

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269b.2
(2064 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
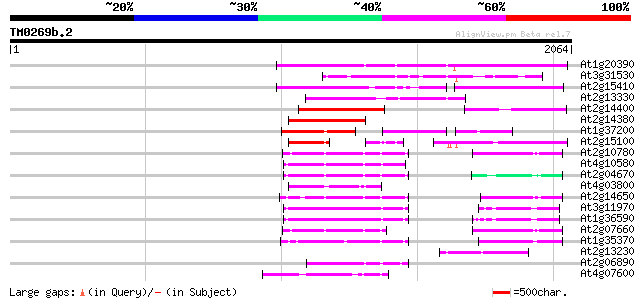
Score E
Sequences producing significant alignments: (bits) Value
At1g20390 hypothetical protein 692 0.0
At3g31530 hypothetical protein 425 e-118
At2g15410 putative retroelement pol polyprotein 362 1e-99
At2g13330 F14O4.9 318 2e-86
At2g14400 putative retroelement pol polyprotein 279 1e-74
At2g14380 putative retroelement pol polyprotein 249 1e-65
At1g37200 hypothetical protein 248 2e-65
At2g15100 putative retroelement pol polyprotein 242 2e-63
At2g10780 pseudogene 239 2e-62
At4g10580 putative reverse-transcriptase -like protein 234 3e-61
At2g04670 putative retroelement pol polyprotein 233 7e-61
At4g03800 229 1e-59
At2g14650 putative retroelement pol polyprotein 225 2e-58
At3g11970 hypothetical protein 223 7e-58
At1g36590 hypothetical protein 223 7e-58
At2g07660 putative retroelement pol polyprotein 219 1e-56
At1g35370 hypothetical protein 212 2e-54
At2g13230 pseudogene 184 5e-46
At2g06890 putative retroelement integrase 183 1e-45
At4g07600 182 1e-45
>At1g20390 hypothetical protein
Length = 1791
Score = 692 bits (1787), Expect = 0.0
Identities = 404/1091 (37%), Positives = 601/1091 (55%), Gaps = 33/1091 (3%)
Query: 980 EEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPI 1039
E V++ + R + A I P ++ ++ LLK FAW ++M G+ + +L +
Sbjct: 712 EMVNIDESDPTRCVGVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNV 771
Query: 1040 KEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVC 1099
KPVKQ R+ P+ + EE+E+LL+ I +Y +WLAN V V KKNGK RVC
Sbjct: 772 DPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVC 831
Query: 1100 IDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCP 1159
+D+ DLN A PKD Y +P + +V++ +G+ LS +D +SGYNQI + ++D KT+F
Sbjct: 832 VDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTD 891
Query: 1160 GALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLR 1219
GTY + VM FGLKNAGATYQR +N + D I ++VYIDD++VKS +DH+ HL
Sbjct: 892 R--GTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLS 949
Query: 1220 KSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQ 1279
K F+ + YG+K+NP KC FGV +G+FLG+VV K+GIE N + +AIL+ P + +++Q
Sbjct: 950 KCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRNAREVQ 1009
Query: 1280 SLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMAS 1339
L G+I L RFI+ ++K F L LK+ F W+ + ++AF++LK YLS+PPI+
Sbjct: 1010 RLTGRIAALNRFISRSTDKCLPFYNL--LKRRAQFDWDKDSEEAFEKLKDYLSTPPILVK 1067
Query: 1340 PIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYF 1399
P G+ + LYI+ +D + S+L +ED ++R IFY S+ L +AETRY +IEK L +
Sbjct: 1068 PEVGETLYLYIAVSDHAVSSVLVREDR-GEQRPIFYTSKSLVEAETRYPVIEKAALAVVT 1126
Query: 1400 SCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQA 1459
S KL+ Y + + V + ++ L P R+ KWA+ L+EY + + P A+K Q
Sbjct: 1127 SARKLRPYFQSHTIAVLTDQP-LRVALHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQV 1185
Query: 1460 IADFLADQTL--PQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKS 1517
+ADFL + L + ++ + W L+ DGSS G+GIG+ +VSP + FR++
Sbjct: 1186 LADFLIELPLQSAERAVSGNRGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRF 1245
Query: 1518 NCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKASN 1577
+NN AEYE L +GL + + + DS+L+ QL+ EY+ +E + Y
Sbjct: 1246 VATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQL 1305
Query: 1578 LLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPS--DLDILVI 1635
+ F+ +L + R DN A+ LA +A + L+ +I V+ PS D + I
Sbjct: 1306 MTKDFENFKLSKIPRGDNAPADALAALA---LTSDSDLRRIIPVESIDKPSIDSTDAVEI 1362
Query: 1636 ----------DNMAPNDWRKPIVDYLQNPVGTTD----RKTKYRAMSYVIMGNELFKKNV 1681
D P DWR I DYL + +D R+ + +A Y +M L K +
Sbjct: 1363 VNTIRSSNAPDPADPTDWRVEIRDYLSDGTLPSDKWTARRLRIKAAKYTLMKEHLLKVSA 1422
Query: 1682 DGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQ 1741
G +L CL + + H+G G H G + L + G YWPT++ DC + +C+
Sbjct: 1423 FGAMLNCLHGTEINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCE 1482
Query: 1742 DCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEA 1801
CQRHA H P L + + P+PF WA+D++G + +SRQ ++I+V DYFTKWVEA
Sbjct: 1483 QCQRHAPTIHQPTELLRAGVAPYPFMRWAMDIVGPMP--ASRQKRFILVMTDYFTKWVEA 1540
Query: 1802 IPLQNVTQDTVIEFI-SSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYY 1860
+ + V F+ I+ R GLP + TD G+ F+ +F SW I+L STP Y
Sbjct: 1541 ESYATIRANDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRY 1600
Query: 1861 AQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQE 1920
Q NGQ EA NKT++S +KK + K W L VLW+YR +P+ +T TPF AYG E
Sbjct: 1601 PQGNGQAEATNKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGME 1660
Query: 1921 AVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNK 1980
A+ P EV S R + P E MMLD L +L+E R L + + AK YN+
Sbjct: 1661 AMAPAEVGYSSLRRSMMVKNP-ELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQ 1719
Query: 1981 KVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGGNR-Q 2039
KV + F +GD V + V N + K NWEG + V K++ Y + + G
Sbjct: 1720 KVHNRHFDVGDLVLRKVFE-NTAEINAGKLGANWEGSYQVSKIVRPGDYELLTMSGTAVP 1778
Query: 2040 MTINGKYLKTY 2050
T N +LK Y
Sbjct: 1779 RTWNSMHLKRY 1789
>At3g31530 hypothetical protein
Length = 831
Score = 425 bits (1093), Expect = e-118
Identities = 276/829 (33%), Positives = 408/829 (48%), Gaps = 115/829 (13%)
Query: 1149 EDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKS 1208
ED KTAF + Y VMPFGLKNAGATY+R +N IF I M+VYIDD++VKS
Sbjct: 99 EDQEKTAFYTEQGIFCYR--VMPFGLKNAGATYERFVNKIFALQIGKTMEVYIDDMLVKS 156
Query: 1209 PSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILD 1268
+ DH+ HLR+ F+++ Y +K+NP KC FGV + GIE N + +A+L
Sbjct: 157 MTEKDHISHLRECFKQLNLYNVKLNPAKCRFGV-----------RSGIEANPKQIEALLG 205
Query: 1269 TSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELK 1328
+ P +K+++Q L G++ L RFI+ +EK +F +LR K+ F W ++AF ELK
Sbjct: 206 MASPQNKREVQCLTGRVAALNRFISRSTEKCLAFYDVLRRNKK--FEWTTRCEEAFQELK 263
Query: 1329 VYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYT 1388
YL++PPI+A P+ G+P LY++ +D + L +ED +++ IFY+S+ AE+RY
Sbjct: 264 KYLATPPILAKPVIGEPQYLYVAVSDTAVSGELVREDR-GEQKLIFYVSQTFTSAESRYP 322
Query: 1389 MIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLT 1448
+EKL L + S KL+ Y + ++V ++ +L P R+ KW + L+EY +
Sbjct: 323 QMEKLALAVVMSAQKLRPYFQSHSIIVMGSMP-LRVILHSPSQSGRLAKWTIELSEYDIE 381
Query: 1449 YAPLKAVKGQAIADFLADQTLPQEIITYVGIQ-PWKLFFDGSSHKNGTGIGMFIVSPRGI 1507
Y K Q +ADF+ + LP + + W L DGSS K G+G+G+ + SP G
Sbjct: 382 YQNKTCAKSQVLADFIVE--LPTKEARENPLDTTWLLHVDGSSSKQGSGVGIRLTSPTG- 438
Query: 1508 PTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISEN 1567
+NN AEYEAL +GL + L DS+L+ Q EY +
Sbjct: 439 ----------EATNNVAEYEALVAGLNLAWGLKIGKTRAFCDSQLIANQFNGEYTTQDKK 488
Query: 1568 LARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNP 1627
+ Y NL FDE L + R +N A+ LA +AS LK +I V+ P
Sbjct: 489 MEAYLIHVQNLAKNFDEFELTRIPRGENTSADALAALAS---TSDTSLKRVIPVEFIEKP 545
