

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.8
(765 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
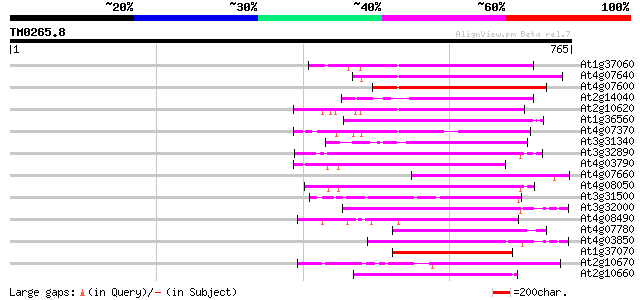
Score E
Sequences producing significant alignments: (bits) Value
At1g37060 Athila retroelment ORF 1, putative 181 2e-45
At4g07640 putative athila transposon protein 178 1e-44
At4g07600 155 1e-37
At2g14040 putative retroelement pol polyprotein 154 1e-37
At2g10620 putative Athila retroelement ORF1 protein 154 1e-37
At1g36560 hypothetical protein 151 1e-36
At4g07370 146 5e-35
At3g31340 Athila ORF 1, putative 144 1e-34
At3g32890 Athila ORF 1, putative 143 4e-34
At4g03790 putative athila-like protein 138 1e-32
At4g07660 putative athila transposon protein 137 2e-32
At4g08050 135 1e-31
At3g31500 hypothetical protein 129 8e-30
At3g32000 unknown protein 128 1e-29
At4g08490 putative athila-like protein 128 1e-29
At4g07780 putative athila transposon protein 125 7e-29
At4g03850 putative transposon protein 124 3e-28
At1g37070 hypothetical protein 124 3e-28
At2g10670 pseudogene 123 3e-28
At2g10660 putative Athila retroelement ORF1 protein 121 1e-27
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 181 bits (458), Expect = 2e-45
Identities = 120/333 (36%), Positives = 179/333 (53%), Gaps = 31/333 (9%)
Query: 408 NPKENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGEN-KKKQKEKEE------ 460
N KE AITLRSG L + N N D D E +N +K EE
Sbjct: 427 NSKEYAHAITLRSGKELPTKESP---NQNTEDSLDQDGEDFCQNGNSAEKAIEEPILHQP 483
Query: 461 ------------EKRKVELEKKFTKVPFP-----PFPTNIAKRRLEKQFSKFISMFKKLR 503
EK K+ +P P PFP K ++K + K L
Sbjct: 484 TRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLE 543
Query: 504 VELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRK-DPGS 562
V +P + L +P K++K+++++ R+ E +V L+ ECS I+Q+K+ PK+ DPGS
Sbjct: 544 VTMPLVDCLALIPDSNKYVKDMITE--RIKEVQGMVVLSHECSAIIQQKIIPKKLGDPGS 601
Query: 563 FTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVV 622
FTLP G + LCDLG+SV+LMPL + ++L + KP + L LADRS+ G++
Sbjct: 602 FTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVRISHGLL 661
Query: 623 EDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKI 681
ED+ V +G E P DFV+++MDE+ K PLILGRPFLA ++A I+V KG I L + D K+
Sbjct: 662 EDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNLGRDLKM 721
Query: 682 GFNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQ 714
F+I + KP + ++F ++ M+ +D+ L++
Sbjct: 722 TFDITNTMKKPTIEGNIFWIEEMDMLADKMLEE 754
>At4g07640 putative athila transposon protein
Length = 866
Score = 178 bits (451), Expect = 1e-44
Identities = 105/291 (36%), Positives = 169/291 (57%), Gaps = 7/291 (2%)
Query: 468 EKKFTKVPFPP---FPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKE 524
EK F P+ P FP K +K + F K++ + +P + L +P KF+K+
Sbjct: 444 EKVFVPPPYKPELPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIPDSHKFLKD 503
Query: 525 ILSKKRRLSEENEIVELTEECSVILQRKLPPKR-KDPGSFTLPVNFGASKQVRALCDLGS 583
++ + R+ E +V L+ CS I+Q+K+ PK+ DPGSFTLP + G R LCDLG+
Sbjct: 504 LIVE--RIQEVQGMVVLSHGCSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGA 561
