLMSILSKSLDAARMNLILTSYDD 272
Query: 513 YKAKQISRDDFVKEMRLIVGDTLL 536
++ +++ R+ V+++R +VGD LL
Sbjct: 273 FRKRKLRREQLVRKIREVVGDNLL 296
>At5g62520 unknown protein
Length = 309
Score = 96.7 bits (239), Expect = 3e-20
Identities = 65/246 (26%), Positives = 121/246 (48%), Gaps = 54/246 (21%)
Query: 292 GDANVRYAWLSFSKGELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARYCDIDG 351
G A V+Y W S SK EL T+ E+G E ++G G++L+ + P + +
Sbjct: 104 GGAKVKYGWCSVSKHELKTIFEYGFS--EPLRNDGSFGRGLYLSPDNSPLDCLKDSASES 161
Query: 352 -NGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNIST 410
+G+R L+LCR+++G E++ P T + PSS E+D+GVDD+ + Y +W+ +++T
Sbjct: 162 EDGMRFLLLCRVLLGKSEIV-PQGSTRSC---PSSPEFDSGVDDLVSTKKYIVWSTHMNT 217
Query: 411 YICPAFVVSFKVSRGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAG 470
++ P F+V K N++R
Sbjct: 218 HVLPEFLVCIKA-------------PFNLTR----------------------------- 235
Query: 471 NGVACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLI 530
+ K +SPWM +LI A++ +PP+ + +I+++++ + ++I+R + ++ +R I
Sbjct: 236 -----SPKRLRSPWMAFPVLIKALSKFLPPSQILVIQKHYKDQQNRRITRSELIQRVRSI 290
Query: 531 VGDTLL 536
GD LL
Sbjct: 291 TGDKLL 296
>At3g47720 unknown protein
Length = 316
Score = 89.4 bits (220), Expect = 5e-18
Identities = 77/305 (25%), Positives = 129/305 (42%), Gaps = 76/305 (24%)
Query: 257 NNDFEIVEIYRCSCASMQVQSKL--FLTQAEITKELHGD-----------ANVRYAWLSF 303
+N FEIV I + + Q+KL F AE + G A V+Y
Sbjct: 72 SNQFEIVSILKNGFQTPLGQAKLKAFQIYAESVAKKSGSCCGNKAAVAEAARVKYGCCGV 131
Query: 304 SKGELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARYCDIDGNGVRHLILCRII 363
K EL ++ +G + L + A + C + C+ DG + L+ RII
Sbjct: 132 EKEELKAILMYGFSNNALCLSPDN-------APLQCMIDPSSSCNEDG--ISFLLFSRII 182
Query: 364 MGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICPAFVVSFKVS 423
MG E++C S Q PSS E+D+GVD + P Y IW+ +++T++ P FVV K
Sbjct: 183 MGKSEVVC-----STSQSYPSSMEFDSGVDSLTSPNKYIIWSTHMNTHVLPEFVVCIKTP 237
Query: 424 RGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAGNGVACTQKIPKSP 483
++ +K PKSP
Sbjct: 238 SILK-------------------------------------------------RKNPKSP 248
Query: 484 WMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTITTL 543
W+ +LI +I+ + + + LI ++++ ++ ++ISR + ++ +R I GD+LL I ++
Sbjct: 249 WISFPVLINSISKFLNQSQIRLIHKHYKEHQDRRISRCELIQRLRSITGDSLLVQIIKSV 308
Query: 544 QFMVH 548
VH
Sbjct: 309 GQKVH 313
>At1g70530 putative protein kinase
Length = 646
Score = 37.7 bits (86), Expect = 0.017
Identities = 42/197 (21%), Positives = 83/197 (41%), Gaps = 26/197 (13%)
Query: 1 MEARTTLKRNETTRCGVHIRGASQQELCQQHFCASPTINFVNRMRLGDYKRKMNILGSGS 60
++ R K+ E + G A++ LC ++ N R DY N LG G
Sbjct: 283 LKKRHAKKQREKKQLGSLFMLANKSNLC---------FSYENLERATDYFSDKNKLGQGG 333
Query: 61 GQSYLRFYLNYKKSARPERMMFYNHGEWVDYPKDIVDLVKK-DFEIKNAAIGIRLNGRDL 119
S + L K+ +R +F+N +WVD+ + V+L+ + D + +G + G +
Sbjct: 334 SGSVYKGVLTNGKTVAVKR-LFFNTKQWVDHFFNEVNLISQVDHKNLVKLLGCSITGPES 392
Query: 120 MLDFLHACQMDLKTGF-----IQPIAWIDKVGGYFFPEVDVASGEESYNFGEQEGMK-FH 173
+L + + L +QP+ W + ++ + + E E+ ++ H
Sbjct: 393 LLVYEYIANQSLHDYLFVRKDVQPLNWAKRF------KIILGTAEGMAYLHEESNLRIIH 446
Query: 174 VEIKMNMV---DEFQLR 187
+IK++ + D+F R
Sbjct: 447 RDIKLSNILLEDDFTPR 463
>At5g43130 unkown protein
Length = 689
Score = 31.2 bits (69), Expect = 1.6
Identities = 15/65 (23%), Positives = 35/65 (53%)