Query: 1628 S-----DLDILVIDNMAPND--------WRKPIVDYLQNPVGTTD----RKTKYRAMSYV 1670
S + +L I A D W +PI+ Y+ +D RK K +A +V
Sbjct: 546 SIELGKEEHVLPIQISADQDDPDDCNSEWMEPIISYISEGKLPSDKWKARKLKAQAARFV 605
Query: 1671 IMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIM 1730
++ +L+K + G L+ C+ + + +H G
Sbjct: 606 LVDTKLYKWRLSGPLMTCVEAEAICKIMKEIHSG-------------------------- 639
Query: 1731 KDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIV 1790
HA H PA L SI P+PF W++D+IG ++P S+Q K ++V
Sbjct: 640 --------------SHAPTIHQPAELLSSIASPYPFMRWSMDIIGPMHP--SKQKKLVLV 683
Query: 1791 AIDYFTKWVEAIPLQNVTQDTVIEFI-SSIVYRFGLPESLTTDQGTVFVGQKVASFAESW 1849
DYF+KW+EA ++ V F+ I+ R G+P + TD G+ F+ + F + W
Sbjct: 684 LTDYFSKWIEAESYASIKDAQVENFVLKHILCRHGIPYEIVTDNGSQFISTRFQGFCDKW 743
Query: 1850 GIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTG 1909
GI+L STP Y Q NGQ EAAN+ L VLW +R +P+ +T
Sbjct: 744 GIRLSKSTPRYPQGNGQAEAANE--------------------LEGVLWLHRTTPRRATR 783
Query: 1910 ATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDE 1958
TPF YG E V+P E+ + S R E D LDEL +DE
Sbjct: 784 ETPFASIYGTECVIPAEMIVPSLRRSLSPE-NDPDNTQTPLDELDLIDE 831
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 362 bits (929), Expect = 1e-99
Identities = 218/626 (34%), Positives = 329/626 (51%), Gaps = 64/626 (10%)
Query: 981 EVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIK 1040
+V++ + R I + EL++ +V L++ FAW ++MPG+ V +L +
Sbjct: 718 QVNIDESDPSRCVGIRIDLPSELQNELVNFLRQNAATFAWSVEDMPGIDSAVTCHELNVD 777
Query: 1041 EDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCI 1100
KP+KQ R+ PD + EE+++LL I RY DWL N V V KKNGK RVCI
Sbjct: 778 PTYKPLKQKRRKLGPDRTKDVNEEVKKLLDAGSIVEVRYPDWLRNPVVVKKKNGKWRVCI 837
Query: 1101 DFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPG 1160
DF DLN A PKD + +P + +V++ AG+E LS +D +SGYNQI + + D KT F
Sbjct: 838 DFTDLNKACPKDSFPLPHIDRLVEATAGNELLSFMDAFSGYNQILMHQNDREKTVFITDQ 897
Query: 1161 ALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRK 1220
GTY + VMPFGLKNAGATY R++N +F D ++ M+VYIDD++VKS ++H+ HLR+
Sbjct: 898 --GTYCYKVMPFGLKNAGATYPRLVNQMFTDQLDHSMEVYIDDMLVKSLRAEEHITHLRQ 955
Query: 1221 SFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQS 1280
F+ + +Y +K+NP KC FGV +G+FLG++V ++GIE N + AI+D P + +++Q
Sbjct: 956 CFQVLNRYNMKLNPSKCTFGVTSGEFLGYLVTRRGIEANPKQISAIIDLPSPRNTREVQR 1015
Query: 1281 LLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASP 1340
L+G+I L RFI+ ++K F LLR K F W+ + ++
Sbjct: 1016 LIGRIAALNRFISRSTDKCLPFYQLLRANKR--FEWDEKCEE------------------ 1055
Query: 1341 IRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFS 1400
G+ + LYI+ + + +L +ED ++ IFY+S+ L+ AE RY +EKL + S
Sbjct: 1056 --GETLYLYIAVSTSAVSGVLVREDR-GEQHPIFYVSKTLDGAELRYPTLEKLAFAVVIS 1112
Query: 1401 CVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAI 1460
KL+ Y K V V ++ ++ +L P R+ KWA+ L+EY + Y K +
Sbjct: 1113 ARKLRPYFKSYTVEVLTNQP-LRTILHSPSQSGRLAKWAVELSEYDIAY------KNRTC 1165
Query: 1461 ADFLADQTLPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCS 1520
A T P R IP + + S
Sbjct: 1166 AKSHRAPTRPYR--------------------------------RSIPCRKVDLARFPAS 1193
Query: 1521 NNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLA 1580
NNEAEYEAL +GL + + K + DS+LV Q + Y+ +E + Y L
Sbjct: 1194 NNEAEYEALIAGLRLAHGIEVKKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVVRELSY 1253
Query: 1581 KFDEARLGHVSRVDNQEANELAQIAS 1606
F+ L + R DN A+ LA +AS
Sbjct: 1254 NFEVFELTKIPRSDNAPADALAVLAS 1279
Score = 258 bits (660), Expect = 2e-68
Identities = 143/406 (35%), Positives = 211/406 (51%), Gaps = 9/406 (2%)
Query: 1636 DNMAPNDWRKPIVDYLQNPVGTTD----RKTKYRAMSYVIMGNELFKKNVDGTLLKCLSE 1691
D+ A +W + I Y+ + + D R+ + R+ Y + L + + G LL CL +
Sbjct: 1369 DSPASTNWIEEIRSYIADRIVPKDKWAARRLRARSAQYTLPHEHLLRWSATGVLLSCLDD 1428
Query: 1692 DDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQH 1751
++A + +H+G G H G + + + G YWPT+ DC ++ RC+ CQ HA H
Sbjct: 1429 EEAQQVMREIHEGAGGNHSGGRALALKIRKHGQYWPTMNADCEKFVARCEKCQGHAPFIH 1488
Query: 1752 VPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDT 1811
+P+ L + + P+PF WA+D+I +SRQ KYI+V DYFTKW EA +
Sbjct: 1489 IPSEVLQTAMPPYPFMRWAMDIIRPFP--ASRQKKYILVMTDYFTKWFEAESYARIQSKE 1546
Query: 1812 VIEFI-SSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAA 1870
V F+ +I+ GLP + TD GT F + F W I+L STP Y Q NGQ EA
Sbjct: 1547 VQNFVWKNIICHHGLPYEIVTDNGTQFTSLQFEGFCAKWKIRLSKSTPRYPQCNGQAEAT 1606
Query: 1871 NKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQ 1930
NKT++ +KK + K W L VLW+YR +P+ ST TPF L YG EA+ P E L
Sbjct: 1607 NKTILDGLKKRLDEKKGAWADELDGVLWSYRTTPRRSTDRTPFSLTYGMEALAPCEAGLP 1666
Query: 1931 SCRIQRQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLG 1990
+ R P+ + M+LD L L+E+R L + + AK YNKKV+ + F G
Sbjct: 1667 TVRRSMLINDPTLND-QMLLDNLDTLEEQRDQALLRIQNYQQAAAKFYNKKVKNRLFAEG 1725
Query: 1991 DYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGG 2036
D V + V N ++ K +WEGP+ + KV+ Y + + G
Sbjct: 1726 DLVLRKVFE-NTVEQDAGKLGAHWEGPYLISKVVKPGVYELLTMDG 1770
>At2g13330 F14O4.9
Length = 889
Score = 318 bits (816), Expect = 2e-86
Identities = 202/592 (34%), Positives = 312/592 (52%), Gaps = 49/592 (8%)
Query: 1090 IKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEE 1149
++KNGK RVCIDFRDLN A PKD + +P + +V++ HE LS +D + GYNQI + +
Sbjct: 288 LEKNGKWRVCIDFRDLNKACPKDSFPLPHIDRLVEATVEHEKLSFMDAFYGYNQILMRRD 347
Query: 1150 DVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSP 1209
KTAF GTY + VMPFGLKNAG TYQR++N +F D + ++VYIDD++VKS
Sbjct: 348 GQEKTAFITDR--GTYYYKVMPFGLKNAGTTYQRLVNRMFVDQLGKTIEVYIDDMLVKSA 405
Query: 1210 SRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDT 1269
DH+ LR+ F+ + K+ +K+NP KC+F V + +FL ++V ++GIE N + A ++
Sbjct: 406 HEKDHVPQLRECFKILIKFEMKLNPEKCSFEVQSREFLEYLVTERGIEANPKQIAAFIEM 465
Query: 1270 SPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKV 1329
P +++Q L G+I L FI+ + K F LR KE F W + ++AF +LK
Sbjct: 466 PSPKMAREVQRLTGRIPALNGFISRSANKCVPFYQPLRKGKE--FDWNKDCEQAFKQLKA 523
Query: 1330 YLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTM 1389
YL+ PPI+A P +G+P+ LY + + R
Sbjct: 524 YLTEPPILAKPEKGEPLYLY-------------------------------TNRDKRCPA 552
Query: 1390 IEKLCLCLYFSCVKLKYYIK--PIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSL 1447
+EKL L + S KL+ Y + PI VM I H L++ R+ KWA+ L EY +
Sbjct: 553 MEKLALTVVMSVRKLRLYFQSHPIVVMTSQPIRTILHSLTQS---GRLAKWAIELREYDI 609
Query: 1448 TYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGI 1507
Y ++K Q +ADF+ L T + W L DGSS++ G+G+G+ + SP G
Sbjct: 610 EYRTRTSLKAQVLADFVIKLPLADLDGTNSN-KKWLLHVDGSSNRQGSGVGIQLTSPTGE 668
Query: 1508 PTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGDSELVIKQLTKEYKCISEN 1567
+ ++ N SNNE+EYEAL +G+++ G + + DS+LV Q EY+ E
Sbjct: 669 VIEQSLQLGFNASNNESEYEALIAGIKLAQEKGIREIHAYSDSQLVTSQFHGEYEAKDER 728
Query: 1568 LARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNP 1627
+ Y L +F+ +L + R +N A+ LA +AS +K +I V E +