Query: 584 SVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDM 643
SV+LM LS+ +RL + K + + L LADRS+ P G++++ + +G E P DFV+++M
Sbjct: 562 SVSLMALSVAKRLGFTQYKSSNISLILADRSVRIPHGLLKNFGITIGAVEIPTDFVVLEM 621
Query: 644 DEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKIGFNIFDLKPKPVEKNDVFLVK 702
DE+ K PLILGRPFLAT+ A I+V KG I L + D ++ F++ D KP + +F +K
Sbjct: 622 DEEPKDPLILGRPFLATAGAMIDVKKGKIDLSLGKDFRMTFDVKDAMKKPTIEGQLFWIK 681
Query: 703 MMEEWSDEKLKQFFLKEKAGASNKKKKEDT*INQTEQVYSMSMVVNSELKE 753
M++ +DE L++ ++ ++ K ED ++ Y + N ++E
Sbjct: 682 EMDQLADELLEELAEEDHLNSALTKSGEDGFLHLETFGYQKLLDSNKAMEE 732
>At4g07600
Length = 630
Score = 155 bits (391), Expect = 1e-37
Identities = 90/239 (37%), Positives = 145/239 (60%), Gaps = 4/239 (1%)
Query: 495 FISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLP 554
F K++ + +P + L + KF+K+++ + R+ E +V L+ ECS I+Q+K+
Sbjct: 2 FAKNIKEVELRIPLVDALALILDTHKFLKDLIVE--RIQEVQGMVVLSHECSAIIQKKIV 59
Query: 555 PKR-KDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADR 613
PK+ DPGSFTLP G R LCDLG+ V+ MPLS+ +RL + K + L LADR
Sbjct: 60 PKKLSDPGSFTLPCFLGTVAFNRCLCDLGALVSPMPLSIAKRLGFTQYKSCNISLILADR 119
Query: 614 SIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTIS 673
S+ G+++++ +R+G E DFVI++MDE+ K PLIL RPFLAT+ A I+V KG I
Sbjct: 120 SVRISHGLLKNLPIRIGAAEISTDFVILEMDEEPKDPLILRRPFLATAGAMIDVKKGKID 179
Query: 674 LRVA-DEKIGFNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKED 731
L + D ++ F+I D KP + +F ++ M++ +DE L++ ++ ++ K +D
Sbjct: 180 LNLGKDFRMTFDIKDAMKKPNIEGQLFWIEEMDQLADELLEELAQEDHLNSALTKTGQD 238
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 154 bits (390), Expect = 1e-37
Identities = 102/262 (38%), Positives = 135/262 (50%), Gaps = 38/262 (14%)
Query: 453 KKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVL 512
K +K+ EEK K + + VP PFP K + EK++S F E++
Sbjct: 2 KLRKQTLEEKAKKTVLPPY--VPKLPFPGRQRKIQREKEYSLF-------------DEIM 46
Query: 513 EKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGAS 572
++ +Y+ ECS ILQ +P KR+DPGSF LP G
Sbjct: 47 RQLQRYSP-----------------------ECSAILQNVIPVKREDPGSFVLPSRIGEY 83
Query: 573 KQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEF 632
R LCDLG+ V+LMP S+ +RL PT M L L DRSI P GV EDV VRVG F
Sbjct: 84 TFDRCLCDLGAGVSLMPFSVAKRLGDTNFTPTKMSLVLGDRSISFPVGVAEDVQVRVGNF 143
Query: 633 EFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGFNIFDLKPKP 692
P DFVII++DE+ + LILGRPFL A I+V K I+LR+ D FN+ + KP
Sbjct: 144 YIPTDFVIIELDEEPRHRLILGRPFLNIVAALIDVRKSKINLRIGDIVQEFNMERIMSKP 203
Query: 693 VEKNDVFLVKMMEEWSDEKLKQ 714
+ F V +M+E +E L +
Sbjct: 204 TTECQTFWVDIMDELVNELLAE 225
>At2g10620 putative Athila retroelement ORF1 protein
Length = 451
Score = 154 bits (390), Expect = 1e-37
Identities = 113/386 (29%), Positives = 180/386 (46%), Gaps = 73/386 (18%)
Query: 387 GTDGQANVRKTPGMFPSDTVINPKENCSAITLRSGATL------------SEPKQKIVEN 434
G + +K P + NPKE AITLRSG L P +K +N
Sbjct: 53 GNSASTSAQKQTSQLPGKAIQNPKEYAHAITLRSGKALPTIEAPKTVTVGKRPSRKTSKN 112
Query: 435 NN----------------VNDEFVG------DIEPLGENKKKQKEKEEEKRKVELEKK-- 470
+ + F G ++ GE+ K++ +++ + L++
Sbjct: 113 DRGRTGRERNSKGTKAKEMGKRFGGVIVTEDSVDHDGEDFSLTKDQADKELEQPLDQSLE 172
