

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.14.1
(63 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
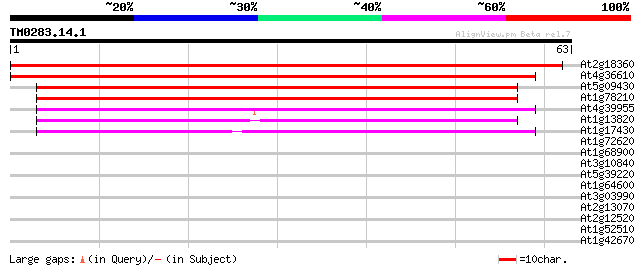
Sequences producing significant alignments: (bits) Value
At2g18360 unknown protein 99 4e-22
At4g36610 unknown protein 94 1e-20
At5g09430 putative hydrolase 52 7e-08
At1g78210 unknown protein 50 2e-07
At4g39955 unknown protein 45 7e-06
At1g13820 hypothetical protein 42 4e-05
At1g17430 unknown protein 38 8e-04
At1g72620 unknown protein 35 0.007
At1g68900 hypothetical protein 32 0.077
At3g10840 alpha/beta hydrolase like protein 30 0.22
At5g39220 unknown protein 29 0.50
At1g64600 unknown protein 27 1.5
At3g03990 unknown protein 26 3.2
At2g13070 pseudogene 26 4.2
At2g12520 pseudogene 25 7.2
At1g52510 unknown protein (At1g52510) 25 9.4
At1g42670 unknown protein 25 9.4
>At2g18360 unknown protein
Length = 313
Score = 99.0 bits (245), Expect = 4e-22
Identities = 44/62 (70%), Positives = 53/62 (84%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
+IHLLWGE+DQIF E AK+MKEQLG+ AT + IKKAGHL HLERPCVYNR LK+F+AS
Sbjct: 250 KIHLLWGESDQIFNLEFAKSMKEQLGENATMESIKKAGHLAHLERPCVYNRRLKKFLASV 309
Query: 61 FA 62
++
Sbjct: 310 YS 311
>At4g36610 unknown protein
Length = 317
Score = 94.0 bits (232), Expect = 1e-20
Identities = 42/59 (71%), Positives = 51/59 (86%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIAS 59
+IH LWGE+DQIF ELA++MKEQ+G+ AT + IKKAGHLV LERPCVYNR LK+F+AS
Sbjct: 248 KIHFLWGESDQIFDLELARDMKEQIGENATIESIKKAGHLVQLERPCVYNRRLKKFLAS 306
>At5g09430 putative hydrolase
Length = 303
Score = 51.6 bits (122), Expect = 7e-08
Identities = 24/54 (44%), Positives = 34/54 (62%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
++WGE DQIF EL +K +G+ A IKKAGH V+LE+ + + LK F+
Sbjct: 246 IIWGEEDQIFPLELGYRLKRHIGESAEIVVIKKAGHAVNLEKSKEFVKHLKSFL 299
>At1g78210 unknown protein
Length = 314
Score = 50.4 bits (119), Expect = 2e-07
Identities = 19/54 (35%), Positives = 35/54 (64%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
++WGE+DQ+F E+ K +++ +GD IK+ GH+ + E+P + + LK F+
Sbjct: 248 IIWGEHDQVFPLEMGKRLEKHVGDNGKLVIIKRTGHIFNFEKPKKFIKLLKSFL 301
>At4g39955 unknown protein
Length = 328
Score = 45.1 bits (105), Expect = 7e-06
Identities = 22/57 (38%), Positives = 34/57 (59%), Gaps = 1/57 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLG-DGATFQGIKKAGHLVHLERPCVYNRCLKRFIAS 59
++WGE DQ+F ELA +K LG D A +KK GH ++ E+P + +K F+ +
Sbjct: 243 MIWGEEDQVFPVELAHRLKRYLGEDRAQLVLLKKTGHAINEEKPKEMYKHMKSFLCT 299
>At1g13820 hypothetical protein
Length = 339
Score = 42.4 bits (98), Expect = 4e-05
Identities = 19/54 (35%), Positives = 31/54 (57%), Gaps = 1/54 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+LWGE+DQI +LA + +L + A + I GHL H+E+P + + F+
Sbjct: 274 ILWGEDDQIISNKLAWRLHGELSN-ARVKQISNCGHLPHVEKPAAVTKLIAEFV 326
>At1g17430 unknown protein
Length = 332
Score = 38.1 bits (87), Expect = 8e-04
Identities = 17/56 (30%), Positives = 31/56 (55%), Gaps = 1/56 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIAS 59
++WG+ D++F E A ++ L + + IK+ GH V++E P N + F+ S
Sbjct: 277 IVWGDKDKVFPLEHAYRLQRHL-QSSRLEIIKETGHAVNIEAPTTLNNFITSFVLS 331
>At1g72620 unknown protein
Length = 331
Score = 35.0 bits (79), Expect = 0.007
Identities = 15/57 (26%), Positives = 32/57 (55%), Gaps = 1/57 (1%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
R ++WG+ D +F E + ++ L + ++ + +K+ GH V++E P N + F+
Sbjct: 272 RTLIVWGDKDNVFPLEHGRRLQRHLPN-SSLEVLKEIGHGVNIEAPTTLNNLIISFV 327