Sbjct: 729 MEAYLELVKTLAQQFESFKLTRIPRGENTSADTLAALAS---TSDPFVKRIIPV-EGIEH 784
Query: 1628 SDLDILVIDNMAPNDWRKPIVDYLQNPVGTTDRK----TKYRAMSYVIMGNE 1675
+ +D+ V + P ++ + RK ++R +S +I+ NE
Sbjct: 785 TSIDLTVKHAGMEPEAPPPQLELSRQLRRQVQRKRSQNKRFRKLSMMIVRNE 836
Score = 32.3 bits (72), Expect = 3.0
Identities = 18/65 (27%), Positives = 34/65 (51%)
Query: 967 VDFSKKMEAQDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMP 1026
+D S++ + + +V + + KR I +D +++ ++ LKE KD FAW +
Sbjct: 225 LDPSQRGTRKSLITQVCIDESDPKRCVGIGHDLDLTVREDLITFLKENKDSFAWSSANLQ 284
Query: 1027 GLSRE 1031
G+S E
Sbjct: 285 GISLE 289
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 279 bits (714), Expect = 1e-74
Identities = 138/316 (43%), Positives = 211/316 (66%), Gaps = 5/316 (1%)
Query: 1061 IKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAE 1120
+ +E+++LL+ IR +Y DWLAN V V KKNGK RVCIDF DLN A PKD + +P +
Sbjct: 480 VNDEVDKLLKIGSIREVQYPDWLANTVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHID 539
Query: 1121 MMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGAT 1180
+V+S AG+E LS +D +SGYNQI + ED KT F G Y + VMPFGL+NAGAT
Sbjct: 540 RLVESTAGNELLSFMDAFSGYNQIMMNPEDQEKTLFITDR--GIYCYKVMPFGLRNAGAT 597
Query: 1181 YQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFG 1240
Y R++N +F + + M+VYIDD+++KS ++DH+ HL + F + +Y +K+NP KC FG
Sbjct: 598 YPRLVNKMFSEHVGKTMEVYIDDMLIKSLKKEDHVKHLEECFAILNQYQMKLNPAKCTFG 657
Query: 1241 VIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTK 1300
V +G+FLG++V K+GIE N N+ A L+ P + K++Q L G+I L RFI+ ++K+
Sbjct: 658 VPSGEFLGYIVTKRGIEANPNQINAFLNMPSPKNFKEVQRLTGRIAALNRFISRSTDKSL 717
Query: 1301 SFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSM 1360
F +L+ KE F W+ + ++AF +LK YL+SPP++ P + + LY+S ++ + +
Sbjct: 718 PFYQILKGNKE--FLWDEKCEEAFGQLKAYLTSPPVLFKPELDEKLYLYVSVSNHAVSGV 775
Query: 1361 LAQEDEDSKERAIFYL 1376
L +ED +++ I+Y+
Sbjct: 776 LIREDR-GEQKPIYYI 790
Score = 230 bits (587), Expect = 6e-60
Identities = 133/378 (35%), Positives = 199/378 (52%), Gaps = 51/378 (13%)
Query: 1674 NELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDC 1733
N L+++ V L + I +S VH+GLCG+H +G M + + + YWPT++ DC
Sbjct: 951 NSLYRRGVSDPYLLSIFGPVVEIVMSEVHEGLCGSHSSGRAMAFKIKKMCYYWPTMITDC 1010
Query: 1734 MEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAID 1793
++YA+RC+ CQ HA + H P+ SI P+PF W++D+IG ++ S+R +Y++V D
Sbjct: 1011 VKYAQRCKRCQLHAPLIHQPSELFSSISAPYPFMRWSMDIIGPLHR-STRGVQYLLVLTD 1069
Query: 1794 YFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKL 1853
YF+KW+EA + +WGIK+
Sbjct: 1070 YFSKWIEA-----------------------------------------EAILLNWGIKV 1088
Query: 1854 LTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPF 1913
ST Y Q NGQ EAANKT++S +KK + W+ L VLWAYR +P+ STG TPF
Sbjct: 1089 SYSTLRYPQGNGQAEAANKTILSNLKKRLNHLKGGWYDELQPVLWAYRTTPRRSTGETPF 1148
Query: 1914 RLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWN--MMLDELVNLDEERLLVLDTLTRQK 1971
L YG +AV+P E+ + R+ E P + N M+ D L ++E R L + +
Sbjct: 1149 SLVYGMKAVVPAELNVPGL---RRTEAPLNEKENSAMLDDSLDTINERRDQALIQIQNYQ 1205
Query: 1972 DRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKVLVNNAYSI 2031
A+ YN KV+++ F +GDYV K V NKK++ K NWEGP+ V +V+ N Y +
Sbjct: 1206 HAAARYYNSKVKSRPFFVGDYVLKRVFD-NKKEEGAGKLGINWEGPYIVTEVVRNGVYRL 1264
Query: 2032 KELGG---NRQMTINGKY 2046
K+L R +NG +
Sbjct: 1265 KDLEDRPVQRPWNVNGHH 1282
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 249 bits (636), Expect = 1e-65
Identities = 131/284 (46%), Positives = 183/284 (64%), Gaps = 2/284 (0%)
Query: 1024 EMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWL 1083
+M G+ EV +L + K VKQ R+ P+ + +E+++LL I +Y +WL
Sbjct: 476 DMVGIDPEVACHELNVDPTFKLVKQKRRKLGPERSKAVNDEVDKLLDAGSIVEVKYPEWL 535
Query: 1084 ANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQ 1143
AN V V KKN K RVCIDF DLN A PKD + +P + MV++ G+E LS +D +SGYNQ
Sbjct: 536 ANPVVVKKKNDKWRVCIDFTDLNKACPKDSFPLPHIDRMVEATTGNELLSFMDAFSGYNQ 595
Query: 1144 IFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDD 1203
I + ++D KT+F GTY + VMPFGLKN GA YQR++N +F + M+VYIDD
Sbjct: 596 IPMHKDDQEKTSFIIDR--GTYCYKVMPFGLKNVGARYQRLVNQMFAPQLGKTMEVYIDD 653
Query: 1204 IVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKA 1263
++VKS DH+ HL+ FE + KY +K+NP KC FGV +G+FLG++V K+GIE N +
Sbjct: 654 MLVKSTRSADHIDHLKACFETLNKYNMKLNPAKCLFGVTSGEFLGYIVTKRGIEANPKQI 713
Query: 1264 KAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLR 1307
+AILD P +KK++Q L G+I L RFIA ++K+ F LLR
Sbjct: 714 RAILDLQSPRNKKEVQRLTGRIAGLNRFIARSTDKSLPFYQLLR 757
>At1g37200 hypothetical protein
Length = 1564
Score = 248 bits (634), Expect = 2e-65
Identities = 124/274 (45%), Positives = 177/274 (64%), Gaps = 6/274 (2%)
Query: 999 IDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVL 1058
+DP +++ + LKE KD FAW + G+S EV +L + +P+KQ R+ P+
Sbjct: 639 LDPAIREDFITFLKENKDSFAWSSANLQGISLEVTSHELNVDPTYRPIKQKRRKLGPERA 698
Query: 1059 VKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPI 1118
+++E++RLL+ IR +Y DWLAN V V KNGK RVCIDF DLN A PKD + +P
Sbjct: 699 KAVQDEVDRLLKIGSIREVKYPDWLANPVVVKNKNGKWRVCIDFMDLNKACPKDSFPLPH 758
Query: 1119 AEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAG 1178
+ +V + AGHE LS +D +SGYNQI + +D KTAF T + VMPFGLKN G
Sbjct: 759 IDRLVKATAGHELLSFMDAFSGYNQILMRPDDQEKTAFI------TDCYKVMPFGLKNTG 812
Query: 1179 ATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCA 1238
ATYQR++N +F D + M++YIDD++VKS DHL LR+ F+ + K+ +K+NP KC+
Sbjct: 813 ATYQRLVNRMFADQLGKTMELYIDDMLVKSAHEKDHLPQLRECFKILNKFEMKLNPEKCS 872
Query: 1239 FGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPP 1272
FGV +G+FLG++V ++GIE N + A +D P
Sbjct: 873 FGVPSGEFLGYLVTQRGIEANPKQIAAFIDMPSP 906
Score = 137 bits (344), Expect = 9e-32
Identities = 79/237 (33%), Positives = 127/237 (53%), Gaps = 2/237 (0%)
Query: 1370 ERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKP 1429
++ I+Y+S+ L DAET Y +EKL L + S KL+ Y++ ++V + I + +L
Sbjct: 931 QKPIYYVSKTLIDAETHYRAMEKLALAVVMSARKLRQYLQSHPIVVMTSQPI-RTILHST 989
Query: 1430 ILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEIITYVGIQPWKLFFDGS 1489
SR+ KWA+ L+EY + Y + K Q +ADF+ + L T + W L DGS
Sbjct: 990 TQSSRLAKWAIELSEYDIEYRTRTSFKAQVLADFVIELPLADLDGTNSN-KKWLLHVDGS 1048
Query: 1490 SHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAKNVVVKGD 1549
S++ G+G G+ + SP G + FR+ N SNNE+EYEAL G+++ + +++ D
Sbjct: 1049 SNRQGSGEGIQLTSPTGEVIEQSFRLGFNASNNESEYEALIDGIKLAQGMRIRDIHAHSD 1108
Query: 1550 SELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIAS 1606
S+LV Q EY+ E + Y L +F+ L + R +N + LA +AS
Sbjct: 1109 SQLVTSQFHGEYEAKDERMEAYLELVKTLTQQFESFELTRIPRGENTSTDALAALAS 1165
Score = 135 bits (339), Expect = 3e-31
Identities = 72/214 (33%), Positives = 112/214 (51%), Gaps = 16/214 (7%)
Query: 1641 NDWRKPIVDYLQNPVGTTD----RKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFI 1696
NDWR PI+DY++ + D RK K ++ Y IM L K++V G + C
Sbjct: 1246 NDWRIPIIDYIERGITPPDKWETRKLKAQSARYCIMEGRLMKRSVAGPYMVCTYGQQTKD 1305
