

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269a.3
(490 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
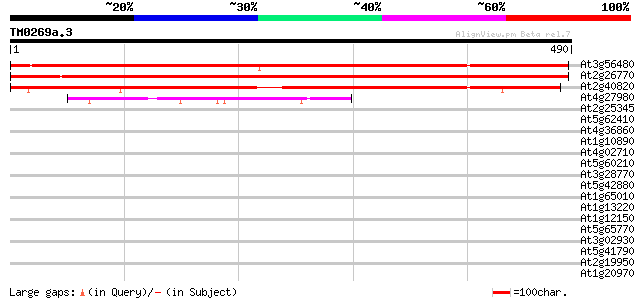
Sequences producing significant alignments: (bits) Value
At3g56480 unknown protein 630 0.0
At2g26770 unknown protein 583 e-167
At2g40820 hypothetical protein 560 e-160
At4g27980 putative protein 51 1e-06
At2g25345 putative protein 42 0.001
At5g62410 chromosomal protein - like 41 0.001
At4g36860 Unknown protein 41 0.001
At1g10890 unknown protein 41 0.001
At4g02710 predicted protein of unknown function 40 0.003
At5g60210 putative protein 40 0.004
At3g28770 hypothetical protein 40 0.004
At5g42880 putative protein 39 0.007
At1g65010 hypothetical protein 38 0.011
At1g13220 putative nuclear matrix constituent protein 38 0.011
At1g12150 hypothetical protein 38 0.011
At5g65770 nuclear matrix constituent protein 1 (NMCP1)-like 38 0.014
At3g02930 unknown protein 38 0.014
At5g41790 myosin heavy chain-like protein 37 0.032
At2g19950 hypothetical protein 37 0.032
At1g20970 hypothetical protein 37 0.032
>At3g56480 unknown protein
Length = 490
Score = 630 bits (1626), Expect = 0.0
Identities = 329/492 (66%), Positives = 402/492 (80%), Gaps = 6/492 (1%)
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MTK+ PEIE + + VP VS DVSF SD FP YKLG D+QI+EE ED +GPS+K+ +E
Sbjct: 1 MTKVCPEIEGTLSLSVVP-VSVDVSFASDHFPTYKLGPDNQIVEEPKEDEKGPSVKETVE 59
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 120
+E+ LSDQHKR+SVRDLA KFDKNLAAA L+NEAK+REVASLEGHV+LKKLRDALEY+
Sbjct: 60 KESELLSDQHKRLSVRDLASKFDKNLAAAVSLANEAKLREVASLEGHVMLKKLRDALEYM 119
Query: 121 RVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 180
R R G+NK+DVE AISMVEALAVKLTQNEGELIQEKFEVKKL NFLKQ SEDAKKLVNQ
Sbjct: 120 RGRTDGQNKQDVETAISMVEALAVKLTQNEGELIQEKFEVKKLGNFLKQTSEDAKKLVNQ 179
Query: 181 EKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQ--DVDGLVEEVQEARRIKLLHQPS 238
EKSF+CAEIE+AR+VVLR+GEA EEQE+ S+AS+ Q DV+ LVEEVQEAR+IK +H P+
Sbjct: 180 EKSFSCAEIETARAVVLRLGEAFEEQERISEASRAQGPDVEKLVEEVQEARQIKRMHHPT 239
Query: 239 KVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALGSYLKVQP 298
KVM M++EL LR++I+EK + S+KL KE+ + KR EE+KS P++L G ++LGS L+++
Sbjct: 240 KVMGMQHELHGLRNRIQEKYMNSVKLHKEIAIIKRVEESKSCPFVLEGKQSLGSCLRIRV 299
Query: 299 RSGE-VPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDIVSNGKKLT 357
+ + P +S CS QWYR + E S RE ISGA +S+YAP+PFDVGRILQ DI+SNG+K T
Sbjct: 300 NAQDNAPDLSNCSIQWYRAACETSRREAISGAIQSMYAPEPFDVGRILQADILSNGQKFT 359
Query: 358 LTT-NPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCRG 416
+TT +P+ SGL S VE+L+RKSN++F VVISQMNG+D++SRS H F VG+ RIKL RG
Sbjct: 360 VTTDDPVDPDSGLPSRVESLMRKSNSEFSVVISQMNGQDYASRS-HVFTVGKTRIKLSRG 418
Query: 417 WITKAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIMIARKYA 476
WITKARE+YS SMQLCGVRG+ KA+FWQ RK L+F+LTFESE ERN AI +ARKYA
Sbjct: 419 WITKARELYSTSMQLCGVRGNIKAPTKAVFWQPRKSLTFILTFESEHERNAAIALARKYA 478
Query: 477 LDCNVVLAGPDD 488
DC+V L GPDD
Sbjct: 479 FDCSVTLLGPDD 490
>At2g26770 unknown protein
Length = 496
Score = 583 bits (1502), Expect = e-167
Identities = 300/489 (61%), Positives = 388/489 (78%), Gaps = 2/489 (0%)
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MT+++ + + + VP VSADV F S RFP Y++GA+ QI + +D + S+K+V+
Sbjct: 1 MTRVTRDFRDSLQRDGVPAVSADVKFASSRFPNYRIGANDQIFD-VKDDPKVMSMKEVVA 59
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 120
+E L DQ KR+SVRDLA KF+K LAAAAKLS EAK++E SLE HVLLKKLRDALE L
Sbjct: 60 RETAQLMDQQKRLSVRDLAHKFEKGLAAAAKLSEEAKLKEATSLEKHVLLKKLRDALESL 119