Query: 471 -----FTKVPFP----------------------------PFPTNIAKRRLEKQFSKFIS 497
FT+ PFP FP K K F
Sbjct: 173 QPLDHFTRPPFPITAPTAVKPVVVKNKEKVFVPPPYKPHPSFPGRHKKALAHKYRVMFAK 232
Query: 498 MFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKR 557
K++ +++P + L + KF+K+++ + R+ E +V L++ECS I+Q+K+ K+
Sbjct: 233 NIKEVELQIPLVDALALIMDSHKFLKDLIVE--RIQELQGMVVLSDECSAIIQKKIIHKK 290
Query: 558 -KDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIV 616
DPGSFTLP + G + LCDLG+S +LMPLS+ +RL + K + L LADRS+
Sbjct: 291 LSDPGSFTLPCSLGPLAFNKCLCDLGASASLMPLSVTKRLGFTQYKSCNISLILADRSVR 350
Query: 617 TPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRV 676
P +E++ +R+ E P DFV+++MDE+ K LILGRPFL T A I + KG I+L +
Sbjct: 351 IPHDFLENIPIRIRAVEIPTDFVVLEMDEEPKDHLILGRPFLTTVGAMIYIKKGKINLNL 410
Query: 677 A-DEKIGFNIFDLKPKPVEKNDVFLV 701
D ++ F++ D KP + + L+
Sbjct: 411 GKDFRMTFDVKDTMKKPTIEGQLLLM 436
>At1g36560 hypothetical protein
Length = 524
Score = 151 bits (382), Expect = 1e-36
Identities = 98/277 (35%), Positives = 152/277 (54%), Gaps = 10/277 (3%)
Query: 456 KEKEEEKRKV--ELEKKFTKVPFPP--FPTNIAKRRLEKQFSKFISMFKKLRVELPFSEV 511
+ EEK + + K+F P P P K + + S ++ +P EV
Sbjct: 14 RSTSEEKAAIIERMVKRFKPTPLPSRALPWTFRKAWMARYKSVAAKQLNEIEAVMPLMEV 73
Query: 512 LEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGA 571
L +P K ++ ++ + ++ +++ +R + K +DPGSFTLP + G+
Sbjct: 74 LNLIPNPHKDVRNLILEWIKMYHDSDDESDATPSRAADKRIVQEKLEDPGSFTLPCSLGS 133
Query: 572 SKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGE 631
+ LCDLG+SV+LMPLS+ +RL + K + L LAD S+ P GV+ED+ +++G+
Sbjct: 134 LTFNKCLCDLGASVSLMPLSVAKRLGFNKYKYCNISLILADGSVRQPHGVLEDLPIKIGK 193
Query: 632 FEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKIGFNIFDLKP 690
E P DF+I++MDE+ K PLILGRPFLAT+ A I+V +G I L + D K+ F+I D
Sbjct: 194 VEVPTDFIILNMDEEPKDPLILGRPFLATAGAIIDVKQGKIDLNMGKDFKMKFDINDAMK 253
Query: 691 KPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKK 727
KP + FLV+ + DE L + LKE A+N K
Sbjct: 254 KPTIEGQTFLVEETDRLVDELLVE--LKE---ANNSK 285
>At4g07370
Length = 531
Score = 146 bits (368), Expect = 5e-35
Identities = 105/339 (30%), Positives = 167/339 (48%), Gaps = 42/339 (12%)
Query: 387 GTDGQANVRKTPGMFPSDTVINPKENCSAITLRSGATLSEPKQKIVENNNVNDEFVGD-- 444
G + K G+ P ++ NPKE R + + + E+ ++N++ V
Sbjct: 54 GESASTSSPKVTGL-PGKSIQNPKEE------RPKSVTEDSAYQDGEDFSLNEDQVDKPT 106
Query: 445 --IEPLGENKKKQKEKEEEKRKVEL------EKKFTKVPFPP---FPTNIAKRRLEKQFS 493
+EP+ + + ++L E F P+ P FP K +K +
Sbjct: 107 EVLEPILDRDTRPTNPLTSSAALKLVAAKNNEIAFIPPPYKPPLPFPGRHKKEVEDKYRA 166
Query: 494 KFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRK- 552
F KK+ + +P ++ L +P KF+K+++ + R+ E + L+ ECS I+Q
Sbjct: 167 MFAKNIKKVELRIPLADALTHIPDSQKFLKDLIME--RIQEVQKTTVLSHECSAIIQEND 224
Query: 553 LPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLAD 612
+ K DPGSFTLP + G+ + LCDLG SVNLMPLS+ A
Sbjct: 225 VSEKLGDPGSFTLPCSLGSLTFNKCLCDLGPSVNLMPLSV------------------AK 266
Query: 613 RSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTI 672
RS+ P G++ED+ +++ E P DFV+++MDE+ K PLILGRPFLAT I+V +G I
Sbjct: 267 RSVRLPHGLLEDLPIKIRNVEVPTDFVVLNMDEEPKDPLILGRPFLATVGVIIDVKQGKI 326