Query: 1697 AISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASE 1756
+ ++H+G CG+H +G ++ I+ + DC+ ++ RC CQRHA H P E
Sbjct: 1306 LMKSMHEGQCGSHYSGRTLRCIM----------LADCIAHSLRCDKCQRHAPTLHQPPEE 1355
Query: 1757 LHSIIKPWPFRGWALDLIGEINPCSSRQH-KYIIVAIDYFTKWVEAIPLQNVTQDTVIEF 1815
+ SI P+PF W++D++ + ++ K ++V DY TKW+E Q VT+ V F
Sbjct: 1356 MSSISSPYPFMKWSMDVVRPMEASGGKKRLKNLLVLTDYSTKWIEGKAFQQVTEKQVEVF 1415
Query: 1816 -ISSIVYRFGLPESLTTDQGTVFVGQKVASFAES 1848
+ +IVYR G+P + TD G +K FA S
Sbjct: 1416 LLENIVYRHGIPYEIVTDNGLTSPREKSKHFATS 1449
Score = 37.4 bits (85), Expect = 0.092
Identities = 23/97 (23%), Positives = 45/97 (45%), Gaps = 3/97 (3%)
Query: 1956 LDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWE 2015
+DE R + + + + YN ++ + F +G+ V + V N ++ K NWE
Sbjct: 1467 IDERRDHAMIQVQNYQQPAVRYYNSNIKIRRFEVGELVLRKVFS-NTRELNAGKLGTNWE 1525
Query: 2016 GPFTVEKVLVNNAYSIKEL--GGNRQMTINGKYLKTY 2050
GP+ + +V+ + Y + ++ G N +LK Y
Sbjct: 1526 GPYRITEVVRDGIYKLVKVFNGVPELRPWNAMHLKKY 1562
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 242 bits (617), Expect = 2e-63
Identities = 162/550 (29%), Positives = 253/550 (45%), Gaps = 76/550 (13%)
Query: 1560 EYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANELAQIASGYMVD-------- 1611
EY+ + Y + +F++ L + R +N AN LA +AS + V
Sbjct: 795 EYEAKDACMEAYLNLVREVSGRFEQFELTRIPRAENSAANALAALASTFEVTLPRVIPVE 854
Query: 1612 -----KCRLKELI---------------------------EVKEKLNPSDL---DILVID 1636
RL E+ E+ + ++P+++ L +
Sbjct: 855 TISQPSIRLDEISFVTTRAMRRRLDAQSAENGLHQLGDDEEISDAVHPTEIVENQSLPDN 914
Query: 1637 NMAP----------NDWRKPIVDYLQNPVGTTD----RKTKYRAMSYVIMGNELFKKNVD 1682
+ AP DWR+PI DY+ N + RK K + I + L+++
Sbjct: 915 HNAPLPDQPPHDWGADWREPIRDYILNGTLPAEKWAARKLKATCARFCIANDILYRRIFS 974
Query: 1683 GTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQD 1742
C+ + + VHDG CG H G + + + + G YWPT++ DC YA++C+
Sbjct: 975 APDAVCIFGEQTRTVMKEVHDGTCGNHTGGRSLAFKVRKYGYYWPTLVADCEAYARKCEQ 1034
Query: 1743 CQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAI 1802
CQ+HA + PA L ++ P+PF W +D++G ++ T+ VEA
Sbjct: 1035 CQKHAPLILQPAELLTTVSAPYPFMKWLMDIVGPLHVS---------------TRGVEAA 1079
Query: 1803 PLQNVTQDTVIEFI-SSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYA 1861
N+T V FI I+ R GLP + TD G+ F+ ++ F E W I+L STP Y
Sbjct: 1080 AYSNITHVQVWNFIWKDIICRHGLPYEIVTDNGSQFISEQFEVFCEEWQIRLSHSTPRYP 1139
Query: 1862 QANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEA 1921
Q NGQ EA NKT+IS +KK + W L VLWA R +P+ +T TPF L YG EA
Sbjct: 1140 QGNGQAEAMNKTIISNLKKKLNAYKGAWFGELQNVLWAVRTTPRRATDETPFSLIYGMEA 1199
Query: 1922 VLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKK 1981
V+P E+ + S R R + +E+ M++D + +DE R L + + A+ YN
Sbjct: 1200 VIPAEIKVPSARRIRNPQNETENN-EMIIDVIDTIDERRNRALARMQNYHNAGARYYNSN 1258
Query: 1982 VRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGGNR-QM 2040
VR +SF +G V + V NK +K K +WEGP+ + V+ N Y + + G +
Sbjct: 1259 VRNRSFEVGTLVLRRVQQ-NKAEKGAGKLGISWEGPYKITHVVRNGVYRLINMEGKTVRR 1317
Query: 2041 TINGKYLKTY 2050
N +LK +
Sbjct: 1318 AWNSMHLKRF 1327
Score = 140 bits (353), Expect = 8e-33
Identities = 75/154 (48%), Positives = 96/154 (61%), Gaps = 2/154 (1%)
Query: 1024 EMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWL 1083
+M G+S EV+ +L + KPVKQ R+ PD + E+ RLL IR +Y +WL
Sbjct: 411 KMTGISTEVISHELNVDPTFKPVKQKRRKLGPDRAQAVNIEVVRLLEVGRIREVKYPEWL 470
Query: 1084 ANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQ 1143
AN V V KKNGK RVC+DF DLN A KD + +P + +V+S GHE LS +D +SGYNQ
Sbjct: 471 ANPVVVKKKNGKWRVCVDFTDLNKACSKDFFPLPHIDRLVESTTGHEMLSFMDAFSGYNQ 530
Query: 1144 IFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNA 1177
I + ED KT+F GTY + VMPFGLKNA
Sbjct: 531 ILMNPEDQEKTSFIT--ECGTYYYKVMPFGLKNA 562
Score = 73.6 bits (179), Expect = 1e-12
Identities = 44/141 (31%), Positives = 76/141 (53%), Gaps = 9/141 (6%)
Query: 1309 KKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDS 1368
K F W ++AF +LK YLS P ++A G+ + LYI+ ++ + + + E S
Sbjct: 624 KISKGFIWNESCEEAFKQLKRYLSEPSVLAKREFGEQLFLYIAVSESAVTGVQVRV-ERS 682
Query: 1369 KERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSK 1428
+R IFY+ +TRY M+EKL L + + KL+ Y + ++V + ++ +L
Sbjct: 683 DKRPIFYV-------KTRYPMMEKLALAVVTAARKLRPYFQSHPIVVLTSLP-LRTILHS 734
Query: 1429 PILHSRIGKWALALTEYSLTY 1449
P R+GKWA+ L+E+ L +
Sbjct: 735 PTQSGRLGKWAIELSEFDLEF 755
>At2g10780 pseudogene
Length = 1611
Score = 239 bits (609), Expect = 2e-62
Identities = 155/467 (33%), Positives = 251/467 (53%), Gaps = 22/467 (4%)
Query: 1002 ELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKI 1061
ELKD + ++ EF D FA P S + ++ P+ + P R P + K+
Sbjct: 621 ELKD--IPIVNEFSDVFAAVSGVPPDRSDPFT---IELEPGTTPISKAPYRMAPAEMAKL 675
Query: 1062 KEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEM 1121
K+++E LL FIR + W A V+ V KK+G R+CID+R LN T K++Y +P +
Sbjct: 676 KKQLEELLDKGFIRPSSS-PWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDE 734
Query: 1122 MVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATY 1181
++D G ++ S +D SGY+QI I DV KTAFR +E+VVMPFGL NA A +
Sbjct: 735 LMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRT--RYDHFEFVVMPFGLTNAPAAF 792
Query: 1182 QRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGV 1241
++MN +F DF++ F+ ++I+DI+V S S + H HLR ER+R++ L KC+F
Sbjct: 793 MKMMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQ 852
Query: 1242 IAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKS 1301
+ FLG V+ +G+ ++ K ++I + P + +++S LG + RRF+ + + +
Sbjct: 853 RSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQ- 911
Query: 1302 FSPLLRLK-KEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSM 1360
PL RL K+ AF W E +K+F ELK L++ P++ P G+P +Y A+ +G +
Sbjct: 912 --PLTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCV 969
Query: 1361 LAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYD 1420
L Q K I Y SR L E Y + + F + Y+ V +++ +
Sbjct: 970 LMQ-----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHK 1024
Query: 1421 IIKHMLSKPILHSRIGKWALALTEYSL--TYAPLKAVKGQAIADFLA 1465
+K++ ++P L+ R +W + +Y+L Y P KA +AD L+
Sbjct: 1025 SLKYIFTQPELNLRQRRWMELVADYNLDIAYHPGKA---NQVADALS 1068
Score = 86.3 bits (212), Expect = 2e-16
Identities = 79/336 (23%), Positives = 141/336 (41%), Gaps = 20/336 (5%)
Query: 1702 HDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSI- 1760
H H KM L R +W + KD + +C CQ VP+ L ++
Sbjct: 1168 HQSKFSIHPGSNKMYRDLKRY-YHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQNLP 1226
Query: 1761 IKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVI--EFISS 1818
I W + +D + + +H + V +D TK + + + +I ++I
Sbjct: 1227 IPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDE 1286
Query: 1819 IVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLI 1878
IV G+P S+ +D+ T F + +F ++ G ++ ST Y+ Q + Q E +TL ++
Sbjct: 1287 IVRLHGIPVSIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQTDEQSERTIQTLEDML 1346
Query: 1879 KKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQE 1938