Query: 121 RVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 180
R R AGRNK+DVE+AI+MVEALAV+LTQ EGEL EK EVKKL +FLKQASEDAKKLV++
Sbjct: 120 RGRVAGRNKDDVEEAIAMVEALAVQLTQREGELFIEKAEVKKLASFLKQASEDAKKLVDE 179
Query: 181 EKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKV 240
E++FA AEIESAR+ V R+ EAL E EQ S+AS QD++ L++EVQEARRIK+LHQPS+V
Sbjct: 180 ERAFARAEIESARAAVQRVEEALREHEQMSRASGKQDMEDLMKEVQEARRIKMLHQPSRV 239
Query: 241 MAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALGSYLKVQPRS 300
M MEYELRALR+Q+ EKS ++LQK+L M ++ EEN S Y + G+EALGS L+V+P S
Sbjct: 240 MDMEYELRALRNQLAEKSKHFLQLQKKLAMCRKSEENISLVYEIDGTEALGSCLRVRPCS 299
Query: 301 GEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDIVSNGKKLTLTT 360
+ P +SKC+ QWYR SS+GS +E+ISGA KS+YAP+PFDVGR+L DI+ +G L+L+T
Sbjct: 300 NDAPDLSKCTIQWYRSSSDGSKKELISGATKSVYAPEPFDVGRVLHADIIYDGHSLSLST 359
Query: 361 -NPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCRGWIT 419
I A+GLGS+VEAL+RK + DF+VV++QM+G+DH+S S H F+VG+MRIKLC+G
Sbjct: 360 VGKIDPAAGLGSYVEALVRKHDVDFNVVVTQMSGEDHTSESIHLFHVGKMRIKLCKGKTV 419
Query: 420 KAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIMIARKYALDC 479
A+E YS +MQLCGVRG + AA+AL+WQA+KG+SFV+ FESE+ERN AIM+AR++A DC
Sbjct: 420 IAKEYYSSAMQLCGVRGGGNAAAQALYWQAKKGVSFVIAFESERERNAAIMLARRFACDC 479
Query: 480 NVVLAGPDD 488
NV LAGP+D
Sbjct: 480 NVTLAGPED 488
>At2g40820 hypothetical protein
Length = 945
Score = 560 bits (1443), Expect = e-160
Identities = 305/516 (59%), Positives = 382/516 (73%), Gaps = 57/516 (11%)
Query: 1 MTKISPEIEIKMPME--AVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDV 58
MTK+ P+ E K + V P+SADVSF S+ FP YKLG D QI++E ED++ PS+KDV
Sbjct: 1 MTKVCPKTEEKRVVSEAVVEPISADVSFASNHFPTYKLGPDDQIVDEPDEDDKVPSVKDV 60
Query: 59 IEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNE--------------AKIREVASL 104
+ +E +LSDQHK++SVRDLACKFDKNLAAA+KL +E AK+ +V SL
Sbjct: 61 VGKETGDLSDQHKKLSVRDLACKFDKNLAAASKLVDEVPLYFSSSYDYVSVAKLNDVTSL 120
Query: 105 EGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLL 164
EGHV+LKKLRDALE +R R GRN+E VEKAISMVEALAVKLTQNEGELI +KFEVKKL
Sbjct: 121 EGHVMLKKLRDALETMRGRMDGRNREAVEKAISMVEALAVKLTQNEGELIHDKFEVKKLA 180
Query: 165 NFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEE 224
+FLK+AS+DAKKLVNQEKSFACAEIESAR++V+++G A EEQE S+AS+ Q
Sbjct: 181 SFLKKASDDAKKLVNQEKSFACAEIESARALVMKLGGAFEEQELCSKASRDQ-------- 232
Query: 225 VQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYML 284
V+ M++ELR L+ QI+EKS +S+KLQ+E+ + K+ E +KS PY+L
Sbjct: 233 -------------GPVIGMQHELRDLKSQIQEKSAYSVKLQREIAIIKKAEGSKSCPYVL 279
Query: 285 HGSEALGSYLKVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRI 344
G+++LGS L+++ S +SKCS WYR +SE S RE ISGA++S+YAP+PFDVGR+
Sbjct: 280 DGAQSLGSCLRIRASSDSGIDISKCSIHWYRAASESSRREAISGANRSVYAPEPFDVGRV 339
Query: 345 LQVDIVSNGKKLTLTTN-PIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHS 403
+Q DIVSNG+K T+TT+ PI TA+GL S VE+LLRKSN++F VVISQMNG+DH+SRS H
Sbjct: 340 IQADIVSNGQKFTVTTDGPINTAAGLQSRVESLLRKSNSEFTVVISQMNGQDHASRS-HV 398
Query: 404 FNVGRMRIKLCRGWITKAREIYSPSM------------------QLCGVRGDASNAAKAL 445
F VG+ RIKL RGWITKAREIYS SM QLCGVRG+A+ AKAL
Sbjct: 399 FTVGKARIKLSRGWITKAREIYSTSMQLYLKFSFSSSLNGLVTFQLCGVRGNANVPAKAL 458
Query: 446 FWQARKGLSFVLTFESEKERNVAIMIARKYALDCNV 481
FWQ RKGL+F+LTFESE+ERN AI++AR YA DCNV
Sbjct: 459 FWQLRKGLTFLLTFESEQERNAAIVLARTYAYDCNV 494
>At4g27980 putative protein
Length = 565
Score = 51.2 bits (121), Expect = 1e-06
Identities = 65/276 (23%), Positives = 122/276 (43%), Gaps = 37/276 (13%)
Query: 51 QGPSLK-DVIEQEATNLSD-----QHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASL 104