Query: 673 SLRVADE-KIGFNIFDLKPKPVEKNDVFLVKMMEEWSDE 710
L + ++ F+I D KP + FLVK ++ + E
Sbjct: 327 DLNLGKNFEMKFDINDAMKKPTIEEQTFLVKEVDRLAGE 365
>At3g31340 Athila ORF 1, putative
Length = 781
Score = 144 bits (364), Expect = 1e-34
Identities = 98/276 (35%), Positives = 136/276 (48%), Gaps = 38/276 (13%)
Query: 431 IVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEK 490
+V V D+ V D EPL E E K+PFP I R+ E+
Sbjct: 444 VVVETLVEDKIVEDDEPLSE---------------EPPPYVPKLPFPGRERQIQSRKREE 488
Query: 491 QFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQ 550
L VE S+ +K Q ++ + K E+ CS I
Sbjct: 489 Y---------ALLVE---SQRQQKEAQLTDVVEVLAGK-----------EMVSTCSAIPP 525
Query: 551 RKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQL 610
+P K DPGSF LP G S R LCDLG+ VNLMPLSM +RL + KP+ + L L
Sbjct: 526 ATIPEKLGDPGSFVLPCRIGKSAFERCLCDLGAGVNLMPLSMSKRLGITNFKPSRISLIL 585
Query: 611 ADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKG 670
ADRS+ P G+ E+V VRVG+F P DFV++++D++ PL LGRPFL T A I+V +
Sbjct: 586 ADRSVRFPVGLAENVHVRVGDFYIPTDFVVLELDKEPHDPLTLGRPFLNTVGAIIDVRRS 645
Query: 671 TISLRVADEKIGFNIFDLKPKPVEKNDVFLVKMMEE 706
TI+L++ D + F++ + P + F V +E
Sbjct: 646 TINLQIGDFALEFDMKGTRKNPTIEGHAFSVDTNDE 681
>At3g32890 Athila ORF 1, putative
Length = 755
Score = 143 bits (360), Expect = 4e-34
Identities = 108/349 (30%), Positives = 174/349 (48%), Gaps = 21/349 (6%)
Query: 389 DGQANVRKTPGM--FPSDTVINPKENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIE 446
+GQ+ TP + P ++ NPKE +A + T+ ++ + +V D GD +
Sbjct: 356 EGQSASTSTPKVTGLPGKSIQNPKEYATAHAI----TICHDRE--LPTRHVPDFITGDSD 409
Query: 447 PLGENKKKQKEKEEEKRKVE-LEKKFTKVPFPP--FPTNIAKRRLEKQFSKFISMFKKLR 503
Q EE+ +E + K+F P P P K +EK S ++
Sbjct: 410 VQDGEASTQSTSEEKAAIIERMVKRFKPTPLPSSALPWTFRKAWMEKYKSVASKQLDEIE 469
Query: 504 VELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSF 563
+P EVL +P K ++ ++ ++ ++ +++ +R + K +DPGSF
Sbjct: 470 AVMPLIEVLNLIPDPHKDVRNLILERIKMYHDSDDESDATPSRASDKRIVQEKLEDPGSF 529
Query: 564 TLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVE 623
TLP + G LCDLG+SV+LMPLS+ RL + KP + L LADRS P+G+++
Sbjct: 530 TLPCSIGELAFSDCLCDLGASVSLMPLSVARRLEFIQYKPCDLTLILADRSFRKPFGMLK 589
Query: 624 DVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADE-KIG 682
D+ V + E P DFV++DM+ + K PLILGRP LA+ A I+V +G ISL + K+
Sbjct: 590 DLPVMINGVEVPTDFVVLDMEVEHKDPLILGRPLLASVGAVIDVREGKISLNLGKHIKLQ 649
Query: 683 FNIFDLKPKPVEK-----NDVFLVKMMEEWSDEKLKQFFLKEKAGASNK 726
F I E ND + + E + E++K+ LK+K+ +K
Sbjct: 650 FGINKTPQGSTEDGRTSGNDRVISR--EGYETERVKE--LKKKSDKQDK 694
>At4g03790 putative athila-like protein
Length = 1064
Score = 138 bits (348), Expect = 1e-32
Identities = 97/306 (31%), Positives = 153/306 (49%), Gaps = 17/306 (5%)
Query: 387 GTDGQANVRKTPGMFPSDTVINPKENCSAITLRSGATLSEPKQKIV-----ENNNVNDEF 441
G +V K G+ P ++ NPKE +A + P + ++ E++ E
Sbjct: 223 GQSASTSVPKVTGL-PGKSIQNPKEYATAHAITICHDRELPTRPVLDLIIGESDVQEGEA 281
Query: 442 VGDIE--------PLGENKKKQKEKEEEKRKVE-LEKKFTKVPFPP--FPTNIAKRRLEK 490
IE G + Q EE+ +E + K+F P P P K +E+
Sbjct: 282 CTQIEVSVIEFNHSAGSHHLIQSTSEEKAAIIERMVKRFKPTPLPSRALPWTFRKAWIER 341