+ V NW + L V +AY NS + S G +P+ YG+ P+ C E
Sbjct: 1347 RACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACRTPL------CWTPVGE 1400
Query: 1939 EIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAY-NKKVRAKSFMLGDYVWKVV 1997
+ +V+ ER+ L ++ K+Y NK+ + F +GD V+
Sbjct: 1401 R-------RLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVYLKA 1453
Query: 1998 LPINKKDK--RYRKWVPNWEGPFTVEKVLVNNAYSI 2031
+ + +K P + GP+ V + + AY +
Sbjct: 1454 MTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKL 1489
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 234 bits (598), Expect = 3e-61
Identities = 148/451 (32%), Positives = 246/451 (53%), Gaps = 19/451 (4%)
Query: 1008 VKLLKEFKDCFAWDYDEMPGLSREVVE-LKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1066
+++++EF+D F + GL + + ++ P+ + P R P + ++K++++
Sbjct: 457 IRVVQEFQDVF----QSLQGLPPSQSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLK 512
Query: 1067 RLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1126
LL FIR + W A V+ V KK+G R+CID+R+LN T K+ Y +P + ++D
Sbjct: 513 DLLGKGFIRPSTS-PWGAPVLFVKKKDGSFRLCIDYRELNRVTVKNRYPLPRIDELLDQL 571
Query: 1127 AGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMN 1186
G S +D SGY+QI IAE DV KTAFR G +E+VVMPFGL NA A + R+MN
Sbjct: 572 RGATCFSKIDLTSGYHQIPIAEADVRKTAFRT--RYGHFEFVVMPFGLTNAPAVFMRLMN 629
Query: 1187 TIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1246
++F +F++ F+ ++IDDI+V S S ++ +HLR+ E++R+ L KC+F F
Sbjct: 630 SVFQEFLDEFVIIFIDDILVYSKSPEEQEVHLRRVMEKLREQKLFAKLSKCSFWQREMGF 689
Query: 1247 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLL 1306
LG +V +G+ ++ K +AI D PT+ +++S LG + RRF+ + + P+
Sbjct: 690 LGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLGWAGYYRRFVKGFASMAQ---PMT 746
Query: 1307 RLKKEDA-FRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQED 1365
+L +D F W E ++ F LK L+S P++A P G+P +Y A+ +G +L Q
Sbjct: 747 KLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQHG 806
Query: 1366 EDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHM 1425
+ I Y SR L E Y + + F+ + Y+ V VF+ + +K++
Sbjct: 807 -----KVIAYASRQLMKHEGNYPTHDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYI 861
Query: 1426 LSKPILHSRIGKWALALTEYSL--TYAPLKA 1454
++P L+ R +W + +Y L Y P KA
Sbjct: 862 FTQPELNLRQRRWMELVADYDLEIAYHPGKA 892
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 233 bits (595), Expect = 7e-61
Identities = 151/462 (32%), Positives = 251/462 (53%), Gaps = 22/462 (4%)
Query: 1008 VKLLKEFKDCFAWDYDEMPGLSREVVE-LKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIE 1066
+++++EF+D F + GL + + ++ P+ + P R P + ++K+++E
Sbjct: 483 IRVIQEFEDVF----QSLQGLPPSRSDPFTIELEPGTAPLSKAPYRMAPAEMTELKKQLE 538
Query: 1067 RLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSA 1126
LL FIR + W A V+ V KK+G R+CID+R LN T K++Y +P + ++D
Sbjct: 539 DLLGKGFIRPSTS-PWGAPVLFVKKKDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQL 597
Query: 1127 AGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMN 1186
G S +D SGY+QI IAE DV KTAFR G +E+VVMPF L NA A + R+MN
Sbjct: 598 RGATCFSKIDLTSGYHQIPIAEADVRKTAFRT--RYGHFEFVVMPFALTNAPAAFMRLMN 655
Query: 1187 TIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDF 1246
++F +F++ F+ ++IDDI+V S S ++H +HLR+ E++R+ L KC+F F
Sbjct: 656 SVFQEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREIGF 715
Query: 1247 LGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLL 1306
LG +V +G+ ++ K +AI D PT+ +++S L + RRF+ + + P+
Sbjct: 716 LGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLRLTGYYRRFVKGFASMAQ---PMT 772
Query: 1307 RLKKEDA-FRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQED 1365
+L +D F W E ++ F LK L+S P++A P G+P +Y A+ +G +L Q
Sbjct: 773 KLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQ-- 830
Query: 1366 EDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHM 1425
+ + I Y SR L E Y + + F+ + Y+ V VF+ + +K++
Sbjct: 831 ---RGKVIAYASRQLRKHEGNYPTHDLEMAVVIFALKIWRSYLYGGKVQVFTDHKSLKYI 887
Query: 1426 LSKPILHSRIGKWALALTEYSL--TYAPLKAVKGQAIADFLA 1465
++P L+ R +W + +Y L Y P KA +AD L+
Sbjct: 888 FNQPELNLRQMRWMELVADYDLEIAYHPGKA---NVVADALS 926
Score = 70.5 bits (171), Expect = 1e-11
Identities = 78/340 (22%), Positives = 133/340 (38%), Gaps = 51/340 (15%)
Query: 1698 ISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASEL 1757
+S H + H KM L R W + +D + C CQ VP +
Sbjct: 1020 LSEAHASMFSIHPGATKMYRDLKRY-YQWVGMKRDVANWVAECDVCQLVKAEHQVPDA-- 1076
Query: 1758 HSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVI--EF 1815
I V +D TK + ++ V+ ++
Sbjct: 1077 ------------------------------IWVIMDRLTKSAHFLAIRKTDGAAVLAKKY 1106
Query: 1816 ISSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLI 1875
+S IV G+P S+ +D+ + F +F G K+ ST Y+ Q +GQ E +TL
Sbjct: 1107 VSEIVKLHGVPVSIVSDRDSKFTFAFWRAFQAKMGTKVQMSTAYHPQTDGQSERTIQTLE 1166
Query: 1876 SLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQ 1935
+++ V +W L V +AY NS + S G PF YG+ P L+ +++
Sbjct: 1167 DMLRMCVLDWGGHWADHLSLVEFAYNNSYQASIGMAPFEALYGRPCWTP----LRWTQVE 1222
Query: 1936 RQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAY-NKKVRAKSFMLGDYVW 1994
+ I DY V ER+ VL ++ ++Y +K+ R F +GD V+
Sbjct: 1223 -ERSIYGADY--------VQETTERIRVLKLNMKEAQARQRSYADKRRRELEFEVGDRVY 1273
Query: 1995 KVVLPINKKDKRY--RKWVPNWEGPFTVEKVLVNNAYSIK 2032
+ + ++ K P + GPF + + + AY ++
Sbjct: 1274 LKMAMLRGPNRSILETKLSPRYMGPFRIVERVGPVAYRLE 1313
>At4g03800
Length = 637
Score = 229 bits (584), Expect = 1e-59
Identities = 123/341 (36%), Positives = 201/341 (58%), Gaps = 25/341 (7%)
Query: 1025 MPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLA 1084
MP + ++ +L + KP+KQ R+ + + +I++LL+ IR +Y DW+A
Sbjct: 321 MPDIDPSIICHELNVDPRFKPLKQKRRKLGVERAKAVNGKIDKLLKIGSIREVQYPDWVA 380
Query: 1085 NVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQI 1144
V V KKNGK RVCIDF DLN A PKD + +P + +V+S AG+E L+ +D + GYNQI
Sbjct: 381 ITVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHIDRLVESTAGNELLTFMDAFLGYNQI 440
Query: 1145 FIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDI 1204
+ ED KT+F + + GATYQ ++N +F++ + M+V IDD
Sbjct: 441 MMNPEDQEKTSF-----------------ITDRGATYQWLVNKMFNEHLRKTMEVSIDDT 483
Query: 1205 VVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAK 1264
+VKS ++DH+ HL + FE + +Y +K+N KC FGV +G+FLG++V K+GIE N N+
Sbjct: 484 LVKSLKKEDHVKHLGECFEILNQYQMKLNLAKCTFGVPSGEFLGYIVTKRGIEANPNQIN 543
Query: 1265 AILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAF 1324
A L T + K++Q L G+I + ++K+ F + LK + F W+ + ++AF
Sbjct: 544 AFLKTPSLRNFKEVQRLTGRI------ASRSTDKSLPFYQI--LKGNNGFLWDEKCEEAF 595
Query: 1325 DELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQED 1365
+LK YL++PP+++ P + + LY+ ++ + +L +ED
Sbjct: 596 RQLKAYLTTPPVLSKPEADEKLYLYVFVSNHAVSGVLVRED 636
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 225 bits (574), Expect = 2e-58
Identities = 153/483 (31%), Positives = 253/483 (51%), Gaps = 38/483 (7%)