QG S++ DV EQ + Q K S+ + + +FD +A+ K ++ E+
Sbjct: 4 QGDSVESDVTEQNSEGYLKILDKIQAKSSSIFEFSLQFDDVVASLEKKEERLRVVEMKEK 63
Query: 105 EGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMV-EALAVKLT---QNEGELIQEKFEV 160
E +L + + L L ++++E + +V EA ++L Q+E + Q K E
Sbjct: 64 EIGLLEESISRRLSVLE-------EKEIETDLRLVIEASIMRLVLEKQSEDLVTQLKTEE 116
Query: 161 KKLLNFLKQASEDAKKLVNQ---EKSFAC------AEIESARSVVLRIGEALEEQEQDSQ 211
KL FL+ ++ ++LV++ K AC E+E A A E+ +
Sbjct: 117 NKLGLFLRSTTKKLEELVSEFDGRKEEACRVSEKLCELEKAEKEFHLKQRAETERRNEES 176
Query: 212 ASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQ---------IREKSIFSI 262
++ +D+ L E V+E + + + M+ E LR++ I+EK++
Sbjct: 177 EAREKDLRALEEAVKEKTAELKRKEETLELKMKEEAEKLREETELMRKGLEIKEKTLE-- 234
Query: 263 KLQKELTMSKRDEENKSHPYMLHGSEALGSYLKVQP 298
K KEL + + + E S P ++ S L+++P
Sbjct: 235 KRLKELELKQMELEETSRPQLVEAESRKRSNLEIEP 270
>At2g25345 putative protein
Length = 600
Score = 41.6 bits (96), Expect = 0.001
Identities = 54/207 (26%), Positives = 86/207 (41%), Gaps = 22/207 (10%)
Query: 90 AKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQN 149
A+L+ E ++E L + D LE RNKE++E+A+ K+ +
Sbjct: 361 ARLNQETAVKEY--------LNRKVDDLEVELETTKQRNKENLEQALMTERQSVTKMQWD 412
Query: 150 EGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQD 209
EL Q+ FE++ L + S D+K N S + ++ + ++ E L + +
Sbjct: 413 MEELRQKTFEMELKLKSKEDGSSDSKTSGNSTISESHELLQEMDATKQQL-EDLSRRYVE 471
Query: 210 SQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELT 269
+A D+ LV EV+ RR M ME EL + EKS LQ+E
Sbjct: 472 LEAKSKADIKVLVREVKSLRR--------SHMEMEKELTR---SLTEKSDTEKLLQQERI 520
Query: 270 MSKRDEENKSHPYMLHGSEALGSYLKV 296
+ + E + Y E L LKV
Sbjct: 521 IVENTLEARRRLY--SDCEILHDRLKV 545
>At5g62410 chromosomal protein - like
Length = 1175
Score = 41.2 bits (95), Expect = 0.001
Identities = 54/215 (25%), Positives = 94/215 (43%), Gaps = 30/215 (13%)
Query: 111 KKLRDALEYLRVRFAGRNKEDVEKAISMVEAL---AVKLTQNEGELIQEKFEVKKLLNFL 167
K+++ + G K EK S+ + + + KL E L+ EK V+K+++ +
Sbjct: 279 KQIKALTQAKEASMGGEVKTLSEKVDSLAQEMTRESSKLNNKEDTLLGEKENVEKIVHSI 338
Query: 168 KQASEDAKKLVNQEKSFACAEIESARSVVLRIGE---ALEEQEQDSQA-----SKPQDVD 219
ED KK V + + E A + R E LEE E++ Q S +
Sbjct: 339 ----EDLKKSVKERAAAVKKSEEGAADLKQRFQELSTTLEECEKEHQGVLAGKSSGDEEK 394
Query: 220 GLVEEVQEAR------RIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKR 273
L +++++A+ +L +K+ E EL+ + Q+ K +I+++ EL K
Sbjct: 395 CLEDQLRDAKIAVGTAGTELKQLKTKIEHCEKELKERKSQLMSKLEEAIEVENELGARKN 454
Query: 274 DEEN-----KSHPYMLHGSEAL----GSYLKVQPR 299
D E+ +S PY EAL G+ L+V R
Sbjct: 455 DVEHVKKALESIPYNEGQMEALEKDRGAELEVVQR 489
>At4g36860 Unknown protein
Length = 542
Score = 41.2 bits (95), Expect = 0.001
Identities = 38/158 (24%), Positives = 74/158 (46%), Gaps = 24/158 (15%)
Query: 128 NKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACA 187
+KE++E AI A+ L++ E +I + + KK++ + + ED ++++ + A
Sbjct: 66 DKEEIECAI------ALSLSEQE-HVIPQDDKGKKIIEYKSETEEDDDDDEDEDEEYMRA 118
Query: 188 EIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYEL 247
++E+A R+ +A E+E+ +A E Q KLL +K E E+
Sbjct: 119 QLEAAEEEERRVAQAQIEEEEKRRA-----------EAQLEETEKLL---AKARLEEEEM 164
Query: 248 RALRDQIREKSIFSIKLQKELTMS---KRDEENKSHPY 282
R + Q+ E + + LQ+ + + + D N PY
Sbjct: 165 RRSKAQLEEDELLAKALQESMNVGSPPRYDPGNILQPY 202
>At1g10890 unknown protein
Length = 592
Score = 41.2 bits (95), Expect = 0.001
Identities = 36/159 (22%), Positives = 76/159 (47%), Gaps = 8/159 (5%)
Query: 127 RNKEDVEKAISMVEALAVKLTQNE-GELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFA 185
R + E + ++E VK + + ++E + +K+ + E+ +K +N+E