Query: 491 QFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQ 550
S ++ +P EVL +P + K ++ ++ +K ++ +++ V+ +
Sbjct: 342 YKSVAAKQLDEIEAVMPLMEVLNLIPDHHKDVRNLILEKIKMYHDSDDESDATPSRVVDK 401
Query: 551 RKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQL 610
R + K +DPGSFTLP + G LCDLG+SV+LMPLSM RL KP + L L
Sbjct: 402 RIVQEKLEDPGSFTLPCSIGELAFSDFLCDLGASVSLMPLSMGRRLEFIHYKPCDLTLIL 461
Query: 611 ADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKG 670
ADRS P+G+++D+ V + E P DFV+++M+ K PLILGRPFLA+ A I++ +G
Sbjct: 462 ADRSSRKPFGMLKDLPVMINGVEVPTDFVVLNMEVKHKDPLILGRPFLASVGAVIDIREG 521
Query: 671 TISLRV 676
ISL +
Sbjct: 522 KISLNL 527
>At4g07660 putative athila transposon protein
Length = 724
Score = 137 bits (345), Expect = 2e-32
Identities = 84/223 (37%), Positives = 132/223 (58%), Gaps = 7/223 (3%)
Query: 548 ILQRKLPPKRK-DPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMM 606
I Q+K+ PK+ DPGSFTLP + G R LCDLG+ V+LMPLS+ +RL + K +
Sbjct: 4 ITQKKIVPKKLIDPGSFTLPCSLGPLAFKRCLCDLGALVSLMPLSVAKRLGFTQYKSCNI 63
Query: 607 MLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKIN 666
L LADRS+ P + E++ +R+G + P DFV+++MDE+ K PLILGRPFLAT+ A +
Sbjct: 64 SLILADRSVRIPHSLFENLPIRIGAVDIPTDFVVLEMDEEPKDPLILGRPFLATAGAMND 123
Query: 667 VGKGTISLRVADE-KIGFNIFDLKPKPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASN 725
V KG I L + ++ F++ D KP K +F ++ +++ +DE L++ ++ ++
Sbjct: 124 VKKGKIDLNLGKYCRMTFDVKDAMKKPTIKGQLFWIEEIDQLADELLEERAEEDHLYSAL 183
Query: 726 KKKKEDT*INQTEQVY-----SMSMVVNSELKEKINTKQKEEM 763
K+ ED ++ Y S + SE E++N + E M
Sbjct: 184 TKRGEDGFLHLETLGYQKLLDSHKAMEESEPFEELNGPETEVM 226
>At4g08050
Length = 1428
Score = 135 bits (339), Expect = 1e-31
Identities = 103/336 (30%), Positives = 164/336 (48%), Gaps = 24/336 (7%)
Query: 402 PSDTVINPKENCSAITLRSGATLSEPKQKIVE----NNNVND-EFVGDIEP--------L 448
P ++ NPKE +A + P + +++ +N+V + E +E
Sbjct: 400 PGKSIQNPKEYATAHAITICHDRELPTRHVLDLITRDNDVQEGEASTQVEASVVEFNHSA 459
Query: 449 GENKKKQKEKEEEKRKVE-LEKKFTKVPFP--PFPTNIAKRRLEKQFSKFISMFKKLRVE 505
G Q EE+ +E + K+F P P P K +E+ S ++
Sbjct: 460 GSRHLTQSTSEEKAAIIERMVKRFKPTPLPLRALPWTFRKAWMERYKSVAAKQLDEIEAV 519
Query: 506 LPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTL 565
+P EVL + K ++ ++ ++ ++ +++ +R + K +DPGSFTL
Sbjct: 520 MPLMEVLNLILDPHKVVRNLILERIKMYHDSDDESDATPSRAADKRIVQEKLEDPGSFTL 579
Query: 566 PVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDV 625
P + G LCDLG+SVNLMPLSM RL + KP + L LADRS +G+++D+
Sbjct: 580 PCSIGEFAFSDCLCDLGASVNLMPLSMARRLEFIQYKPCDLTLILADRSSRKHFGMLKDL 639
Query: 626 LVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADE-KIGFN 684
V + E P DFV++DM+ + K PLILGRPFLA+ A I+V +G I L + K+ FN
Sbjct: 640 PVMINGVEVPTDFVVLDMEVEHKDPLILGRPFLASVGAVIDVKEGKIGLNLGKHIKLQFN 699
Query: 685 IFDLKPKPVEK-----NDVFLVKMMEEWSDEKLKQF 715
I E ND + + EE+ E++K+F
Sbjct: 700 INRTPQGSTEDGRTSGNDRVISR--EEYETERVKEF 733
>At3g31500 hypothetical protein
Length = 591
Score = 129 bits (323), Expect = 8e-30
Identities = 90/294 (30%), Positives = 157/294 (52%), Gaps = 26/294 (8%)
Query: 410 KENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEK 469