Query: 991 RPTYISALIDPELKDRMVK----LLKEFKDCFAWDYDEMPGLSREVVE-LKLPIKEDKKP 1045
RPT S +I ++M++ EF+D F + GL + + ++ P
Sbjct: 423 RPTSGSLVISAVQAEKMIEKGCEAYLEFEDVF----QSLQGLPPSRSDPFTIELEPGTAP 478
Query: 1046 VKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDL 1105
+ + P R P + ++K+++E LL A V+ V KK+G R+CID+R L
Sbjct: 479 LSKAPYRMAPAEMAELKKQLEDLLG-------------APVLFVKKKDGSFRLCIDYRGL 525
Query: 1106 NAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTY 1165
N T K++Y +P + ++D G S +D SGY+ I IAE DV KTAFR G +
Sbjct: 526 NWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADVRKTAFRT--RYGHF 583
Query: 1166 EWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERM 1225
E+VVMPFGL NA A + R+MN++F + ++ F+ ++IDDI+V S S ++H +HLR+ E++
Sbjct: 584 EFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHEVHLRRVMEKL 643
Query: 1226 RKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKI 1285
R+ L KC+F FLG +V +G+ ++ K +AI D PT+ +++S LG
Sbjct: 644 REQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTPTNATEIRSFLGLA 703
Query: 1286 NFLRRFIANLSEKTKSFSPLLRLKKEDA-FRWEAEHQKAFDELKVYLSSPPIMASPIRGK 1344
+ RRF+ + + P+ +L +D F W E ++ F LK L+S P++A P G+
Sbjct: 704 GYYRRFVKGFASMAQ---PMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGE 760
Query: 1345 PMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKL 1404
P +Y A+ +G +L Q + + I Y SR L E Y + + F+
Sbjct: 761 PYMVYTDASGVGLGCVLMQ-----RGKVIAYASRQLRKHEGNYPTHDLEMAAVIFALKIW 815
Query: 1405 KYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSL--TYAPLKAVKGQAIAD 1462
+ Y+ +V VF+ + +K++ ++P L+ R +W + +Y L Y P KA +AD
Sbjct: 816 RSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLEIAYHPGKA---NVVAD 872
Query: 1463 FLA 1465
L+
Sbjct: 873 ALS 875
Score = 77.8 bits (190), Expect = 6e-14
Identities = 78/309 (25%), Positives = 127/309 (40%), Gaps = 23/309 (7%)
Query: 1731 KDCMEYAKRCQDCQRHAGIQHVPASELHSI-IKPWPFRGWALDLIGEINPCSSRQHKYII 1789
+D + C CQ VP L S+ I W + +D + + SR I
Sbjct: 941 EDVANWVAECDVCQLVKAEHQVPGGMLQSLPIPEWKWDFITIDFV--VGLPVSRTKDAIW 998
Query: 1790 VAIDYFTKWVEAIPLQNVTQDTVI--EFISSIVYRFGLPESLTTDQGTVFVGQKVASFAE 1847
V +D TK + ++ V+ +++S IV G+P S+ +D+ + F +F
Sbjct: 999 VIVDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQA 1058
Query: 1848 SWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKES 1907
G K+ ST Y+ Q GQ E +TL +++ V +W L V +AY NS S
Sbjct: 1059 EMGTKVQMSTAYHPQTYGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPAS 1118
Query: 1908 TGATPFRLAYGQEAVLPVEVYLQSCRIQ-RQEEIPSEDYWNMMLDELVNLDEERLLVLD- 1965
G PF Y + P+ C Q + I DY V ER+ VL
Sbjct: 1119 IGMAPFEALYERPCRTPL------CLTQVGERSIYGADY--------VQETTERIRVLKL 1164
Query: 1966 TLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRY--RKWVPNWEGPFTVEKV 2023
+ +DR +K+ R F +GD V+ + + ++ K P + GPF + +
Sbjct: 1165 NMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSISETKLSPRYMGPFRIVER 1224
Query: 2024 LVNNAYSIK 2032
+ AY ++
Sbjct: 1225 VGPVAYRLE 1233
>At3g11970 hypothetical protein
Length = 1499
Score = 223 bits (569), Expect = 7e-58
Identities = 150/459 (32%), Positives = 241/459 (51%), Gaps = 18/459 (3%)
Query: 1009 KLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERL 1068
++L E+ D F + +P RE K+ + E PV Q P R+ +I + +E L
Sbjct: 567 EVLNEYPDIFI-EPTALPPF-REKHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDL 624
Query: 1069 LRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAG 1128
L ++ A + + VV V KK+G R+C+D+R+LN T KD + +P+ E ++D G
Sbjct: 625 LTNGTVQ-ASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGG 683
Query: 1129 HEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTI 1188
S +D +GY+Q+ + +D+ KTAF+ G +E++VMPFGL NA AT+Q +MN I
Sbjct: 684 AVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHS--GHFEYLVMPFGLTNAPATFQGLMNFI 741
Query: 1189 FHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLG 1248
F F+ F+ V+ DDI+V S S ++H HL++ FE MR L KCAF V ++LG
Sbjct: 742 FKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLG 801
Query: 1249 FVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRL 1308
+ +GIE + K KA+ + PT+ KQL+ LG + RRF+ + PL L
Sbjct: 802 HFISAQGIETDPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIA---GPLHAL 858
Query: 1309 KKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDS 1368
K DAF W A Q+AF++LK L P+++ P+ K + A IG++L QE
Sbjct: 859 TKTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEG--- 915
Query: 1369 KERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSK 1428
+ Y+SR L + ++ EK L + F+ K ++Y+ ++ + +K++L +
Sbjct: 916 --HPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQ 973
Query: 1429 PILHSRIGKWALALTE--YSLTYAPLKAVKGQAIADFLA 1465
+ +W L E Y + Y + K +AD L+
Sbjct: 974 RLNTPIQQQWLPKLLEFDYEIQY---RQGKENVVADALS 1009
Score = 80.5 bits (197), Expect = 9e-15
Identities = 70/306 (22%), Positives = 136/306 (43%), Gaps = 28/306 (9%)
Query: 1725 YWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRG--WA---LDLIGEINP 1779
YW ++KD Y + C CQ+ + PA+ +++P P W+ +D I + P
Sbjct: 1111 YWKGMIKDIQAYIRSCGTCQQ---CKSDPAAS-PGLLQPLPIPDTIWSEVSMDFIEGL-P 1165
Query: 1780 CSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIE--FISSIVYRFGLPESLTTDQGTVF 1837
S + I+V +D +K I L + + ++ ++ G P S+ +D+ VF
Sbjct: 1166 VSGGK-TVIMVVVDRLSKAAHFIALSHPYSALTVAHAYLDNVFKLHGCPTSIVSDRDVVF 1224
Query: 1838 VGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVL 1897
+ F G+ L ++ Y+ Q++GQ E N+ L + ++ +P+ W + L
Sbjct: 1225 TSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAE 1284
Query: 1898 WAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLD 1957
+ Y + S+ TPF + YGQ V PV + +P E ++ L +
Sbjct: 1285 YWYNTNYHSSSRMTPFEIVYGQ--VPPVHL----------PYLPGESKVAVVARSLQERE 1332
Query: 1958 EERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKD---KRYRKWVPNW 2014
+ L + L R + R+ + ++ + F +GDYV+ + P ++ + +K P +
Sbjct: 1333 DMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDYVYVKLQPYRQQSVVMRANQKLSPKY 1392
Query: 2015 EGPFTV 2020
GP+ +
Sbjct: 1393 FGPYKI 1398
>At1g36590 hypothetical protein
Length = 1499
Score = 223 bits (569), Expect = 7e-58
Identities = 150/459 (32%), Positives = 241/459 (51%), Gaps = 18/459 (3%)
Query: 1009 KLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERL 1068
++L E+ D F + +P RE K+ + E PV Q P R+ +I + +E L
Sbjct: 567 EVLNEYPDIFI-EPTALPPF-REKHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDL 624
Query: 1069 LRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAG 1128
L ++ A + + VV V KK+G R+C+D+R+LN T KD + +P+ E ++D G
Sbjct: 625 LTNGTVQ-ASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGG 683
Query: 1129 HEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTI 1188
S +D +GY+Q+ + +D+ KTAF+ G +E++VMPFGL NA AT+Q +MN I
Sbjct: 684 AVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHS--GHFEYLVMPFGLTNAPATFQGLMNFI 741
Query: 1189 FHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLG 1248
F F+ F+ V+ DDI+V S S ++H HL++ FE MR L KCAF V ++LG