Sbjct: 70 RKRRQREAELKLIEEETVKRVEEAIRKKVEESLQSEKIKMEILTLLEEGRKRLNEE---V 126
Query: 186 CAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEY 245
A++E + L + EE+EQ + + + + ++ V+EA+R + + + K
Sbjct: 127 AAQLEEEKEASLIEAKEKEEREQQEKEERERIAEENLKRVEEAQRKEAMERQRKEEERYR 186
Query: 246 ELRALRDQIREKSIFSIKLQKE---LTMSKRDEENKSHP 281
EL L+ Q +E+++ K ++E L K +NKS P
Sbjct: 187 ELEELQRQ-KEEAMRRKKAEEEEERLKQMKLLGKNKSRP 224
>At4g02710 predicted protein of unknown function
Length = 1111
Score = 40.0 bits (92), Expect = 0.003
Identities = 52/226 (23%), Positives = 101/226 (44%), Gaps = 28/226 (12%)
Query: 91 KLSNEAKIREVASLEGHVLLKKLRDALEYLRV--RFAGRNKEDVE-KAISMVEALAVKLT 147
KL +E +E + L+ H L+K+ D + L V + AG E + +++ +LA T
Sbjct: 279 KLESE---KESSFLQYHKCLQKIADLEDGLSVAHKEAGERASKAETETLALKRSLAKAET 335
Query: 148 QNEGELIQEKFEVKKLLNF---LKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALE 204
E LIQ + + + N L++A EDA+ L+N+ A E+E+ + V ++
Sbjct: 336 DKETALIQYRQCLNTISNLEERLRKAEEDAR-LINERAEKAGVEVENLKQTVSKL----- 389
Query: 205 EQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKL 264
+D +AS+ Q L + + ++KL H + ++ +E+ E + +K
Sbjct: 390 --IKDKEASELQFQQCL--NIIASLKVKLHHAQEETQSLSHEI--------EDGVAKLKF 437
Query: 265 QKELTMSKRDEENKSHPYMLHGSEALGSY-LKVQPRSGEVPQVSKC 309
+E + H + E LG+ K+ + E+ ++ C
Sbjct: 438 SEEKCLLLERSNQNLHSELDSLLEKLGNQSQKLTEKQTELVKLWSC 483
>At5g60210 putative protein
Length = 629
Score = 39.7 bits (91), Expect = 0.004
Identities = 44/184 (23%), Positives = 82/184 (43%), Gaps = 15/184 (8%)
Query: 94 NEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGEL 153
N+ K E++ E L + LE + + K D KA+ + +AV+L Q++ +
Sbjct: 245 NQLKDCEISEAETEALATETLRQLENAK-KAVEELKSDGTKAVESYKKMAVELEQSKSRM 303
Query: 154 IQEKFEVKKLLNFLKQAS------EDAKKLVNQEKSFACAEIESARSVVLRIGEALE--- 204
+ + V KL N +D + L E + E+ S R V R+ ALE
Sbjct: 304 VWLEALVNKLQNNPADLENHEILLKDYESLRRGESNEMDEEVSSLRCEVERLRAALEASD 363
Query: 205 --EQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSI 262
+QE + +AS + ++ + + ++ +++M E EL+ + + E+ FS+
Sbjct: 364 KKDQEGNVEASSRLRIQAELQSELKIAKSEIDELKARLMDKETELQFISE---ERDNFSM 420
Query: 263 KLQK 266
KL K
Sbjct: 421 KLMK 424
>At3g28770 hypothetical protein
Length = 2081
Score = 39.7 bits (91), Expect = 0.004
Identities = 49/225 (21%), Positives = 99/225 (43%), Gaps = 18/225 (8%)
Query: 40 SQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIR 99
S+ EE ++ + K E+++ + ++ RDL K + K S K +
Sbjct: 1020 SKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSK 1079
Query: 100 EVASLEGHV---LLKKLRDALEYLRVRFA-GRNKEDVEKAISMVEALAVKLTQNEGELIQ 155
+ + H +KK D E + + R KE+ +K + +E QN + +
Sbjct: 1080 KKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLE------DQNSNKKKE 1133
Query: 156 EKFEVKKL--LNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQAS 213
+K E KK + +K+ S+ +K N+EKS EIES++S + + ++ +D Q
Sbjct: 1134 DKNEKKKSQHVKLVKKESDKKEKKENEEKS-ETKEIESSKSQKNEVDKKEKKSSKDQQKK 1192
Query: 214 KPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKS 258
K +++ + E +++K + K E + ++ +EK+
Sbjct: 1193 KEKEM-----KESEEKKLKKNEEDRKKQTSVEENKKQKETKKEKN 1232
>At5g42880 putative protein
Length = 751
Score = 38.9 bits (89), Expect = 0.007
Identities = 65/292 (22%), Positives = 122/292 (41%), Gaps = 38/292 (13%)
Query: 20 VSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLA 79
V A++ + K K+ A S + E + Q L++ ++E+T L+ + + + +L
Sbjct: 428 VKANIEKAASEVKKLKIIAGS-LQSELGRERQ--DLEETKQKESTGLARTNDKDAGEELV 484
Query: 80 CKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRD----ALEYLRVRFAGRNKE-DVEK 134
+ K L A K + +AK AS + + K+L + + + R KE + +