KE C AI ++ E ++IV V V +E L E+K + ++E VE
Sbjct: 234 KEQCYAIMIQ------EELREIVVAKQVETNVVV-VETLVEDKIVE---DDEPLSVEPPP 283
Query: 470 KFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKK 529
K+PFP I + +K++++F + K+L V LPF +++ +P Y ++K ILS K
Sbjct: 284 YVPKLPFPGRERQIQR---QKEYARFDEIMKQLYVRLPFLQLVLHVPSYRSYLKYILSNK 340
Query: 530 RRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMP 589
R + E +++ EE + +++ + +++ T V A K++ C + P
Sbjct: 341 RSIEEGVKLISKGEEHAQLVESQ---RQQKEAQLTNVVEMLAGKEMVNTCSA-----IPP 392
Query: 590 LSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKI 649
++ ++L + KP+ + L LADRS+ P G+ E+V RVG F P +FV++++D++
Sbjct: 393 ATIPKKLGITNFKPSRISLILADRSVQFPMGLAENVHARVGNFYIPTNFVVLELDKEPHD 452
Query: 650 PLILGRPFLATSQAKINVGKGTISLRVADEKIGFNIFDLKPKP-----VEKNDV 698
PL LGRPFL T +A I+V + TI+L++ D + F+I + P +E +DV
Sbjct: 453 PLNLGRPFLNTVEAIIDVRRSTINLQIGDYALEFDIKGTRKNPTIEDCIELSDV 506
>At3g32000 unknown protein
Length = 839
Score = 128 bits (322), Expect = 1e-29
Identities = 99/316 (31%), Positives = 155/316 (48%), Gaps = 26/316 (8%)
Query: 455 QKEKEEEKRKVE-LEKKFTKVPFPP--FPTNIAKRRLEKQFSKFISMFKKLRVELPFSEV 511
Q EE+ +E + K+F P P P K +E+ S ++ +P EV
Sbjct: 470 QSTSEEKAAIIERMVKRFKPTPLPSRALPWTFRKSWMERYKSVAAKQPDEIEAVMPLMEV 529
Query: 512 LEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGA 571
L +P K ++ ++ ++ ++ +++ +R + K +DPGSF+LP + G
Sbjct: 530 LNLIPDPHKDVRNLILERIKMYHDSDDESDATPSRAADKRIVQEKLEDPGSFSLPCSIGE 589
Query: 572 SKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGE 631
LCDLG+ V+LMP S+ RL + KP + L LADRS P+G+++D+ V +
Sbjct: 590 FAFSDCLCDLGAFVSLMPCSVARRLEFIQYKPCDLTLILADRSSRKPFGMLKDLPVMING 649
Query: 632 FEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADE-KIGFNIFDLKP 690
E P DFV++DM+ + K PLILGRPFLA+ A I+V +G I L + K+ F I
Sbjct: 650 VEVPTDFVVLDMEVEHKDPLILGRPFLASVGAVIDVREGKIGLNLGKHIKLQFGINKTPQ 709
Query: 691 KPVEK-----NDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKEDT*INQTEQVYSMSM 745
E ND V E + E++K+ KK+ D Q E + ++
Sbjct: 710 GSTEDGKTSGND--RVISGEGYETERVKEL-----------KKRSD---KQDETIEKLAH 753
Query: 746 VVNSELKEKINTKQKE 761
V EL+ K+N QKE
Sbjct: 754 TV-EELRSKLNQLQKE 768
>At4g08490 putative athila-like protein
Length = 587
Score = 128 bits (321), Expect = 1e-29
Identities = 98/349 (28%), Positives = 167/349 (47%), Gaps = 54/349 (15%)
Query: 393 NVRKTPGMFPSDTVINPKENCSAITLRSGATL-----SEPKQKIVENNNVN-DEFVGDIE 446
++++ G P + NP+++C+AI +R G + +E + ++ V+ D +
Sbjct: 51 SIKRQEGFLPGNPDANPRKSCNAILIREGDDVWEELDTEDELELAVAEMVSTDTLMYRST 110
Query: 447 PLGENKKKQKEKE-------------------EEKRKVELEKKFTKVPFPPFPTNIAKRR 487
P G + + + E + ++LE +T P P+P +RR
Sbjct: 111 PYGMLFSEMTQYDGVDRYPSSIDRHWIRTAPCEVQVPMQLEPVYT--PHVPYP----RRR 164
Query: 488 LEKQ---FSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSK---------------- 528
KQ +K ++ +K+ LP +++K+++
Sbjct: 165 RSKQEIHVAKCTTIMEKILHILPTDASETSSASLNRYVKQLVDNGISSDEAKLLTRDICA 224
Query: 529 ---KRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSV 585
+ E+ + V++ E I K DPGSF L + S+ R+LCDLGSS+
Sbjct: 225 IMLPKAKKEKAQRVDIAEYIRTITPGLTTEKLPDPGSFVLDCSISTSRFSRSLCDLGSSI 284