Sbjct: 742 FKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLG 801
Query: 1249 FVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRL 1308
+ +GIE + K KA+ + PT+ KQL+ LG + RRF+ + PL L
Sbjct: 802 HFISAQGIETDPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIA---GPLHAL 858
Query: 1309 KKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDS 1368
K DAF W A Q+AF++LK L P+++ P+ K + A IG++L QE
Sbjct: 859 TKTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEG--- 915
Query: 1369 KERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSK 1428
+ Y+SR L + ++ EK L + F+ K ++Y+ ++ + +K++L +
Sbjct: 916 --HPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQ 973
Query: 1429 PILHSRIGKWALALTE--YSLTYAPLKAVKGQAIADFLA 1465
+ +W L E Y + Y + K +AD L+
Sbjct: 974 RLNTPIQQQWLPKLLEFDYEIQY---RQGKENVVADALS 1009
Score = 78.6 bits (192), Expect = 4e-14
Identities = 75/329 (22%), Positives = 145/329 (43%), Gaps = 36/329 (10%)
Query: 1702 HDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSII 1761
H G HQ ++K + + +GM +KD Y + C CQ+ + PA+ ++
Sbjct: 1096 HSGRDVTHQ---RVKGLFYSKGM-----IKDIQAYIRSCGTCQQ---CKSDPAAS-PGLL 1143
Query: 1762 KPWPFRG--WA---LDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIE-- 1814
+P P W+ +D I + P S + I+V +D +K I L + +
Sbjct: 1144 QPLPIPDTIWSEVSMDFIEGL-PVSGGK-TVIMVVVDRLSKAAHFIALSHPYSALTVAQA 1201
Query: 1815 FISSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTL 1874
++ ++ G P S+ +D+ VF + F G+ L ++ Y+ Q++GQ E N+ L
Sbjct: 1202 YLDNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCL 1261
Query: 1875 ISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRI 1934
+ ++ +P+ W + L + Y + S+ TPF + YGQ V PV +
Sbjct: 1262 ETYLRCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQ--VPPVHL------- 1312
Query: 1935 QRQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVW 1994
+P E ++ L ++ L + L R + R+ + ++ + F +GDYV+
Sbjct: 1313 ---PYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDYVY 1369
Query: 1995 KVVLPINKKD---KRYRKWVPNWEGPFTV 2020
+ P ++ + +K P + GP+ +
Sbjct: 1370 VKLQPYRQQSVVMRANQKLSPKYFGPYKI 1398
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 219 bits (559), Expect = 1e-56
Identities = 138/387 (35%), Positives = 213/387 (54%), Gaps = 17/387 (4%)
Query: 1002 ELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKI 1061
ELKD + ++ EF D FA P S + ++ P+ + P R P + ++
Sbjct: 116 ELKD--ILIVNEFSDVFAAVSGVPPDRSDPFT---IELEPGTTPISKAPYRMAPAEMAEL 170
Query: 1062 KEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEM 1121
K+++E LL FIR + W A V+ V KK+G R+CID+R LN T K++Y +P +
Sbjct: 171 KKQLEELLAKGFIRPSSS-PWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDE 229
Query: 1122 MVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATY 1181
++D G ++ S +D SGY+QI I DV KTAFR G +E+VVMPFGL NA A +
Sbjct: 230 LMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRT--RYGHFEFVVMPFGLTNAPAAF 287
Query: 1182 QRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGV 1241
++MN +F DF++ F+ ++IDDI+V S S + H HLR ER+R++ L K +F
Sbjct: 288 MKMMNGVFRDFLDEFVIIFIDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFSFWQ 347
Query: 1242 IAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKS 1301
+ FLG V+ +G+ ++ K ++I + P + +++S LG + RRF+ + + +
Sbjct: 348 RSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQ- 406
Query: 1302 FSPLLRLK-KEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSM 1360
PL RL K+ AF W E +K+F ELK L + P++ P G+P +Y A+ +G +
Sbjct: 407 --PLTRLTGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCV 464
Query: 1361 LAQEDEDSKERAIFYLSRVLNDAETRY 1387
L Q K I Y SR L E Y
Sbjct: 465 LMQ-----KGSVIAYASRQLRKHEKNY 486
Score = 88.6 bits (218), Expect = 3e-17
Identities = 78/336 (23%), Positives = 142/336 (42%), Gaps = 20/336 (5%)
Query: 1702 HDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSI- 1760
H H KM L R +W + KD + +C CQ VP+ L ++
Sbjct: 560 HQSKFSIHPGSNKMYRDLKRY-YHWVGMRKDVARWVAKCPTCQLVKAEHQVPSGLLQNLP 618
Query: 1761 IKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVI--EFISS 1818
I W + +D + + +H + V +D TK + + + +I ++I
Sbjct: 619 ISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDE 678
Query: 1819 IVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLI 1878
I+ G+P S+ +D+ T F + +F ++ G ++ ST Y+ Q +GQ E +TL ++
Sbjct: 679 IMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLSTAYHPQTDGQSERTIQTLEDML 738
Query: 1879 KKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQE 1938
+ V NW + L + +AY NS + S G +P+ YG+ P+ C E
Sbjct: 739 RACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALYGRACRTPL------CWTPVGE 792
Query: 1939 EIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAY-NKKVRAKSFMLGDYVWKVV 1997
+ +V+ ER+ L ++ K+Y NK+ + F +GD V+
Sbjct: 793 R-------RLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVYLKA 845
Query: 1998 LPINKKDK--RYRKWVPNWEGPFTVEKVLVNNAYSI 2031
+ + +K P + GP+ V + + AY +
Sbjct: 846 MTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKL 881
>At1g35370 hypothetical protein
Length = 1447
Score = 212 bits (540), Expect = 2e-54
Identities = 150/477 (31%), Positives = 246/477 (51%), Gaps = 33/477 (6%)
Query: 995 ISALIDPELKDRMVK-LLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRF 1053
ISAL +++ +V+ +++EF D FA D P RE + K+ + E PV Q P R+
Sbjct: 531 ISALTSDVVEESVVQNIVEEFPDVFAEPTDLPP--FREKHDHKIKLLEGANPVNQRPYRY 588
Query: 1054 HPDVLVKIKEEIERLLRCKFIRTARYVD---WLANVVPVIKKNGKMRVCIDFRDLNAATP 1110
+V K+EI+++++ V + + VV V KK+G R+C+D+ +LN T
Sbjct: 589 ----VVHQKDEIDKIVQDMIKSGTIQVSSSPFASPVVLVKKKDGTWRLCVDYTELNGMTV 644
Query: 1111 KDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVM 1170
KD + +P+ E ++D G S +D +GY+Q+ + +D+ KTAF+ G +E++VM
Sbjct: 645 KDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDPDDIQKTAFKTHN--GHFEYLVM 702
Query: 1171 PFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGL 1230
FGL NA AT+Q +MN++F DF+ F+ V+ DDI++ S S ++H HLR FE MR + L
Sbjct: 703 LFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSSIEEHKEHLRLVFEVMRLHKL 762
Query: 1231 KMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRR 1290
F + + LG + + IE + K +A+ + PT+ KQ++ LG + RR
Sbjct: 763 --------FAKGSKEHLGHFISAREIETDPAKIQAVKEWPTPTTVKQVRGFLGFAGYYRR 814
Query: 1291 FIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYI 1350
F+ N PL L K D F W E Q AFD LK L + P++A P+ K +
Sbjct: 815 FVRNFGVIA---GPLHALTKTDGFCWSLEAQSAFDTLKAVLCNAPVLALPVFDKQFMVET 871
Query: 1351 SATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKP 1410
A I ++L Q K + Y+SR L + ++ EK L F+ K ++Y+ P
Sbjct: 872 DACGQGIRAVLMQ-----KGHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLLP 926
Query: 1411 IDVMVFSHYDIIKHMLSKPILHSRIGKWALALTE--YSLTYAPLKAVKGQAIADFLA 1465
++ + +K++L + + +W L E Y + Y + K +AD L+
Sbjct: 927 SHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQY---RQGKENLVADALS 980
Score = 77.0 bits (188), Expect = 1e-13
Identities = 69/322 (21%), Positives = 138/322 (42%), Gaps = 28/322 (8%)