Sbjct: 485 -ETAKKLEQATKEAEDAKALATASRDELRMAKELSEQAKRGMSTIESRLVEAKKEMEAAR 543
Query: 135 AISMVEALAVKLTQNEGELIQEKFE------------VKKLLNFLKQASEDAKKLVNQEK 182
A + A+K Q ++FE V++ KQA E ++ N
Sbjct: 544 ASEKLALAAIKALQETES--SQRFEEINNSPRSIIISVEEYYELSKQALESEEE-ANTRL 600
Query: 183 SFACAEIESARSVVLRIGEALEEQEQDSQASKPQ--DVDGLVEEVQEARRIKLLHQPSKV 240
S ++IE A+ RI E LEE ++ K + + +G E+ ++ +
Sbjct: 601 SEIVSQIEVAKEEESRILEKLEEVNREMSVRKAELKEANGKAEKARDGK----------- 649
Query: 241 MAMEYELRALRDQI-REKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALG 291
+ ME ELR R + + ++ + +K T S + NK + + S A G
Sbjct: 650 LGMEQELRKWRSENGKRRTDEGREPEKSPTRSSTEGRNKENGFGQSKSFAFG 701
>At1g65010 hypothetical protein
Length = 1318
Score = 38.1 bits (87), Expect = 0.011
Identities = 49/215 (22%), Positives = 101/215 (46%), Gaps = 38/215 (17%)
Query: 85 NLAAAAKLSNEA--KIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEAL 142
+L A+K +NE K+ E A E + L+ ++ ++ F +K + ++
Sbjct: 466 SLKLASKETNEKYEKMLEDARNE----IDSLKSTVDSIQNEFENSKAGWEQKELHLMGC- 520
Query: 143 AVKLTQNEGELIQEKFEVKKLLNFLKQASEDA-------------KKLVNQEKSFACAEI 189
VK ++ E QE EV +L+N LK++ EDA K+ E + +
Sbjct: 521 -VKKSEEENSSSQE--EVSRLVNLLKESEEDACARKEEEASLKNNLKVAEGEVKYLQETL 577
Query: 190 ESARSVVLRIGEALEEQEQD-----SQASKPQDVDG-LVEEVQEARRIK--LLHQPSKVM 241
A++ +++ E+L ++E+D ++ S ++ +G ++E+++E ++K L+ + +K+
Sbjct: 578 GEAKAESMKLKESLLDKEEDLKNVTAEISSLREWEGSVLEKIEELSKVKESLVDKETKLQ 637
Query: 242 AMEYELRALR-------DQIREKSIFSIKLQKELT 269
++ E L+ QI E S + L E T
Sbjct: 638 SITQEAEELKGREAAHMKQIEELSTANASLVDEAT 672
Score = 38.1 bits (87), Expect = 0.011
Identities = 51/241 (21%), Positives = 102/241 (42%), Gaps = 14/241 (5%)
Query: 39 DSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKI 98
D L T E+ Q + + +A N + H + + +K A++L +
Sbjct: 155 DISALLSTTEELQRVKHELSMTADAKNKALSHAEEATKIAEIHAEKAEILASELGRLKAL 214
Query: 99 ----REVASLEGHVLLKKLRDALEYLR--VRFAGRNKEDVEKAISMVEALAVKLTQNEGE 152
E ++EG+ ++ KL+ +E LR + + +++ +VE L V L +
Sbjct: 215 LGSKEEKEAIEGNEIVSKLKSEIELLRGELEKVSILESSLKEQEGLVEQLKVDLEAAKMA 274
Query: 153 LIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQA 212
V++ N + + ++ ++ N+ KS A +ES + + L E + D+ A
Sbjct: 275 ESCTNSSVEEWKNKVHELEKEVEE-SNRSKSSASESMESVMKQLAELNHVLHETKSDNAA 333
Query: 213 SKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSK 272
K + L+E+ EA+R L +V + E L E + SIK + E++ +
Sbjct: 334 QK--EKIELLEKTIEAQRTDLEEYGRQVCIAKEEASKL-----ENLVESIKSELEISQEE 386
Query: 273 R 273
+
Sbjct: 387 K 387
>At1g13220 putative nuclear matrix constituent protein
Length = 1128
Score = 38.1 bits (87), Expect = 0.011
Identities = 50/232 (21%), Positives = 109/232 (46%), Gaps = 25/232 (10%)
Query: 59 IEQEATNLSDQHKRISVRDLAC--KFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDA 116
+E++ + +++ R+ A KFD+ ++ EAK++ + E + ++ R +
Sbjct: 411 LERQKVEIDHSEEKLEKRNQAMNKKFDR--VNEKEMDLEAKLKTIKEREKIIQAEEKRLS 468
Query: 117 LEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKK 176
LE ++ +KE +E +E + ++T+ E E+I+E+ K L K+ E+ +
Sbjct: 469 LEKQQLL---SDKESLEDLQQEIEKIRAEMTKKE-EMIEEEC---KSLEIKKEEREEYLR 521
Query: 177 LVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQA-SKPQDVDGLVEEVQEARRIKLLH 235
L ++ KS +IE +R + + +E +Q+ + K ++ + V RI++
Sbjct: 522 LQSELKS----QIEKSRVHEEFLSKEVENLKQEKERFEKEWEILDEKQAVYNKERIRISE 577
Query: 236 QPSKV--------MAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKS 279
+ K ++ E ALR QI ++ + I+LQ+E + + E +
Sbjct: 578 EKEKFERFQLLEGERLKKEESALRVQIMQE-LDDIRLQRESFEANMEHERSA 628
>At1g12150 hypothetical protein
Length = 548
Score = 38.1 bits (87), Expect = 0.011