Query: 586 NLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDE 645
NLM S+ ERL + + +PT + L ADRS G++EDV V+VG P DFV++D ++
Sbjct: 285 NLMSKSVVERLGMTQYRPTRITLLFADRSKRISEGILEDVPVKVGNSIIPADFVVLDYEK 344
Query: 646 DSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGFNIFDLK-PKPV 693
+ K PLILGR FLAT+ A+ +V +G I L+V D ++ F + L+ KP+
Sbjct: 345 EPKNPLILGRAFLATAAARFDVKRGRIFLKVCDSEMEFGMDSLELTKPI 393
>At4g07780 putative athila transposon protein
Length = 446
Score = 125 bits (315), Expect = 7e-29
Identities = 77/211 (36%), Positives = 115/211 (54%), Gaps = 12/211 (5%)
Query: 523 KEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLG 582
K+ +K + + E++ ++ + K DPGSFTLP + G R LCDLG
Sbjct: 247 KKFKIEKLKKDQNQEMISRVSTQETYKKKFIQEKLDDPGSFTLPCSLGPLTFNRCLCDLG 306
Query: 583 SSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIID 642
+SV+LMPLS +RL + E K + L L D S+ P G++ ++ V++G E P DFV++D
Sbjct: 307 ASVSLMPLSTAKRLGIVEYKFCNLALLLPDGSVAHPHGLIGNLPVKIGNDEIPTDFVVLD 366
Query: 643 MDEDSKIPLILGRPFLATSQAKINVGKGTISLRVAD-EKIGFNIFDLKPKPVEKNDVFLV 701
DE+ K PLILGRPFLA++ A I+VG G I L + ++ FNI K + F +
Sbjct: 367 TDEEGKDPLILGRPFLASAGAVIDVGNGKIDLNLEKCIEMRFNISKASGKSTTRGQSFGI 426
Query: 702 KMMEEWSDEKLKQFFLKEKAGASNKKKKEDT 732
++M+ + E+ A K KKE T
Sbjct: 427 QVMD-----------VDEETEAVKKIKKEFT 446
>At4g03850 putative transposon protein
Length = 334
Score = 124 bits (310), Expect = 3e-28
Identities = 91/280 (32%), Positives = 143/280 (50%), Gaps = 23/280 (8%)
Query: 488 LEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSV 547
+E+ S ++ +P EVL + K ++ ++ ++ ++ ++
Sbjct: 1 MERYKSVAAKQLDEIEAVMPLIEVLNLIHDPHKNVRNLILERIKMYHDSNDDSDATPSRA 60
Query: 548 ILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMM 607
+R + K DPGSFTLP + G LCDLG+SV+LMPLSM RL + +P +
Sbjct: 61 ADKRIVQEKFGDPGSFTLPCSIGELAFSDCLCDLGASVSLMPLSMARRLEFIQYRPCDLT 120
Query: 608 LQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINV 667
L LADR+ P+G+++D+ V + E P+DFV++DM+ + K PLILGRPFLA+ A I+V
Sbjct: 121 LILADRASRKPFGMLKDLPVMINGVEVPIDFVVLDMEVEHKDPLILGRPFLASVGAVIDV 180
Query: 668 GKGTISLRVADEKIGFNIFDLKPKPVEKND------VFLVKMMEEWSDEKLKQFFLKEKA 721
+G ISL + K FD+ P E + V E++ EK+K+
Sbjct: 181 REGKISLNLG--KHIMLQFDINKTPQESKEDEKTSGDDRVIPGEKYETEKVKEL------ 232
Query: 722 GASNKKKKEDT*INQTEQVYSMSMVVNSELKEKINTKQKE 761
KK+ D Q E + ++ V EL+ K+N QKE
Sbjct: 233 -----KKRSD---KQEETIERLAHSV-EELRSKLNQLQKE 263
>At1g37070 hypothetical protein
Length = 590
Score = 124 bits (310), Expect = 3e-28
Identities = 67/163 (41%), Positives = 99/163 (60%)
Query: 523 KEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLG 582
K+ +K + + E++ ++ + K DPGSFTLP + G R LCDLG
Sbjct: 220 KKFKIEKLKKDQNQEMISSASTQETYKKKIIQEKLDDPGSFTLPCSLGPLTFNRCLCDLG 279
Query: 583 SSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIID 642
+SV+LMPLS +RL + E K + L LAD S+ P G++E++ V++ E P DFV++D
Sbjct: 280 ASVSLMPLSTAKRLGIMEYKFCNLALLLADGSVAHPHGLIENLPVKIENVEIPTDFVVLD 339
Query: 643 MDEDSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGFNI 685
+DE K PLILGRPFLA++ A I+V G I+L + K+ F+I
Sbjct: 340 VDEKGKDPLILGRPFLASAGAVIDVRNGKINLNLEGIKMKFDI 382