Query: 1725 YWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGW---ALDLIGEINPCS 1781
YW ++KD + + C CQ+ L + P P + W ++D I + +
Sbjct: 1082 YWKGMVKDIQAFIRSCGTCQQCKSDNAAYPGLLQPL--PIPDKIWCDVSMDFIEGLP--N 1137
Query: 1782 SRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIE--FISSIVYRFGLPESLTTDQGTVFVG 1839
S I+V +D +K + L + + F+ ++ G P S+ +D+ +F
Sbjct: 1138 SGGKSVIMVVVDRLSKAAHFVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTS 1197
Query: 1840 QKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWA 1899
F + G++L S+ Y+ Q++GQ E N+ L + ++ +P W++ L +
Sbjct: 1198 DFWKEFFKLQGVELRMSSAYHPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYW 1257
Query: 1900 YRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEE 1959
Y + S+ TPF L YGQ P+ + +P + ++ L +
Sbjct: 1258 YNTNYHSSSQMTPFELVYGQAP--PIHL----------PYLPGKSKVAVVARSLQERENM 1305
Query: 1960 RLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYR---KWVPNWEG 2016
L + L R + R+ + ++ ++F +GD+V+ + P ++ R K P + G
Sbjct: 1306 LLFLKFHLMRAQHRMKQFADQHRTERTFDIGDFVYVKLQPYRQQSVVLRVNQKLSPKYFG 1365
Query: 2017 PFTV----EKVLVNNAYSIKEL 2034
P+ + +V+V N + +L
Sbjct: 1366 PYKIIEKCGEVMVGNVTTSTQL 1387
>At2g13230 pseudogene
Length = 353
Score = 184 bits (467), Expect = 5e-46
Identities = 115/352 (32%), Positives = 186/352 (52%), Gaps = 36/352 (10%)
Query: 1581 KFDEARLGHVSRVDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPS-DLD-------I 1632
+F E +L + R +N A+ LA +AS ++ +I V+ PS +LD I
Sbjct: 14 QFHEFKLTRIPRGENTSADALAALAS---TSDLNMRRVIPVEFIEKPSIELDKVEHAFPI 70
Query: 1633 LVI-------DNM----APNDWRKPIVDYLQNPVGTTD----RKTKYRAMSYVIMGNELF 1677
V+ DN ++W +PI +++ D RK K A +V++ +LF
Sbjct: 71 QVVAGYEDASDNSPSLGCDSEWIEPIRNFISKRKVRLDKWEARKLKAEAARFVLVEEKLF 130
Query: 1678 KKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYA 1737
K + G L+ C+ ++ A + VH G CG + G + + R G +WPT++KDC
Sbjct: 131 KWRLSGPLMTCVEKEAARKVMKEVHGGSCGNNSGGRALAIKIKRLGYFWPTMIKDC---- 186
Query: 1738 KRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTK 1797
+ CQR+A H+PA L SI P+P W++D+IG ++ +S+Q K ++V DYF+K
Sbjct: 187 ---EKCQRNAPTIHLPAELLSSIASPYPLMRWSMDIIGPMH--ASKQKKLVLVLTDYFSK 241
Query: 1798 WVEAIPLQNVTQDTVIEFI-SSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTS 1856
EA ++ + V F+ I+ R G+P + TD G+ F+ + F + WGI+L S
Sbjct: 242 LREAESYASIKEAQVESFMWKHILCRHGVPYEIVTDNGSQFISTRFQGFCDKWGIRLSKS 301
Query: 1857 TPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKEST 1908
TP Y Q NGQ +AANKT++ +KK + K +W L VLW++R +P+ +T
Sbjct: 302 TPRYLQGNGQADAANKTILDGLKKRLTAKKGSWVDELEGVLWSHRTTPRRAT 353
>At2g06890 putative retroelement integrase
Length = 1215
Score = 183 bits (464), Expect = 1e-45
Identities = 121/380 (31%), Positives = 195/380 (50%), Gaps = 16/380 (4%)
Query: 1091 KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEED 1150
+K+G R+C D R +N T K + +P + M+D G S +D SGY+QI + E D
Sbjct: 464 QKDGSWRMCFDCRAINNVTVKYCHPIPRLDDMLDELHGSSIFSKIDLKSGYHQIRMNEGD 523
Query: 1151 VSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPS 1210
KTAF+ G YEW+VMPFGL +A +T+ R+MN + FI F+ VY DDI+V S S
Sbjct: 524 EWKTAFKTKH--GLYEWLVMPFGLTHAPSTFMRLMNHVLRAFIGIFVIVYFDDILVYSES 581
Query: 1211 RDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTS 1270
+H+ HL +RK L N KC F FLGFVV G+++++ K KAI D
Sbjct: 582 LREHIEHLDSVLNVLRKEELYANLKKCTFCTDNLVFLGFVVSADGVKVDEEKVKAIRDWP 641
Query: 1271 PPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVY 1330
P + +++S G F RRF + S + + +KK+ F+WE ++AF LK
Sbjct: 642 SPKTVGEVRSFHGLAGFYRRFFKDFSTIVAPLTEV--MKKDVGFKWEKAQEEAFQSLKDK 699
Query: 1331 LSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMI 1390
L++ P++ K ++ A+ IG++L Q+ ++ I + S L A Y
Sbjct: 700 LTNAPVLILSEFLKTFEIECDASGIGIGAVLMQD-----QKLIAFFSEKLGGATLNYPTY 754
Query: 1391 EKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYA 1450
+K L + + ++Y+ P ++ + ++ +KH+ + L+ R +W E+ T+A
Sbjct: 755 DKELYALVRALQRWQHYLWPKVFVIHTDHESLKHLKGQQKLNKRHARW----VEFIETFA 810
Query: 1451 ---PLKAVKGQAIADFLADQ 1467
K K +AD L+ +
Sbjct: 811 YVIKYKKGKDNVVADALSQR 830
Score = 37.4 bits (85), Expect = 0.092
Identities = 33/130 (25%), Positives = 55/130 (41%), Gaps = 27/130 (20%)
Query: 1911 TPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQ 1970
+PF++ YG + P ++ IP + L E V++D ++ L +
Sbjct: 988 SPFQIVYGFNPISPFDL------------IP------LPLSERVSIDGKKKAELVQQIHE 1029
Query: 1971 KDR---------IAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVE 2021
R AK NK R + F +GD VW + +R K +P +GPF +
Sbjct: 1030 NARRNIEEKTKLYAKQANKGRREQIFEVGDMVWIHLRKERFPAQRKSKLMPRIDGPFKII 1089
Query: 2022 KVLVNNAYSI 2031
K + +NAY +
Sbjct: 1090 KRINDNAYQL 1099
>At4g07600
Length = 630
Score = 182 bits (463), Expect = 1e-45
Identities = 149/472 (31%), Positives = 219/472 (45%), Gaps = 61/472 (12%)
Query: 931 IEENEQSA----EECGFEDLCNLRLDCIYDDG-----PLGFEKPVVDFSKKMEAQDPLEE 981
IEE +Q A EE ED N L DG LG++K ++D K ME P EE
Sbjct: 208 IEEMDQLADELLEELAQEDHLNSALTKTGQDGFLHLKVLGYQK-LLDSHKAMEESKPFEE 266
Query: 982 VDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKE 1041
++ G + + + +++ S + +LK P
Sbjct: 267 LN-GPATKVMVMNEEGSTQVQPAHSRTYSTNHSTSTVSNSGEQIIPTSDDWSKLKAP--- 322
Query: 1042 DKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCID 1101
PV+ +P++ V+ K+E+ +P+ G +R+CID
Sbjct: 323 KVSPVQCIPKKGGITVIKNEKDEL---------------------IPIRTITG-LRMCID 360
Query: 1102 FRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGA 1161
+R+LNAA+ D + +P + M++ A H Y LDGYSG+ QI I D KT F CP
Sbjct: 361 YRNLNAASRNDHFPLPFTDQMLERLANHPYYCFLDGYSGFFQIPIHPNDHEKTTFTCP-- 418
Query: 1162 LGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKS 1221
GT+ + MPFGL NA AT+QR M +IF D IE ++V++DD V PS LL+L +
Sbjct: 419 YGTFAYERMPFGLCNAPATFQRCMTSIFSDIIEEMVEVFMDDFSVYGPSFSSCLLNLGRV 478
Query: 1222 FERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSL 1281
R + L +N KC F V G LG + +KGIE++K
Sbjct: 479 LTRCEETNLVLNWEKCHFMVKEGIMLGHKISEKGIEVDKG------------------CF 520
Query: 1282 LGKINFLRRFIANLSEKTKSFSPLLRLK-KEDAFRWEAEHQKAFDELKVYLSSPPIMASP 1340
LG F RRFI + S K PL RL KE F ++ + K+F +K L S P++ +
Sbjct: 521 LGHAGFYRRFIKDFS---KIARPLTRLLCKETEFEFDEDCPKSFHTIKEALVSAPVVRAT 577
Query: 1341 IRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEK 1392
P ++ A D +G++L Q+ D K I+Y SR L++A+ RY EK
Sbjct: 578 NWDYPFEIMCDALDYAVGAVLGQK-IDKKLHVIYYASRTLDEAQGRYATTEK 628
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.144 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,698,268
Number of Sequences: 26719
Number of extensions: 1773468
Number of successful extensions: 7106
Number of sequences better than 10.0: 117
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 6698
Number of HSP's gapped (non-prelim): 255
length of query: 2064
length of database: 11,318,596
effective HSP length: 115
effective length of query: 1949
effective length of database: 8,245,911
effective search space: 16071280539
effective search space used: 16071280539
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0269b.2