Identities = 58/287 (20%), Positives = 119/287 (41%), Gaps = 39/287 (13%)
Query: 17 VPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVR 76
V +S ++S + D + KL A + E N D+ E T + + K++ V
Sbjct: 203 VNELSKEISDMKDAIHQLKLAAAQNLQEHA---NIVKEKDDLRECYRTAVEEAEKKLLV- 258
Query: 77 DLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAI 136
L +++ L+ EAK+ E S E VL ++++ A E +
Sbjct: 259 -LRKEYEPELSRTL----EAKLLETTS-EIEVLREEMKKA------------HESEMNTV 300
Query: 137 SMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVN--QEKSFACAEIESARS 194
++ + T E ++ ++ L+N L+ ED ++ Q+K EIE +
Sbjct: 301 KIITNELNEATMRLQEAADDECSLRSLVNSLRMELEDLRREREELQQKEAERLEIEETKK 360
Query: 195 VVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAM-----EYELRA 249
+ EAL+++ + K + ++ E R+I+ L + ++ + E L
Sbjct: 361 L-----EALKQESLKLEQMKTEAIEARNEAANMNRKIESLKKETEAAMIAAEEAEKRLEL 415
Query: 250 LRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALGSYLKV 296
+ ++ E K+++E+ M + +E+K E+ GS +K+
Sbjct: 416 VIREVEEAKSAEEKVREEMKMISQKQESKKQD-----EESSGSKIKI 457
>At5g65770 nuclear matrix constituent protein 1 (NMCP1)-like
Length = 1042
Score = 37.7 bits (86), Expect = 0.014
Identities = 55/258 (21%), Positives = 109/258 (41%), Gaps = 32/258 (12%)
Query: 42 ILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEA-KIRE 100
+ EE + + +++V+ + L + + +L CK K++ + A ++RE
Sbjct: 362 VAEEKIASKESELIQNVLANQEVILRKRKSDVEA-ELECK-SKSVEVEIESKRRAWELRE 419
Query: 101 VASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEV 160
V + L+ + LE A + K+ EK+ + L + E L+ + ++
Sbjct: 420 VDIKQREDLVGEKEHDLEVQSRALAEKEKDITEKSFN--------LDEKEKNLVATEEDI 471
Query: 161 KKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDG 220
+ L+ E +KL ++E +S+ +LE++ + S Q ++
Sbjct: 472 NRKTTMLEDEKERLRKL----------DLELQQSLT-----SLEDKRKRVD-SATQKLEA 515
Query: 221 LVEEVQEARRI--KLLHQPSKVMAMEYELRALRDQIR-EKSIFSIKLQKELTMSKRDEEN 277
L E E + KL + + A + E+ A D+++ EK+ F + + E KR+E
Sbjct: 516 LKSETSELSTLEMKLKEELDDLRAQKLEMLAEADRLKVEKAKF--EAEWEHIDVKREELR 573
Query: 278 KSHPYMLHGSEALGSYLK 295
K Y+ EA YLK
Sbjct: 574 KEAEYITRQREAFSMYLK 591
Score = 35.8 bits (81), Expect = 0.055
Identities = 46/178 (25%), Positives = 82/178 (45%), Gaps = 38/178 (21%)
Query: 44 EETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVAS 103
E+ +E+ + SLK++ E+E ++ + KR+ L K D+ E + RE A
Sbjct: 680 EKKLEEERIQSLKEMAEKELEHVQVELKRLDAERLEIKLDR----------ERREREWA- 728
Query: 104 LEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKL 163
+L+D++E L+V+ +E +E M+ A E+ E E+KKL
Sbjct: 729 --------ELKDSVEELKVQ-----REKLETQRHMLRA-------ERDEIRHEIEELKKL 768
Query: 164 LNFLKQASEDAK----KLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQD 217
N LK A +D +L N E+S+ ++ + + V+ + L+ Q S S +D
Sbjct: 769 EN-LKVALDDMSMAKMQLSNLERSW--EKVSALKQKVVSRDDELDLQNGVSTVSNSED 823
>At3g02930 unknown protein
Length = 806
Score = 37.7 bits (86), Expect = 0.014
Identities = 42/156 (26%), Positives = 71/156 (44%), Gaps = 12/156 (7%)
Query: 129 KEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKL-LNFLKQASEDAKKLVNQEKSFACA 187
KE E + + EAL + E I EKFEV + + +++ E+ KK + K+ +
Sbjct: 122 KEAEEASEKLDEALEAQKKSLENFEI-EKFEVVEAGIEAVQRKEEELKKELENVKNQHAS 180
Query: 188 EIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEY-E 246
E + +L + + LE Q+ +K L ++ + + ++++ E
Sbjct: 181 E----SATLLLVTQELENVNQELANAKDAKSKALCRADDASKMAAIHAEKVEILSSELIR 236
Query: 247 LRALRDQIREKSIFS-----IKLQKELTMSKRDEEN 277
L+AL D REK I S +KL E+ KRD EN
Sbjct: 237 LKALLDSTREKEIISKNEIALKLGAEIVDLKRDLEN 272
Score = 35.8 bits (81), Expect = 0.055