Query: 485 MPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTITTLQ 544
+P L+ + +++ + ++ + K +I ++ F + M+ IVGD +LR ++ LQ
Sbjct: 80 VPFAALLPTLMNQLDKDRALQLRTLYARLKKNEIPKEGFTRHMKDIVGDQMLRMAVSKLQ 139
Query: 545 FMVHN 549
+ +N
Sbjct: 140 QVNYN 144
>At5g16280 putative protein
Length = 1265
Score = 31.2 bits (69), Expect = 1.6
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 3/50 (6%)
Query: 190 IGESNALVKDAQVKVRNMMSIMDCGY---IGEEIAKNEELGLSAYTGYVH 236
IG+ A+V V VRNM+ ++DCGY +EI + + TG H
Sbjct: 510 IGQWYAIVGMHDVAVRNMLKVLDCGYQSKATQEIFLRDFFDIVKKTGMKH 559
>At3g51370 protein phosphatase 2C like protein
Length = 379
Score = 31.2 bits (69), Expect = 1.6
Identities = 20/84 (23%), Positives = 44/84 (51%), Gaps = 2/84 (2%)
Query: 156 ASGEESYNFGEQEGMKFHVEIKMNMVDEFQLRECIGESNALVKDAQVKVRNMMSIMDCGY 215
+SG+ S + G+Q+G+ ++ + ++V EF + + ++N L++D +S +D G
Sbjct: 18 SSGKSSDSTGKQDGLLWYKDFGQHLVGEFSM--AVVQANNLLEDQSQVESGPLSTLDSGP 75
Query: 216 IGEEIAKNEELGLSAYTGYVHGNL 239
G I + G + +V+ +L
Sbjct: 76 YGTFIGIYDGHGGPETSRFVNDHL 99
>At5g47380 putative protein
Length = 598
Score = 30.8 bits (68), Expect = 2.1
Identities = 22/92 (23%), Positives = 40/92 (42%), Gaps = 7/92 (7%)
Query: 421 KVSRGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAGNGVACT-QKI 479
K R + + C +D + R S+ + D H SG+ +++ + + + V T K
Sbjct: 12 KQRRNGDSWHC--LDSHKHGRSKSASSERDLHTSGNGASQSANNFTRMQASSVQTTANKR 69
Query: 480 PKSPWMPLHMLIAAINDKVPPNDMSLIKENFE 511
PK PLH + V ND + ++ + E
Sbjct: 70 PK----PLHNCQMLTKNNVSSNDRASLERDVE 97
>At3g47500 H-protein promoter binding factor-2a
Length = 448
Score = 30.0 bits (66), Expect = 3.6
Identities = 27/79 (34%), Positives = 36/79 (45%), Gaps = 18/79 (22%)
Query: 421 KVSRGVEGYFCGTVDKNNVSR-----DNSSGAN---SDSHC-------SGDPVQSASSAY 465
KVS G F G D+ V+R D SSG++ S++H SG V++ +
Sbjct: 220 KVSNGARNRFHGLADQRLVARVENGDDCSSGSSVTTSNNHSVDESRAQSGSVVEAQMNNN 279
Query: 466 NEIAGNGVACTQKIPKSPW 484
N NG AC IP PW
Sbjct: 280 NNNNMNGYAC---IPGVPW 295
>At3g48770 hypothetical protein
Length = 1899
Score = 29.3 bits (64), Expect = 6.1
Identities = 29/101 (28%), Positives = 41/101 (39%), Gaps = 19/101 (18%)
Query: 38 INFVNRMRLGDYKRKMNILGSGSGQSYLRFYLNYKKSARPERMMFYNHGEW--------- 88
IN NR L + N L S G S+LR +N RP FY+ W
Sbjct: 964 INKCNRHSL-----RSNFLNSVRGGSWLRTTMNGVSDYRPPSQSFYHTSSWGSILQNGSI 1018
Query: 89 -VDYPKDIVDLVKKDFEIKNAAIGIRLNGRDLMLDFLHACQ 128
VD P +VD EI+ +++ G +M +F C+
Sbjct: 1019 LVDIP--LVDRSYYGNEIEKYKEELKIAG--VMFEFSEVCR 1055
>At3g07440 unknown protein
Length = 235
Score = 29.3 bits (64), Expect = 6.1
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 2/59 (3%)
Query: 477 QKIPKSPWMPLHMLIAAINDKVPPNDMSLI-KENFELYKAKQISRDDFVKEMRLIVGDT 534
+K KS W + + I D VPP + I ++N + KQ+ RD F++ RL++ ++
Sbjct: 46 KKEEKSEWWIVDGEMHEIGDHVPPRERFTIPRDNIPNKRRKQL-RDQFMRRTRLVLKES 103
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,818,428
Number of Sequences: 26719
Number of extensions: 565387
Number of successful extensions: 1210
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1183
Number of HSP's gapped (non-prelim): 18
length of query: 561
length of database: 11,318,596
effective HSP length: 104
effective length of query: 457
effective length of database: 8,539,820
effective search space: 3902697740
effective search space used: 3902697740
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0283.6