>At1g68900 hypothetical protein
Length = 656
Score = 31.6 bits (70), Expect = 0.077
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 8/63 (12%)
Query: 2 IHLLWGENDQIFK-------RELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLK 54
I L++GE D +K RE++K+ K+ + + I +AGH VHLE P L+
Sbjct: 572 ISLVFGEKDVKYKQIATRMYREMSKS-KKSVNNIIEIVEIPEAGHAVHLESPLRVILALR 630
Query: 55 RFI 57
+F+
Sbjct: 631 KFL 633
>At3g10840 alpha/beta hydrolase like protein
Length = 429
Score = 30.0 bits (66), Expect = 0.22
Identities = 16/58 (27%), Positives = 31/58 (52%), Gaps = 1/58 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASFF 61
++ G+ D+I A+ + + G+ F+ IKK GHL E+P + + +F+ + F
Sbjct: 356 IVTGDTDRIVPAWNAERLARAI-PGSVFEVIKKCGHLPQEEKPDEFISIVAKFLGNAF 412
>At5g39220 unknown protein
Length = 330
Score = 28.9 bits (63), Expect = 0.50
Identities = 14/39 (35%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 8 ENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
ENDQI +L+ + +L + A + + +GHL H+E P
Sbjct: 277 ENDQIVSNQLSVKLLCELAN-AVLREVPDSGHLPHVENP 314
>At1g64600 unknown protein
Length = 537
Score = 27.3 bits (59), Expect = 1.5
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 8/50 (16%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYN 50
R H+LW E KR+L K K+ DG +K H+V PC ++
Sbjct: 262 RSHILWME-----KRKLRKLEKKMKKDGKEVLDLKSGAHIV---APCPHD 303
>At3g03990 unknown protein
Length = 267
Score = 26.2 bits (56), Expect = 3.2
Identities = 12/39 (30%), Positives = 20/39 (50%)
Query: 17 LAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKR 55
+A+ ++ LG T + +K GHL L P + L+R
Sbjct: 225 VAEYLRSHLGGDTTVETLKTEGHLPQLSAPAQLAQFLRR 263
>At2g13070 pseudogene
Length = 239
Score = 25.8 bits (55), Expect = 4.2
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 8/56 (14%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRF 56
+IH L +++ KR + + T QG KKA H LE Y+R L F
Sbjct: 68 QIHRLEQRREELSKRVMDLTL--------TAQGAKKAVHDAELELAIAYSRLLAGF 115
>At2g12520 pseudogene
Length = 356
Score = 25.0 bits (53), Expect = 7.2
Identities = 15/40 (37%), Positives = 21/40 (52%), Gaps = 3/40 (7%)
Query: 14 KRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCL 53
+ EL+K + E +T QG KKA H +E Y+R L
Sbjct: 193 REELSKRVMELT---STVQGAKKAVHDAKVELAAAYSRLL 229
>At1g52510 unknown protein (At1g52510)
Length = 380
Score = 24.6 bits (52), Expect = 9.4
Identities = 12/43 (27%), Positives = 21/43 (47%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
L WG D+ + +A+ ++Q + I+ AGHL + P
Sbjct: 327 LAWGIADKYLPQSIAEEFEKQNPQNVKLRLIEGAGHLPQEDWP 369
>At1g42670 unknown protein
Length = 579
Score = 24.6 bits (52), Expect = 9.4
Identities = 9/32 (28%), Positives = 19/32 (59%)
Query: 32 QGIKKAGHLVHLERPCVYNRCLKRFIASFFAS 63
+G++K GH+ +R N + +F+ S+ A+
Sbjct: 385 KGVEKRGHICCKQRWSKVNDAVNKFVGSYLAA 416
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.142 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,417,771
Number of Sequences: 26719
Number of extensions: 43547
Number of successful extensions: 151
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 136
Number of HSP's gapped (non-prelim): 17
length of query: 63
length of database: 11,318,596
effective HSP length: 39
effective length of query: 24
effective length of database: 10,276,555
effective search space: 246637320
effective search space used: 246637320
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0283.14.1