Identities = 66/283 (23%), Positives = 125/283 (43%), Gaps = 42/283 (14%)
Query: 14 MEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQG-----PSLKDVIEQ-----EA 63
++A+ + + IS KLGA+ L+ +E+ + L+ +IEQ EA
Sbjct: 237 LKALLDSTREKEIISKNEIALKLGAEIVDLKRDLENARSLEAKVKELEMIIEQLNVDLEA 296
Query: 64 TNLSDQHKRISV---RDLACKFDKNLAAAAKLSNEAKIR--------EVASLEGHVL--- 109
+++ + ++ A + +K L A KL A + EV++ H +
Sbjct: 297 AKMAESYAHGFADEWQNKAKELEKRLEEANKLEKCASVSLVSVTKQLEVSNSRLHDMESE 356
Query: 110 LKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQ 169
+ L++ +E L + A + K D+EK+ KL E E + + E +KL N L+
Sbjct: 357 ITDLKEKIELLEMTVASQ-KVDLEKSEQ-------KLGIAEEESSKSEKEAEKLKNELET 408
Query: 170 ASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEAR 229
VN+EK+ A + + A S V R+ E ++ + ++SK ++ ++ E+
Sbjct: 409 --------VNEEKTQALKKEQDATSSVQRLLEEKKKILSELESSKEEEEKS--KKAMESL 458
Query: 230 RIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSK 272
L S+ ++ +L + DQ E I +KL + T +K
Sbjct: 459 ASALHEVSSESRELKEKLLSRGDQNYETQIEDLKLVIKATNNK 501
Score = 31.6 bits (70), Expect = 1.0
Identities = 47/248 (18%), Positives = 109/248 (43%), Gaps = 15/248 (6%)
Query: 39 DSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKI 98
D ++ E N+ +L ++EA + K +RD + + + + EAK
Sbjct: 549 DEEVSSMGKEMNRLGNLVKRTKEEAD--ASWEKESQMRDCLKEVEDEVIYLQETLREAKA 606
Query: 99 REVASLEGHVLLKKLR-----DALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGEL 153
E L+G +L K+ + LRV+ K+ E + + EALA K + GEL
Sbjct: 607 -ETLKLKGKMLDKETEFQSIVHENDELRVKQDDSLKKIKELSELLEEALAKKHIEENGEL 665
Query: 154 IQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQAS 213
+ + + L ++ + E+ + ++ S ++E+ + +++ E E++E+ ++
Sbjct: 666 SESEKDYDLLPKVVEFSEENGYRSAEEKSS----KVETLDGMNMKLEEDTEKKEKKERSP 721
Query: 214 KPQ--DVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMS 271
+ + +V+ + E + + ++ H+ S E +L + + + + + EL
Sbjct: 722 EDETVEVEFKMWESCQIEKKEVFHKES-AKEEEEDLNVVDQSQKTSPVNGLTGEDELLKE 780
Query: 272 KRDEENKS 279
K ++ K+
Sbjct: 781 KEKKKKKT 788
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 36.6 bits (83), Expect = 0.032
Identities = 47/199 (23%), Positives = 88/199 (43%), Gaps = 29/199 (14%)
Query: 44 EETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKF-DKNLAAAAKLSNEA----KI 98
EET+E SL++ +E + + ++IS ++ + ++ L ++ E +
Sbjct: 1088 EETME-----SLRNELEMKGDEIETLMEKISNIEVKLRLSNQKLRVTEQVLTEKEEAFRK 1142
Query: 99 REVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKA---ISMVEALAVKLTQNEGELIQ 155
E LE LL+K L + G KE +K + ++++ KLT+ +G +
Sbjct: 1143 EEAKHLEEQALLEK---NLTMTHETYRGMIKEIADKVNITVDGFQSMSEKLTEKQGRYEK 1199
Query: 156 EKFEVKKLL----NFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQ 211
E K+L N++ + N EK EIE + ++G + E E++ +
Sbjct: 1200 TVMEASKILWTATNWVIER--------NHEKEKMNKEIEKKDEEIKKLGGKVREDEKEKE 1251
Query: 212 ASKPQDVDGLVEEVQEARR 230
K + + GL EE +EA R
Sbjct: 1252 MMK-ETLMGLGEEKREAIR 1269
Score = 34.3 bits (77), Expect = 0.16
Identities = 62/269 (23%), Positives = 100/269 (37%), Gaps = 44/269 (16%)
Query: 38 ADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAK 97
A I E E Q V E+E T L D H+ R+ + + + L KL +
Sbjct: 444 AQKTIQEHMSESEQLKESHGVKERELTGLRDIHETHQ-RESSTRLSE-LETQLKLLEQRV 501
Query: 98 IREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVK---LTQNEGEL- 153
+ ASL KK +L + + K+ K +V LA LTQ E EL
Sbjct: 502 VDLSASLNAAEEEKK---SLSSMILEITDELKQAQSKVQELVTELAESKDTLTQKENELS 558
Query: 154 -----------------------IQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIE 190
++ E K LN +SE+ KK+++Q+ S +I+
Sbjct: 559 SFVEVHEAHKRDSSSQVKELEARVESAEEQVKELNQNLNSSEEEKKILSQQISEMSIKIK 618