>At2g10670 pseudogene
Length = 929
Score = 123 bits (309), Expect = 3e-28
Identities = 105/365 (28%), Positives = 171/365 (46%), Gaps = 41/365 (11%)
Query: 393 NVRKTPGMFPSDTVINPKE--NCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGE 450
+V P P + NPK+ AIT+ L + V +N D + + E +
Sbjct: 437 SVNNNPCQLPGKAIQNPKQYSTAHAITITHDRELPT---RYVPTSNTEDSVILEGEDFYQ 493
Query: 451 NKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSE 510
+ EE + P PF ++ E +K I V P+
Sbjct: 494 DDVLADNPIEEPILTSQPTRPQAPPATPFA-----KKTEAAKTKDIDF-----VPPPYKP 543
Query: 511 VLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFG 570
+L P +F K +++K R L E++ +C ++ +
Sbjct: 544 LL---PFPGRFKKVLVAKYRALLEKHIKDIPLVDCLALIHDE------------------ 582
Query: 571 ASKQV---RALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLV 627
KQ+ LCDLG+SV+LMPLS+ ++L KP + L LADR+ TP+G++EDV V
Sbjct: 583 -HKQLTFNNCLCDLGASVSLMPLSVVKKLGFVHYKPCDLTLILADRTSKTPFGLLEDVPV 641
Query: 628 RVGEFEFPVDFVIIDMDEDSKIPLILGRPFLATSQAKINVGKGTISLRVA-DEKIGFNIF 686
+ E P DFV+++MD +SK PLILGRPFLA+ A I+V +G I+L + D K+ F+I
Sbjct: 642 MINGVEVPTDFVVLEMDGESKDPLILGRPFLASVGAVIDVKQGKINLNLGEDVKMKFDIR 701
Query: 687 DLKPKPVEKNDVFLVKMMEEWSDEKLKQFFLKEKAGASNKKKKEDT*INQTEQVYSMSMV 746
D KP + FLV+ M++ +E L++ ++ + + E ++ Y MS+
Sbjct: 702 DAMKKPTIEGQTFLVEEMDQLGNELLEEIVDRDHLQTTLTESSEAGFLSSETLSYEMSLD 761
Query: 747 VNSEL 751
++E+
Sbjct: 762 SHNEV 766
>At2g10660 putative Athila retroelement ORF1 protein
Length = 1303
Score = 121 bits (304), Expect = 1e-27
Identities = 75/226 (33%), Positives = 122/226 (53%), Gaps = 3/226 (1%)
Query: 469 KKFTKVPFPP--FPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEIL 526
K+F P P P K +E+ S ++ + +VL +P K ++ +
Sbjct: 338 KRFKPAPLPSRALPWKFRKAWIERYNSLAEKQLDEMEAVMLLIKVLNLIPDPHKDVRNSI 397
Query: 527 SKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVN 586
++ ++ +++E V ++R + K +DPGSFTLP + G LCDLG+SV+
Sbjct: 398 LERIKMYQDSEDECDANPSRVTVKRSVQEKLEDPGSFTLPCSIGQLVFSNCLCDLGASVS 457
Query: 587 LMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDED 646
LMPLS+ +L + +P + L LADRS P+G+++ + + + E P DFV++ M+ +
Sbjct: 458 LMPLSVARKLEFTQYRPCDLSLILADRSSRKPFGLLKHLPLMINGVEVPTDFVVLGMEVE 517
Query: 647 SKIPLILGRPFLATSQAKINVGKGTISLRVADEKIGFNIFDLKPKP 692
K PLILGRPFLA+ A I+V G ISL + + NI +P+P
Sbjct: 518 PKDPLILGRPFLASVGAMIDVKDGRISLNLGKTTVEENI-RAQPQP 562
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.337 0.147 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,444,672
Number of Sequences: 26719
Number of extensions: 708267
Number of successful extensions: 5167
Number of sequences better than 10.0: 101
Number of HSP's better than 10.0 without gapping: 48
Number of HSP's successfully gapped in prelim test: 54
Number of HSP's that attempted gapping in prelim test: 4869
Number of HSP's gapped (non-prelim): 222
length of query: 765
length of database: 11,318,596
effective HSP length: 107
effective length of query: 658
effective length of database: 8,459,663
effective search space: 5566458254
effective search space used: 5566458254
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0265.8