Query: 191 SARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRAL 250
A S + + E+ + S A K ++ L R I HQ + +LR L
Sbjct: 619 RAESTIQELSSE-SERLKGSHAEKDNELFSL-------RDIHETHQ----RELSTQLRGL 666
Query: 251 RDQIREKSIFSIKLQKELTMSKRDEENKS 279
Q+ ++L + L ++ + S
Sbjct: 667 EAQLESSEHRVLELSESLKAAEEESRTMS 695
Score = 29.6 bits (65), Expect = 3.9
Identities = 45/228 (19%), Positives = 90/228 (38%), Gaps = 29/228 (12%)
Query: 48 EDNQGPSLK-----DVIEQEATNLSDQHKRISVRDLACKFDKN----LAAAAKLSNEAKI 98
E+N+ SLK DVI+Q T + + + K ++ L K
Sbjct: 6 EENKSLSLKVSEISDVIQQGQTTIQELISELGEMKEKYKEKESEHSSLVELHKTHERESS 65
Query: 99 REVASLEGHV---------LLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQN 149
+V LE H+ + L +A E + + ++ I + +L
Sbjct: 66 SQVKELEAHIESSEKLVADFTQSLNNAEE--EKKLLSQKIAELSNEIQEAQNTMQELMSE 123
Query: 150 EGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQD 209
G+L + ++ L L+ E ++ + S A++ES++ V + +L+ E++
Sbjct: 124 SGQLKESHSVKERELFSLRDIHEIHQRDSSTRASELEAQLESSKQQVSDLSASLKAAEEE 183
Query: 210 SQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREK 257
++A ++V+ + KL + + + EL L+D REK
Sbjct: 184 NKAISSKNVETM---------NKLEQTQNTIQELMAELGKLKDSHREK 222
>At2g19950 hypothetical protein
Length = 713
Score = 36.6 bits (83), Expect = 0.032
Identities = 52/268 (19%), Positives = 107/268 (39%), Gaps = 19/268 (7%)
Query: 3 KISPEIEIKMPMEA-VPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIEQ 61
K+ I ++ E VP S ++ DR + S +++ +E+ QG
Sbjct: 234 KVGTSINLEKEQEPKVPVTSTNLKREQDR----RADTTSMKIQDQLEEAQGLL------- 282
Query: 62 EATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLR 121
+AT + Q K + + L + A++ E+ + E L K ++ +L+
Sbjct: 283 KATVSTGQSKEARLARVCAGLSSRLQEIK--AENAQLEELLTAEQE-LTKSYEASIRHLQ 339
Query: 122 VRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQE 181
+ E + SMVEALA K +E E + + K L + + + + +E
Sbjct: 340 KDLSAAKSEVTKVESSMVEALAAK--NSEIETLVSAMDALKNQAALNEGKLSSLQALREE 397
Query: 182 KSFACAEIESARSV--VLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSK 239
+ E RS ++ E+E + +A ++ + + R K+ K
Sbjct: 398 LATTERRAEEERSAHNATKMAAMERERELEHRAVDASTALVRIQRIADERTAKVADFEQK 457
Query: 240 VMAMEYELRALRDQIREKSIFSIKLQKE 267
V +E E +L ++++ + + + QK+
Sbjct: 458 VALLEAECTSLNQELQDMEVRARRGQKK 485
>At1g20970 hypothetical protein
Length = 1498
Score = 36.6 bits (83), Expect = 0.032
Identities = 32/131 (24%), Positives = 63/131 (47%), Gaps = 20/131 (15%)
Query: 129 KEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAE 188
KE++EK+ + E KLT+ E ELI+++ E K+ ++A K+ Q + E
Sbjct: 1194 KEEIEKSATPEEEEPPKLTKEEEELIKKEEE--------KRKQKEAAKMKEQHR---LEE 1242
Query: 189 IESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIK--LLHQPSKVMAMEYE 246
I A+ EA+E +++ + +K + V +E +E ++K H ++
Sbjct: 1243 IAKAK-------EAMERKKKREEKAKARAVLKAQKEAEEREKVKAYFCHSVIQIGFRSVS 1295
Query: 247 LRALRDQIREK 257
L R++ +E+
Sbjct: 1296 LARTREEAKEE 1306
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.131 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,103,143
Number of Sequences: 26719
Number of extensions: 413775
Number of successful extensions: 1695
Number of sequences better than 10.0: 121
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 111
Number of HSP's that attempted gapping in prelim test: 1617
Number of HSP's gapped (non-prelim): 170
length of query: 490
length of database: 11,318,596
effective HSP length: 103
effective length of query: 387
effective length of database: 8,566,539
effective search space: 3315250593
effective search space used: 3315250593
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0269a.3


